Abstract
Purification of specific DNA–protein complexes is a challenging task, as the involved interactions can be both electrostatic/H-bond and hydrophobic. The chromatographic stringency needed to obtain reasonable purifications uses salts and detergents. However, these components elicit the removal of proteins unspecifically bound to the chromatographic support itself, thus contaminating the purification products. In this work, a photocleavable linker connected the target oligonucleotidic sequence to the chromatographic beads so as to allow the irradiation-based release of the purified DNA–protein complexes off the beads. Our bioanalytical conditions were validated by purifying the tetracycline repressor protein onto a specific oligonucleotide. The purification factor was unprecedented, with a single contaminant. The robustness of our method was challenged by applying it to the purification of multiprotein assemblies forming onto DNA damage-mimicking oligonucleotides. The purified components were identified as well-known DNA repair proteins, and were shown to retain their enzymatic activities, as seen by monitoring DNA ligation products. Remarkably, kinase activities, also monitored, were found to be distinct on the beads and on the purified DNA–protein complexes, showing the benefits to uncouple the DNA–protein assemblies from the beads for a proper understanding of biochemical regulatory mechanisms involved in the DNA–protein assemblies.
INTRODUCTION
Nucleic acid–protein complexes based on specific interactions have recently been the matter of a great number of contributions in analytical biochemistry (1). To study these interactions, the purification of nucleic acid–protein complexes is generally carried over by mixing standard chromatography techniques and specific affinity methodologies. Often, the preliminary chromatographical steps are conventional and aim at removing entire classes of undesirable analytes. For example, Yaneva and Tempst (2) used a first phosphocellulose fractionation of nuclear extracts to eliminate negatively charged molecular species (saccharides, proteins and nucleic acids). The remaining chromatographic steps are based on the specific or non-specific interaction of proteins with oligonucleotidic target sequences (2,3). This sequence of chromatographical steps makes the purification of nucleic acids interacting proteins time-consuming and very large amounts of the initial sample might be needed.
Affinity chromatography procedures most often involve the coupling of an appropriate specificity determinant molecule (an oligonucleotide bearing a specific target sequence, for example) to a chromatographic support (CNBr-activated or streptavidin-coated agarose beads, for example) in order to craft an affinity chromatography resin. Following washes of the resin, the retained molecules are eluted directly from the chromatographic phase. These methodologies were used to set up so-called one-step purification procedures (4,5). The authors prepared their affinity chromatography phase with biotinylated oligonucleotides bearing the target sequence that were used to functionalize streptavidin-coated beads. Because the chromatographic support (agarose or polyacrylamide beads, for example) is of huge dimensions with respect to the affinity determinant (the oligonucleotide), it lends itself favourably to non-specific interactions with the analytes in the sample thus leading to high contamination levels upon elution of the analytes of interest by applying either salts or detergents (or both) onto the whole chromatographic phase. The analytes of interest are thus less well purified.
In order to limit this adverse effect, we reasoned along with others (6–12) that the uncoupling of the affinity determinant (along with potential bound molecular species) from the chromatographic support itself would yield much more useful purifications of analytes present in the initial sample at very low concentrations. A number of systems have been devised in order to allow the uncoupling of the DNA–protein assemblies from the chromatographic support according to biologically compatible mechanisms. Shimkus et al. (6) chose a disulfide bond-containing linker to couple the oligonucleotidic target sequence to a biotin moiety that was later attached to streptavidin-coated agarose beads. The uncoupling of the nucleic acid–protein assembly from the chromatographic support was triggered by incubating the chromatographic phase with a reducing agent. Bachler et al. (9) and Hartmuth et al. (10) devised a competitive elution strategy based on the use of aminoglycosidic antibiotic-coated chromatographic supports onto which an oligonucleotide linked to an aptamer specifically binding to the antibiotic was attached. The uncoupling of the oligonucleotide from the chromatographic support was achieved by adding excess amounts of the antibiotic which competed for the aptamer. Martinez et al. (11) described a procedure by which the target nucleic acids (a double-strand RNA eicosamer) was linked to a biotin via a photocleavable linker. The chromatographic phase was prepared by attaching the biotin-conjugated target RNA duplex to modified avidin-coated beads. Upon UV irradiation of the chromatographic phase the oligonucleotide is detached from the chromatographic support. Strategies based on the use of reducing agents or antibiotics have some drawbacks such as the addition of molecules in the purification medium that can disturb the subsequent analyses, which prompted us to opt for the system employing the photocleavable linker.
In this report, we describe a simplification of this method, specifically aimed at generalizing it to any situation involving the affinity-based purification of DNA-interacting proteins. We simplified experimental conditions that allowed—without loosing any purification efficiency—the use of a single buffer composition throughout the whole purification process. Our bioanalytical conditions were validated by affinity purifying the tetracycline repressor protein (TetR) expressed in quite low amounts in eukaryotic cells (13) onto an oligonucleotide bearing its cognate sequence (TetO). The purification efficiency was unprecedented with a single contaminant protein being co-purified with the repressor protein. The robustness of our method was challenged with a highly complex DNA–protein system that has the advantage of being well studied: the DNA repair machinery which comprises a large number of proteins that assemble onto DNA damages (double strand breaks, in our case) in supramolecular structures (14). We succeeded in purifying these protein assemblies: the purified components were identified as well-known DNA repair proteins. Our work shows that these proteins retained their enzymatic activities throughout the purification, as seen by monitoring DNA ligation products. Further, kinase activities, also monitored in our experiments, were found to be distributed distinctly either on the beads or on the purified DNA–protein complexes. These results showed the major benefits of the uncoupling of the purified DNA–protein assemblies from the beads as far as a detailed and accurate understanding of the biochemical regulatory mechanisms involved in the assembly/disassembly of DNA–protein complexes is concerned.
MATERIALS AND METHODS
Cell culture and cellular extracts
T-Rex HeLa cells (Invitrogen, Cergy Pontoise, France) stably express TetR under the control of the human CMV promoter (15). These cells were cultured to subconfluency in minimal essential medium (MEM; Invitrogen) supplemented with 10% fetal bovine serum and non-essential amino acids (Sigma, Lyon, France) with 100 U/ml penicillin and 100 μg/ml streptomycin. Cells were grown at 37°C in a 5% CO2 atmosphere.
Cellular extracts were obtained according to Baron et al. (16). Briefly, cells were harvested in the following lysis buffer: 10 mM HEPES pH 7.2, 1.5 mM MgCl2, 10 mM KCl, 0.5 mM dithiothreitol and 1 mM phenylmethylsulfonyl fluoride supplemented with the Complete Protease Inhibitor Cocktail (Roche Molecular Biochemicals, Mannheim, Germany) and subjected to freeze/thaw cycles to disrupt their membrane. Cell lysates were cleared by centrifugation at 20 000 g at 4°C for 30 min. The protein concentration in the resulting supernatant was determined according to the Bradford protein assay (Bio-Rad Laboratories, Marnes-la-Coquette, France), bovine serum albumin was used as the standard. The cellular extracts were brought to a final protein concentration of 10 mg/ml with lysis buffer.
HeLa nuclear protein extracts were prepared according to Dignam et al. (17) (Cilbiotech, Mons, Belgium).
Conjugated oligonucleotide
A photocleavable moiety was coupled to a biotin moiety to form the photocleavable biotin (PCB) linker which was conjugated to the oligonucleotide according to the chemistry described in (18,19). Briefly, it was covalently linked through its 5′phosphate end to the 1-(2 nitrophenyl)ethyl photo-reactive group that is itself bound to the biotin moiety via a 6-aminocaproic acid linker (Figure 1A). It was synthesized by Eurogentec (Seraing, Belgium) using commercially available phosphoramidite derivatives. The PCB-TetO was a 58 bp double-strand DNA 
 conjugated to the photocleavable biotin label sequence. The DNA sequence comprised the TetO sequence (20) and a
conjugated to the photocleavable biotin label sequence. The DNA sequence comprised the TetO sequence (20) and a  sequence stretch that was added to the 5′P oligonucleotidic end thus increasing the distance between the TetO target sequence and the chromatographic support so as to diminish the steric hindrance that might hamper the interaction between the proteins of interest and the TetO sequence (21).
sequence stretch that was added to the 5′P oligonucleotidic end thus increasing the distance between the TetO target sequence and the chromatographic support so as to diminish the steric hindrance that might hamper the interaction between the proteins of interest and the TetO sequence (21).
Figure 1.
Overview of the nucleic acid-binding protein purification strategy. (A) Schematic representation of the chemistry involved in the preparation of the chromatographic resin. The 5′P of the oligonucleotide is conjugated to the photo-reactive group coupled itself to a biotin through a C6 spacer. The oligonucleotide is incubated with streptadivin-coated magnetic beads to yield the chromatographic solid phase. (B) Purification procedure. Protein extracts are incubated with the chromatographic resin prepared in (A). After magnetic harvesting of the beads, washes are performed. Photocleavage of the linker, with near UV-light irradiation, releases the DNA–protein complexes, thus minimizing their contamination by the proteins unspecifically bound to the beads.
Antibodies
Anti-TetR antibody polyclonal rabbit was from Mobitec (Goettingen, Germany). Anti-DNA-PKcs (clone 18-2), anti-Ku70 (clone N3H10) and anti-Ku80 (clone 111) monoclonal mouse antibodies were from Neomarkers (Fremont, USA) and anti-PARP-1 antibody was from AbD Serotec (Cergy Saint-Christophe, France). Anti-XRCC4 polyclonal rabbit antibody was from Calbiochem (La Jolla, USA).
Electrophoretic mobility shift assay
The PCB-TetO was radiolabeled using the T4 polynucleotide kinase (New England Biolabs, Saint-Quentin-en-Yvelines, France) and [γ−32P]ATP (GE Healthcare). Binding reactions were performed for 20 min at room temperature with 50 μg of extracts from T-Rex HeLa cells and 25 fmol of 5′P radiolabeled PCB-TetO in a final volume of 10 μl. Binding buffer was 10 mM Tris–HCl pH 7.5, 5% glycerol and 2 mg/ml sheared genomic DNA competitor (fish sperm DNA, Roche), with or without tetracycline (1 μg/ml). Complexes were resolved by non-denaturing electrophoresis on 5% polyacrylamide/0.13% bisacrylamide gels containing 5% glycerol running in buffer 50 mM Tris base, 45 mM boric acid and 0.5 mM EDTA at room temperature and at 15 V/cm (16). The oligonucleotide was radiolabeled after the purification (see Figure 2B) only if a western blot analysis was performed in parallel (see Figure 3A). Total 1/20 of the purification product was radiolabeled and analyzed by EMSA. The quantification of radioactivity was performed using the Image Quant software (GE Healthcare).
Figure 2.
EMSA analysis of fractions obtained from nucleic acid-binding protein purification. (A) To observe the band shift corresponding to the TetR-TetO interaction, whole cell extracts were mixed with the PCB-TetO bait in absence or in presence of tetracycline (Tet; lanes 2 and 3, respectively). To perform the purification, whole cell extract was incubated with the PCB-TetO immobilized on beads, then the presence of the complex was analyzed in the purification product (lane 4). (B) Tetracycline was mixed with the purification product to confirm the specificity of the DNA–protein interaction. The asterisk indicates that only 1/2 of the corresponding fraction was loaded.
Figure 3.
Purification of the tetracycline repressor protein from whole human cell extracts by affinity for its cognate sequence. Whole cell extracts were mixed with the chromatographic phase followed by washes. (A) Supernatant and pellet fractions obtained from irradiated and non-irradiated samples were analyzed by SDS-PAGE gel electrophoresis and proteins were blotted onto a membrane and revealed by Sypro blot-staining (left panel) and subsequently anti-TetR antibody-staining (right panel). (B) 2D-gel electrophoresis of proteins released after detergent treatment of the whole chromatographic material (left panel) and of DNA–protein complexes recovered in the supernatant after irradiation (right panel). The proteins were stained with Sypro Ruby. The arrows point to the TetR spot. The asterisk indicate that only 1/3 of the pellet fractions was loaded. MW, molecular weight markers.
DNA affinity purification of protein complexes
In order to purify the TetR protein from T-Rex HeLa cells, the PCB-TetO duplex was immobilized on streptavidin-coated magnetic beads (Roche) according to the protocol provided by the manufacturer. To bind 0.5 pmol of the PCB-TetO oligonucleotide onto the beads, 1.25 pmol of duplex was incubated with 42 μg of magnetic beads followed by two equilibration steps with 84 μl of buffer A containing the lysis buffer supplemented with one volume of 2× buffer B (20 mM Tris–HCl pH 7.5 and 10% glycerol). Incubation of 1 mg protein extract with the bait-coated beads was performed at room temperature for 20 min in buffer A supplemented with 2 mg/ml fish sperm DNA (Roche). Five rounds of washes were performed by alternatively sedimenting the resin with a magnet and resuspending it with buffer A. The resuspended beads (50 μl of buffer A) were irradiated 10 min with a UXM-200HO xenon-mercury lamp (λ > 300 nm, 90 mW/cm2 at 365 nm, 5% light transmittance at 295 nm) (Lot-Oriel, Palaiseau, France). The beads were pelleted with a magnet; the supernatant contained the released oligonucleotidic bait along with the proteins bound to it. The different purification fractions (10 μl) were analyzed by electrophoretic mobility shift assay (EMSA).
When the purification products were to be analyzed either by 1D- or by 2D-gel electrophoresis, the experiment above was scaled-up by a factor 5 or 10, respectively. The supernatant volume was reduced to 5 μl by filtration with Microcon YM-10 devices (Millipore, Saint-Quentin-en-Yvelines, France) and mixed either with an equal volume of 2× Laemmli buffer (1D-gel electrophoresis) or with 120 μl of loading buffer (2D-gel electrophoresis).
In order to purify the proteins involved in DNA-end joining, 40 pmol of the PCB-TetO were immobilized onto the chromatographic support by mixing 100 pmol of duplex with 500 μg of magnetic beads. The chromatographic phase was equilibrated with 1000 μl of buffer C (40 mM HEPES-KOH pH 7.8, 5 mM MgCl2, 60 mM KCl, 0.5 mM dithiothreitol, 0.4 mM EDTA, 3.4% glycerol and 0.01% Nonidet P-40 substitute) and was later incubated with 400 μg of HeLa nuclear protein extracts in the absence of DNA competitor at 30°C for 30 min in a final volume of 170 μl. After five cycles of washes with buffer C, the beads were resuspended in 500 μl of buffer C and were irradiated 10 min; the supernatant contained the oligonucleotide and its associated proteins.
1D-gel electrophoresis and western blotting
Protein mixtures (either proteins bound to the DNA or proteins bound to the chromatographic phase) from irradiated or non-irradiated samples were separated on acrylamide denaturing electrophoresis gels. The proteins bound to the beads (irradiated or non-irradiated) were stripped off with a denaturing buffer and the proteins in the supernatants were filtration-concentrated as above. Proteins from TetR or DNA repair complexes purified fractions were mixed with the 2× Tris–glycine SDS sample buffer (Novex, Invitrogen) supplemented with 10% β-mercaptoethanol or Laemmli buffer, respectively and were later separated on 14% acrylamide denaturing Tris–glycine electrophoresis gels (Novex, Invitrogen) or 8% SDS-PAGE gels, respectively. Electrophoresed proteins were then transferred from the gel onto a nitrocellulose membrane (Hybond ECL, GE Healthcare). Protein bands were visualized by incubation of the membrane with Sypro Ruby protein blot stain (Invitrogen; visualization device: Typhoon 9410 fluorescent scanner, GE Healthcare). The same membrane was used for western blotting, revealed using the ECL Plus kit (GE Healthcare) and imaged using the same Typhoon 9410 apparatus. The BenchMark Protein ladder (Invitrogen) or the Perfect Protein marker (Novagen, Wisconsin, USA) was used as molecular weight markers.
2D-gel electrophoresis and mass spectrometry
Home-made 2D electrophoretic gels were prepared and run as described previously (22). The concentrated supernatant was mixed with 120 μl of 2D-gel loading buffer (8.75 M urea, 2 M thiourea, 6% CHAPS, 20 mM dithiothreitol). Proteins from the non-irradiated chromatographic phase were eluted with the same buffer. Samples were then loaded onto pH [3–10] 7 cm-long Immobiline DryStrips (GE Healthcare) for the first dimension and on 11% SDS-PAGE gels for the second dimension. The gels were stained with Sypro Ruby protein gel stain (Invitrogen).
Spots of interest were digested in-gel with trypsin and the peptidic mixture was analyzed by MALDI-TOF mass spectrometry as previously described (22,23) except for the reduction and alkylation steps that were performed before the denaturing electrophoresis, during the Immobiline DryStrips equilibration. Sequence editing and mass spectral data simulations were performed using the GNU polyxmass Free Software package (24).
Gas-phase fragmentation experiments were performed in the positive-ion mode using a hybrid quadrupole time-of-flight mass spectrometer equipped with a Protana source (nanoESI MS, Q-Star Pulsar i, MDS Sciex-Applied Biosystems). Data acquisition and storage were performed using the Analyst QS software package shipped with the mass spectrometer. Ions were selected based on their [M+2H]2+ m/z value, in the mass unit mode, using the Analyst QS software package.
Protein kinase assay
The protein kinase activities associated with purification fractions were detected by monitoring the phosphorylation level of the XRCC4 protein, a component of the DNA repair protein complexes that assembled onto our DNA damage-mimicking oligonucleotide. The purification of proteins involved in DNA repair was performed as described above, starting from HeLa nuclear extracts depleted in ATP with 8 U· ml−1 hexokinase and 2 mM glucose for 10 min at 30°C. Following incubation of the protein extracts with the chromatographic resin, five rounds of washes were performed and the chromatographic phase was resuspended in 20 μl of buffer C for further kinase assays. Kinase assay conditions were: 1 mM ATP, incubation for 2h at 30°C. When the kinase activities were monitored in presence of the beads (i.e. without irradiation), the chromatographic phase was incubated without or with ATP, and only after were the DNA–protein assemblies released from the beads by irradiation. When the kinase activities were monitored on the released DNA–protein complexes, the irradiation step was performed and the supernatant was collected for kinase assay. In either case, the proteins were migrated on SDS-PAGE gels for further western blot analysis.
When indicated, 10 μM wortmannin (Sigma), a kinase inhibitor, was incubated with the sample assayed for kinase activities on ice for 30 min right before the addition of ATP (25). The samples were desalted in a microchromatography device prepared according to Rusconi et al. (26) with small modifications: Poros resin (kind gift from Dr Carole Feltaille, Applied Biosystems, Coutabœuf, France) was packed in a microloader pipette tip (3 μl packed resin bead) and was equilibrated with 500 μl of 1% formic acid. The sample (20 μl) was deposited on top of the resin bead and acidified in place with 2 μl of pure formic acid. Desalting was performed by passing 500 μl of 1% formic acid and the proteins were eluted with 500 μl of 80% acetonitrile–1% formic acid (v/v). The proteins were lyophilized, electrophoresed (14% SDS-PAGE) and later transferred onto a nitrocellulose membrane. Western blotting was performed with anti-XRCC4 antibody. To determine the phosphorylated forms of the XRCC4 protein, 10 or 5 U of calf intestinal phosphatase (New England Biolabs) were incubated at 37°C for 60 min with 30 μg HeLa nuclear extracts or the purification products, respectively.
Oligonucleotide ligation assay
The purification of proteins involved in DNA repair (double-strand DNA breaks) was performed as described in the protein kinase assay paragraph. The proteins associated to the target oligonucleotide were incubated with 1 mM ATP for 2 h at 30°C before or after the UV-irradiation of the chromatographic slurry. The purification products were then incubated with 200 μg ml−1 proteinase K (Ambion, Courtabœuf, France) at 55°C for 60 min. The proteins were removed by phenol-chloroform-isoamyl alcohol treatment and the recovered oligonucleotides were precipitated. The samples were electrophoresed on 8% polyacrylamide/0.42% bisacrylamide gel running in 50 mM Tris base, 45 mM boric acid and 0.5 mM EDTA buffer. Denaturation of the nucleic acids was performed by incubating the gel in 1.5 M NaOH for 30 min; it was then neutralized by soaking it in a 1 M Tris–HCl pH 7.5; 3 M NaCl solution. The Southern blotting was performed in the experimental conditions described in (27). The hybridization with the 5′[γ−32P] radiolabeled non-photocleavable strand of the PCB-TetO duplex was at 55°C. The positive ligation control was obtained by incubating 25 pmol of the TetO oligonucleotide with 0.4 U of T4 DNA ligase (New England Biolabs) for 120 min at room temperature in the buffer provided by the manufacturer.
RESULTS
General purification procedure
The schematic in Figure 1 shows that our strategy involves the use of a photocleavable linker molecule between the oligonucleotide that will serve as the bait for the purification and the chromatographic support (magnetic streptavidin-coated beads). The functionalization of the magnetic beads with the oligonucleotidic bait is actually made possible because the linker molecule is itself bound to a biotin moiety. All the chemical components involved in the production of the chromatographic phase are available in the commerce, thus making the method straightforward.
Once the chromatographic resin had been produced, cellular extracts were incubated batch-wise with it and several washes were performed to eliminate most contaminants. Usual procedures involve the use of washing buffers of increasing stringency, with the unwanted effect that analytes bound to the affinity determinant might detach along with unspecifically bound molecules. In our strategy, the washing buffer was actually the same as the incubation buffer, thus we did not expect to loose any analyte bound to the oligonucleotide via labile yet specific interactions.
After the washing steps and prior to the elution of the proteins bound to the oligonucleotidic bait, we UV-irradiated the chromatographic phase so as to detach the oligonucleotide–protein complexes off the magnetic beads. Following pelleting of the beads, the supernatant contained the DNA–protein complexes of interest that could thus be obtained in their native state, since the photocleavage step did not require any denaturing component.
We first applied this method to a simple case: the purification of the tetracycline repressor protein (TetR) from crude cellular extracts. The TetR homodimer is known to regulate bacterial genes responsible for tetracycline resistance by binding to the tetracycline operator sequence (TetO). TetR and its derivatives are widely used to modulate the expression of ectopic genes in eukaryotic cells (13). T-Rex HeLa cells stably express TetR under the control of the human cytomegalovirus promoter (15). Using an anti-TetR antibody and different amounts of the pure TetR protein we estimated by western blot that the TetR protein represents < 0.008% of total proteins in the protein extract from this cell line (data not shown).
DNA–protein complex formation
In a preliminary experiment, the ability of the TetR protein expressed in the T-Rex HeLa cells to bind to the oligonucleotidic cognate sequence was established by incubating—in presence or absence of tetracycline—whole cell extracts with the radiolabeled oligonucleotide. The formation of DNA–protein complexes was monitored by the electrophoretic mobility shift assay (EMSA, Figure 2A, lanes 1–3). The band shift observed in lane 2 indicates that at least one protein did bind to the oligonucleotide. Addition of tetracycline in the incubation mixture abolished that observed band shift, suggesting that the oligonucleotide had been bound by the tetracycline repressor (lane 3). Indeed, tetracycline inhibits the interaction between the TetR protein and its cognate TetO sequence (28).
The result from a typical chromatographic experiment is shown in Figure 2A, lane 4, where the chromatographic phase was first assembled by incubating the biotin-labeled oligonucleotide with the streptavidin-coated magnetic beads. The T-Rex HeLa cellular extracts were then batch-wise incubated with the chromatographic resin. The beads were pelleted and the supernatant (equivalent to a flowthrough) was collected prior to performing five washes of the resin. The chromatographic slurry was UV-irradiated, thus releasing the oligonucleotide along with any protein bound to it. The band shift observed in lane 4 shows that the oligonucleotide had effectively been recognized by a protein. Further, when the purification product was incubated with tetracycline, that band shift was abolished, showing that the oligonucleotide was bona fide recognized by the tetracycline repressor (Figure 2B). The ratio between the free oligonucleotide and the bound oligonucleotide in the product (Figure 2A, lane 4) was 60% lower than the one obtained in the solution (Figure 2A, lane 2). This observation might be explained by the fact that the binding of the oligonucleotide onto the beads might sterically hamper the interaction between the TetR protein and the DNA sequence. Inspired by others (9,11,29), we performed a purification experiment in which proteins were first incubated with the oligonucleotide and the formed DNA–protein complexes were attached to the beads only thereafter. In our hands the pre-incubation of the free oligonucleotide with the cellular extracts brought no significant advantage with respect to our first methodology, confirming the hypothesis of steric hindrance. We thus standardized our experiments with the procedure in which the chromatographic resin is prepared prior to its incubation with the cellular extracts.
Benefits of the photocleavage
The previous experiments allowed us to monitor and quantify the binding of a protein onto its cognate sequence in a radiolabeled oligonucleotide. In order to get a better view of the separative performance of our chromatographic phase we analyzed—by 1D- or 2D-gel electrophoresis—the purification products obtained with different chromatographic methodologies.
Cellular extracts were incubated with the chromatographic phase and, following thorough washing of the beads, the chromatographic slurry was either UV-irradiated or mixed with denaturing Laemmli sample buffer. In each case, the beads were magnet-pelleted and the obtained fractions—supernatant and pellet—were further resolved by SDS-PAGE. The proteins were transferred onto a membrane (Figure 3A) that was first stained with the Sypro blot stain (left panel) and then used to probe the presence of the tetracycline repressor protein with a polyclonal antibody (right panel). The left panel shows that the vast majority of the proteins are found attached to the beads, in the pellet fractions, with or without previous UV-irradiation of the chromatographic phase (lanes 2 and 3). The supernatants (lanes 4 and 5) only show a faint band at ca 40 kDa. The presence of that band in the supernatant fractions—obtained with or without irradiation—indicated that it did not correspond to a molecular species specifically bound to the oligonucleotide but that it had probably leaked from the beads themselves. The western blot shown in the right panel confirmed that the tetracycline repressor protein effectively interacted with the beads (lane 8). Indeed, when no UV-irradiation was performed, the supernatant did not contain the repressor (lane 10), which was found in the pellet fraction instead (lane 8). Upon UV-irradiation of the chromatographic slurry, 75% of the tetracycline repressor protein remained on beads after washes was recovered in the supernatant (lane 9), showing that most of the repressor protein actually interacted with the beads via the oligonucleotide.
Because the repressor protein band appeared somewhat thick, the same experiment was performed but was followed by a 2D-gel electrophoresis in order to further resolve potentially overlapping protein variants in that band. Equally significant is the fact that 2D-gel electrophoreses do concentrate proteins in spots that are more suitable for further analysis of their contents by mass spectrometry. The 2D-gel obtained without irradiation (Figure 3B, left) corresponds to the biological material recovered on the beads in the same way as for lane 3 of Figure 3A. When the chromatographic slurry was irradiated, the supernatant did contain only two molecular species visible as spots at ca 40 kDa and 25 kDa (Figure 3B, right). The lower spot is visible on both gels and is pointed to by an arrow. We suspected this spot to contain the tetracycline repressor protein, which was later confirmed by mass spectrometry. The two detected spots were excised from the gel and subjected to in-gel trypsinolysis. The peptidic mixtures were analyzed by matrix-assisted laser desorption/ionization mass spectrometry. The peptide mass fingerprint results are shown in Table 1 for the lower spot, indicating that the protein contained therein was the tetracycline repressor protein (30% sequence coverage; 8 matching peptides). The ∼ 40 kDa protein was identified as human β-actin (34% sequence coverage; 13 matching peptides; data not shown). Actin is one of the most abundant proteins in eukaryotic cells, as can be seen on 2D gels where actin makes one of the largest spots at roughly 40 kDa. It is thus possible that this protein leaked from the beads even after the numerous washing steps performed during the purification. The definitive confirmation of the identity of the protein contained in the ∼ 25 kDa spot was brought by nanospray tandem mass spectrometry with gas-phase fragmentation of the [M+2H]2+ parent ion at m/z 734.34. The obtained sequence is shown in Table 2.
Table 1.
MALDI-TOF analysis of proteins recovered from 2D gel after purification on PCB-TetO beads
| Measured mass (M) | Computed mass (M) | ΔM | Residues [start to end] | Peptide sequence | Chemical modification |
|---|---|---|---|---|---|
| 855.507 | 855.438 | 0.069 | [88–94] | CALLSHR | cyst* |
| 1136.679 | 1136.630 | 0.049 | [99–108] | VHLGTRPTEK | |
| 1226.680 | 1226.618 | 0.062 | [88–98] | CALLSHRDGAK | cyst* |
| 1442.803 | 1442.780 | 0.024 | [50–62] | ALLDALAIEMLDR | |
| 1458.793 | 1458.774 | 0.019 | [50–62] | ALLDALAIEMLDR | metox |
| 1472.744 | 1472.717 | 0.027 | [159–171] | ETPTTDSMPPLLR | metox |
| 1568.889 | 1568.834 | 0.055 | [34–46] | LGVEQPTLYWHVK | |
| 1886.896 | 1886.903 | −0.008 | [156–171] | EERETPTTDSMPPLLR | metox |
cyst*= carbamidomethyl cysteinyl residue, metox = oxidized methionyl residue and (M) = monoisotopic mass.
‘Profound’ = software available at http://129.85.19.192/profound_bin/WebProFound.exe
The peptide masses obtained after trypsin digestion of the 25 kDa spot were searched in NCBI database using the ‘Profound’ software with a tolerance of 0.08 Da and found to match (8 peptides) to TetR protein (30% coverage).
Table 2.
Sequence-based identity confirmation of the TetR protein
| E | T | P | T | T | D | S | M | P | P | L | L | R | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| y11 | y10 | y9 | y8 | y7 | y6 | y5 | y4 | y3 | y2 | y1 | |||
| Measured y fragment masses | 1243.59 | n.d. | 1045.48 | 944.44 | 829.43 | 742.39 | 595.36 | 498.32 | 401.26 | 288.19 | 175.11 | ||
| Calculated y fragment masses* | 1243.64 | 1146.58 | 1045.54 | 944.49 | 829.46 | 742.43 | 595.39 | 498.34 | 401.29 | 288.2 | 175.12 |
*Sulfoxide taken into account in the calculation of fragment masses. n.d. : not determined. Database interrogations were performed using the ‘Mascot’ software (41); detailed mass spectral analyses and interpretations were performed using the ‘GNU polyxmass’ mass spectrometric software authored by Filippo Rusconi. ‘Mascot’ : software available at http://www.matrixscience.com. ‘ GNU polyxmass’ : software freely available at http://www.polyxmass.org.
MALDI-TOF mass spectrometric analysis of the peptide mixture yielded one molecular species of m/z 1472.74 ([M+H+]+) which was identified as the ETPTTDSMPPLLR tryptic TetR peptide with a putatively oxidized methionyl residue. Subsequent mass spectrometric analysis of the mixture on an ESI-qQTOF (Q-Star; Applied Biosystems) mass spectrometer afforded an ion at m/z 737.34 ([M+2H+]2+) corresponding to that same peptide. Fragmentation of this ion afforded fragment ions in the y-series (y1 through y9 and y11). Noteworthy, fragments y6 through y9 and y11 showed masses with an increment of 16 amu due to the sulfoxide formation on the methionyl residue.
These results showed that the tetracycline repressor protein could be purified with an unprecedented enrichment factor and that the uncoupling of the affinity determinant (the oligonucleotide) from the chromatographic support (the streptavidin-coated beads) is responsible for that achievement. The specificity of the interaction between the tetracycline receptor protein and its cognate oligonucleotidic sequence is not questionable, as shown by the disruption of that interaction in the presence of tetracycline (Figure 2B). Equally significant is the fact that the DNA–protein complex is purified in its native form, even after the UV-irradiation. Finally, because the chromatography steps are compatible with 2D-gel electrophoresis, the protein was successfully identified by mass spectrometry.
Purification of DNA-binding multiprotein complexes
The results obtained with the TetR/TetO system showed the benefits of the photocleavage approach with respect to conventional elution methods. The TetR/TetO system was rather simple as it involved a binary interaction between a protein and its cognate sequence. Therefore, to challenge the robustness of our method, we applied it to the purification of more complex protein assemblies involving both protein–protein and protein–DNA interactions. The experimental system that we chose was the DNA repair machinery that recognizes double-strand breaks on DNA. Two distinct factors motivated this choice: first, on the protein side, the repair machinery is well studied and a number of proteins taking part to the multiprotein repair assemblies are well known and characterized; second, on the nucleic acids side, the DNA double-strand break to be repaired is easily implemented in the test tube because a duplex with free double-strand ends mimicks a double-strand break and as such is able to recruit the DNA repair machinery (30,31). Two protein complexes are known to be required for DNA double-strand break repair: the DNA-PK holoenzyme and the DNA ligase IV-XRCC4 complex (14). DNA-PK comprises three subunits: the DNA-dependent protein kinase catalytic subunit (DNA-PKcs), Ku70 and Ku80. This heterotrimer binds to the DNA ends and recruits the DNA ligase IV-XRCC4 complex that accomplishes the ligation reaction (31,32).
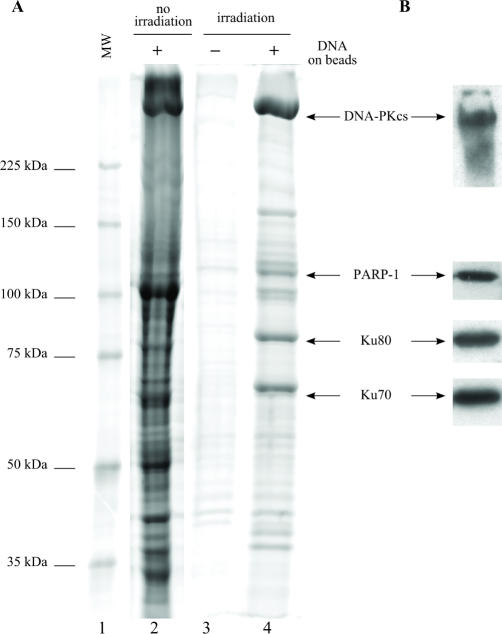
In order to purify these protein assemblies, we coupled to the chromatographic support the PCB-TetO oligonucleotide of which the free blunt ends mimicked double-strand breaks (see sequence in Materials and Methods section). The sequence length of the PCB-TetO oligonucleotide was sufficient to recruit the DNA repair proteins (33). The chromatographic phase was incubated with HeLa nuclear extracts. After washes, the proteins were recovered either without irradiation or after irradiation of the chromatographic resin, resolved by SDS-PAGE and stained (Figure 4A). Without irradiation, proteins from both the beads and the target oligonucleotide were deposited in lane 2, showing a heavy staining pattern. If the DNA–protein complexes were first cleaved off the beads with an irradiation step and the beads removed, the observed staining pattern was much lighter (lane 4), clearly showing that the beads did retain a huge amount of proteins unspecifically bound to them. Interestingly, we performed a purification procedure without DNA on beads and after the irradiation of the chromatographic slurry, the proteins in the supernatant were recovered and stained (lane 4). A very small amount of proteins was released off the chromatographic support, which indicated that the proteins recovered from the purification with DNA were specifically bound to the oligonucleotidic bait (lane 4, compare with lane 3). These results confirmed our initial results with the TetR/TetO binary system, underlining the benefits of the photocleavage step in our method. In lane 3, four major proteins were detected with apparent molecular weights suggesting their identity: Ku70 at 70 kDa, Ku80 at 80 kDa, PARP-1 at 120 kDa (34) and probably DNA-PKcs > 225 kDa. These hypotheses were confirmed by western blotting the gel with antibodies against each one of these proteins (Figure 4B and Supplementary Figure 1). Because saline conditions were identical throughout the purification process, weak protein–protein and DNA–protein interactions could be preserved, letting us hope that the purified material would have retained its native structure and enzymatic activities.
Figure 4.
Affinity purification of the DNA ends-binding proteins. The proteins were purified from HeLa nuclear extracts using the PCB-TetO target sequence immobilized on beads (lanes 2 and 4) and without oligonucleotide on beads as control (lane 3). (A) The chromatographic slurry was either treated with detergents (lane 2) or irradiated (lanes 3 and 4), then the released proteins were separated on an 8% SDS-PAGE gel and stained with Sypro Ruby protein gel stain. (B) The purification product obtained after irradiation was analyzed by western blot with antibodies against the DNA ends-binding proteins as indicated. MW, molecular weight markers.
Enzymatic activities of DNA-binding multiprotein assemblies
Functional studies were performed on the purified DNA–protein complexes by analyzing enzymatic activities known to operate in vitro for such complexes: we first monitored the actual DNA repair via oligonucleotide end-joining and second the phosphorylation of XRCC4 (35,36). In the following section, these different enzymatic activities operating either on the DNA component or on the proteinaceous components of the DNA–protein complex were systematically compared before and after the irradiation step.
In the first experiment, the oligonucleotide ligation was tested either with the DNA–protein assemblies still coupled to the beads (no irradiation) or after their irradiation-mediated release in the supernatant (after removal of the beads). The ligation products were analyzed by Southern blot (Figure 5). No ligation product could be detected when the DNA–protein assemblies were still coupled to the beads, suggesting that the beads may have reduced the ligation rate too much for its products to be detectable (lane 3). Conversely, the TetO oligonucleotide dimer was detected when the DNA–protein complexes were detached from the beads (lane 4) albeit with a low yield of ligation (∼ 1%), as was expected for blunt DNA ends (37).
Figure 5.
Oligonucleotide ligation by DNA ends-binding protein complexes. After washes, the DNA–protein assemblies were incubated with ATP to allow the ligation of the TetO sequence. In lane 3, the ligation was performed on the beads in presence of ATP then the nucleic sequence was released after irradiation of the chromatographic slurry. In lane 4, the DNA–protein assemblies were recovered after the irradiation step then incubated with ATP. The oligonucleotides were resolved on an 8% non-denaturing polyacrylamide gel and revealed by Southern blot with the radiolabeled non-photocleavable strand of the PCB-TetO duplex. Lane 1 shows the intact TetO oligonucleotide and lane 2 the oligonucleotide ligated by the T4 DNA ligase protein.
In the next experiments, we analyzed kinase activities that resulted in the phosphorylation of the XRCC4 protein. We focused our attention on XRCC4 because it is a known target of kinases (amongst which DNA-PKcs) (38), with the advantage that the non-phosphorylated and the phosphorylated species can easily be resolved on SDS-PAGE gels and later detected by western blot (Figure 6). To prevent phosphorylation from occurring throughout the purification process, our starting material was prepared by depleting HeLa nuclear extracts of their ATP content. No phosphatase inhibitors were added during this step. Figure 6A, lane 1 shows the XRCC4 variants in the starting material in the absence of ATP that were detected as a doublet of bands. The heavier variant could be eliminated by incubating the sample with calf intestine phosphatase (CIP), indicating that it was most probably due to phosphorylation (Figure 6B, lanes 8 and 9).
Figure 6.
Analysis of the phosphorylation of the XRCC4 protein. (A) HeLa nuclear extracts were depleted in ATP with hexokinase and glucose then incubated with the oligonucleotide immobilized on beads. After washes, the chromatographic slurry was incubated with ATP for 2 h at 30°C then irradiated (lanes 4 and 6) or first irradiated then incubated with ATP (lanes 5 and 7). To determine the involved kinases, the proteins were pre-incubated with 10 μM of wortmannin (lanes 6 and 7) right before the incubation with ATP. Controls were performed in absence of ATP (lanes 2 and 3). The proteins from HeLa nuclear extracts depleted in ATP and from the purification products were resolved on an 8% SDS-PAGE gel then transferred on a nitrocellulose membrane and immunoblotted with a polyclonal rabbit anti-XRCC4 antibody. (B) HeLa nuclear extract depleted in ATP (lanes 1, 8 and 9) and the purification product (lanes 2–7, 10 and 11) incubated with ATP were treated with calf intestine phosphatase (CIP) to detect the phosphorylated forms of the XRCC4 protein. An asterisk indicates a possible proteolytic product of the XRCC4 protein. Symbols (filled and empty circles, filled square) indicate XRCC4 phosphorylated variants.
For the next series of experiments, the purification of protein assemblies involved in DNA repair was performed in the same manner as before. However, in order to monitor the molecular species involved in the phosphorylation of XRCC4 and infer their main localization (on the beads or on the oligonucleotide), ATP was added at two different steps: either before or after the irradiation-based release of the DNA–protein complexes off the beads. When ATP was added before the irradiation of the chromatographic phase (i.e. when the DNA–protein complexes were still attached to the beads), a number of XRCC4 variants were found (lane 4) of which the ones migrating in the band labeled with a filled circle are the heaviest. When ATP was added to the DNA–protein complexes alone (i.e. after irradiation, so as to detach and remove them from the beads), XRCC4 variants could also be detected, albeit with lower apparent molecular weights than in the previous experiment (lane 5, filled square). In each of these cases, if no ATP was added, no heavy XRCC4 variants could be detected (lanes 2 and 3). In order to ascertain that the molecular variants detected in these experiments reflected the existence of phosphorylated XRCC4, another experiment was performed in which the DNA–protein complexes—without the beads—were incubated with ATP first (same as lane 5) and subsequently incubated with CIP (or without as a control). The results are shown in Figure 6B, lanes 10 and 11, which demonstrate that the heavy molecular variants were indeed XRCC4 phosphorylation variants, as the slow-migrating bands (lane 10) were converted into bands migrating at the same rate as the ones in lane 9 (no incubation with ATP). The same results were shown with the DNA–protein assemblies coupled to the beads and incubated with ATP (same as lane 4, data not shown). Overall, these results demonstrated that with or without irradiation, kinases copurifying with the DNA repair complex (either on beads or on the bait oligonucleotide) retained their activity with or without irradiation. Further, one intriguing result obtained in this series of experiments is that the phosphorylation status of XRCC4 varied significantly depending on the presence or absence of the beads during the incubation of the purification products with ATP. To appreciate this, one can compare lanes 4 and 5 (Figure 6A): if ATP is added when the DNA–protein complexes are still attached to the beads (lane 4), the XRCC4 phosphorylated variants appear conspicuously heavier than those observed in lane 5 (no beads). This result might suggest that kinase activities adsorbed on the beads do phosphorylate XRCC4 in a distinct manner than do the activities interacting with the DNA oligonucleotide.
One might think that the phosphorylation events catalyzed by kinases unspecifically adsorbed on the beads are irrelevant from a strict DNA repair regulation perspective. To challenge that hypothesis, we performed the same kind of experiment as above but with an additional component added to the incubation mixtures: wortmannin, an inhibitor of the PI3-K kinase family (DNA-PKcs is a component of that family). When wortmannin and ATP were added to DNA–protein complexes still attached to the beads (lane 6), the slow-migrating band that was observed in lane 4 (filled circle) disappeared in favor of a band having an intermediate migration rate (lane 6, empty circle). The same experiment was performed but without beads present in the incubation mixture: the XRCC4 phosphorylation status was brought to a level comparable to that observed in control experiments with no ATP incubation (lane 7, compare with lane 3). Overall, these results establish firmly that kinase activities did adsorb onto the beads and phosphorylated XRCC4. These kinase activities could be only partly inhibited using a wide-spectrum inhibitor. Removal of the beads prior to making the phosphorylation test allowed us to show that the oligonucleotide did effectively retain kinase activities distinct from the previous ones, as these were totally inhibited with wortmannin.
DISCUSSION
DNA-binding proteins are generally purified by affinity purification using an oligonucleotide as a bait. Salts and/or detergents are generally added to the washing or elution buffers to recover proteins specifically interacting with the target oligonucleotide. A compromise has to be found between a high protein background and the loss of components or activities of interest. In fact, numerous proteins from whole cell extract interact with the chromatographic support that accounts for a huge interaction surface with respect to that of the target oligonucleotide. To circumvent this problem, a number of strategies were devised in the past and were based on the separation of the DNA–protein assemblies from the chromatographic support before elution. The insertion of a disulfide bond between the target oligonucleotide and the chromatographic support (6,7) (see Introduction Section), was appealing at first sight. However, it might not be widely applicable as reducing conditions are often required for the proper interaction of proteins with their cognate DNA sequence (39). In the case of competition-based strategies used to uncouple DNA–protein assemblies from the chromatographic support (9,10,12), one serious problem is that the aptamer usually has non-negligible length (in the range of 20–30 nucleotides) and that, as such, it might recruit proteins impeding it to bind to the chromatographic phase (10). In all the strategies above, the uncoupling of the nucleic acid–protein complex is achieved by adding chemicals (either small or large molecules) to the purification medium. This might prove detrimental to downstream analytical steps, like the direct analysis of nucleic acid–protein complexes by mass spectrometry or the isoelectrofocalization of the purified proteins, for example. The near-UV-irradiation strategy used in this work clearly had none of the drawbacks described above. This approach avoids to set physico-chemical conditions optimized to eliminate contaminants from the chromatographic support, since the same weak saline conditions are maintained throughout the purification process. In the case of the purification of DNA ends-binding proteins, we obtained a profile very similar to the ones already published (30,40). The proscription of stringent conditions (such as high salts and/or presence of detergents) preserved weak interactions (protein–protein and DNA–protein) and allowed to recover DNA–protein assemblies in their native state, as demonstrated by the detection of biologically relevant enzymatic activities copurifying with the DNA–protein complexes. Enzymatic tests performed before or after the UV-irradiation step (i.e. the purified material was respectively in presence or absence of the beads), underlined the fact that photocleaving the DNA–protein assemblies off the beads was highly beneficial for two reasons: first, significant amounts of contaminant enzymatic activities not relating to target oligonucleotide were found to be unspecifically adsorbed onto the beads, leading to spurious phosphorylation of one molecular partner of the purified DNA–protein assemblies; second, monitoring of the ligation activity operating on tethered DNA–protein complexes proved unsuccessful, while it could be achieved if the complexes were first uncoupled from the beads. Taken as a whole, these results demonstrated that the photocleavage step allowed a good compromise between the yield and the purity of the purified proteins.
Finally, this method presents the advantage of being fast and easy to carry out as it is a one-step procedure, thus reducing to a minimum the loss of molecular partners from the assemblies of interest. The photocleavable biotin can be incorporated by chemical synthesis in solid phase at the end of the DNA-probe. Short PC-biotin modified primers can be also used to synthesize a longer DNA sequence by PCR (concatemers made of an affinity sequence repetition, for example), or else to hybridize a complementary sequence linked to the nucleic probe of interest (adapter strategy). These different approaches were tested in the case of TetR purification and all gave similar results as the basic duplex PCB-TetO (data not shown). In conclusion, the procedures developed in this work are generalizable to any nucleic acid-binding proteins and permit as well the identification of these proteins as the study of their functions.
SUPPLEMENTARY DATA
Supplementary Data are available at NAR Online.
ACKNOWLEDGEMENTS
We thank Dr Carine Giovannangeli for advice and helpful discussions, and Lionel Dubost (mass spectrometry facility, MNHN) for technical assistance with MALDI-TOF mass spectrometry. Our studies are supported by grants from INSERM, CNRS, MNHN and INTAS 03-51-5281. N.H. benefits from a pre-doctoral fellowship from the Ministère délégué à l'enseignement supérieur et à la recherche (France) and from the Ligue nationale contre le cancer (France). Funding to pay the Open Access publication charges for this article was provided by INSERM.
Conflict of interest statement. None declared.
REFERENCES
- 1.Rusconi F, Guillonneau F, Praseuth D. Contributions of mass spectrometry in the study of nucleic acid-binding proteins and of nucleic acid-protein interactions. Mass Spectrom. Rev. 2002;21:305–348. doi: 10.1002/mas.10036. [DOI] [PubMed] [Google Scholar]
- 2.Yaneva M, Tempst P. Affinity capture of specific DNA-binding proteins for mass spectrometric identification. Anal. Chem. 2003;75:6437–6448. doi: 10.1021/ac034698l. [DOI] [PubMed] [Google Scholar]
- 3.Yaneva M, Kippenberger S, Wang N, Su Q, McGarvey M, Nazarian A, Lacomis L, Erdjument-Bromage H, Tempst P. PU.1 and a TTTAAA element in the myeloid defensin-1 promoter create an operational TATA Box that can impose cell specificity onto TFIID function. J. Immunol. 2006;176:6906–6917. doi: 10.4049/jimmunol.176.11.6906. [DOI] [PubMed] [Google Scholar]
- 4.Nordhoff E, Krogsdam AM, Jorgensen HF, Kallipolitis BH, Clark BF, Roepstorff P, Kristiansen K. Rapid identification of DNA-binding proteins by mass spectrometry. Nat. Biotechnol. 1999;17:884–888. doi: 10.1038/12873. [DOI] [PubMed] [Google Scholar]
- 5.Drewett V, Molina H, Millar A, Muller S, vonHesler F, Shaw PE. DNA-bound transcription factor complexes analysed by mass-spectrometry: binding of novel proteins to the human c-fos SRE and related sequences. Nucleic Acids Res. 2001;29:479–487. doi: 10.1093/nar/29.2.479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Shimkus M, Levy J, Herman T. A chemically cleavable biotinylated nucleotide: usefulness in the recovery of protein-DNA complexes from avidin affinity columns. Proc. Natl Acad. Sci. USA. 1985;82:2593–2597. doi: 10.1073/pnas.82.9.2593. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ruby SW, Abelson J. An early hierarchic role of U1 small nuclear ribonucleoprotein in spliceosome assembly. Science. 1988;242:1028–1035. doi: 10.1126/science.2973660. [DOI] [PubMed] [Google Scholar]
- 8.Soukup GA, Cerny RL, Maher LJ. Preparation of oligonucleotidebiotin conjugates with cleavable linkers. Bioconjug. Chem. 1995;6:135–138. doi: 10.1021/bc00031a017. [DOI] [PubMed] [Google Scholar]
- 9.Bachler M, Schroeder R, vonAhsen U. StreptoTag: a novel method for the isolation of RNA-binding proteins. RNA. 1999;5:1509–1516. doi: 10.1017/s1355838299991574. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Hartmuth K, Urlaub H, Vornlocher H-P, Will CL, Gentzel M, Wilm M, Lührmann R. Protein composition of human prespliceosomes isolated by a tobramycin affinity-selection method. Proc. Natl Acad. Sci. USA. 2002;99:16719–16724. doi: 10.1073/pnas.262483899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Martinez J, Patkaniowska A, Urlaub H, Lührmann R, Tuschl T. Single-stranded antisense siRNAs guide target RNA cleavage in RNAi. Cell. 2002;110:563–574. doi: 10.1016/s0092-8674(02)00908-x. [DOI] [PubMed] [Google Scholar]
- 12.Dangerfield JA, Windbichler N, Salmons B, Günzburg WH, Schröder R. Enhancement of the StreptoTag method for isolation of endogenously expressed proteins with complex RNA binding targets. Biochem. J. 2006;27:1874–1877. doi: 10.1002/elps.200500709. [DOI] [PubMed] [Google Scholar]
- 13.Berens C, Hillen W. Gene regulation by tetracyclines. Constraints of resistance regulation in bacteria shape TetR for application in eukaryotes. Eur. J. Biochem. 2003;270:3109–3121. doi: 10.1046/j.1432-1033.2003.03694.x. [DOI] [PubMed] [Google Scholar]
- 14.Burma S, Chen BPC, Chen DJ. Role of non-homologous end joining (NHEJ) in maintaining genomic integrity. DNA Repair (Amst) 2006;5:1042–1048. doi: 10.1016/j.dnarep.2006.05.026. [DOI] [PubMed] [Google Scholar]
- 15.Yao F, Svensjö T, Winkler T, Lu M, Eriksson C, Eriksson E. Tetracycline repressor, tetR, rather than the tetR-mammalian cell transcription factor fusion derivatives, regulates inducible gene expression in mammalian cells. Hum. Gene Ther. 1998;9:1939–1950. doi: 10.1089/hum.1998.9.13-1939. [DOI] [PubMed] [Google Scholar]
- 16.Baron U, Gossen M, Bujard H. Tetracycline-controlled transcription in eukaryotes: novel transactivators with graded transactivation potential. Nucleic Acids Res. 1997;25:2723–2729. doi: 10.1093/nar/25.14.2723. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Dignam JD, Lebovitz RM, Roeder RG. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res. 1983;11:1475–1489. doi: 10.1093/nar/11.5.1475. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Olejnik J, Sonar S, Krzymañska-Olejnik E, Rothschild KJ. Photocleavable biotin derivatives: a versatile approach for the isolation of biomolecules. Proc. Natl Acad. Sci. USA. 1995;92:7590–7594. doi: 10.1073/pnas.92.16.7590. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Olejnik J, Krzymanska-Olejnik E, Rothschild KJ. Photocleavable biotin phosphoramidite for 5′-end-labeling, affinity purification and phosphorylation of synthetic oligonucleotides. Nucleic Acids Res. 1996;24:361–366. doi: 10.1093/nar/24.2.361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Gossen M, Bujard H. Tight control of gene expression in mammalian cells by tetracycline-responsive promoters. Proc. Natl Acad. Sci. USA. 1992;89:5547–5551. doi: 10.1073/pnas.89.12.5547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Gadgil H, Taylor WL, Jarrett HW. Comparative studies on discrete and concatemeric DNA-sepharose columns for purification of transcription factors. J. Chromatogr. A. 2001;917:43–53. doi: 10.1016/s0021-9673(01)00647-1. [DOI] [PubMed] [Google Scholar]
- 22.Guillonneau F, Guieysse AL, Caer JPL, Rossier J, Praseuth D. Selection and identification of proteins bound to DNA triple-helical structures by combination of 2D-electrophoresis and MALDI-TOF mass spectrometry. Nucleic Acids Res. 2001;29:2427–2436. doi: 10.1093/nar/29.11.2427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Guillonneau F, Labas V, Auvin C, Praseuth D. A reliable and simple method for two-dimensional electrophoresis and identification of HeLa nuclear alkaline nucleic acid-binding proteins using immobilized pH gradient. Electrophoresis. 2001;22:4391–4403. doi: 10.1002/1522-2683(200112)22:20<4391::AID-ELPS4391>3.0.CO;2-8. [DOI] [PubMed] [Google Scholar]
- 24.Rusconi F. GNU polyxmass: a software framework for mass spectrometric simulations of linear (bio-)polymeric analytes. BMC Bioinformatics. 2006;7:226. doi: 10.1186/1471-2105-7-226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Budman J, Chu G. Processing of DNA for nonhomologous end-joining by cell-free extract. EMBO J. 2005;24:849–860. doi: 10.1038/sj.emboj.7600563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Rusconi F, Schmitter JM, Rossier J, Le Maire M. hromatographic separation and sample preparation in one step for maldi mass spectrometric analysis of subpicomole amounts of heterogeneous protein samples. Anal. Chem. 1998;70:3046–3052. [Google Scholar]
- 27.Guieysse AL, Praseuth D, Giovannangeli C, Asseline U, Hélène C. Psoralen adducts induced by triplex-forming oligonucleotides are refractory to repair in HeLa cells. J. Mol. Biol. 2000;296:373–383. doi: 10.1006/jmbi.1999.3466. [DOI] [PubMed] [Google Scholar]
- 28.Hillen W, Berens C. Mechanisms underlying expression of Tn10 encoded tetracycline resistance. Annu. Rev. Microbiol. 1994;48:345–369. doi: 10.1146/annurev.mi.48.100194.002021. [DOI] [PubMed] [Google Scholar]
- 29.Gadgil H, Jarrett HW. Oligonucleotide trapping method for purification of transcription factors. J. Chromatogr. A. 2002;966:99–110. doi: 10.1016/s0021-9673(02)00738-0. [DOI] [PubMed] [Google Scholar]
- 30.Calsou P, Delteil C, Frit P, Drouet J, Salles B. Coordinated assembly of Ku and p460 subunits of the DNA-dependent protein kinase on DNA ends is necessary for XRCC4-ligase IV recruitment. J. Mol. Biol. 2003;326:93–103. doi: 10.1016/s0022-2836(02)01328-1. [DOI] [PubMed] [Google Scholar]
- 31.Chen L, Trujillo K, Sung P, Tomkinson AE. Interactions of the DNA ligase IV-XRCC4 complex with DNA ends and the DNA-dependent protein kinase. J. Biol. Chem. 2000;275:26196–26205. doi: 10.1074/jbc.M000491200. [DOI] [PubMed] [Google Scholar]
- 32.Nick McElhinny SA, Snowden CM, McCarville J, Ramsden DA. Ku recruits the XRCC4-ligase IV complex to DNA ends. Mol. Cell Biol. 2000;20:2996–3003. doi: 10.1128/mcb.20.9.2996-3003.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.West RB, Yaneva M, Lieber MR. Productive and nonproductive complexes of Ku and DNA-dependent protein kinase at DNA termini. Mol. Cell Biol. 1998;18:5908–5920. doi: 10.1128/mcb.18.10.5908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Audebert M, Salles B, Calsou P. Involvement of poly(ADP-ribose) polymerase-1 and XRCC1/DNA ligase III in an alternative route for DNA double-strand breaks rejoining. J. Biol. Chem. 2004;279:55117–55126. doi: 10.1074/jbc.M404524200. [DOI] [PubMed] [Google Scholar]
- 35.Leber R, Wise TW, Mizuta R, Meek K. The XRCC4 gene product is a target for and interacts with the DNA-dependent protein kinase. J. Biol. Chem. 1998;273:1794–1801. doi: 10.1074/jbc.273.3.1794. [DOI] [PubMed] [Google Scholar]
- 36.Drouet J, Delteil C, Lefrançois J, Concannon P, Salles B, Calsou P. DNA-dependent protein kinase and XRCC4-DNA ligase IV mobilization in the cell in response to DNA double strand breaks. J. Biol. Chem. 2005;280:7060–7069. doi: 10.1074/jbc.M410746200. [DOI] [PubMed] [Google Scholar]
- 37.Baumann P, West SC. DNA end-joining catalyzed by human cell-free extracts. Proc. Natl Acad. Sci. USA. 1998;95:14066–14070. doi: 10.1073/pnas.95.24.14066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Yu Y, Wang W, Ding Q, Ye R, Chen D, Merkle D, Schriemer D, Meek K, Lees-Miller SP. DNA-PK phosphorylation sites in XRCC4 are not required for survival after radiation or for V(D)J recombination. DNA Repair (Amst) 2003;2:1239–1252. doi: 10.1016/s1568-7864(03)00143-5. [DOI] [PubMed] [Google Scholar]
- 39.Marsich E, Bandiera A, Tell G, Scaloni A, Manzini G. A chicken hnRNP of the A/B family recognizes the single-stranded d(CCCTAA)(n) telomeric repeated motif. Eur. J. Biochem. 2001;268:139–148. doi: 10.1046/j.1432-1327.2001.01860.x. [DOI] [PubMed] [Google Scholar]
- 40.Ruscetti T, Lehnert BE, Halbrook J, Trong HL, Hoekstra MF, Chen DJ, Peterson SR. Stimulation of the DNA-dependent protein kinase by poly(ADP-ribose) polymerase. J. Biol. Chem. 1998;273:14461–14467. doi: 10.1074/jbc.273.23.14461. [DOI] [PubMed] [Google Scholar]
- 41.Perkins DN, Pappin DJ, Creasy DM, Cottrell JS. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis. 1999;20:3551–3567. doi: 10.1002/(SICI)1522-2683(19991201)20:18<3551::AID-ELPS3551>3.0.CO;2-2. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.