Abstract
TFIIH has been implicated in several fundamental cellular processes, including DNA repair, cell cycle progression, and transcription. In transcription, the helicase activity of TFIIH functions to melt promoter DNA; however, the in vivo function of the Cdk7 kinase subunit of TFIIH, which has been hypothesized to be involved in RNA polymerase II (Pol II) phosphorylation, is not clearly understood. Using temperature-sensitive and null alleles of cdk7, we have examined the role of Cdk7 in the activation of Drosophila heat shock genes. Several in vivo approaches, including polytene chromosome immunofluorescence, nuclear run-on assays, and, in particular, a protein-DNA cross-linking assay customized for adults, revealed that Cdk7 kinase activity is required for full activation of heat shock genes, promoter-proximal Pol II pausing, and Pol II-dependent chromatin decondensation. The requirement for Cdk7 occurs very early in the transcription cycle. Furthermore, we provide evidence that TFIIH associates with the elongation complex much longer than previously suspected.
Accurate transcription of most eukaryotic genes requires the coordinated recruitment of RNA polymerase II (Pol II), along with a host of general transcription factors (GTFs), sequence-specific transcription factors, and chromatin-remodeling factors. The assembly of these factors culminates in the formation of a Pol II promoter complex that initiates transcription, begins elongation, and matures into an elongationally competent state. This maturation is characterized by the phosphorylation of the carboxy-terminal domain (CTD) in the largest subunit of Pol II and the association of a set of elongation and RNA-processing factors.
One of the GTFs, TFIIH, has been implicated in both ATP-dependent promoter melting and phosphorylation of the Pol II CTD at serine 5 (Ser5) within the heptapeptide repeat. TFIIH is comprised of two subcomplexes, core IIH and CAK (Cdk-activating kinase) (41). Core IIH contains six subunits, two of which are ATP-dependent DNA helicases. One of these helicases, ERCC3, has been shown to be required for promoter melting in vitro (41) and in vivo (13). CAK is a tripartite complex composed of MAT-1, cyclin H, and the CTD kinase, Cdk7. In higher eukaryotes, CAK also occurs as a free complex, which is primarily involved in cell cycle regulation (19).
Cdk7 has been proposed to be one of possibly several kinases responsible for phosphorylating the CTD and thus contributing to the formation of the mature, elongating form of polymerase. Indeed, in vitro assays have demonstrated that Cdk7 kinase activity is required for transcription from the dihydrofolate reductase (DHFR) promoter (1, 41). However, this requirement may depend on the promoter that is used and the purity of the transcription system. For example, Makela et al. (30) found that a kinase-deficient TFIIH supported transcription from the adenovirus major late promoter. More recently, Tirode et al. (41) confirmed these results but found that the mere physical presence of the CAK complex could stimulate basal transcription from the major late promoter, even when CAK contained a catalytically inactive Cdk7. Chen and Zhou (6) have shown that HIV-1 transcription is independent of TFIIH kinase in vitro. Therefore, most of the promoters tested in vitro do not absolutely require Cdk7 kinase activity.
To date, most in vivo studies of Cdk7 have been performed in the yeast Saccharomyces cerevisiae. The yeast homologue of cdk7, KIN28, lacks CAK activity, leading to the postulate that it functions only in transcription (8). Using a temperature-sensitive allele of KIN28, Valay et al. have found that transcription is dramatically reduced at the nonpermissive temperature (42). This transcription impairment correlates with a striking decrease in CTD phosphorylation, indicating that in yeast, KIN28 is generally required for transcription. Still, exceptions do exist. The copper-induced CUP1 gene and yeast heat-shock genes are at least partially inducible even in yeasts that are dramatically depleted of KIN28 (22, 32).
Studies of higher eukaryotes aimed at elucidating the role of Cdk7 in transcription have also yielded unclear—and seemingly contradictory—results. Rossi et al. (38) have generated a loss-of-function allele of MAT1 in mice which severely diminished levels of Cdk7 protein, but not core TFIIH. In these mutant cells, CTD phosphorylation of both serine 5 and serine 2 was reduced, but no transcriptional defects were observed. Although transcription was measured indirectly (and nonquantitatively) by visualizing the expression of green fluorescent protein from a microinjected reporter plasmid, these results suggest that an intact CAK complex is not absolutely required for transcription in mammalian cells. It is quite possible that another CTD kinase could supply the minimal critical phosphorylations needed to activate Pol II. In Drosophila, as in yeast, Cdk7 is an essential protein. Larochelle et al. created both null and temperature-sensitive alleles of cdk7 and found that lethality occurred as a result of cell cycle arrest that was not operating simply by blocking cyclin expression, as the levels of cyclin protein were unaffected (18). In apparent contradiction, Leclerc et al., using an ectopically expressed mutant cdk7 allele, found that the onset of zygotic transcription of the ftz gene in Drosophila was delayed, while cell cycle timing was not appreciably affected (21). Interestingly, the authors did not observe any irregularities in the level of CTD phosphorylation. A genetic study of Drosophila by Merino et al. shows that the development of body structures that require high levels of transcription is affected by a temperature-sensitive mutation in cdk7 (33). Wallenfang and Seydoux isolated a temperature-sensitive cdk7 allele in Caenorhabditiselegans and showed that a partial loss of cdk7 activity affects both transcription and cell cycle progression in embryos. They also noted a general decrease in CTD phosphorylation (43).
Taken together, these in vivo results do not provide a unified view of Cdk7 function and indicate that metazoans may not have as strict a requirement for Cdk7 in general transcription as is seen in yeast. This may be the consequence of additional CTD kinases in metazoans that can be recruited to transcriptionally active Pol II and that have some degree of functional redundancy with TFIIH. One such candidate is P-TEFb (31), a Cdk9-cyclin T complex whose critical role in general transcription has recently been verified by Chao and Price (5). They found that they could drastically decrease both CTD phosphorylation and Pol II transcript accumulation in vivo using a highly specific inhibitor of P-TEFb, suggesting that P-TEFb is globally required for transcription. In vivo studies by Lis et al. have demonstrated that P-TEFb is present at transcriptionally active loci in Drosophila (24), where it is located over the extent of the entire gene (2). This distribution contrasts with that reported for TFIIH, which, at least in vitro, is known to dissociate from the elongation complex soon after initiation (46).
The lack of conclusive in vivo evidence linking Cdk7 to transcription in higher organisms, coupled with the more recent emergence of P-TEFb, has called into question the precise role of Cdk7 in transcriptional elongation. To date, most in vivo studies aimed at elucidating the role of Cdk7 in this process have examined transcriptional effects indirectly by measuring accumulated mRNA or protein. In this study, we have directly examined the dependence of several Drosophila genes on Cdk7 at the level of Pol II elongation in vivo using three independent methods: immunofluorescence staining of polytene chromosomes, nuclear run-on assays, and chromatin cross-linking and immunoprecipitation (ChIP) assays. Using temperature-sensitive and null alleles of cdk7, we found that the heat shock genes require Cdk7 for full activation. Inactivating Cdk7 results in a decrease in the density of transcriptionally active Pol II along heat shock genes. The level of paused Pol II is also reduced on the uninduced hsp70 gene in the absence of Cdk7. Furthermore, we find that Cdk7 associates with the elongation complex across the entire transcription unit in vivo. We propose that the kinase activity of Cdk7 is required at a step early in elongation, prior to the formation of the paused polymerase. Interestingly, transcription of the histone genes appears to be unaffected by loss of Cdk7. This work, therefore, indicates that Cdk7 is required for maximum transcription of a subset of genes in vivo and that it associates with Pol II elongation complexes well beyond the promoter-proximal region.
MATERIALS AND METHODS
Fly stocks.
Cdk7ts flies (line 15) are homozygous for two point mutations in cdk7: S164A and T170A. This line was propagated at 18°C on standard yeast-glucose medium prior to experimentation. For experiments involving Cdk7ts mutants, an isogenic fly line (line 47) was used as a control. It contains the inert cdk7 mutation, S180A. Cdk7null flies were balanced over FM7a, marked with green fluorescent protein, and propagated at room temperature. Transgenic flies containing two integrated copies of an hsp70 hybrid gene at 61A and 9E were described previously (28).
Recombinant protein purification and antibody production.
Full-length cdk7 was cloned into a six-His expression vector (pQE32) by X. Zhang. DH5α cells containing the construct were induced with 1 mM IPTG (isopropyl-β-d-thiogalactopyranoside). This protein was insoluble and had to be extracted from inclusion bodies using CAPS detergent (Novagen) as described by the manufacturer. Purified protein was sent to Pocono Rabbit Farm and Laboratory for antibody production. Recombinant Cdk7 was injected into a rabbit and a guinea pig. The guinea pig antiserum from the second or third bleed was used for immunoprecipitation and polytene immunofluorescence assays. Rabbit anti-heat shock factor (HSF) antibodies were produced as previously described (2). Rabbit anti-cyclin T antibodies were a gift from David Price. The RNA polymerase CTD antibodies H14 and H5, which recognize phosphorylated serine 5 and serine 2, respectively, were obtained from Covance (Richmond, Calif.). 8WG16, which recognizes unphosphorylated CTD, was also from Covance.
Indirect immunofluorescence assays.
Salivary gland polytene chromosomes from third-instar larvae were dissected and immunostained as previously described (24). Primary antibodies were used at the following concentrations: Cdk7, 1:100; HSF, 1:50; H14, 1:50; cyclin T, 1:20. Appropriate secondary antibodies raised in donkey and conjugated to either rhodamine or fluorescein isothiocyanate labels (Jackson ImmunoResearch Laboratories, West Grove, Pa.) were used at 1:100, and images were collected.
Northern blotting.
RNA was extracted from adult flies that had been heat shocked for 1 h at 36.5°C using Trizol reagent (Invitrogen); 10 μg of RNA per lane was loaded onto a 1.5% agarose gel containing formaldehyde, and standard Northern analysis was performed as described previously (3a). For quantifying signals, 0.1 and 0.01 μg of total RNA were diluted into loading buffer and loaded on the gel to give 1:100 and 1:1,000 dilutions, respectively. These lanes were probed for rRNA and used as internal standards. The probes were radiolabeled using the RadPrime DNA-labeling system (Life Technologies). The probe used to detect rRNA was a HindIII-EcoRI restriction fragment from the plasmid DmrY22. For hsp70, full-length cDNA was liberated from clone 1B8 by PstI digestion (27). The following PCR-amplified fragments were used as probes: hsp83, +1625 to +2039; hsp26, +955 to +1380; H3, +75 to +491; H2A, +1 to +190. For α-β repeats, a 1.5-kb HindIII restriction fragment from the plasmid Dm 83.23 was used. For U6, the entire gene was used as a probe.
Nuclear run-on assay.
Isolation of nuclei from flies was performed as previously described (23). The nuclear run-on assay was also done as described, but volumes were scaled up twofold to enhance signals. PCR-amplified DNA fragments were purified with a QiaQuick PCR purification kit (Qiagen, Valencia, Calif.) and slot blotted to Hybond-XL positively charged nylon membranes (Amersham Pharmacia Biotech, Little Chalfont, England) at 2 μg of DNA per slot. For ribosomal DNA, 4 μg of DNA was added per slot. The labeled RNAs were hybridized for 48 h at 65°C in Church buffer (0.5 M Na3PO4 [pH 7.1], 7% [vol/vol] sodium dodecyl sulfate [SDS], 0.1 mM EDTA [pH 8.0]). The membranes were washed in two changes of 2× SSC (1× SSC is 0.15 M NaCl plus 0.015 M sodium citrate)-0.2% SDS for 15 min apiece at room temperature. They were then washed in 0.1× SSC-0.1% SDS for 30 min at room temperature and once for 30 min at 55°C. Signals were detected by a Storm 840 PhosphorImager and analyzed with ImageQuant software. The values reported for average decreases (n-fold) in Pol II density represent the averages of three independent experiments.
Formaldehyde cross-linking of Drosophila adults.
(This technique may be applicable to other Drosophila life stages, although we have only tested adults.) Between 1.5 and 2 g of Cdk7ts flies were placed in bottles containing media and incubated for 24 h at 29°C to inactivate Cdk7. The isogenic control flies (line 47) were treated similarly. The flies were then heat shocked at 36.5°C for 20 min in a water bath. Next, they were collected in a prechilled bottle and transferred to an Omni mixer (DuPont) containing ice-cold buffer A (0.3 M sucrose, 2 mM MgOAc, 3 mM CaCl2, 10 mM Tris-Cl [pH 8], 0.3% Triton X-100, 0.5 mM dithiothreitol). One protease inhibitor cocktail tablet (Roche) was added. The flies were ground in buffer A at full speed for 1.5 min, and the mixture was quickly filtered through two layers of 100-μm-pore-size nylon mesh into a prechilled glass-Teflon homogenizer. The mixture was stroked on ice 35 times with pestle B. Next, the homogenate was filtered through two layers of 35-μm-pore-size nylon mesh into a 50-ml conical tube (prechilled on ice). Exactly 1 volume of cross-linking buffer (0.1 M NaCl, 1 mM EDTA, 0.5 mM EGTA, 50 mM Tris [pH 8]) at 40°C was added to bring the temperature up to 20°C. Formaldehyde was immediately added to a final concentration of 0.3% and swirled gently for 1 min. Then, 2.5 M glycine was added to a final concentration of 125 mM to stop the reaction. The mixture was spun down for 3 min in a clinical centrifuge at 2,110 × g, and the supernatant was discarded. The pellet was resuspended in 10 ml of RIPA buffer (140 mM NaCl, 10 mM Tris-Cl [pH 8.0], 1 mM EDTA, 1% Triton, 0.3% SDS, 0.1% sodium deoxycholate), and another protease inhibitor cocktail tablet was added. This solution was sonicated on ice in a Branson digital sonifier for a total time of 8 min. Each burst of the sonifier lasted 30 s and was followed by a 1-min cool off. The sonicated material was spun down at 20,000 × g, and the supernatant was aliquoted into 500-μl portions and flash frozen in liquid nitrogen.
Immunoprecipitation of protein-DNA complexes.
To each 500-μl aliquot of frozen material, 1 ml of cold RIPA buffer (lacking SDS) was added. The lysates were precleared by adding 50 μl of protein A-agarose beads (Invitrogen) that had been washed three times in RIPA buffer lacking SDS. Preclearing was done for 30 min on a rotating platform at 4°C. Following this, the beads were spun down and the lysate was removed to a new tube. The immunoprecipitating antibody was then added. The following amounts of antibodies were added per immunoprecipitation: HSF, 2 μl; Cdk7, 5 μl; cyclin T, 1 μl; H14, 3 μl; H5, 3 μl; 8WG16, 2 μl. Incubation usually lasted overnight and was done at 4°C on a rotating platform. Next, we added 50 μl of protein A-agarose (or donkey anti-immunoglobulin M-agarose; Sigma) that had been washed in RIPA (minus SDS) and blocked with salmon sperm DNA overnight at a concentration of 0.2 μg of DNA/μl of bead suspension volume. Immune complexes were allowed to form for 1 h at 4°C on a rotating platform. Then, the beads were spun down in a microcentrifuge at 110 × g, and the supernatant was removed. The beads were washed for 3 min on a rotisserie in 1 ml of the following ice-cold solutions: three times in low-salt wash buffer (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mM Tris-Cl [pH 8.0], 150 mM NaCl), once in high-salt wash buffer (0.1% SDS, 1% Triton X-100, 2 mM EDTA, 20 mM Tris-Cl [pH 8.0], 500 mM NaCl), once in LiCl wash buffer (0.25 M LiCl, 1% NP-40, 1% deoxycholate, 1 mM EDTA, 10 mM Tris-Cl [pH 8.0]), and twice in Tris-EDTA. Protein-DNA conjugates were eluted from the beads by adding 250 μl of elution buffer (1% SDS, 0.1 M NaHCO3) and rotating the mixture for 15 min at room temperature. The beads were then spun down, the supernatant was collected, and the elution was repeated with another 250 μl of elution buffer. To the combined eluates totaling 500 μl, NaCl was added to 0.2 M, and the mixture was incubated at 65°C overnight to reverse the protein-DNA cross-links. As an input control, 20% of the original 1.5 ml of starting material was saved, and the cross-links were also reversed by this method. The following day, 1 μl of 15-mg/ml GlycoBlue (Ambion) was added to the samples, and then they were extracted twice with buffered phenol and once with phenol-chloroform-isoamyl alcohol. The DNA was precipitated with ethanol and resuspended in 50 μl of 10 mM Tris.
Determination of protein-DNA interactions by real-time PCR.
Real-time PCR was performed in 384-well plates with the ABPrism 7900 sequence detection system, and the data were analyzed with SDS software version 2.0 (Applied Biosystems). (See Fig. 6 for the hsp70 gene fragments that were analyzed.)
FIG. 6.
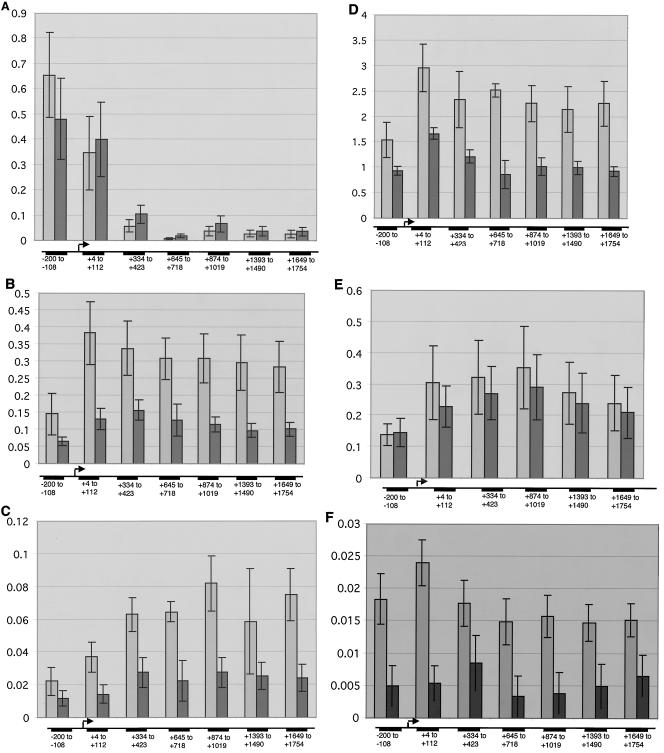
ChIP analysis of the induced hsp70 gene. Results from three or four independent cross-linking experiments for the wild type (lightly shaded bars) and the Cdk7ts mutant (darkly shaded bars) are shown. The y axis indicates cross-linking efficiency as a percentage of input, and the error bars indicate standard error. Antibodies were against the following proteins: HSF (A), P-Ser5 Pol II CTD (B), Pol II largest subunit (C), P-Ser2 Pol II CTD (D), cyclin T of P-TEFb (E), and Cdk7 (F). Background levels of cross-linking were determined by omitting the precipitating antibody and were then subtracted from the signals. The hsp70 gene fragments (base pair coordinates relative to the start site) that were analyzed are shown at the bottom of each panel.
RESULTS
Cdk7 recruitment to activated heat shock loci.
Immunofluorescent staining of formaldehyde-fixed salivary gland polytene chromosomes allows the visualization of the global distribution of a protein in vivo. Using an antibody against full-length Drosophila Cdk7, we examined the distribution of this protein and compared it to those of various forms of Pol II. The actively elongating form of polymerase contains a hyperphosphorylated CTD (Pol IIo), while the CTD of newly recruited and promoter-proximal polymerases is hypophosphorylated (Pol IIa) (20, 29). If Cdk7 kinase activity is required for general transcription, then the Cdk7 protein is likely to colocalize with one or both forms of Pol II. A genomic view of Cdk7 and polymerase distributions is shown in Fig. 1A. There, Cdk7 does indeed colocalize strongly with Pol IIo, as detected by an antibody that specifically recognizes phosphoserine 5 within the heptapeptide repeat. A high-magnification view of chromosome 3R is shown at the bottom of Fig. 1A to highlight the close correspondence between Cdk7 and Pol IIo.
FIG. 1.
(A) Cdk7 colocalizes with phosphorylated Pol II. Non-heat-shocked polytene chromosomes from wild-type larvae were stained with monoclonal antibodies recognizing phosphoserine 5 of the polymerase CTD (green) and Cdk7 (red). At the bottom is a high magnification of chromosome 3R in which the chromosome images from the different antibody stains were split down their axes using Photoshop software and aligned. (B) Cdk7 is recruited to native and transgenic heat shock loci. Bg9 larvae were immunostained with HSF (green) and Cdk7 (red) antibodies after a 20-min heat shock. Site 61A contains an hsp70 hybrid transgene. 87A and 87C contain two and three copies of native hsp70, respectively. 67B contains a cluster of the small heat shock genes hsp22, -23, -26, and -27. (C) Cdk7 and P-TEFb colocalize at heat shock puffs. Bg9 larvae were heat shocked and immunostained for the cyclin T subunit of P-TEFb (green) and Cdk7 (red). The yellow areas in the merge (right) indicate regions of colocalization.
Heat shock triggers the rapid recruitment of HSF to the promoters of heat shock genes, which results in a 150-fold increase in transcription at major heat shock loci. In polytene chromosome spreads, these highly active loci form striking puffs of decondensed chromatin. After a 20-min heat shock treatment, the heat shock genes are fully occupied with elongating Pol II molecules, and the puffs are at their maximum size. Consistent with a role in heat shock gene transcription, Cdk7 is recruited to puffs housing the major hsp70 genes at 87A and -C, as well as a single-copy hsp70-lacZ transgene at 61A (Fig. 1B). Interestingly, although Cdk7 is recruited to loci containing induced genes, it does not appear to coincide exactly with HSF. Rather, it has a broader distribution within the puff that is not confined to the same region as the promoter-bound HSF. This is especially evident at 87A, which contains two divergently transcribed copies of hsp70 whose promoters are ∼1.7 kb apart. As expected, HSF occupies this promoter region but is flanked by Cdk7, which is likely to represent the transcribed portion of each gene. A similar pattern can be seen at 61A, which contains the hsp70-lacZ transgene. Here, HSF is confined to the base of the puff (which represents the promoter of the transgene), while the Cdk7 signal extends into the body of the puff. As seen in Fig. 1C, Cdk7 appears to be more coincident with P-TEFb, which is known to localize with Pol II in the bodies of heat shock genes (3).
Cdk7 is required for full activation of heat shock genes.
The general colocalization of Cdk7, Pol II, and other transcription elongation factors by polytene immunofluorescence suggests that Cdk7 plays a role in Pol II transcription. To test this, we measured the level of various mRNAs in a Drosophila line containing a temperature-sensitive cdk7 allele This allele of cdk7 contains two mutations within the T loop that inhibit phosphorylation events critical to the stabilization of the Cdk7-cyclin H-Mat1 complex. In the absence of T-loop phosphorylation, the complex becomes thermolablile, thereby eliminating the kinase activity of Cdk7 (17). Both Cdk7ts flies and an isogenic control line were incubated at the nonpermissive temperature of 29°C for 24 h to fully inactivate Cdk7. To test the dependency of inducible genes on Cdk7, the flies were then heat shocked, whole RNA was harvested, and heat shock gene transcripts were measured by Northern blotting. The constitutively active histone genes were also examined (11, 35). Figure 2 shows that the three heat shock genes tested exhibit between two- and fourfold reductions in mRNA levels, while the histone genes show no such reduction. In order to study this defect more directly, we examined the densities of elongating polymerase molecules on both heat shock genes and histone genes by nuclear run-on analysis (Fig. 3A). These data show that the density of polymerase is reduced approximately ninefold on heat shock genes, while the effect on histone genes is negligible.
FIG. 2.
Heat shock gene mRNA accumulation is reduced in Cdk7ts flies. RNAs from the isogenic control strain line 47 (WT) and Cdk7ts flies were used for Northern blotting. Signals from each gene were normalized to the rRNA 1:1,000 standard to account for differences in total RNA between WT and Cdk7ts flies.
FIG. 3.
RNA polymerase density is decreased on heat shock genes in Cdk7ts flies. (A) Polymerase densities for hsp70, hsp83, hsp26, and histone 2A and -3 (H2A and H3). The decreases (n-fold) represent the means of three independent experiments. (B) Polymerase pausing is decreased in the Cdk7ts mutant. Paused molecules were released by the addition of 0.6% Sarkosyl to the nuclear run-on reactions containing non-heat-shocked nuclei. The mean of two independent experiments is shown. N.D., quantification not determined; WT, wild type; TS, Cdk7ts.
The uninduced heat shock gene is poised for rapid activation in response to stress. It contains a transcriptionally engaged polymerase molecule that is “paused” between +20 and +40 relative to the transcription start site (12). Previous work demonstrated that the escape of the paused polymerase into productive elongation is a rate-limiting step in heat shock gene induction and is therefore subject to regulation by a variety of factors, including kinases. To test whether such a polymerase molecule is present at the uninduced heat shock gene in the absence of a functional Cdk7, nuclear run-on experiments were performed on flies that had been incubated at the nonpermissive temperature but not subjected to heat shock stress. The addition of the detergent Sarkosyl to the run-on reaction allows the paused polymerase (should it be present) to escape into productive elongation. As Fig. 3B shows, the level of pausing is reduced relative to that of the isogenic control, indicating that Cdk7 contributes to the formation or maintenance of the paused polymerase.
CTD phosphorylation and heat shock puffs in Cdk7ts mutant.
We were interested in determining if the defect in heat shock gene activation was due to a loss of Pol II CTD phosphorylation in the Cdk7ts mutant. However, we first wanted to ascertain that the CAK complex was indeed disrupted and therefore depleted from heat shock loci by our conditions. To address this, polytene chromosomes were prepared from third-instar larvae that had been incubated at 29°C for 24 h prior to heat shock treatment. The chromosomes were immunostained against Cdk7, as well as HSF as a control. Chromosome sites 87A and 87C contain two and three copies of hsp70 genes, respectively, which stain intensely with anti-HSF antibody. Figure 4A and B show that the Cdk7ts mutant has lower levels of the Cdk7 protein at heat shock loci than an isogenic control. Despite this, the mutant appears to have levels of phosphorylated serine 5 that are difficult to distinguish from those of the wild type (Fig. 4C and D). To ensure that the Ser5 phosphorylation is not due to residual Cdk7 activity in the temperature-sensitive mutant at the nonpermissive temperature, we also examined polytene chromosomes from Cdk7null larvae, which lack the Cdk7 protein by Western analysis (data not shown). These larvae die in the late third-instar stage. Here again, the phosphoserine 5 signal is not dramatically reduced (Fig. 4E). These results indicate that either the decrease in serine 5 phosphorylation is too subtle to be detected by immunofluorescent staining or another kinase can phosphorylate serine 5 in the absence of functional Cdk7.
FIG. 4.
Induced heat shock genes in Cdk7ts mutant contain phosphorylated serine 5 but little Cdk7 protein and display abnormal puff pattern. (A and B) Isogenic control (WT) (A) and Cdk7ts mutant (B) showing recruitment of HSF (green) and Cdk7 (red) to hsp70 genes at 87A and 87C. The 87A puff is smaller in the Cdk7ts mutant and displays weak recruitment of the Cdk7 protein. (C and D) HSF and phosphoserine 5 (P-ser5) at 87A and 87C in the WT and Cdk7ts mutant, respectively. (E) Serine 5 phosphorylation persists in the Cdk7null mutant.
Interestingly, we also noticed that the heat shock puff at 87A was significantly smaller in the mutant than in the isogenic strain, while the 87C puff was mostly unchanged. Because high transcriptional activity at a particular locus can contribute to puffing (40), we reasoned that the small puff was reflective of lower levels of hsp70 transcription in the mutant. The nearly normal puff size at 87C could be explained by the transcription of 5 to 11 copies of the heat-inducible α-β sequences that are housed at 87C but not 87A (26, 27). Alternatively, the 87C puff may not be strictly a result of transcription. Rather, it could be caused by localized chromatin decondensation mediated by the binding of HSF to the promoters of the three hsp70 genes, as well as the several α-β gene promoters, which also contain binding sites for HSF (Fig. 5A). To distinguish between these two possibilities, we performed a Northern blot to determine if the α-β transcripts can still be induced in the Cdk7ts mutant. Figure 5B shows that the level of α-β mRNA is reduced in the mutant by 2.5-fold, similar to the reduction in hsp70 mRNA. These measurements were made using Pol I-derived rRNA transcripts and Pol III-derived U6 transcripts as internal standards. Based on this analysis, it is likely that both hsp70 genes and α-β genes are sensitive to the Cdk7 mutation and that the binding of HSF to several heat shock elements dispersed over a 38-kb region within the 87C locus is sufficient to induce puffing. Indeed, in HeLa cells, HSF has been shown to promote the disruption of chromatin at least 400 bp into the gene in the presence of the transcriptional inhibitor alpha-amanitin (4).
FIG. 5.
α-β transcription is reduced in the Cdk7ts mutant. (A) Arrangement of hsp70 and α-β genes. (B) α-β transcription is sensitive to the Cdk7ts mutation. Northern blots probed αβ for RNA and standards rRNA and U6 snRNA (see Materials and Methods). WT, wild type; ts, Cdk7ts.
High-resolution protein-DNA mapping by ChIP in adults.
To examine the protein distribution at higher resolution in vivo, we developed a simple in vivo ChIP assay for adult flies. This assay, which is a modification of a preexisting ChIP protocol for cultured cells, affords us a high-resolution view of proteins associated with virtually any segment of DNA in whole flies. Because nearly all of the known available Drosophila mutants are in the context of whole flies rather than cultured cells, this technique should have broad utility. The only requirement is that the flies reach adulthood; therefore, conditional mutants such as Cdk7ts are ideal. Immunoprecipitated DNA fragments were analyzed by TaqMan real-time PCR.
We applied our fly ChIP assay to study transcription factor distribution along the hsp70 gene under induced conditions and compared it to that in the Cdk7ts mutant. As Fig. 6A shows, HSF is strongly recruited to the promoter regions in both the wild type and mutant. Therefore, Cdk7 does not influence the early event of HSF recruitment to the hsp70 upstream regulatory region. As shown in Fig. 6B and C, we examined the effect of the Cdk7 mutation on CTD phosphorylation. In wild-type flies, Pol II molecules carrying the phosphoserine 5 epiptope (Fig. 6B) occur most strongly in the 5′ region of the gene and are slightly reduced in downstream regions. The cdk7 mutation significantly reduces serine 5 phosphorylation across the entire transcription unit. Serine 2 phosphorylation in wild-type flies (Fig. 6C) is more pronounced near the middle of the gene and persists to the 3′ end. Interestingly, serine 2 phosphorylation is also significantly reduced in the Cdk7ts mutant. As shown in Fig. 6E, we found that cyclin T, the CTD kinase subunit of P-TEFb, is present throughout the gene in both wild-type and mutant flies, indicating that its recruitment is not significantly affected by the cdk7 mutation. Our analysis of the Cdk7 protein (Fig. 6F) revealed that in wild-type flies, Cdk7 does not appear to be restricted to the promoter-proximal region, as suggested by earlier in vitro assays with mammalian nuclear extracts (46). Instead, its distribution resembles that of Pol II containing phosphorylated serine 5. This is consistent with the polytene immunofluorescence patterns (Fig. 1B and C), where the Cdk7 signal appears to occupy the entire heat shock puff and generally colocalizes with phosphoserine 5. Also, we have noted a similar immunofluorescence pattern using an antibody directed against the p62 subunit of core TFIIH (data not shown). In the Cdk7ts mutant (Fig. 6F), the level of Cdk7 protein cross-linked to the hsp70 gene is decreased dramatically. In conclusion, the primary effect of the Cdk7ts mutation is a reduction in the levels of recruited Cdk7 and in the number of mature (i.e., phosphorylated) elongation complexes that escape into the body of the gene.
DISCUSSION
In this study, we provide in vivo evidence that the TFIIH kinase, Cdk7, is required for the full induction of heat shock genes in Drosophila. First, we demonstrate by indirect immunofluorescence on polytene chromosomes that Cdk7 is associated with sites of active transcription, as it colocalizes strongly with the phosphorylated, elongationally competent form of Pol II. Upon heat shock, Cdk7 is recruited to both endogenous and transgenic heat shock genes. Second, using a temperature-sensitive Cdk7 mutant, we show that the density of elongating polymerase, as measured by nuclear run-on and ChIP assays along the induced hsp70, is reduced in the absence of Cdk7 activity, which results in a reduction in hsp70 mRNA as measured by Northern blotting. One key feature of the uninduced heat shock gene (and many other genes as well) is a Pol II molecule that is paused between +20 and +40 relative to the transcription start site (12). Third, we show that Cdk7 contributes to this paused Pol II because inactivating Cdk7 causes a significant decrease in Sarkosyl-releasable polymerase by the nuclear run-on assay in the Cdk7ts mutant.
In this study, we also define the consequences of a Cdk7 mutant on the phosphorylation of Pol II at activated heat shock loci. Although it has been firmly established that Cdk7, in the context of holo-TFIIH, can phosphorylate the serine 5 residue of the Pol II CTD, a significant decrease in phosphoserine 5 (or phosphoserine 2) was not readily apparent by immunofluorescence analysis of heat-shocked polytene chromosomes in the Cdk7ts mutant. However, interpreting this result is complicated by the reduced decondensation (smaller heat shock puff) in the Cdk7ts mutant, which effectively concentrates the phosphoserine 5 signal. Moreover, immunofluorescence analysis has inherently more variation than does the quantitative ChIP assay, which does show reduced Ser2 and Ser5 phosphorylation. Consistent with the nuclear run-on analysis, we also noted a general decrease in total levels of Pol II when ChIP was performed with an antibody that recognizes all Pol II molecules, regardless of phosphorylation state (Fig. 6D). We did, however, note a two- to threefold decrease in phosphoserine 5 on the hsp70 gene with the temperature-sensitive mutant, using a highly quantitative assay, real-time ChIP analysis. The remaining phosphorylation of serine 5 is not simply a consequence of the leaky temperature-sensitive allele, because the Cdk7null mutant also appeared to have significant levels of serine 5 phosphorylation at heat shock loci. This mutant has no Cdk7 protein detectable by Western blotting. Therefore, we propose that another kinase, possibly P-TEFb, contributes to serine 5 phosphorylation. Preliminary experiments indicate that P-TEFb can phosphorylate both serine 5 and serine 2 in the context of an activated heat shock gene (B. E. Schwartz, unpublished results). Also, Zhou et al. have demonstrated that P-TEFb, in the presence of the activator Tat, can phosphorylate serine 5, as well as serine 2, of the CTD (47). Because the Cdk7-mediated phosphorylation of serine 5 appears to occur very early in the transcription cycle, a kinase-deficient TFIIH mutant may effectively limit the number of early elongation complexes that escape into the body of the gene. P-TEFb, which is rapidly recruited to activated heat shock genes (24), may be able to partially compensate for this defect by phosphorylating some serine 5 residues within the CTD.
It was apparent that inactivating Cdk7 prior to heat shock treatment resulted in lower levels of elongating polymerase and a decrease in heat shock mRNA production, yet the ninefold decrease in polymerase density as measured by nuclear run-on was more severe than the three- to fourfold decrease in mRNA measured by Northern blotting. One possible, albeit unlikely, explanation is that heat shock mRNA is preferentially stabilized in the mutant to partially compensate for the defect in transcription. However, we believe that this quantitative difference could simply be due to variation between the two types of experiments. For example, in the nuclear run-on analysis, the polymerase density on the histone genes is, on average, slightly lower in the Cdk7ts mutant, yet according to the Northern analysis, histone mRNA synthesis appears to be slightly up-regulated. If the hsp70 signals are normalized to the histone signals for both analyses, then the decrease in polymerase density on hsp70 in the Cdk7ts mutant is exactly the same as the decrease in hsp70 mRNA, fivefold.
Because TFIIH functions as an integral component of the Pol II transcriptional machinery, the Pol I-driven rRNA genes were chosen as internal controls in many of our experiments. Recently emerging data from other groups, however, are challenging the notion that TFIIH is restricted to Pol II transcription. Iben et al. reported that TFIIH is involved in rRNA transcription, yet the mechanism is still unclear (14). Although TFIIH is required in their in vitro system, hydrolyzable ATP is not. Therefore, it is likely that neither the helicase or the kinase activity of TFIIH is required for productive rRNA transcription (14). We, too, find that rRNA transcription is not significantly affected by the Cdk7ts mutation. As shown in Fig. 5B, we used the Pol III-driven U6 gene as a control for Pol I transcription and found that the steady-state level of rRNA was unchanged in the Cdk7ts mutant. In nuclear run-on assays, where changes in transcriptional activity can be assessed immediately after shifting to the nonpermissive temperature, we again saw no decrease in the amount of radiolabeled run-on rRNAs from mutant flies compared to the same number of wild-type flies. We therefore conclude that in Drosophila, rRNA measurements serve as a valid internal standard when analyzing mutations that affect TFIIH kinase activity.
An intriguing phenotype of both Cdk7ts and Cdk7null fly lines was the unusual morphology of the heat shock puffs in polytene chromosomes. We found that the 87C puff, which contains three copies of hsp70, was consistently reduced in size relative to the wild type, while the 87A puff, containing two copies, was virtually eliminated. The 87C locus is distinguished from 87A by having 5 to 11 copies of α-β and γ repeats (Fig. 5A) (25). Each of these α-β repeats is a transcription unit that has a copy of an hsp70 promoter, γ, and is therefore heat inducible. Because high levels of transcription can contribute to puffing, we wondered if α-β transcription, unlike hsp70 gene transcription, was insensitive to the Cdk7 mutations. Northern analysis suggested that α-β transcription was, in fact, decreased by the Cdk7ts mutation to an extent similar to that of hsp70. We speculate that HSF binding, even in the absence of high levels of transcription, is sufficient to elicit the puff at 87C. A similar puff pattern (puffing at 87C but not 87A at the hsp70 locus) was achieved by Winegarden et al., who used sodium salicylate to elicit HSF binding that was not accompanied by an increase in hsp70 transcription (45). Certain developmental loci have been reported to form puffs when transcription is inhibited, and these too could be, like 87C, the consequence of sequence-specific factors eliciting decondensation over a region of chromatin (9).
Initial polytene immunofluorescence analysis revealed that the TFIIH core subunit p62 colocalized with polymerase within heat puffs associated with both native and transgenic hsp70 genes, as well as ecdysone-induced developmental puffs (Schwartz, unpublished). Additional experiments using polyclonal antibodies generated against Cdk7 yielded similar results (Fig. 1A and B), suggesting that TFIIH may remain associated with the elongation complex beyond +60, a position indicated in previous in vitro experiments as the point where TFIIH leaves the elongation complex (46). Another Cdk7 antibody—the monoclonal antibody 20H5—displays an immunofluorescence pattern similar to that of the polyclonal antibody (data not shown). Finally, our ChIP analysis of wild-type, heat-shocked flies shows a low level of Cdk7 cross-linked throughout the transcription unit. These results may imply a sustained requirement in vivo for one or more enzymatic activities of TFIIH as the elongation complex progresses through the gene. Recently, Cheng and Sharp found that the p62 subunit of core TFIIH preferentially cross-linked to the promoter-proximal region of the mammalian DHFR gene in vivo; however, on that gene, Pol II was detected only on the promoter-proximal region (7). It will be interesting to see how TFIIH distributes on more actively transcribed genes.
Based on these experiments, we can begin to build a more precise model for the role of TFIIH in Drosophila heat shock gene activation. The uninduced heat shock gene is “potentiated” for a rapid response to heat stress. The most notable feature of this potentiated state is the polymerase molecule that is paused near the promoter after having synthesized 20 to 40 bases of RNA transcript. On average, this paused polymerase molecule escapes into productive elongation every 10 min and must be regenerated. According to several in vitro and in vivo reports, this process is certain to require TFIIH helicase activity for promoter melting and promoter escape (10, 13). We propose that Cdk7 activity is also required for serine 5 phosphorylation of the CTD and that this initial phosphorylation allows polymerase to elongate to the pause site. Past studies have shown a lack of serine 2 phosphorylation on the paused polymerase (34), while more recent work by Boehm et al. reveals that the paused Pol II molecule is indeed phosphorylated at serine 5 (36). In yeast, serine 5 phosphorylation serves to recruit capping machinery, and because the RNA transcripts associated with the paused polymerase in Drosophila are also capped, this process may be conserved (37, 44). Based on our nuclear run-on assays under non-heat shock conditions, it appears that Cdk7 does in fact contribute to the formation of the paused polymerase. Regulation of transcription through a promoter-proximal paused polymerase molecule is a phenomenon that is not confined to Drosophila heat shock genes (39). Indeed, several mammalian genes, including c-myc, c-fos (2, 15, 16, 36), and, more recently, the constitutively active DHFR and γ-actin genes, have also been shown to display higher concentrations of Pol II in promoter-proximal regions than in interior gene regions (7). Therefore, Pol II pausing is likely to be a more general means of regulation than previously thought.
After heat shock, TFIIH is recruited to heat shock genes, as determined by polytene immunofluorescence. Under these conditions, the increased rate of polymerase recruitment and transcription initiation would require more TFIIH for promoter melting and CTD phosphorylation. These Cdk7-mediated phosphorylation events are likely to occur early in the transcription cycle, possibly soon after promoter clearance, since our nuclear run-on assays reveal a uniform decrease in polymerase density across the gene. If transcribing polymerase molecules encountered an elongation defect later in the cycle, we would instead expect to see polymerase densities progressively decrease along the gene. Therefore, we postulate that CTD serine 5 phosphorylation represents one of the first modifications that polymerase undergoes and that it may be just one of many events that influence the ability of RNA polymerase to escape into and remain in productive elongation.
Acknowledgments
We thank Eric Alani and Volker Vogt for critical reading of the manuscript and Jian Chen for helpful advice.
This work was supported by National Institutes of Health grant GM25232 to J.T.L.
REFERENCES
- 1.Akoulitchev, S., T. P. Makela, R. A. Weinberg, and D. Reinberg. 1995. Requirement for TFIIH kinase activity in transcription by RNA polymerase II. Nature 377:557-560. [DOI] [PubMed] [Google Scholar]
- 2.Albert, T., J. Mautner, J. O. Funk, K. Hoertnagel, A. Pullner, and D. Eick. 1997. Nucleosomal structures of c-myc promoters with transcriptionally engaged RNA polymerase II. Mol. Cell. Biol. 17:4363-4371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Andrulis, E. D., E. Guzman, P. Doring, J. Werner, and J. T. Lis. 2000. High-resolution localization of Drosophila Spt5 and Spt6 at heat shock genes in vivo: roles in promoter proximal pausing and transcription elongation. Genes Dev. 14:2635-2649. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3a.Ausubel, F. M., R. Brent, R. E. Kingston, D. D. Moore, J. G. Seidman, J. A. Smith, and K. Struhl (ed.). 1997. Current protocols in molecular biology. John Wiley & Sons, New York, N.Y.
- 3b.Boehm, A. K., A. Saunders, J. Werner, and J. T. Lis. Transcription factor and polymerase recruitment, modification, and movement on dhsp70 in vivo in the minutes following heat shock. Mol. Cell Biol. in press. [DOI] [PMC free article] [PubMed]
- 4.Brown, S. A., and R. E. Kingston. 1997. Disruption of downstream chromatin directed by a transcriptional activator. Genes Dev. 11:3116-3121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Chao, S. H., and D. H. Price. 2001. Flavopiridol inactivates P-TEFb and blocks most RNA polymerase II transcription in vivo. J. Biol. Chem. 276:31793-31799. [DOI] [PubMed] [Google Scholar]
- 6.Chen, D., and Q. Zhou. 1999. Tat activates human immunodeficiency virus type 1 transcriptional elongation independent of TFIIH kinase. Mol. Cell. Biol. 19:2863-2871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Cheng, C., and P. A. Sharp. 2003. RNA polymerase II accumulation in the promoter-proximal region of the dihyhydrofolate reductase and γ-actin genes. Mol. Cell. Biol. 23:1961-1967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Cismowski, M. J., G. M. Laff, M. J. Solomon, and S. I. Reed. 1995. KIN28 encodes a C-terminal domain kinase that controls mRNA transcription in Saccharomyces cerevisiae but lacks cyclin-dependent kinase-activating kinase (CAK) activity. Mol. Cell. Biol. 15:2983-2992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Crowley, T. E., and E. M. Meyerowitz. 1984. Steroid regulation of RNAs transcribed from the Drosophila 68C polytene chromosome puff. Dev. Biol. 102:110-121. [DOI] [PubMed] [Google Scholar]
- 10.Dvir, A., R. C. Conaway, and J. W. Conaway. 1997. A role for TFIIH in controlling the activity of early RNA polymerase II elongation complexes. Proc. Natl. Acad. Sci. USA 94:9006-9010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Gilmour, D. S., and J. T. Lis. 1985. In vivo interactions of RNA polymerase II with genes of Drosophila melanogaster. Mol. Cell. Biol. 5:2009-2018. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Gilmour, D. S., and J. T. Lis. 1986. RNA polymerase II interacts with the promoter region of the noninduced hsp70 gene in Drosophila melanogaster cells. Mol. Cell. Biol. 6:3984-3989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Guzman, E., and J. T. Lis. 1999. Transcription factor TFIIH is required for promoter melting in vivo. Mol. Cell. Biol. 8:5652-5658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Iben, S., H. Tschochner, M. Bier, D. Hoogstraten, P. Hozak, J. M. Egly, and I. Grummt. 2002. TFIIH plays an essential role in RNA polymerase I transcription. Cell 109:297-306. [DOI] [PubMed] [Google Scholar]
- 15.Krumm, A., T. Meulia, M. Brunvand, and M. Groudine. 1992. The block to transcriptional elongation within the human c-myc gene is determined in the promoter-proximal region. Genes Dev. 6:2201-2213. [DOI] [PubMed] [Google Scholar]
- 16.Krumm, A., L. B. Hickey, and M. Groudine. 1995. Promoter-proximal pausing of RNA polymerase II defines a general rate-limiting step after transcription initiation. Genes Dev. 9:559-572. [DOI] [PubMed] [Google Scholar]
- 17.Larochelle, S., J. Chen, R. Knights, J. Pandur, P. Morcillo, H. Erdjument-Bromage, P. Tempst, B. Suter, and R. P. Fisher. 2001. T-loop phosphorylation stabilizes the CDK7-cyclin H-MAT1 complex in vivo and regulates its CTD kinase activity. EMBO J. 20:3749-3759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Larochelle, S., J. Pandur, R. Fisher, H. Salz, and B. Suter. 1998. Cdk7 is essential for mitosis and for in vivo Cdk-activating kinase activity. Genes Dev. 12:370-381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Larochelle, S., and R. P. Fisher. 2002. Drosophila CDK7: a paradigm for CAK in metazoans, p. 39-53. In P. Kaldis (ed.), The CDK-activating kinase (CAK). Landes Bioscience Publishers, Georgetown, Tex.
- 20.Laybourn, P. J., and M. E. Dahmus. 1990. Phosphorylation of RNA polymerase IIA occurs subsequent to interaction with the promoter and before the initiation of transcription. J. Biol. Chem. 265:13165-13173. [PubMed] [Google Scholar]
- 21.Leclerc, V., S. Raisin, and P. Leopold. 2000. Dominant-negative mutants reveal a role for the Cdk7 kinase at the mid-blastula transition in Drosophila embryos. EMBO J. 19:1567-1575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Lee, D., and J. T. Lis. 1998. Transcriptional activation independent of TFIIH kinase and the RNA polymerase II mediator in vivo. Nature 393:389-392. [DOI] [PubMed] [Google Scholar]
- 23.Lee, H., K. W. Kraus, M. F. Wolfner, and J. T. Lis. 1992. DNA sequence requirements for generating paused polymerase at the start of hsp70. Genes Dev. 6:284-295. [DOI] [PubMed] [Google Scholar]
- 24.Lis, J. T., P. Mason, J. Peng, D. H. Price, and J. Werner. 2000. P-TEFb kinase recruitment and function at heat shock loci. Genes Dev. 14:792-803. [PMC free article] [PubMed] [Google Scholar]
- 25.Lis, J. T., D. Ish-Horowicz, and S. M. Pinchin. 1981. Genomic organization and transcription of the alpha beta heat shock DNA in Drosophila melanogaster. Nucleic Acids Res. 9:5297-5310. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Lis, J. T., L. Prestidge, and D. S. Hogness. 1978. A novel arrangement of tandemly repeated genes at a major heat shock site in D. melanogaster. Cell 14:901-919. [DOI] [PubMed] [Google Scholar]
- 27.Lis, J. T., W. Neckameyer, R. Dubensky, and N. Costlow. 1981. Cloning and characterization of nine heat-shock-induced mRNAs of Drosophila melanogaster. Gene 15:67-80. [DOI] [PubMed] [Google Scholar]
- 28.Lis, J. T., J. A. Simon, and C. A. Sutton. 1983. New heat shock puffs and beta-galactosidase activity resulting from transformation of Drosophila with an hsp70-lacZ hybrid gene. Cell 35:403-410. [DOI] [PubMed] [Google Scholar]
- 29.Lu, H., O. Flores, R. Weinmann, and D. Reinberg. 1991. The nonphosphorylated form of RNA polymerase II preferentially associates with the preinitiation complex. Proc. Natl. Acad. Sci. USA 88:10004-10008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Makela, T. P., J. D. Parvin, J. Kim, L. J. Huber, P. A. Sharp, and R. A. Weinberg. 1995. A kinase-deficient transcription factor TFIIH is functional in basal and activated transcription. Proc. Natl. Acad. Sci. USA 92:5174-5178. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Marshall, N. F., J. Peng, Z. Xie, and D. H. Price. 1996. Control of RNA polymerase II elongation potential by a novel carboxyl-terminal domain kinase. J. Biol. Chem. 271:27176-27183. [DOI] [PubMed] [Google Scholar]
- 32.McNeil, J. B., H. Agah, and D. Bentley. 1998. Activated transcription independent of the RNA polymerase II holoenzyme in budding yeast. Genes Dev. 12:2510-2521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Merino, C., E. Reynaud, M. Vazquez, and M. Zurita. 2002. DNA repair and transcriptional effects of mutations in TFIIH in Drosophila development. Mol. Biol. Cell. 13:3246-3256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.O'Brien, T., S. Hardin, A. Greenleaf, and J. T. Lis. 1994. Phosphorylation of RNA polymerase II C-terminal domain and transcriptional elongation. Nature 370:75-77. [DOI] [PubMed] [Google Scholar]
- 35.O'Brien, T., and J. T. Lis. 1993. Rapid changes in Drosophila transcription after an instantaneous heat shock. Mol. Cell. Biol. 13:3456-3463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Plet, A., D. Eick, and J. M. Blanchard. 1995. Elongation and premature termination of transcripts initiated from c-fos and c-myc promoters show dissimilar patterns. Oncogene 10:319-328. [PubMed] [Google Scholar]
- 37.Rasmussen, E., and J. T. Lis. 1993. In vivo transcriptional pausing and cap formation on three Drosophila heat shock genes. Proc. Natl. Acad. Sci. USA 90:7923-7927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Rossi, D. J., A. Londesborough, N. Korsisaari, A. Pihlak, E. Lehtonen, M. Henkemeyer, and T. P. Makela. 2001. Inability to enter S phase and defective RNA polymerase II CTD phosphorylation in mice lacking Mat1. EMBO J. 20:2844-2856. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Rougvie, A., and J. T. Lis. 1990. Postinitiation transcription control in Drosophila melonagaster. Mol. Cell. Biol. 10:6041-6045. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Simon, J. A., C. A. Sutton, R. B. Lobell, R. L. Glaser, and J. T. Lis. 1985. Determinants of heat shock-induced chromosome puffing. Cell 40:805-817. [DOI] [PubMed] [Google Scholar]
- 41.Tirode, F., D. Busso, F. Coin, and J. M. Egly. 1999. Reconstitution of the transcription factor TFIIH: assignment of functions for the three enzymatic subunits, XPB, XPD, and Cdk7. Mol. Cell 3:87-95. [DOI] [PubMed] [Google Scholar]
- 42.Valay, J. G., M. Simon, M. F. Dubois, O. Bensaude, C. Facca, and G. Faye. 1995. The KIN28 gene is required both for RNA polymerase II mediated transcription and phosphorylation of the Rpb1p CTD. J. Mol. Biol. 249:535-544. [DOI] [PubMed] [Google Scholar]
- 43.Wallenfang, M., and G. Seydoux. 2002. cdk-7 is required for mRNA transcription and cell cycle progression in Caenorhabditis elegans embryos. Proc. Natl. Acad. Sci. USA 99:5527-5532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Wen, Y., and A. J. Shatkin. 1999. Transcription elongation factor hSPT5 stimulates mRNA capping. Genes Dev. 13:1774-1779. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Winegarden, N. A., K. S. Wong, M. Sopta, and J. T. Westwood. 1996. Sodium salicyclate decreases intracellular ATP, induces both heat shock factor binding and chromosomal puffing, but does not induce hsp70 gene transcription in Drosophila. J. Biol. Chem. 271:26971-26980. [DOI] [PubMed] [Google Scholar]
- 46.Zawel, L., K. P. Kumar, and D. Reinberg. 1995. Recycling of the general transcription factors during RNA polymerase II transcription. Genes Dev. 9:1479-1490. [DOI] [PubMed] [Google Scholar]
- 47.Zhou, M., M. A. Halanski, M. F. Radonovich, F. Kashanchi, J. Peng, D. H. Price, and J. N. Brady. 2000. Tat modifies the activity of CDK9 to phosphorylate serine 5 of the RNA polymerase II carboxyl-terminal domain during human immunodeficiency virus type 1 transcription. Mol. Cell. Biol. 20:5077-5086. [DOI] [PMC free article] [PubMed] [Google Scholar]