Abstract
Background
The Kit gene encodes a receptor tyrosine kinase involved in various biological processes including melanogenesis, hematopoiesis and gametogenesis in mice and human. A large number of Kit mutants has been described so far showing the pleiotropic phenotypes associated with partial loss-of-function of the gene. Hypomorphic mutations can induce a light coat color phenotype while complete lack of KIT function interferes with embryogenesis. Interestingly several intermediate hypomorphic mutations induced in addition growth retardation and post-natal mortality.
Results
In this report we investigated the post-natal role of Kit by using a panel of chemically-induced hypomorphic mutations recently isolated in the mouse. We found that, in addition to the classical phenotypes, mutations of Kit induced juvenile steatosis, associated with the downregulation of the three genes, VldlR, Lpin1 and Lpl, controlling lipid metabolism in the post-natal liver. Hence, Kit loss-of-functions mimicked the inactivation of genes controlling the hepatic metabolism of triglycerides, the major source of energy from maternal milk, leading to growth and viability defects during neonatal development.
Conclusion
This is a first report involving KIT in the control of lipid metabolism in neonates and opening new perspectives for understanding juvenile steatosis. Moreover, it reinforces the role of Kit during development of the liver and underscores the caution that should be exerted in using KIT inhibitors during anti-cancer treatment.
Background
KIT is a well-known receptor tyrosine kinase controlling melanogenesis, hematopoiesis, gametogenesis, hearing, mast cells and the survival of Cajal cells from the intestine [1-7]. Furthermore Kit is expressed in several epithelial endothelium or endocrine cells but also in cholangiocytes during development and in the adult [8,9]. The binding of KITL, induces the dimerization of the KIT molecule, followed by a change in the configuration of the intracellular domain and the autophosphorylation of the receptor, opening several docking sites for signal transduction molecules containing SH2 binding domains. Hence the KIT signal is transduced by several pathways including that of the SRC family kinases, PLC-γ, Ras/MAPK, PI3K-AKT-mTor, JAK/STAT and also through the activation of the SNAI2 and MITF transcription factors [for review see [10-12]]. KIT belongs to the class III of the receptor tyrosine kinase family with CSF1R, PDGFR-a,-b, KDR, FLT-1, -3 and -4, characterized by the presence of an autoinhibitory juxtamembrane (JM) domain. Without phosphorylation, the JM domain is inserted in between the N- and C-lobes blocking the DFG (Asp-Phe-Gly) conserved motif of the activation loop in an inactive conformation. Upon phosphorylation, changes in the conformation of the JM domain opens up the ATP binding site and repositions correctly the activation loop in the kinase domain [13,14]. Mutations that affect the activation loop are found in human mastocytose or leukemia, while deletions or insertions in the JM domain are observed in gastro intestinal tumor (GIST) and often correspond to gain-of-function [15]. The treatment of the Kit-induced tumors had taken a promising perspective with the use of STI-571 (Imatinib, Gleevec Novartis, Basel, Switzerland), that binds to the ATP-binding site, stabilizing the autoinhibitory structure with an inactive tyrosine kinase domain [14].
In human, Kit loss-of-function is linked to Piebald trait [16,17] and a large panel of mutations is reported [for a synthetic panel see [18]]. Similar pigmentation defects are observed in mouse models [2] and the analysis of KIT functions was facilitated by a large number of hypomorphic murine mutants [4,12]. Some correspond to point mutations, others to large chromosomal rearrangements obtained as spontaneous or induced by chemicals or radiation, while a few were engineered [2,8,19,20]. Complete lack of Kit interferes with post-natal viability [8] while hypomorphic mutations could induce not only a variable degree of defects during melanogenesis, hematopoieisis and gametogenesis but also growth retardation and juvenile mortality [4,18].
In this report we used a panel of mutants isolated from genome-wide mutagenesis screens to investigate the post-natal role of KIT. These mutations correspond to a new allelic series of hypomorphic Kit alleles that alter differentially the melanogenesis, hematopoiesis and gametogenesis. In two such mutants, we observed an impaired viability of the homozygotes before weaning. Clearly, growth retardation and premature death of neonates in both Kit mutants were associated with juvenile steatosis during the suckling period. Further analysis pointed out that Kit is expressed in the post-natal liver and Kit mutants were suffering from an early defect in triglycerides (TG) export from the liver correlated with the downregulation of key genes participating in the control of hepatic lipid metabolism. This series of evidence established the major role of Kit in controlling liver triglycerides metabolism in neonates.
Results
Characterization of a new allelic series of mutants affecting the Kit gene
To analyze the role of Kit in neonates, we used new semi-dominant ENU-induced mutant mouse lines named Sco1, Sco5 ("Spotted coat"), Sow3 ("Sister of W") and Whc1 ("White coat"), displaying white head blazes and belly spots, that were isolated in genome wide mutagenesis programs [21,22] (Figure 1A). The corresponding mutations were mapped on the mouse chromosome 5 near the Kit locus (Figure 1B; data not shown). With the sequencing of the Kit gene from the Sco1, Sco5 and Whc1 homozygotes, unique missense mutations were found in the intracellular tyrosine kinase domain that affect M623L for Sco1, V667A for Sco5 and F809L for Whc1 (Figure 1C–1D). These changes were absent in the original C3HeB/FeJ and other strains. No mutation was found in the exonic and the splice junction sequences of the Kit gene from the Sow3 mouse mutant line, suggesting that Sow3 affects the regulation of the Kit gene as described in other mouse mutants [19,23,24]. Nonetheless, a series of complementation assays with other known alleles including KitW, KitW-v and Kittm1Alf confirmed that Kit function was altered in Sow3 (Figure 1E). Therefore, the data support that the mutants described here represent a new allelic series at the Kit locus, with three missense mutations (Sco1, Sco5 and Whc1) and one probable mutation affecting the regulation of the Kit gene (Sow3).
Figure 1.
New ENU-induced alleles at the Kit locus. (A) Heterozygous individuals carrying the Sco1, Sco5, Sow3 and Whc1 mutations display white spotting on the belly and a white forehead blaze while homozygotes (-/-) have a white coat color with black eyes. (B) Haplotype analysis of 114Sow3/+ mutant mice derived from the F1C3FeB6 × B6 strategy. Markers are shown with their position (Mb) from the centromere of chromosome 5 (position based on Ensembl v33) and the position of the Kit gene is shown by an arrowhead. The black and white boxes represent respectively the B6 and the C3Fe alleles. The cumulative number of mice sharing the same haploid genotype is noted at the top of each column. (C) The mutated amino-acids found in the Sco1, Sco5 alleles are located in the N-terminal hinge part of the intracellular tyrosine kinase domain of the KIT protein, at the vicinity of the T660 amino acid, mutated in the KitW-v allele, whereas the F809 modified in Whc1 is found at the level of the Mg2+ binding loop of the activation segment. (D) Sequence chromatograph spanning the Sco1, Sco5 and Whc1 mutation sites respectively in exons 12, 14 and 17 of the Kit gene compared with that of a wild-type control from the same genetic background (C3Fe). The amino-acid substitution is labeled in red. (E) Transheterozygous animals carrying the Sow3 and the KitW-v or the KitW allele exhibit a white coat color and pigmented eyes.
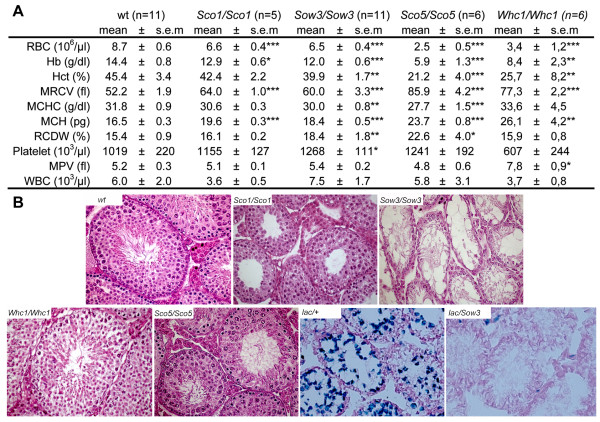
Homozygous animals generated for the four mutations displayed similar white coat pigmentation with black eyes (Figure 1A). However, Sco1 homozygotes showed a few grey hairs and pigmented ears suggesting that this allele alters later stages of pigmentation [25]. Blood analysis showed changes characteristic of macrocytic anemia, with a significant reduction of hematocrit and red blood cell count, together with an increase in mean red cell volume and in mean cell hemoglobin content for all the adult mutant mice (Figure 2A). Hematopoietic defects were more severe in the Sco5 and Whc1 homozygotes than in Sow3 and Sco1 mutants with however some little variations between males and females. Finally, histological analysis of testis isolated from adult males homozygous for Sco1, Sco5 and Whc1 showed different stages of germ cells differentiation while Sow3 homozygotes revealed a complete lack of spermatogenesis (Figure 2B). These results were confirmed by the absence of β-galactosidase staining in the seminiferous tubules of Kittm1Alf/Sow3 transheterozygotes (Figure 2B) and assessed by a fertility test. Only the Sco1 and Sco5 homozygotes were able to reproduce but no progeny was ever obtained from crosses between Sow3/Sow3 males and wild-type females (data not shown). Thus, Sco1, Sco5, Sow3 and Whc1 are hypomorphic mutations of Kit, with the expected effects, although variable, on melanogenesis, hematopoiesis and gametogenesis.
Figure 2.
Blood and germ cells phenotypes in mice homozygous for the Sco1, Sco5 and Sow3 alleles. (A) Specific parameters including Red Blood Cells (RBC), Hemoglobin (Hb), hematocrit (Hct), Mean Red Cell volume (MRCV), Mean Cell Haemoglobin Concentration (MCHC), Mean Cell Hemoglobin (MCH), Red Cell Distribution Width (RCDW), Platelet, Mean Platelet Volume (MPV) and White Blood Cells (WBC) were determined for homozygous males (Sco1/Sco1, n = 5; Sow3/Sow3, n = 11; Sco5/Sco5, n = 6; Whc1/Whc1, n = 6) and compared to wild-type (wt, n = 11) individuals. Results are expressed as mean ± sem. (B) Testis were sectioned and stained with Hematoxylin and Eosin showing a normal appearance and organization in the Sco1 and Sco5 mutants. On the contrary, the seminiferous tubules of Sow3 adult mutant individuals are lacunar with a complete absence of all the layers corresponding to the differentiation of spermatogonial stem cells. The β-galactosidase pattern of expression reveals that Kittm1Alf/Sow3 mutant mice showed no pattern of spermatogenesis compared to Kittm1Alf/+.
Sco5 mutant analysis unravels a function of Kit during post-natal development of the liver
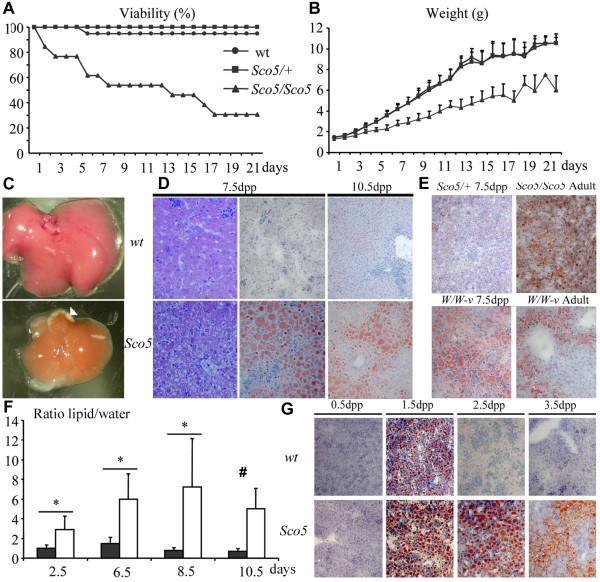
An impaired viability affected KitSco5 homozygotes before weaning, with only 21 mutants recovered out of 241 offspring from KitSco5/+ intercrosses. Although Sco5/Sco5 genotypes scored at birth with a Mendelian ratio, almost 50% of them died before 7.5 days post partum (dpp), and only 25% survived at weaning (Figure 3A). In addition, KitSco5/Sco5 homozygotes displayed a growth defect at 3.5 dpp, with a weight difference reaching up to 50% of the control at 21 dpp (Figure 3B). Comparable phenotypes were also noted in KitWhc1/Whc1 and KitW/W-v animals, thus we analyzed further this post-natal lethal phenotype.
Figure 3.
Growth and liver defects in KitSco5 homozygotes during post-natal development. (A) Viability and (B) body weight curves of Sco5/Sco5 individuals generated from heterozygous Sco5/+ intercrosses (Sco5/Sco5 n = 13; Sco5/+ n = 28; wt n = 16) reveal that mutant homozygotes are strongly affected during post-natal development compared to control littermates. (C) At 7.5 dpc, livers of mutants have a more yellowish color which was associated with the appearance of some white areas (white arrowhead). These alterations were never observed in wild-type (wt) littermates. (D) Hematoxylin and eosin, and Oil-red-O stainings reveal the swelling of hepatocytes with formation of large lipid-containing vesicles in Sco5 homozygotes versus wild-type mouse littermates at 7.5 and 10.5 dpp (magnification ×40). (E) Staining of Sco5/+ individuals at 7.5 dpp shows a relative increase in lipid droplets inside hepatocytes compared to wild-type (D). A similar defect is observed in the liver of 7.5 dpp old mice, surviving adult Sco5 homozygotes and W/W-v compound Kit mutants. (F) The lipid accumulation in the liver was studied by NMR during post-natal development at various ages on wt (n = 5; black) and mutant (n = 9,8,8,3; white) individuals. No statistical analysis was possible at 10.5 dpp because of the death of 6 mutants out of 9 (#), but the lipid/water ratio is still increased in mutant mice, at 10.5 dpp, in coherence with mortality scale and histology. (G) At 0.5 dpp, the Oil-red-O staining of liver sections from Sco5 homozygous and control mice which had not yet suckled, are identical showing no accumulation of lipids. However, after breast feeding, hepatocytes from control newborns appear to stock lipids inside microvesicles at 1.5 dpp, a phenomenon that is not observed at later stages. On the contrary, in mutant mice which have suckled milk from their mother, lipids accumulation is still found in hepatic cells after 1.5 dpp.
Necropsy analysis revealed that the liver from KitSco5/Sco5 animals at 7.5 dpp had a tan yellowish color (Figure 3C) although it had a normal relative organ weight (data not shown). Anemia was detected in mutants at this early age and could partly account for the liver color alteration. However, the spotted aspect and the presence of large white areas in the hepatic lobes in all KitSco5/Sco5 analyzed at 7.5 dpp (n > 10) were associated with an abnormal structure of hepatocytes (Figure 3D). Indeed, lipid microvesicular accumulation, or steatosis, was detected in mutant hepatocytes by specific staining at 7.5 and 10.5 dpp with Oil-red-O (Figure 3D). Similar lipid inclusions were observed in the liver of KitWhc1/Whc1 neonates (data not shown), to a lesser extent in KitSco5/+ animals and also in few KitSco5/Sco5 individuals that survived until two months of age (Figure 3E). A comparable phenotype was detected in 7.5 dpp and adult KitW/W-v mice (Figure 3E). By contrast, KitSow3 and KitSco1 homozygotes developed normally and did not display any hepatic phenotype. Thus the liver phenotype was not specific to the Sco5 allele of Kit, as it was also found in a number of compound Kit mutants.
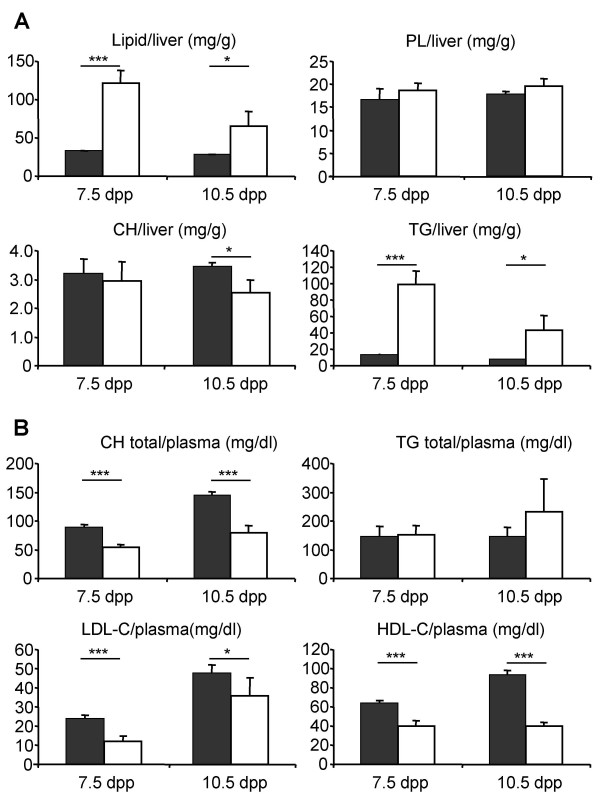
Then, we investigated the onset of the phenotype by following the lipid/water ratio in the liver of newborn mice using nuclear magnetic resonance spectroscopy (NMR). Accumulation of hepatic lipids in KitSco5/Sco5 mutants was detected at 2.5 dpp (Figure 3F) suggesting that defect started earlier. Moreover the lipid/water ratio increased until 8.5 dpp when most of the homozygotes begun to die (5 out of 8 studied) as predicted from the original analysis of viability. Nevertheless, the lipid/water ratio was still high in surviving mutants compared to controls at 10.5 dpp (Figure 3F). To further define the onset of the phenotype, liver sections from groups of mutants and control littermates (n = 3) were analyzed during post-natal development. No change was observed immediately after birth in neonates that had not yet suckled any milk (Figure 3G). However, microvesicular accumulations of lipids were found at 1.5 dpp in the liver of wild-type suckling animals from the C3HeB/FeJ, C57BL/6J and 129S2 inbred lines. In the Sco5/Sco5 mutant, a similar accumulation of lipids appeared at 1.5 dpp but remained later on, showing that lipids, derived mainly from the milk, were efficiently absorbed by the intestine, transported in the blood but abnormally stored in the liver. The characterization of the hepatic lipid content revealed a dramatic increase of triglycerides with no variation in phospholipids but a slight decrease of the cholesterol in the liver of KitSco5/Sco5 mice at 7.5 and 10.5 dpp (Figure 4A). This hepatic accumulation was not followed by an increase of the concentration of plasma triglycerides as observed in obesity or in liver diseases (Figure 4B), but the mutant mice displayed lower concentrations of free and HDL cholesterol as well as LDL cholesterol at both ages. Thus, lipid transport in plasma and exchange between tissues were deficient or not yet mature in Sco5/Sco5 suckling neonates, leading to the persistence of the steatosis at 2.5 dpp and later on. Therefore, these results suggest that the lethality induced by Kit hypomorphic alleles was concomitant with the impaired lipid metabolism observed in mutant animals during the suckling period.
Figure 4.
Quantification of lipids in the liver and plasma of Sco5/Sco5 mutants. Lipids were quantified from the liver (A; 5 individuals per genotype) and from the plasma (B; 10 individuals per genotype) of mutant and wild-type littermates at 7.5 and 10.5 dpp. Concentrations are indicated as a mean ± s.e.m and expressed as mg/g of tissue or mg/dl respectively. PL, CH, TG for plasma lipids, cholesterol, triglycerides. HDL-C and LDL-C for cholesterol associated respectively with HDL and LDL.
Kit is expressed in the liver of neonates and adults
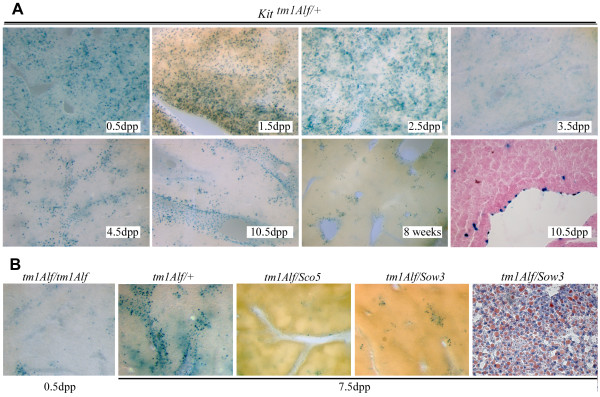
Consistent with the phenotype, Kit gene expression was detected during post-natal development in the liver by using the lacZ reporter gene inserted in the first exon of Kit from the Kittm1Alf/+ mouse [8]. Kit expression was detected in a large number of cells in the liver from 0.5 to 2.5 dpp, as a reminiscence of fetal hematopoiesis (Figure 5A). Thereafter, Kit expression was confined to a few cells located in the hepatic parenchyma and was also found in the epithelial cells of the biliary canals at later stages and in adults. As expected, the number of Kit+ cells was dramatically reduced at 0.5 dpp in moribund Kittm1Alf homozygotes [8]. In trans-heterozygotes Kittm1Alf/Sco5 and Kittm1Alf/Sow3, a reduced number of Kit+ cells were detected at 7.5 dpp while steatosis was observed (Figure 5B and data not shown). Thus, lipid accumulation was induced by several combinations of Kit mutations during the post-natal period and its appearance was clearly associated with a decrease in the number of KIT expressing cells in the liver.
Figure 5.
Hepatic expression of Kit correlates with liver phenotype observed in several mutants during the post-natal development and in adult mice. (A) Kit expression was detected in the liver of Kittm1Alf/+ mice by β-galactosidase activity on vibratome sections (40×) from 0.5 to 10.5 dpp, in adult, and on an eosin-stained sections (last panel). From birth to 2.5 dpp, β-galactosidase staining is found in a large number of hepatic cells, a reminiscence of the haematopoietic liver activity that takes place during late stage of embryonic development and ends after birth. As a consequence, the number of dispersed positive cells is reduced at 3.5 dpp. However, Kit expressing cells are clearly observed in the liver from 4.5 dpp to later stages and in adults. In Kittm1Alf/tm1Alf moribund newborns at 0.5 dpp, β-galactosidase staining is weaker due to the defect in hematopoiesis during fetal development but still remained in few cells. (B) At 7.5 dpp lacZ expression is detected in Kittm1Alf/+ liver whereas no expression is observed in transheterozygous Kittm1Alf/Sco5 and Kittm1Alf/Sow3 liver. Such compound heterozygotes suffer from a strong lipid steatosis as shown here for Kittm1Alf/Sow3 liver stained with oil-red-O (last panel). A similar staining was detected in Kittm1Alf/Sco5 liver (not shown).
Molecular characterisation of the Kit-induced steatosis during post-natal development
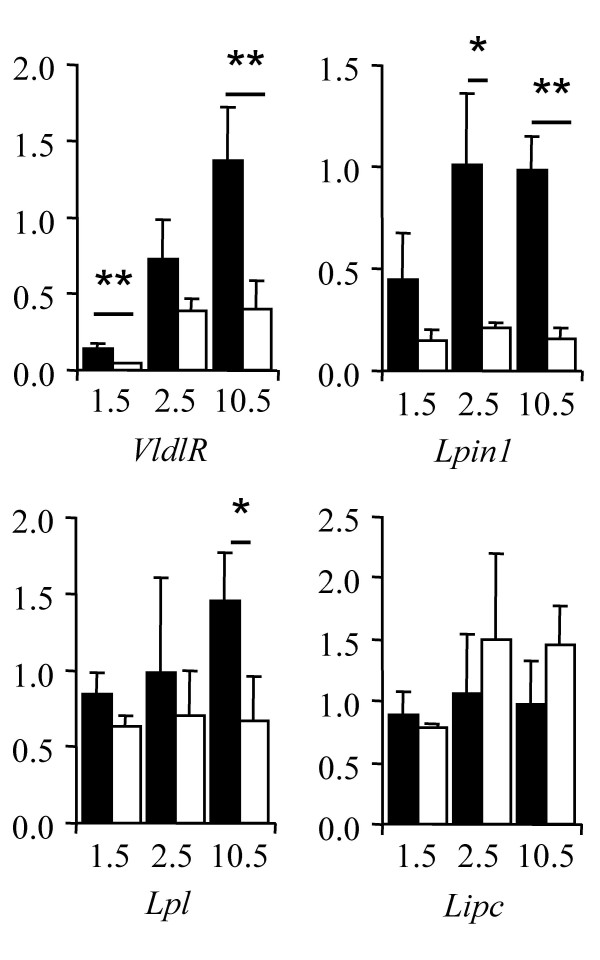
To decipher the molecular basis of Kit-dependent steatosis, we determined the expression profiles of key genes involved in lipid hepatic metabolism: such as apoliproteins (Apoa1, ApoB), lipoprotein receptors (LdlR, VldlR, Scarb1, Lrp1), lipase (Lipc, LipH, Lpl) and others implicated in hepatic lipidogenesis (Scap, Srebf1, Srebf2), lipid secretion (Pltp, Mttp, Abca1), bile acid synthesis (Cyp8b1, Cyp7a1), lipid transport (Slc10a1, Abcb11, Abcb1a, Abcc2) and a lipodystrophy gene, Lipin 1 (Lpin1), encoding a phosphatidate phosphatase enzyme with transcription activity [26-28]. Expression of most of the genes remained unchanged in wild-type and mutant individuals at 10.5 dpp (not shown) except for the very low density lipoprotein receptor (Vldlr), the Lpin1 and the lipoprotein lipase (Lpl) genes that were downregulated in Sco5/Sco5 mutants compared to wild-type littermates (Figure 6). In addition Vldlr expression appeared to be significantly retarded at 1.5 dpp just during the transient steatosis while the expression level reached a plateau at 2.5 dpp with the same expression level found at 10.5 dpp. The downregulation of Lpin1 started to be detected at 1.5 dpp and was significant at 2.5 dpp when homozygote Sco5 mutants suffer from the clearance of microvesicular accumulated lipids. Interestingly, the Lpic gene transcription is not significantly affected in the Sco5 mutant at these ages. Altogether, these changes at the transcriptional level of key genes involved in hepatic lipid uptake (VldlR) or modification (Lpin1, Lpl) are helpful to understand the molecular mechanism of the Kit-induced defect in the liver.
Figure 6.
Misregulation of VldlR, Lipin1 and Lpl, in the liver of Sco5 mutants compared to controls. At 2.5 dpp, a significant dowregulation of Lpin1 is observed in mutant (open rectangle) compare to wild type (filled rectangle) while Vldlr is significantly downregulated at 1.5 dpp. At 10.5 dpp, the expression level of VldlR, Lipin1 and Lpl is low in mutant compare to wild-type livers whereas the expression of Lipc is not affected. All the data were normalized as described in the Materials and Methods and were subjected to statistical analysis (Fischer and Student t tests, * for p < 0.05 and ** for p < 0.01).
Discussion
A new allelic series of Kit as a tool for deciphering downstream signaling pathways
In this paper, we described three new point mutations inducing loss-of-function of the KIT protein (Sco1, Sco5 and Whc1) and one potential mutation affecting the regulation of the Kit gene (Sow3). Several other mutants affecting the level of Kit expression have been described in the literature [2,19,23,29], unraveling that quiet a few dispersed regulatory elements control Kit gene expression. Most of them correspond to large chromosomal rearrangement whereas Sow3 is presumably a point mutation induced by ENU affecting a regulatory element controlling Kit during melanogenesis and gametogenesis. The missense mutations described in Sco1 and Sco5, at position M623 and V667 respectively, are found in the amino-terminal lobe near the ATP-binding site, whereas for Whc1, F809 is located in the DFG motif of the Mg2+ binding loop in the activation segment of the tyrosine kinase domain[14,30]. These residues are highly conserved in the KIT sequence deduced from various species, from human to zebrafish, and also in other mouse receptor tyrosine kinases from the same subfamily such as CSFR1, PDGFR, VEGFR, and FGFR (data not shown). Interestingly, the two residues mutated in Sco5 and Sco1 are located in the amino-terminal lobe composed of β-strands, nearby the T660, that is mutated in the W-v allele, in the structure of the active KIT kinase [13] (Figure 1C). Overall the mutations described here are presumably hypomorphic because complete loss-of-function such as in the lacZ knock-in or in the W allele are homozygous lethal [8,31]. Of particular interest is the consequence of the F809L mutation located in the activation loop. This region of the tyrosine kinase domain undergoes a conformational change after the transphosphorylation of the autoinhibitory JM domain after KITL binding. Such a change will leave open the ATP-binding site and the Mg2+ binding loop for the receptor kinase activity. In addition, the anti-cancer drug STI-571 interacts notably with the F809 and blocks the tyrosine kinase domain in an inactive conformation [14]. Remarkably, the phenotypes observed in the Whc1 homozygotes are similar to the Sco5 allele with a strong impact on pigmentation and peripheral blood cells but not on germ cells. The four Kit mutants display pleiotropic phenotypes affecting with variable degrees the melanogenesis, hematopoiesis and gametogenesis in homozygotes. Interestingly, Sco1, Sco5 and Whc1 displayed essentially normal gametogenesis suggesting that the M623, V667 in the ATP binding site and the F809 in the activation loop domain are not as important as the Y719 bound by the phosphatidylinositol 3'-kinase (PI3K) that is essential for Kit function in testis [20]. Therefore the model of the inhibitory effect of the JM domain should be reconsidered for the germ cells to reconcile the absence of major germline defect in the Whc1 mutant. These data stress out the complexity of the downstream signaling pathways and the targets that are controlled by the KIT protein in different organs.
The impaired viability of Kit mutants is associated with an altered hepatic lipid metabolism
Similar to the KitW/Wv and the KitWads/Wads mutants recently described [18], the post-natal viability of the Whc1 and of the Sco5 homozygotes is strongly affected. Further detailed analysis identifies a major defect in hepatic lipid metabolism with steatosis at 7.5 dpp in animals carrying various combinations of hypomorphic Kit alleles, including W/W-v, Sco5/lac, Sow3/lac, Sco5/Sco5 and Whc1/Whc1. Furthermore the appearance of steatosis is correlated with the defect in growth of the mutant initiated at 3.5 dpp. The determination of hepatic lipid content correlated the steatosis with a deregulated accumulation of TG just after suckling. During mammalian post-natal development, TG originate predominantly from the maternal milk and represent the major source of energy available for the neonates [32-34]. Ingested TG are assembled in chylomicrons particles and delivered to the developing organs. Following the local action of Lipoprotein Lipase (LPL), TG-derived free fatty acids (FFA) are absorbed by tissues while the remaining of the chylomicrons, the remnants, is taken-up by the liver. During the fasting period, the liver participates in the uptake, oxidation, de novo synthesis, assembly and secretion of TG in very low density lipoproteins particules (VLDL; [35]. TG in VLDL are then used by the tissues depending on their LPL activity. The abnormal level of circulating LDL-C observed in Kit mutants suggests a defect in the liver to export and metabolize lipids from maternal milk. In addition, the maintenance of the "transient steatosis" state in the Kit mutants, observed in wild-type animals in the initial suckling phase just before the third post-natal day (this work; [36]. We suggest that the transient steatosis observed in wild-type animals corresponds to a transitory accumulation of lipids while the liver organ of neonates adapts its physiology to the triglyceride-rich regime of the suckling period. This was further supported by the expression analysis of genes such as VldlR and Lpin1 that start to be expressed at 2.5 dpp (this study). In Kit mutants, a defective hepatic metabolism, with the abnormal low expression of VldlR and Lpin1, leads to the maintenance of this "transient steatosis".
Expression profiling for key genes participating to the metabolism, the transport, the modification, lipidogenesis and biliary acid synthesis in the liver revealed that only a few genes, namely Vldlr, Lpin1 and Lpl were downregulated in Kit mutants. Consistent with the Kit-induced hepatic phenotype, interference with lipase function induces type I hyperlipoproteinemia in human (OMIN238600) and more severe hyperchylomicronemia affecting post-natal viability in Lpl mutant mice [37] or in Combined Lipase deficiency (Cld), which alters both Lpl and Hepatic lipase (Lipc) activities [38-40]. In addition, VLDLR plays a major role in the metabolism of postprandial lipoproteins by enhancing LPL-mediated TG hydrolysis [41] and the Vldlr mutant mice suffer from growth retardation during the suckling period [37]. Lpin1 reduced expression is presumably a major event leading to the abnormal metabolism of lipid in KitSco5/Sco5 mice. Indeed, Lpin1 loss-of-function induces steatosis and juvenile lethality in the "fatty liver dystrophy" (fld) mice, a model of human lipodystrophy [27,42]. Recently, Lpin1 has been shown to complement the phosphatidate phosphatase yeast deficiency [28]. In yeast its homolog regulates the level of triacylglycerol and FFA and represses the expression of genes involved in phospholipid synthesis. Similarly the hepatic fatty acid oxidation is reduced in fld/fld mice [43] and Lpin1 is regulated by the transcription factor PGC-1α that regulates several metabolic pathways. In addition LIPIN1 can activate the transcription of Pparα and coactivates with PGC-1α and PPARα a large panel of other genes controlling lipid metabolism [44]. Therefore, downregulation of Lpin1 in Kit mutants will lead to a reduced oxidation of lipids and steatosis in a context of increased lipid delivery during suckling. Similar to fld/fld mice, some KitSco5/Sco5 animals could overcome steatosis after weaning but the most affected ones continued to display it. Consistent with our data, hyperlipidemia was observed in a fraction of adult KitW/W-v mice and correlated with an abnormal plasmatic lipoprotein profile and reduced lipolytic activities [45]. In addition to the Lpin1 downregulation, repression of Lpl and Vldlr would also contribute to the hepatic Kit-induced phenotype. Interestingly, Lpl homozygous mutants died within 48 h after birth with severe hypertriglyceridemia [37]. Thus, linked to the steatosis observed in Kit hypomorphic mutants were changes in the expression of genes that are known to induce hepatic lipid metabolism anomalies in mutant mice. Such a change might explain why the hepatic deficiency is found in the surviving Kit homozygote adults. The absence of Kit expressing-cells and/or reduced Kit activity during development might have an indirect effect. In such a way, Kit loss-of-function mimicked the downregulation of Lpin1, Lpl and Vldlr leading to juvenile steatosis and growth retardation. Consistently, KIT and KITL were found to be expressed in the rat cholangiocytes [9] and several mouse mutants of Kitl displayed comparable low post-natal viability [46,47].
Involvement of the Kit-dependent steatosis and anemia in post-natal lethality
The origin of the post-natal deficiency observed in Kit mutants is commonly associated with anemia. The mutant post-natal phenotype is improved by the injection of mouse embryonic liver cells or bone marrow derived cells, 24 h after birth, respectively in 26% and 32% of W/W-v mutants [48]. The rescue is even better if the transplantation of embryonic liver cells is done through the placental circulation by day 11 of gestation [49]. Similarly the rescue of Kit mutant mice could be achieved by erythropoietin [50]. Interestingly in such rescue, the RBC counts and the percentage of homozygous mutants that survived the post-natal crisis are similar to our results confirming that another KIT-dependent factor influences the survival of the pups after birth. Thus we propose that the liver defect induced by Kit partial loss-of-function is one of these additional factors.
We described some molecular events affecting the neonatal hepatic function, but other changes in the liver of Kit mutants can not be ruled out. Additional analysis will help to describe in details the cellular mechanisms underlying the liver phenotype induced by Kit loss-of-function. Thus, even though Kit-induced anemia might participate to the post-natal crisis observed in the mutant mice, we propose that Kit-induced steatosis is a major factor at the origin of the post-natal mortality. Only a controlled inactivation of Kit in blood or liver compartments will help to clearly discriminate the impact of anemia and liver defect in the Kit-dependent impaired post-natal viability.
KIT and the signaling pathways controlling lipid metabolism in the liver
KIT is a receptor protein kinase controlling several signaling pathways. Some of them are known to control liver metabolism, such as the PI3K-AKT-mTOR, JAK/STAT [51,52], or to induce growth defect like for example in the Snai2 mutant [12,53]. Thus another alternative to explain the importance of KIT in liver homeostasis could be due to the alteration of signaling pathways leading to impaired lipid metabolism. Alternatively, KIT could participate directly to the signaling pathway controlled by lipoprotein receptors. PDGFRB, another member of the same receptor family, is a co-receptor of the LDL receptor related protein [54,55]. Similarly, KIT could cooperate with members of the lipoprotein receptors family, such as VLDLR, or could control the expression of Lpin1, regulating the hepatic triglyceride metabolism and oxidative phosphorylation through its key position in the PPARa/PGC-1a regulatory pathway [44]. Interestingly, altering the binding of PI3K with the KIT protein, and reducing the activation of AKT to 10%, does not affect the weight of mutant animals [20], suggesting that, during the post-natal development, the KIT-dependent metabolism of triglycerides is not fully associated with the PI3K pathway but results from another signalling pathway.
Kit and the Fetal liver stem cells
KIT is a cellular marker of fetal liver stem cells, is involved in the differentiation/proliferation of the bipotent hepato-biliary progenitor cell [56,57]. In addition Kit expression is used to isolate population of hepatocytes during in vitro differentiation of ES cells [58]. Consistently, KITL is an important factor controlling hepatocyte proliferation after hepatectomy in IL6 mutant mice [59]. Thus, the steatosis could be an indirect consequence of a defect in the differentiation or the proliferation of hepatic progenitor cells leading to the loss of KIT+ cells observed in Sco5/lac or Sow3/lac animals. Decreasing the activity of KIT to a certain steatosis threshold will modify the pool of stem cells that will affect either the development of the liver and the post-natal hepatic homeostasis. Consequently, the defect in lipid metabolism of Sco5/Sco5 adults might result from a consequence of a post-natal defect that was not compensated during the juvenile development. This hypothesis is in agreement with the pattern of expression of the genes tested. Kit expression is restricted in the juvenile and adult liver. Thus we should foresee an indirect effect of Kit on hepatocytes functions observed in this study.
Conclusion
Taken together, these data demonstrate the key function of Kit in neonates liver physiology and further explain the early hepatic defect induced by Kit loss-of-function. Now, KIT should be considered as one of the endogenous factors that could lead to steatosis [60]. A very low number of animal models are available to study the consequences of impaired lipid metabolism during suckling, reminiscent of some human conditions (OMIN 212140, 228100). These data open new perspectives for the molecular understanding of juvenile steatosis, the analysis of Kit function during liver development and support the need to follow-up hepatic function in patients during anti-cancer treatment with the inhibitor of KIT activity, STI-571, reported to affect the weight and the viability of rat neonates [61].
Methods
Mice
The Sco1, Sco5, Sow3 mutant alleles were isolated from the chemical mutagenesis program ongoing in the Institute of Experimental Genetic at the GSF (Neuherberg, Germany) whereas the Whc1 was recovered in a similar mutagenesis screen in Orleans. Investigations were achieved in the same genetic background C3HeB/FeJ (C3Fe) for the phenotypic analysis. Outcross and backcross on C57BL/6 (B6) were performed for the genetic mapping except for Whc1 which was generated in a mixed C3Fe.B6 background. The B6 KitW and KitW-v alleles obtained from the TJL (Bar Harbor, Maine, USA) carry respectively a deletion of the transmembrane domain and a missense mutation at T660 M. The 129S2 Kittm1Alf mouse line corresponds to the disruption of the Kit gene sequence due to the insertion of a lacZ reporter gene in the first exon [8]. Standard diet (SDS) and water were provided ad libitum. All the mice were bred under SPF conditions and experiments were carried out according to the guidelines of the French Ministry of Agriculture for experiments with laboratory animals (law 87848 and YH accreditation 45–31).
Genetic and comparative gene sequence analysis
A panel of 60 MIT markers polymorphic between C3Fe and B6 was used for a genome-wide linkage analysis as described previously [21]. For complementation analysis, heterozygous mutant mice carrying the Sco5, Sco1, Sow3 or the Whc1 alleles were mated separately with KitW, KitW-v and Kittm1Alf heterozygous animals. Animals with a white coat color were classified as trans-heterozygous individuals.
For the sequence determination and the genomic DNA analysis, specific primer pairs were designed in the flanking sequences of each exon, except for the large exon 21, for which a series of overlapping PCR products were generated (Primer sequence available upon request). The amplicons generated from DNA isolated from two littermates corresponding to each phenotype (wild-type, heterozygous or homozygous mutants) were directly sequenced using the Big-dye terminator kit and analyzed on an ABI 310 automated sequencer. The nucleotide changes noticed in Sco1, Sco5 and Whc1 alleles were confirmed to be specific to the mutant lines and not found in the founder strain C3HeB/Fe or C57BL/6J and others control strains including DBA, CBA, BALB/c, RIII/Dmob, RIIIS/J, 129, A/J, AKR, PWK, IS/CamRk, IS/Cam Ei, PWD, Cast, NOD.
Phenotypic analysis
Groups of age- and sex-matched mutant and wild-type littermate mice were subjected to a series of tests following the recommendations from the standard operating procedures developed inside the EUMORPHIA network [62]. Several parameters were automatically scored for blood cell analysis by taking 100 μl of blood from suborbital sinus of 6–8 week-old anaesthetized mice (isoflurane), collecting them in EDTA tubes (VWR) and running them through a Technicon H1 hematology analyser (BAYER). Fertility tests were carried out for a period of 4 months using 5 homozygous males mated with two wild-type females from the same genetic background. Females were checked for pregnancy every week and changed every two months.
For histological analysis, the liver was fixed overnight in Bouin's solution, embedded in paraffin wax and subjected to classical procedures for hematoxylin-eosin (H&E) staining. Frozen tissue sections were post-fixed with formalin solution (Accustain, Sigma) before staining with Oil-red-O (ORO) and hematoxylin. Sections were observed under a light microscope (Leica). For lacZ staining, sections of whole fresh liver were made on a vibratome or a cryotome (Leica) and post-fixed with 5% paraformaldehyde. The β-galactosidase activity was revealed using a standard procedure [63]. Pictures were taken with a Leica digital camera and processed with the Adobe Photoshop (v.7) software.
The in vivo NMR experiments were carried out on a 9.4 T horizontal imaging spectrometer (Bruker Biospec, USR, Wissembourg, France) equipped with a 950 mT/m gradient coil and a 20 mm inner diameter 1H custom made surface coil. The mice were anesthetized with 1% isoflurane (TEM, Bordeaux) and the animal's temperature and respiratory signal were controlled during anesthesia. After liver imaging, for spectroscopy, 1D 1H PRESS spectra were recorded on 2 × 2 × 2 mm3 voxels located inside the liver (TR = 4 s, TE = 12 ms, na = 64, acquisition time = 6 min) after manual or semi automatic shimming. The total duration time was about 45 min per mouse. These spectra were used to quantify the lipids inside the liver compared with the water signal. The water and lipids peaks areas were integrated using the Xwinnmr Bruker software.
The lipid composition of the liver was evaluated using the method described in Boehler et al. [64]. Briefly, frozen livers were thawed and homogenized in propanol-2 using an ultra-turrax apparatus (Janke & Kunkel Gmb, Staufen, Germany). After incubation at 60°C for 1 h and centrifugation for 5 min at 3000 g, the supernatant was collected and the pellet was re-extracted with propanol-2. Phospholipids, triglycerides, and total cholesterol were measured enzymatically on the pooled propanolic extracts using appropriate kits: triglycerides PAP 150, phospholipids PAP 150 and Cholesterol RTU (Biomérieux S.A., 69280 Marcy l'étoile, France). Blood chemistry analysis was carried out with plasma isolated from Li-heparin-treated blood on an Olympus AU400 autoanalyzer (Olympus, Hamburg, Germany).
Expression analysis by quantitative RT-PCR
Total RNA was extracted from the liver using Trizol reagent (Life Technologies). Random hexamers were used for priming the first cDNA strand synthesis with 1 μg of total RNA. We tested the expression of a panel of genes involved in hepatic lipid metabolism and known to be expressed in hepatocytes (LdlR, Lrp1, [65], ApoA1 [66], ApoB [67], Cyp8b1, Cyp7a1 [68], Abca1 [65], Ntcp, Scap, Srebf1, Srebf2 [69,70]), in canicular membrane (Abcb11, Abcb1a, Abcc2, Abcb4 [65,69,70]), in both cell type (Scarb1 [71,72], Abcb4 [65,69]), in Kupffer cells (Lpl [73]) or in a non specified hepatic area (LipH [74]; VldlR [75], Pltp [76], Mttp [77], Lipin1 [27]). The gene specific primer pairs for Ldlr, Vldlr, ApoeR, Lpdl, Lpin1, ApoB, Mttp, Abca1, Scap, Srebf1, Srebf2, Cyp7a1, Abcc2, β2-microglobulin were obtained from Quiagen (France) and the others from Primerbank [78]. They were used to amplify the specific transcripts from cDNA obtained from at least five different individuals, and quantified using Sybergreen on MX4000 apparatus (Stratagene, La Jolla, USA) using the 18S and β2m RNAs as internal standards. Data were normalized for each gene using the 2-ΔΔCt method in controls and Sco5 homozygotes littermates (n > 5 at 10.5 dpp or n > 4 at 2.5 dpp per group) after standardization with the 18S rRNA and β2-m mRNA.
Statistical analysis
All the results are expressed by the mean ± s.e.m (standard error of the mean). Statistical analysis was carried using the Fischer's test and the Student's t-test and the null hypothesis was rejected for P < 0.05 unless otherwise indicated.
Abbreviations
ENU: N-Ethyl-N-NitrosoUrea
C3Fe: C3HeB/FeJ
B6: C57BL/6J
NMR: Nucleic Magnetic Resonance
Authors' contributions
LM conceived and performed most of the experiments and did the histological analysis for the Sco5, Sco1 and Sow3. MCC collected the samples and did the QRT-PCR analysis on the liver of mutant and control mice. VN and SA generated the genetic mapping of the Sco5, Sco1, Sow3 and Whc1. SR performed the post-natal survey and collected the samples for blood analysis. HF isolated and established the Sco5, Sco1 and Sow3 mutant lines MK did the blood chemistry analysis. PP analysed the samples from the Whc1 mutant. MR performed the lipid analysis of the liver. BTD and JCB performed the MRI experiments. JJP collected the data for the expression analysis in the liver. AP controlled the Kit sequence polymorphism in several mouse lines. MH helped draft the manuscript. YH drafted the final manuscript. All authors read and approved the final manuscript.
Acknowledgments
Acknowledgements
We thank Jean-Louis Guénet, Valérie Quesniaux, Véronique Brault, members of the laboratory for their helpful discussion and comments on the manuscript, Elodie Desale, Karine Vallon-Geoffroy and Candice Hackney for taking care of the animals, Angele Guilbot, M. Lathrop and colleagues from the CNG in Evry for their support and members of the Eumorphia consortium for their recommendations regarding the phenotype analysis. The mutant mice are available for distribution through the EMMA network ("European Mouse Mutant Archive"). This work was funded by grants from the French National Centre for Scientific Research (CNRS), the EUROCOMP (QLRI-CT1999-00877) and the EUMORPHIA projects (QLG2-CT-2002-00930) supported by the European commission under FP5.
Contributor Information
Laetitia Magnol, Email: magnol@cnrs-orleans.fr.
Marie-Clémence Chevallier, Email: chevalli@cnrs-orleans.fr.
Valérie Nalesso, Email: nalesso@cnrs-orleans.fr.
Stéphanie Retif, Email: sleroux@cnrs-orleans.fr.
Helmut Fuchs, Email: hfuchs@gsf.de.
Martina Klempt, Email: Klempt@gsf.de.
Patricia Pereira, Email: lopes@cnrs-orleans.fr.
Michel Riottot, Email: Michel.Riottot@ibaic.u-psud.fr.
Sandra Andrzejewski, Email: sandra.andrzejewski@cng.fr.
Bich-Thuy Doan, Email: doan@cnrs-orleans.fr.
Jean-Jacques Panthier, Email: panthier@jouy.inra.fr.
Anne Puech, Email: anne.puech@cng.fr.
Jean-Claude Beloeil, Email: beloeil@cnrs-orleans.fr.
Martin Hrabe de Angelis, Email: hrabe@gsf.de.
Yann Hérault, Email: herault@cnrs-orleans.fr.
References
- Geissler EN, McFarland EC, Russell ES. Analysis of pleiotropism at the dominant white-spotting (W) locus of the house mouse: a description of ten new W alleles. Genetics. 1981;97:337–361. doi: 10.1093/genetics/97.2.337. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geissler EN, Ryan MA, Housman DE. The dominant-white spotting (W) locus of the mouse encodes the c-kit proto-oncogene. Cell. 1988;55:185–192. doi: 10.1016/0092-8674(88)90020-7. [DOI] [PubMed] [Google Scholar]
- Chabot B, Stephenson DA, Chapman VM, Besmer P, Bernstein A. The proto-oncogene c-kit encoding a transmembrane tyrosine kinase receptor maps to the mouse W locus. Nature. 1988;335:88–89. doi: 10.1038/335088a0. [DOI] [PubMed] [Google Scholar]
- Besmer P, Manova K, Duttlinger R, Huang EJ, Packer A, Gyssler C, Bachvarova RF. The kit-ligand (steel factor) and its receptor c-kit/W: pleiotropic roles in gametogenesis and melanogenesis. Dev Suppl. 1993:125–137. [PubMed] [Google Scholar]
- Huizinga JD, Thuneberg L, Kluppel M, Malysz J, Mikkelsen HB, Bernstein A. W/kit gene required for interstitial cells of Cajal and for intestinal pacemaker activity. Nature. 1995;373:347–349. doi: 10.1038/373347a0. [DOI] [PubMed] [Google Scholar]
- Cable J, Jackson IJ, Steel KP. Mutations at the W locus affect survival of neural crest-derived melanocytes in the mouse. Mech Dev. 1995;50:139–150. doi: 10.1016/0925-4773(94)00331-G. [DOI] [PubMed] [Google Scholar]
- Cable J, Huszar D, Jaenisch R, Steel KP. Effects of mutations at the W locus (c-kit) on inner ear pigmentation and function in the mouse. Pigment Cell Res. 1994;7:17–32. doi: 10.1111/j.1600-0749.1994.tb00015.x. [DOI] [PubMed] [Google Scholar]
- Bernex F, De Sepulveda P, Kress C, Elbaz C, Delouis C, Panthier JJ. Spatial and temporal patterns of c-kit-expressing cells in WlacZ/+ and WlacZ/WlacZ mouse embryos. Development. 1996;122:3023–3033. doi: 10.1242/dev.122.10.3023. [DOI] [PubMed] [Google Scholar]
- Omori M, Evarts RP, Omori N, Hu Z, Marsden ER, Thorgeirsson SS. Expression of alpha-fetoprotein and stem cell factor/c-kit system in bile duct ligated young rats. Hepatology. 1997;25:1115–1122. doi: 10.1002/hep.510250512. [DOI] [PubMed] [Google Scholar]
- Roskoski R., Jr. Structure and regulation of Kit protein-tyrosine kinase--the stem cell factor receptor. Biochem Biophys Res Commun. 2005;338:1307–1315. doi: 10.1016/j.bbrc.2005.09.150. [DOI] [PubMed] [Google Scholar]
- Ronnstrand L. Signal transduction via the stem cell factor receptor/c-Kit. Cell Mol Life Sci. 2004;61:2535–2548. doi: 10.1007/s00018-004-4189-6. [DOI] [PubMed] [Google Scholar]
- Perez-Losada J, Sanchez-Martin M, Rodriguez-Garcia A, Sanchez ML, Orfao A, Flores T, Sanchez-Garcia I. Zinc-finger transcription factor Slug contributes to the function of the stem cell factor c-kit signaling pathway. Blood. 2002;100:1274–1286. doi: 10.1182/blood-2002-03-0701. [DOI] [PubMed] [Google Scholar]
- Mol CD, Lim KB, Sridhar V, Zou H, Chien EY, Sang BC, Nowakowski J, Kassel DB, Cronin CN, McRee DE. Structure of a c-kit product complex reveals the basis for kinase transactivation. J Biol Chem. 2003;278:31461–31464. doi: 10.1074/jbc.C300186200. [DOI] [PubMed] [Google Scholar]
- Mol CD, Dougan DR, Schneider TR, Skene RJ, Kraus ML, Scheibe DN, Snell GP, Zou H, Sang BC, Wilson KP. Structural basis for the autoinhibition and STI-571 inhibition of c-Kit tyrosine kinase. J Biol Chem. 2004;279:31655–31663. doi: 10.1074/jbc.M403319200. [DOI] [PubMed] [Google Scholar]
- Longley BJ, Reguera MJ, Ma Y. Classes of c-KIT activating mutations: proposed mechanisms of action and implications for disease classification and therapy. Leuk Res. 2001;25:571–576. doi: 10.1016/S0145-2126(01)00028-5. [DOI] [PubMed] [Google Scholar]
- Fleischman RA, Saltman DL, Stastny V, Zneimer S. Deletion of the c-kit protooncogene in the human developmental defect piebald trait. Proc Natl Acad Sci U S A. 1991;88:10885–10889. doi: 10.1073/pnas.88.23.10885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spritz RA, Giebel LB, Holmes SA. Dominant negative and loss of function mutations of the c-kit (mast/stem cell growth factor receptor) proto-oncogene in human piebaldism. Am J Hum Genet. 1992;50:261–269. [PMC free article] [PubMed] [Google Scholar]
- Ruan HB, Zhang N, Gao X. Identification of a novel point mutation of mouse proto-oncogene c-kit through N-ethyl-N-nitrosourea mutagenesis. Genetics. 2005;169:819–831. doi: 10.1534/genetics.104.027177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hough RB, Lengeling A, Bedian V, Lo C, Bucan M. Rump white inversion in the mouse disrupts dipeptidyl aminopeptidase-like protein 6 and causes dysregulation of Kit expression. Proc Natl Acad Sci U S A. 1998;95:13800–13805. doi: 10.1073/pnas.95.23.13800. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blume-Jensen P, Jiang G, Hyman R, Lee KF, O'Gorman S, Hunter T. Kit/stem cell factor receptor-induced activation of phosphatidylinositol 3'-kinase is essential for male fertility. Nat Genet. 2000;24:157–162. doi: 10.1038/72814. [DOI] [PubMed] [Google Scholar]
- Besson V, Nalesso V, Herpin A, Bizot JC, Messaddeq N, Romand R, Puech A, Blanquet V, Herault Y. Training and aging modulate the loss-of-balance phenotype observed in a new ENU-induced allele of Otopetrin1. Biol Cell. 2005;97:787–798. doi: 10.1042/BC20040525. [DOI] [PubMed] [Google Scholar]
- Hrabe de Angelis M, Strivens M. Large-scale production of mouse phenotypes: the search for animal models for inherited diseases in humans. Brief Bioinform. 2001;2:170–180. doi: 10.1093/bib/2.2.170. [DOI] [PubMed] [Google Scholar]
- Duttlinger R, Manova K, Berrozpe G, Chu TY, DeLeon V, Timokhina I, Chaganti RS, Zelenetz AD, Bachvarova RF, Besmer P. The Wsh and Ph mutations affect the c-kit expression profile: c-kit misexpression in embryogenesis impairs melanogenesis in Wsh and Ph mutant mice. Proc Natl Acad Sci U S A. 1995;92:3754–3758. doi: 10.1073/pnas.92.9.3754. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kluppel M, Nagle DL, Bucan M, Bernstein A. Long-range genomic rearrangements upstream of Kit dysregulate the developmental pattern of Kit expression in W57 and Wbanded mice and interfere with distinct steps in melanocyte development. Development. 1997;124:65–77. doi: 10.1242/dev.124.1.65. [DOI] [PubMed] [Google Scholar]
- Wilson YM, Richards KL, Ford-Perriss ML, Panthier JJ, Murphy M. Neural crest cell lineage segregation in the mouse neural tube. Development. 2004;131:6153–6162. doi: 10.1242/dev.01533. [DOI] [PubMed] [Google Scholar]
- Peterfy M, Phan J, Reue K. Alternatively spliced lipin isoforms exhibit distinct expression pattern, subcellular localization, and role in adipogenesis. J Biol Chem. 2005;280:32883–32889. doi: 10.1074/jbc.M503885200. [DOI] [PubMed] [Google Scholar]
- Peterfy M, Phan J, Xu P, Reue K. Lipodystrophy in the fld mouse results from mutation of a new gene encoding a nuclear protein, lipin. Nat Genet. 2001;27:121–124. doi: 10.1038/83685. [DOI] [PubMed] [Google Scholar]
- Han GS, Wu WI, Carman GM. The Saccharomyces cerevisiae Lipin homolog is a Mg2+-dependent phosphatidate phosphatase enzyme. J Biol Chem. 2006;281:9210–9218. doi: 10.1074/jbc.M600425200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berrozpe G, Timokhina I, Yukl S, Tajima Y, Ono M, Zelenetz AD, Besmer P. The W(sh), W(57), and Ph Kit expression mutations define tissue-specific control elements located between -23 and -154 kb upstream of Kit. Blood. 1999;94:2658–2666. [PubMed] [Google Scholar]
- Nolen B, Taylor S, Ghosh G. Regulation of protein kinases; controlling activity through activation segment conformation. Mol Cell. 2004;15:661–675. doi: 10.1016/j.molcel.2004.08.024. [DOI] [PubMed] [Google Scholar]
- Bernstein A, Forrester L, Reith AD, Dubreuil P, Rottapel R. The murine W/c-kit and Steel loci and the control of hematopoiesis. Semin Hematol. 1991;28:138–142. [PubMed] [Google Scholar]
- Manson WG, Weaver LT. Fat digestion in the neonate. Arch Dis Child Fetal Neonatal Ed. 1997;76:F206–11. doi: 10.1136/fn.76.3.f206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Staggers JE, Fernando-Warnakulasuriya GJ, Wells MA. Studies on fat digestion, absorption, and transport in the suckling rat. II. Triacylglycerols: molecular species, stereospecific analysis, and specificity of hydrolysis by lingual lipase. J Lipid Res. 1981;22:675–679. [PubMed] [Google Scholar]
- Fernando-Warnakulasuriya GJ, Staggers JE, Frost SC, Wells MA. Studies on fat digestion, absorption, and transport in the suckling rat. I. Fatty acid composition and concentrations of major lipid components. J Lipid Res. 1981;22:668–674. [PubMed] [Google Scholar]
- den Boer M, Voshol PJ, Kuipers F, Havekes LM, Romijn JA. Hepatic Steatosis: A Mediator of the Metabolic Syndrome. Lessons From Animal Models. Arterioscler Thromb Vasc Biol. 2004;24:644–649. doi: 10.1161/01.ATV.0000116217.57583.6e. [DOI] [PubMed] [Google Scholar]
- Hughes DE, Stolz DB, Yu S, Tan Y, Reddy JK, Watkins SC, Diehl AM, Costa RH. Elevated hepatocyte levels of the Forkhead box A2 (HNF-3beta) transcription factor cause postnatal steatosis and mitochondrial damage. Hepatology. 2003;37:1414–1424. doi: 10.1053/jhep.2003.50253. [DOI] [PubMed] [Google Scholar]
- Weinstock PH, Bisgaier CL, Aalto-Setala K, Radner H, Ramakrishnan R, Levak-Frank S, Essenburg AD, Zechner R, Breslow JL. Severe hypertriglyceridemia, reduced high density lipoprotein, and neonatal death in lipoprotein lipase knockout mice. Mild hypertriglyceridemia with impaired very low density lipoprotein clearance in heterozygotes. J Clin Invest. 1995;96:2555–2568. doi: 10.1172/JCI118319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davis RC, Ben-Zeev O, Martin D, Doolittle MH. Combined lipase deficiency in the mouse. Evidence of impaired lipase processing and secretion. J Biol Chem. 1990;265:17960–17966. [PubMed] [Google Scholar]
- Hill JS, Davis RC, Yang D, Wen J, Philo JS, Poon PH, Phillips ML, Kempner ES, Wong H. Human hepatic lipase subunit structure determination. J Biol Chem. 1996;271:22931–22936. doi: 10.1074/jbc.271.49.31290. [DOI] [PubMed] [Google Scholar]
- Boedeker JC, Doolittle MH, White AL. Differential effect of combined lipase deficiency (cld/cld) on human hepatic lipase and lipoprotein lipase secretion. J Lipid Res. 2001;42:1858–1864. [PubMed] [Google Scholar]
- Goudriaan JR, Espirito Santo SM, Voshol PJ, Teusink B, van Dijk KW, van Vlijmen BJ, Romijn JA, Havekes LM, Rensen PC. The VLDL receptor plays a major role in chylomicron metabolism by enhancing LPL-mediated triglyceride hydrolysis. J Lipid Res. 2004;45:1475–1481. doi: 10.1194/jlr.M400009-JLR200. [DOI] [PubMed] [Google Scholar]
- Langner CA, Birkenmeier EH, Ben-Zeev O, Schotz MC, Sweet HO, Davisson MT, Gordon JI. The fatty liver dystrophy (fld) mutation. A new mutant mouse with a developmental abnormality in triglyceride metabolism and associated tissue-specific defects in lipoprotein lipase and hepatic lipase activities. J Biol Chem. 1989;264:7994–8003. [PubMed] [Google Scholar]
- Rehnmark S, Giometti CS, Slavin BG, Doolittle MH, Reue K. The fatty liver dystrophy mutant mouse: microvesicular steatosis associated with altered expression levels of peroxisome proliferator-regulated proteins. J Lipid Res. 1998;39:2209–2217. [PubMed] [Google Scholar]
- Finck BN, Gropler MC, Chen Z, Leone TC, Croce MA, Harris TE, Lawrence JC, Jr., Kelly DP. Lipin 1 is an inducible amplifier of the hepatic PGC-1alpha/PPARalpha regulatory pathway. Cell Metab. 2006;4:199–210. doi: 10.1016/j.cmet.2006.08.005. [DOI] [PubMed] [Google Scholar]
- Hatanaka K, Tanishita H, Ishibashi-Ueda H, Yamamoto A. Hyperlipidemia in mast cell-deficient W/WV mice. Biochim Biophys Acta. 1986;878:440–445. doi: 10.1016/0005-2760(86)90254-7. [DOI] [PubMed] [Google Scholar]
- Rajaraman S, Wood LK, Willhite DK, Russell LB, Bedell MA. Effects of spontaneous KitlSteel mutations on survival and red blood cells of mice. Mamm Genome. 2003;14:168–174. doi: 10.1007/s00335-002-2193-4. [DOI] [PubMed] [Google Scholar]
- Rajaraman S, Davis WS, Mahakali-Zama A, Evans HK, Russell LB, Bedell MA. An allelic series of mutations in the Kit ligand gene of mice. II. Effects of ethylnitrosourea-induced Kitl point mutations on survival and peripheral blood cells of Kitl(Steel) mice. Genetics. 2002;162:341–353. doi: 10.1093/genetics/162.1.341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Capel B, Hawley R, Covarrubias L, Hawley T, Mintz B. Clonal contributions of small numbers of retrovirally marked hematopoietic stem cells engrafted in unirradiated neonatal W/Wv mice. Proc Natl Acad Sci U S A. 1989;86:4564–4568. doi: 10.1073/pnas.86.12.4564. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fleischman RA, Mintz B. Prevention of genetic anemias in mice by microinjection of normal hematopoietic stem cells into the fetal placenta. Proc Natl Acad Sci U S A. 1979;76:5736–5740. doi: 10.1073/pnas.76.11.5736. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waskow C, Terszowski G, Costa C, Gassmann M, Rodewald HR. Rescue of lethal c-KitW/W mice by erythropoietin. Blood. 2004;104:1688–1695. doi: 10.1182/blood-2004-04-1247. [DOI] [PubMed] [Google Scholar]
- Pang Q, Fagerlie S, Christianson TA, Keeble W, Faulkner G, Diaz J, Rathbun RK, Bagby GC. The Fanconi anemia protein FANCC binds to and facilitates the activation of STAT1 by gamma interferon and hematopoietic growth factors. Mol Cell Biol. 2000;20:4724–4735. doi: 10.1128/MCB.20.13.4724-4735.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hadjur S, Ung K, Wadsworth L, Dimmick J, Rajcan-Separovic E, Scott RW, Buchwald M, Jirik FR. Defective hematopoiesis and hepatic steatosis in mice with combined deficiencies of the genes encoding Fancc and Cu/Zn superoxide dismutase. Blood. 2001;98:1003–1011. doi: 10.1182/blood.V98.4.1003. [DOI] [PubMed] [Google Scholar]
- Jiang R, Lan Y, Norton CR, Sundberg JP, Gridley T. The Slug gene is not essential for mesoderm or neural crest development in mice. Dev Biol. 1998;198:277–285. [PubMed] [Google Scholar]
- Boucher P, Liu P, Gotthardt M, Hiesberger T, Anderson RG, Herz J. Platelet-derived growth factor mediates tyrosine phosphorylation of the cytoplasmic domain of the low Density lipoprotein receptor-related protein in caveolae. J Biol Chem. 2002;277:15507–15513. doi: 10.1074/jbc.M200428200. [DOI] [PubMed] [Google Scholar]
- Newton CS, Loukinova E, Mikhailenko I, Ranganathan S, Gao Y, Haudenschild C, Strickland DK. Platelet-derived growth factor receptor-beta (PDGFR-beta) activation promotes its association with the low density lipoprotein receptor-related protein (LRP). Evidence for co-receptor function. J Biol Chem. 2005;280:27872–27878. doi: 10.1074/jbc.M505410200. [DOI] [PubMed] [Google Scholar]
- Nava S, Westgren M, Jaksch M, Tibell A, Broome U, Ericzon BG, Sumitran-Holgersson S. Characterization of cells in the developing human liver. Differentiation. 2005;73:249–260. doi: 10.1111/j.1432-0436.2005.00019.x. [DOI] [PubMed] [Google Scholar]
- Wang X, Foster M, Al-Dhalimy M, Lagasse E, Finegold M, Grompe M. The origin and liver repopulating capacity of murine oval cells. Proc Natl Acad Sci U S A. 2003;100 Suppl 1:11881–11888. doi: 10.1073/pnas.1734199100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gouon-Evans V, Boussemart L, Gadue P, Nierhoff D, Koehler CI, Kubo A, Shafritz DA, Keller G. BMP-4 is required for hepatic specification of mouse embryonic stem cell-derived definitive endoderm. Nat Biotechnol. 2006;24:1402–1411. doi: 10.1038/nbt1258. [DOI] [PubMed] [Google Scholar]
- Ren X, Hogaboam C, Carpenter A, Colletti L. Stem cell factor restores hepatocyte proliferation in IL-6 knockout mice following 70% hepatectomy. J Clin Invest. 2003;112:1407–1418. doi: 10.1172/JCI200317391. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koteish A, Diehl AM. Animal models of steatosis. Semin Liver Dis. 2001;21:89–104. doi: 10.1055/s-2001-12932. [DOI] [PubMed] [Google Scholar]
- Hensley ML, Ford JM. Imatinib treatment: specific issues related to safety, fertility, and pregnancy. Semin Hematol. 2003;40:21–25. doi: 10.1053/shem.2003.50038. [DOI] [PubMed] [Google Scholar]
- EUmorphia http://www.eumorphia.org
- Herault Y, Beckers J, Kondo T, Fraudeau N, Duboule D. Genetic analysis of a Hoxd-12 regulatory element reveals global versus local modes of controls in the HoxD complex. Development. 1998;125:1669–1677. doi: 10.1242/dev.125.9.1669. [DOI] [PubMed] [Google Scholar]
- Boehler N, Riottot M, Ferezou J, Souidi M, Milliat F, Serougne C, Smith JL, Lutton C. Antilithiasic effect of beta-cyclodextrin in LPN hamster: comparison with cholestyramine. J Lipid Res. 1999;40:726–734. [PubMed] [Google Scholar]
- Chiang JY. Bile acid regulation of gene expression: roles of nuclear hormone receptors. Endocr Rev. 2002;23:443–463. doi: 10.1210/er.2000-0035. [DOI] [PubMed] [Google Scholar]
- Haas MJ, Reinacher D, Li JP, Wong NC, Mooradian AD. Regulation of apoA1 gene expression with acidosis: requirement for a transcriptional repressor. J Mol Endocrinol. 2001;27:43–57. doi: 10.1677/jme.0.0270043. [DOI] [PubMed] [Google Scholar]
- Glickman RM, Rogers M, Glickman JN. Apolipoprotein B synthesis by human liver and intestine in vitro. Proc Natl Acad Sci U S A. 1986;83:5296–5300. doi: 10.1073/pnas.83.14.5296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang J, Olin M, Rozell B, Bjorkhem I, Einarsson C, Eggertsen G, Gafvels M. Differential hepatocellular zonation pattern of cholesterol 7alpha-hydroxylase (Cyp7a1) and sterol 12alpha-hydroxylase (Cyp8b1) in the mouse. Histochem Cell Biol. 2007;127:253–261. doi: 10.1007/s00418-006-0239-5. [DOI] [PubMed] [Google Scholar]
- Geier A, Wagner M, Dietrich CG, Trauner M. Principles of hepatic organic anion transporter regulation during cholestasis, inflammation and liver regeneration. Biochim Biophys Acta. 2007;1773:283–308. doi: 10.1016/j.bbamcr.2006.04.014. [DOI] [PubMed] [Google Scholar]
- Meier PJ, Stieger B. Bile salt transporters. Annu Rev Physiol. 2002;64:635–661. doi: 10.1146/annurev.physiol.64.082201.100300. [DOI] [PubMed] [Google Scholar]
- Sehayek E, Wang R, Ono JG, Zinchuk VS, Duncan EM, Shefer S, Vance DE, Ananthanarayanan M, Chait BT, Breslow JL. Localization of the PE methylation pathway and SR-BI to the canalicular membrane: evidence for apical PC biosynthesis that may promote biliary excretion of phospholipid and cholesterol. J Lipid Res. 2003;44:1605–1613. doi: 10.1194/jlr.M200488-JLR200. [DOI] [PubMed] [Google Scholar]
- Miquel JF, Moreno M, Amigo L, Molina H, Mardones P, Wistuba. Rigotti A. Expression and regulation of scavenger receptor class B type I (SR-BI) in gall bladder epithelium. Gut. 2003;52:1017–1024. doi: 10.1136/gut.52.7.1017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Camps L, Reina M, Llobera M, Bengtsson-Olivecrona G, Olivecrona T, Vilaro S. Lipoprotein lipase in lungs, spleen, and liver: synthesis and distribution. J Lipid Res. 1991;32:1877–1888. [PubMed] [Google Scholar]
- Jin W, Broedl UC, Monajemi H, Glick JM, Rader DJ. Lipase H, a new member of the triglyceride lipase family synthesized by the intestine. Genomics. 2002;80:268–273. doi: 10.1006/geno.2002.6837. [DOI] [PubMed] [Google Scholar]
- Gafvels ME, Paavola LG, Boyd CO, Nolan PM, Wittmaack F, Chawla A, Lazar MA, Bucan M, Angelin BO, Strauss JF., 3rd Cloning of a complementary deoxyribonucleic acid encoding the murine homolog of the very low density lipoprotein/apolipoprotein-E receptor: expression pattern and assignment of the gene to mouse chromosome 19. Endocrinology. 1994;135:387–394. doi: 10.1210/en.135.1.387. [DOI] [PubMed] [Google Scholar]
- Jiang X, Francone OL, Bruce C, Milne R, Mar J, Walsh A, Breslow JL, Tall AR. Increased prebeta-high density lipoprotein, apolipoprotein AI, and phospholipid in mice expressing the human phospholipid transfer protein and human apolipoprotein AI transgenes. J Clin Invest. 1996;98:2373–2380. doi: 10.1172/JCI119050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gordon DA, Wetterau JR, Gregg RE. Microsomal triglyceride transfer protein: a protein complex required for the assembly of lipoprotein particles. Trends Cell Biol. 1995;5:317–321. doi: 10.1016/S0962-8924(00)89054-6. [DOI] [PubMed] [Google Scholar]
- Wang X, Seed B. A PCR primer bank for quantitative gene expression analysis 10.1093/nar/gng154. Nucl Acids Res. 2003;31:e154. doi: 10.1093/nar/gng154. [DOI] [PMC free article] [PubMed] [Google Scholar]