Abstract
The MTG1 gene of Saccharomyces cerevisiae, corresponding to ORF YMR097c on chromosome XIII, codes for a mitochondrial protein essential for respiratory competence. A human homologue of Mtg1p capable of partially rescuing the respiratory deficiency of a yeast mtg1 mutant has also been localized in mitochondria. Mtg1p is a member of a family of GTPases with largely unknown functions. The respiratory deficiency of mtg1 mutants stems from a defect in mitochondrial protein synthesis. Mutations in the 21S rRNA locus are able to suppress the translation defect of mtg1 null mutants. This points to the 21S rRNA or the large ribosomal subunit as the most likely target of Mtg1p action. The presence of mature size 15S and 21S mitochondrial rRNAs in mtg1 mutants excludes Mtg1p from being involved in transcription or processing of these RNAs. More likely, Mtg1p functions in assembly of the large ribosomal subunit. This is consistent with the peripheral localization of Mtg1p on the matrix side of the inner membrane and the results of in vivo mitochondrial translation assays in a temperature-sensitive mtg1 mutant.
INTRODUCTION
Of the 400–500 nuclear genes estimated to be required for the propagation of respiratory competent mitochondria, approximately one-fourth code for components of the mitochondrial protein synthetic system. This is a large pool of genetic information, considering the fact that the organelle only makes a handful of proteins. Most of the nuclear genes dedicated to maintaining mitochondrial translation encode ribosomal proteins, aminoacyl-tRNA synthetases, and initiation, elongation and termination factors. However, other genes products identified through mutant screens code for proteins that function in biogenesis of the translational machinery rather than in translation itself. Examples of the latter class of proteins include nucleases involved in processing and modification of the mitochondrial rRNAs and tRNAs (Morales et al., 1992; Sirum-Connolly and Mason, 1995).
The mitochondrial genetic system has diverged in significant ways from its prokaryotic ancestor and its present day cytoplasmic counterpart. This is evidenced not only by the deviations of the mitochondrial from the universal genetic code and a reduction in the number of tRNAs but also in the appearance of new ribosomal proteins that have no homologues in bacterial and eukaryotic cytoplasmic ribosomes (Graack and Wittmann-Liebold, 1998). Assembly of mitochondrial ribosomes presents special problems because of the dual genetic source of the RNA and protein components. In yeast, all but one of the ribosomal proteins (Terpstra et al., 1979) are derived from nuclear genes, whereas the two ribosomal RNAs are transcribed from mitochondrial genes. The factors that regulate a coordinate output of these compartmentally separated genes are not known.
In an effort to better understand how mitochondrial ribosomes are formed, we have screened for mutants blocked in mitochondrial translation. The first step in this screen was to identify genes that affect translation but do not directly participate in the process. The mtg1 mutants reported in this communication meet this criterion. MTG1 codes for a mitochondrial protein, which does not fractionate with mitochondrial ribosomes and is not homologous to any protein previously described to function in translation. The probable involvement of Mtg1p in ribosome biogenesis is supported by the identification of suppressor mutations in the 21S rRNA gene in mtDNA. We also present evidence that Mtg1p is present in human mitochondria and is capable of partially rescuing a yeast mtg1 null mutant. The wide phylogenetic distribution of Mtg1p suggests that it plays a fundamental role in expression of the mitochondrial translational machinery.
MATERIALS AND METHODS
Yeast Strains and Media
The genotypes and sources of the Saccharomyces cerevisiae strains used in this study are listed in Table 1. The respiratory-deficient mutant N472 was derived from D273-10B/A1 by mutagenesis with nitrosoguanidine. The compositions of the growth media have been described elsewhere (Myers et al., 1985).
Table 1.
Genotypes and sources of yeast strains
| Strain | Genotype | Source |
|---|---|---|
| D273-10B/A1 | α met6 | Tzagoloff et al. (1976) |
| CB11 | a ade1 | ten Berge et al. (1974) |
| N472 | α met6 mtg1-1 | This study |
| aN472 | a met6 mtg1-1 | N472×CB11 |
| W303-1A | a ade2-1 his3-1,15 leu2-3,112 trp1-1 ura3-1 | R. Rothsteina |
| W303-1B | α ade2-1 his3-1,15 leu2-3,112 trp1-1 ura3-1 | R. Rothsteina |
| W303ΔMTG1 | α ade2-1 his3-1,15 leu2-3,112 trp1-1 ura3-1 mtg1::HIS3 | This study |
| aW303ΔMTG1 | a ade2-1 his3-1,15 leu2-3,112 trp1-1 ura3-1 mtg1::HIS3 | This study |
| W303ΔMTG1ts | α ade2-1 his3-1,15 leu2-3,112 trp1-1 ura3-1 mss51::HIS3 mtg1ts | This study |
| W303ΔMSS51ts | α ade2-1 his3-1,15 leu2-3,112 trp1-1 ura3-1 mss51::HIS3 mss51ts | This study |
Dr. Rodney Rothstein, Department of Development and Human Genetics, Columbia University
Cloning of MTG1
The mtg1 mutant N472/U9 (MATα ura3 mtg1-1) was transformed by the method of Schiestl and Gietz (1989) with ∼10 μg of a plasmid library consisting of Sau3A fragments of yeast nuclear DNA averaging 10 kb cloned in the 2-μm plasmid YEp24 (Botstein and Davis, 1982). This library was a generous gift of Dr. Marian Carlson, Department of Development and Human Genetics, Columbia University. The transformation yielded several uracil-independent clones that grew on glycerol as the carbon source. The respiratory competent phenotype of one clone was found to cosegregate with the uracil independence, indicating rescue by a plasmid. This plasmid (pG132/T1) was amplified in Escherichia coli and used to subclone MTG1.
Construction of an mtg1 Null Allele
The subclone pG132/ST3 (Figure 2) was used to delete the 570-base pair long sequence between the SphI and BglII sites internal to the gene. The gapped plasmid was ligated to a 1.80-kb SphI-BamH1 fragment containing the yeast HIS3 gene. The resultant construct was used to obtain a linear 2.5-kb PvuII fragment with the mtg1::HIS3 allele. The null allele was introduced into the respiratory competent haploid strain W303-1B by the one-step gene replacement procedure (Rothstein, 1983).
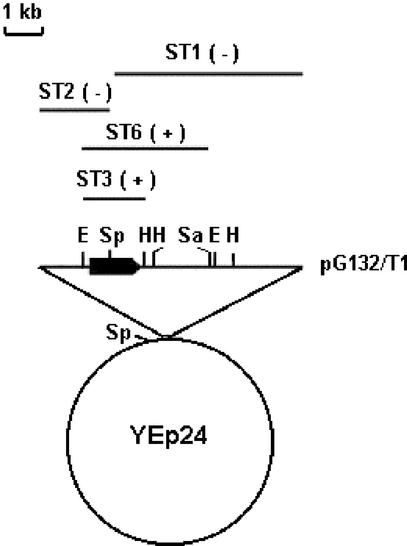
Figure 2.
Cloning of MTG1. Restriction maps of pG132/T1 and of subclones. The locations of the restriction sites for EcoRI (E), HindIII (H), SacI (Sa), and SphI (Sp) are shown above the nuclear DNA insert in pG132/T1. The regions of this insert subcloned in YEp24 (Botstein and Davis, 1982) or YEp352 (Hill et al., 1986) are represented by the solid bars in the top part of the figure. The plus and minus signs indicate complementation or lack thereof, respectively, of the mtg1 mutant N472/U9. The MTG1 reading frame and the direction of the transcription of the gene are indicated by the solid arrow in the pG132/T1 insert.
mtg1 ts Mutants
Temperature-sensitive (ts) mtg1 alleles were obtained by PCR mutagenesis of the wild-type gene (Staples and Dieckmann, 1993). The primers used for the synthesis were as follows: 5′-TGGCCACTGCAGTAGTAG and 5′-CATTTTGCCGCGGATCCAAGAACACG. Synthesis of the gene was carried out in four separate reactions containing 0.25 mM MnCl2, 1.5 mM MgCl2, and 0.2 mM dITP. In each reaction the concentration of one of the four deoxynucleotides was reduced from 0.2 to 0.02 mM. To construct the mutant library, the four 1.5-kb products were pooled and cloned in the centromeric plasmid pRS315 containing the LEU2 marker (Sikorski and Hieter, 1989). Leucine-independent clones obtained by transformation of W303ΔMTG1 with the PCR-generated library were replicated on YPEG and scored for growth at 30 and 37°C. Two mutants were isolated that grew very poorly at 37°C.
Mitochondrial Protein Synthesis
Mitochondrially encoded proteins were labeled in whole cells with [35S]methionine (7 mCi/mmol, Amersham, Piscataway, NJ) in the presence of cycloheximide (Barrientos et al., 2002). The conditions for extraction and separation the radiolabeled proteins on a 17.5% polyacrylamide gel have been described (Hell et al., 2000). The radiolabeled proteins were transferred to nitrocellulose and visualized by exposure to Kodak X-OMAT film. Deviations from this procedure are described in some figure legends.
Cloning and Expression of Human MTG1
The human cDNA clone (IMAGE 3638994) codes for a homologue of MTG1 (hMTG1). The cDNA sequence in the clone pOTB7-hMTG1 was amplified with primers hG132–1: 5′-GGCGAGCTCAATTCGGCCGAGGGCGGC and hG132-HA: 5′-CCGGAAGCTTTCAAGCGTAGTCTGGGACGTATGGGTAGGGCAAAGTCTCAGCCG to express hMtg1p with a hemagluttin (HA) epitope at the carboxy terminus. The cDNA coding for the HA tagged protein was cloned as a SacI-HindIII fragment in the vector pEGFP1-N1 (Clontech, Palo Alto, CA), regulated by a cytomegalovirus immediate early promoter to obtain the construct phG132-ST4H. For immunohistochemistry, human osteosarcoma 143B cells grown on glass slides were transiently transfected with the construct phG132-ST4H, using FuGENE6 Transfection Reagent (Roche, Indianapolis, IN) according to the manufacturer's protocol. After 48 h, immunostaining was carried out with mouse monoclonal anti-HA (Santa Cruz Biotechnology, Santa Cruz, CA) and Cox1 antibodies (Molecular Probes, Eugene, OR) as described (Sciacco and Bonilla, 1996). We used secondary anti-rabbit Cy2 or anti-mouse Cy3 (Jackson Immunochemicals, West Grove, PA) for immunodetection and visualized immunofluorescence in a Zeiss Confocal microscope. Selected digital images were visualized with different pseudocolors for HA or Cox1, as appropriate, and merged in RGB format for evaluation of colocalization. For expression of the human gene in yeast cells, the amplified hMTG1 gene was also cloned as a SacI-HindIII fragment into the yeast expression vector YEp351 (Hill et al., 1986) to obtain the construct pG132/ST5H.
Miscellaneous Procedures
Standard procedures were used for the preparation and ligation of DNA fragments and for transformation and recovery of plasmid DNA from E. coli (Maniatis et al., 1982). The entire insert of pG132/ST6 was sequenced by the method of Maxam and Gilbert (1977) using single-stranded restriction fragments labeled at their 5′ ends with 32P-ATP in the presence of polynucleotide kinase. Proteins were separated by PAGE in the buffer system of Laemmli (1970), and Western blots were treated with antibodies against the appropriate proteins followed by a second reaction with anti-rabbit IgG conjugated to horseradish peroxidase (Sigma, St. Louis, MO). The SuperSignal chemiluminescent substrate kit (Pierce, Rockford, IL) was used for the final detection. For the detection of Mtg1p, the blot was reacted with 125I-labeled protein A according to the protocol of Schmidt et al. (1984). Unless otherwise indicated, mitochondria were prepared by the method of Faye et al. (1974) except that Zymolyase 20T (ICN Laboratories, Aurora, OH) was used instead of Glusulase for the conversion of cells to spheroplasts. Protein concentrations were estimated by the Lowry procedure (Lowry et al., 1951).
RESULTS
Phenotype of mtg1 Mutants
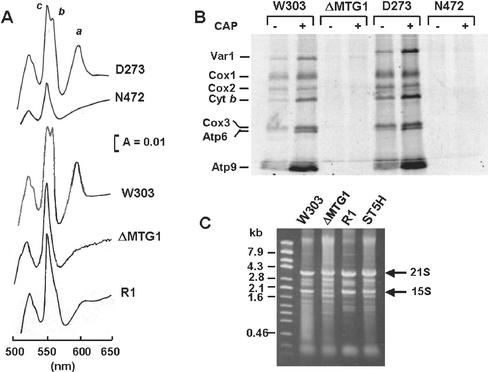
N472 is one of two independent isolates previously assigned to complementation group G132 of a collection of mutants selected for their inability to grow on respiratory carbon sources (Tzagoloff and Dieckmann, 1990). Both mutants are complemented by a ρo tester, indicating that the respiratory defect stems from recessive mutations in a nuclear gene. The mutants are pleiotropically deficient in cytochromes a, a3, and cytochrome b (Figure 1A) and lack cytochrome oxidase, NADH-cytochrome c reductase, and oligomycin-sensitive ATPase activity (Table 2). This phenotype is generally observed in mutants impaired in mitochondrial translation. This was confirmed by assays of mitochondrial protein synthesis in whole cells in the presence of cycloheximide, which showed only trace incorporation of [35S]methionine into mitochondrial gene products (Figure 1B). Preincubation of mutant cells in chloramphenicol did not improve labeling of the proteins during a subsequent pulse with [35S]methionine. The chloramphenicol treatment has been shown to increase translational efficiency, presumably by drawing newly synthesized mitochondrial products into their respective assembly pathways (Tzagoloff, 1971). In agreement with earlier studies, the chloramphenicol preincubation stimulated [35S]methionine incorporation in the two wild-type strains, W303-1A and D273-10B/A1 (Figure 1B).
Figure 1.
Characterization of mtg1 mutants. (A) Cytochrome spectra. Mitochondria were prepared from the wild-type strains D273-10B/A1 (D273) and W303-1A (W303), from the mtg1 mutants N472 and W303ΔMTG1 (ΔMTG1), and from the revertant aW303ΔMTG1/R1 (ΔMTG1/R1). They were extracted at a protein concentration of 5 mg/ml with potassium deoxycholate under conditions that quantitatively solubilize all the cytochromes (Tzagoloff et al., 1975). Difference spectra of the reduced (sodium dithionite) vs. oxidized (potassium ferricyanide) extracts were recorded at room temperature. The α absorption bands corresponding to cytochromes a and a3 have maxima at 603 nm (a). The maxima for cytochrome b (b) and for cytochrome c and c1 (c) are 560 nm and 550 nm, respectively. (B) In vivo labeling of mitochondrial gene products. The wild-type strains W303-1A and D273-10B/A1, and the mtg1 mutants N472 and W303ΔMTG1 (ΔMTG1) were grown in minimal 2% galactose medium supplemented with the appropriate prototrophic requirements. One half of the culture was incubated in the presence (+) of 2 mg/ml chloramphenicol during the last 2 h of growth. Cells were harvested from both media and washed two times with a solution containing 40 mM potassium phosphate plus 2% galactose before labeling with [35S]methionine at 30°C for 15 min in the presence of cycloheximide as described (Barrientos et al., 2002). Equivalent amounts of total cellular proteins were separated by SDS-PAGE on a 17.5% polyacrylamide gel, transferred to a nitrocellulose membrane, and exposed to an x-ray film. The mitochondrially translated ribosomal protein Var1, subunits 1 (Cox1), subunit 2 (Cox2), subunit 3 (Cox3) of cytochrome oxidase, cytochrome b (Cyt b), and subunit 6 (Atp6) and subunit 9 (Atp9) of the oligomycin-sensitive ATPase are identified in the margin. (C) Mitochondrial rRNAs. The wild-type strain W303-1A, the mutant W303ΔMTG1 (ΔMTG1), the revertant aW303ΔMTG1/R1 (R1), and the transformant W303ΔMTG1/ST5H (ST5H) harboring the human hMTG1 gene were grown to early stationary phase in YPGal (2% galactose, 1% yeast extract, 2% peptone). Mitochondria were prepared by the method of Glick and Pon (1995) and total RNA was extracted (Myers et al., 1985) from equivalent amounts of mitochondria. The RNA extracts were separated on a 1% agarose gel and stained with ethidium bromide. The migration of the mitochondrial 21S and 15S rRNAs is indicated in the margin.
Table 2.
Respiratory and ATPase activities in wild-type and mtg1 revertants
|
ATPase
|
||||
|---|---|---|---|---|
| Strain | NADH-c red. | Cyt. oxidase | - Oligomycin | + Oligomycin |
| D273-10B/A1 | 3.120±0.120 | 4.750±0.150 | 4.75 | 0.10 |
| N472 | 0.110±0.020 | 0.023±0.002 | 0.96 | 0.40 |
| W303-1A | 4.900±0.400 | 2.720±0.140 | 5.64 | 0.48 |
| aW303ΔMTG1 | 0.095±0.005 | 0.014±0.010 | 0.9 | 0.5 |
| aW303ΔMTG1/R1 | 3.000±0.300 | 0.870±0.140 | 1.89 | 0.29 |
| aW303ΔMTG1/R2 | 2.370±0.130 | 0.480±0.050 | 1.95 | 0.46 |
Values are specific activities expressed as μmol/min/mg. NADH-cytochrome c reductase and cytochrome c oxidase were measured at 23°C and ATPase at 37°C. The NADH-cytochrome c reductase and cytochrome c oxidase values are averages of two assays
Both the 21S and 15S rRNAs are present in mtg1 mutants (Figure 1C). This excludes transcription and/or processing of the precursor RNAs as a cause of the translational defect. The lowered ratio of 15S to the 21S rRNA observed in the mutant, however, suggests that the mutation affects ribosome assembly. In other studies we have noted that mutations in ribosomal proteins frequently lead to a preferential decrease in the steady state concentration of the 15S rRNA (Myers and Tzagoloff, 1987; unpublished studies).
Cloning and Disruption of MTG1
The respiratory competent clone N472/U9/T1 was obtained by transformation of the mutant with a yeast genomic library. A plasmid (pG132/T1) capable of rescuing the respiratory defect was amplified in E. coli and used to subclone the gene. The smallest subclone that conferred respiration to the mutant contained a 1.4-kb EcoRI-HindIII fragment (pG132/ST3; Figure 2). The complete sequence of this fragment revealed the presence of a single reading frame identical to ORF YMR097c on chromosome XIII. The correct identity of the gene was supported by the failure of subclones pG132/ST1 and pG132/ST2, containing the amino and the carboxyl coding regions of the gene, respectively, to complement the mutant. This gene has been named MTG1 (mitochondrial GTPase).
A mutant allele with a partial deletion of MTG1 (Δmtg1::HIS3) was obtained by replacing the sequence between the SphI and BglII sites of the gene with a 1.7-kb BamH1-SphI fragment containing the yeast HIS3 gene. A respiratory-deficient and histidine-independent clone (W303ΔMTG1), obtained by transformation of the respiratory competent haploid strain W303-1B with a linear fragment of DNA containing the partially deleted mtg1 mutation, was verified by Southern analysis to have the null mutation. The respiratory defect of the null mutant was complemented by a ρ° mutant with a wild-type copy of MTG1 but not by aN472, suggesting that the two mutations are allelic. This was supported by the phenotype of W303ΔMTG1, which is deficient in cytochrome oxidase, NADH-cytochrome c reductase and oligomycin-sensitive ATPase as a consequence of its failure to translate the mitochondrially encoded subunits of these complexes (Figure 1, Table 2).
MTG1 Codes for a Mitochondrial Protein
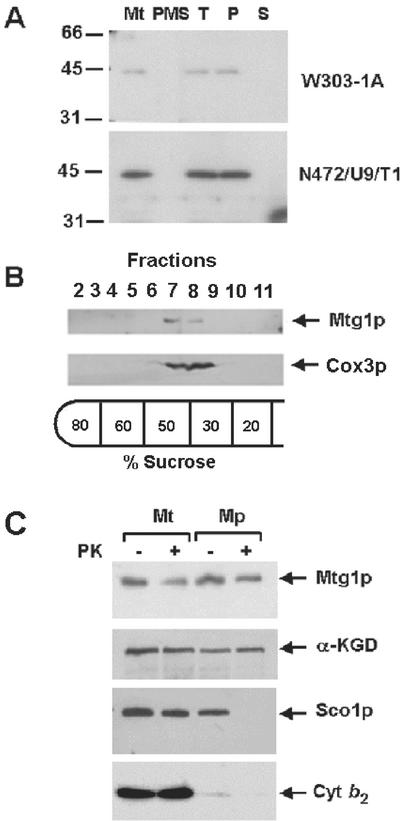
Mtg1p is located exclusively in mitochondria. An antibody against an Mtg1p/trpE fusion protein detected a 45-kDa protein in mitochondria but not in the postmitochondrial supernatant fraction (Figure 3A). The protein is more abundant in a transformant harboring MTG1 on a high-copy plasmid (Figure 3A) and is absent in the mtg1 null mutant (unpublished data).
Figure 3.
Mitochondrial localization of Mtg1p. (A) Mitochondria and the postmitochondrial supernatant fractions were prepared from the wild-type W303-1A and from N472/U8/T1, the mtg1 point mutant transformed with MTG1 on a high-copy plasmid. A sample of mitochondria was sonically irradiated and centrifuged at 100,000 × gav for 30 min. The pellet, consisting of submitochondrial particles, was suspended in the starting volume of buffer. Equivalent volumes of mitochondria (Mt), sonicated mitochondria (T), submitochondrial particles (P), and the supernatant obtained after centrifugation of the sonicated mitochondria (S) were separated on a 12% polyacrylamide gel. The amount of mitochondrial and postmitochondrial supernatant proteins (PMS) loaded on the gel was 40 μg. Proteins were transferred to nitrocellulose paper, and the Western blot was treated with antiserum against Mtg1p followed by a second incubation with 125I-labeled protein A (Schmidt et al., 1984). The antibody-antigen complexes were visualized by exposure of the blot to Kodak x-ray film. (B) Mitochondria of wild-type yeast, at a protein concentration of 10 mg/ml, were converted to submitochondrial particles by sonic disruption as in A. After centrifugation, the particles were resuspended in the starting volume of 10 mM Tris-Cl, pH 7.5, and 0.5 ml was layered on 5 ml of a discontinuous sucrose gradient prepared in 10 mM Tris-Cl, pH 7.5. The gradient was centrifuged at 260,000 × gav for 2 h. The gradient was fractionated into 12 equal fractions. Fractions 2 through 11 were separated on a 12% polyacrylamide gel, transfer to nitrocellulose, and treated with antibodies against cytochrome oxidase subunit 3 (Cox3p) and Mtg1p. Proteins were visualized with anti-rabbit IgG peroxidase-conjugated secondary antibody (Sigma, St. Louis, MO) using the Super Signal chemiluminescent substrate kit (Pierce, Rockford, IL). (C) Mitochondria were prepared by the method of Glick and Pon (1995) from W303-1A and the overexpressor N472/U8/T1. The mitochondria were suspended at a protein concentration of 8 mg/ml in 0.6 M sorbitol, 20 mM HEPES, pH 7.5. To prepare mitoplasts (Mp) the mitochondrial suspension was diluted with 8 volumes of 20 mM HEPES, pH 7.5. For controls, mitochondria (Mt) were diluted with 8 volumes of 0.6 M sorbitol, 20 mM HEPES, pH 7.5. Proteinase K (prot K) was added to one half of each sample at a final concentration of 100 μg/ml and incubated for 60 min on ice. The reaction was stopped by addition of phenylmethylsulfonyl fluoride to a final concentration of 2 mM and the mitochondria and mitoplasts were recovered by centrifugation at 100,000 × gav. The pellets were suspended in, 0.6 M sorbitol, 20 mM HEPES, pH 7.5, and proteins were precipitated by addition of 0.1 volume of 50% trichloroacetic acid and heated for 10 min at 65°C. Mitochondrial and mitoplast proteins from wild-type (80 μg) and the transformant (40 μg) were separated by SDS-PAGE on a 12% polyacrylamide gel, transferred to nitrocellulose, and probed with antibody against Mtg1p, Sco1p, α-ketoglutarate dehydrogenase (αKGD), and cytochrome b2 (Cyt b2). Antibody-antigen complexes were visualized as in B.
Even though the Mtg1p protein sequence is largely hydrophilic (unpublished data), disruption of mitochondria from wild-type or from a high-copy transformant by sonic irradiation failed to solubilize the protein (Figure 3A). The recovery of Mtg1p in the membrane fraction after disruption of mitochondria was not due to cosedimentation as a high-molecular-weight complex. This was confirmed by sedimentation of the membrane fraction on an isopycnic gradient. Mtg1p banded at the same density as the submitochondrial particles and not at the 60–80% sucrose interface where large protein complexes are expected to band (Figure 3B).
The membrane localization and topology of Mtg1p was probed by testing its sensitivity to proteinase K digestion in intact mitochondria and in mitoplasts prepared by hypotonic swelling of mitochondria. Mtg1p was protected against proteinase K in intact mitochondria and in mitoplasts (Figure 3C). Sco1p, an inner membrane protein previously shown to face the intermembrane space (Beers et al., 1997) was digested in the mitoplasts but not in mitochondria (Figure 3C). As expected, the hypotonic conditions used to disrupt the outer membrane resulted in the loss of cytochrome b2, a soluble protein marker of the intermembrane space. α-Ketoglutarate dehydrogenase, a soluble matrix protein, was protected against proteinase K in mitochondria and mitoplasts (Figure 3C). These results indicate that Mtg1p faces the matrix side of the inner membrane. Approximately 50% of Mtg1p is extracted from mitochondria with alkaline carbonate suggestive of a peripheral association with the inner membrane. The solubility properties of Mtg1p in these experiments mimicked that of the peripherally associated F1 component of the ATPase (unpublished data).
Mtg1p Is Not Associated with Ribosomes
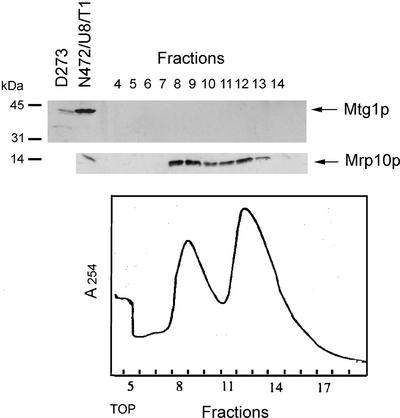
The peripheral association of Mtg1p with the inner membrane could indicate that it is a component of mitochondrial ribosomes. This was excluded by analysis of the large and small subunits of mitochondrial ribosomes. The ribosomal subunits were separated on a sucrose gradient and analyzed immunochemically for the presence of Mtg1p and Mrp10p, a constituent of the small ribosomal subunit (Jin et al., 1997). The results of this experiment failed to show Mtg1p in the fractions containing the small or large ribosomal subunits (Figure 4). The same gradient showed an enrichment of Mrp10p in the small ribosomal subunit (Jin et al., 1997).
Figure 4.
Analysis of yeast mitochondrial ribosomes. Mitochondria of the wild-type strain W303-1A were lysed with 1% potassium deoxycholate and clarified by centrifugation at 14,000 × gav for 15 min. Mitochondrial ribosomes were enriched by centrifugation of the extract through a 50% sucrose cushion (Myers et al., 1987). The ribosomal pellet was suspended in AMT (500 mM ammonium chloride, 10 mM MgCl2, 10 mM Tris-Cl, pH 7.5, 6 mM β-mercaptoethanol) and was layered on a 5-ml column of a 10–30% linear sucrose gradient in AMT buffer. After centrifugation at 65,000 rpm in a Beckman SW65Ti rotor for 100 min, the gradient was fractionated into 18 fractions. Only fractions containing the large and small subunits were analyzed as in Figure 3 with antibody against Mtg1p and Mrp10p (Jin et al., 1997).
Mitochondrial Translation in a Mutant with a ts Mutation in MTG1
Temperature-sensitive mtg1 mutants were obtained by low-stringency PCR amplification of the gene. The mtg1 null mutant was transformed with a library consisting of the PCR products cloned in a low-copy CEN plasmid. Several clones were isolated that were temperature sensitive for growth on rich glycerol medium (YPEG). One of the ts mutants (W303ΔMTG1ts) was further characterized by assessing the effect of the restrictive temperature on mitochondrial rRNAs and translational efficiency.
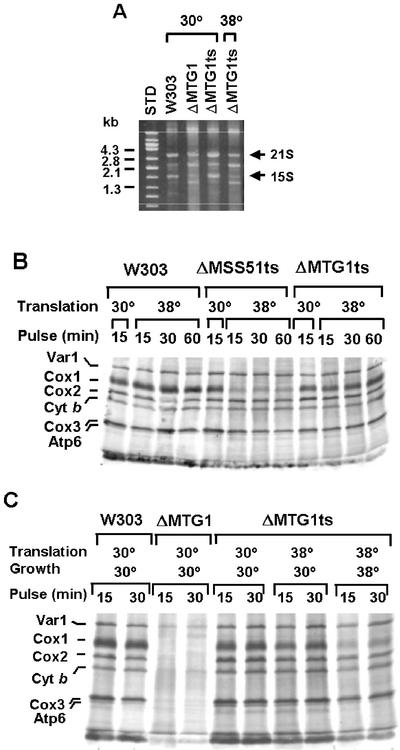
The concentrations of the 21S and 15S rRNAs in the ts transformant were comparable to that of wild-type when cells were grown at 30°C (Figure 5A). Growth at 38°C, however, resulted in a partial loss of the two rRNAs. This was especially evident for the 15S rRNA whose concentration was decreased to a level similar to that of the null mutant grown at 30°C (Figures 1C and 4A).
Figure 5.
Characterization of the mtg1 ts mutant. (A) The respiratory competent strain W303-1A (W303) and the mtg1 null mutant aW303ΔMTG1 (ΔMTG1) were grown at 30°C, and the mtg1 ts mutant aW303ΔMTG1ts (ΔMTG1ts) was grown at 30 and 38°C in YPGal to early stationary phase. Mitochondria were prepared (Hell et al., 1998) and total mitochondrial RNAs, extracted (Myers et al., 1985) from equivalent amounts of mitochondria were separated on a 1% agarose gel and stained with ethidium bromide. The migration of DNA molecular size standards (STD) and the bands corresponding to the 15S and 21S rRNAs are identified in the margins. (B) The wild-type W303-1A (W303), the mtg1 and mss51 ts mutants aW303ΔMTG1ts (ΔMTG1ts), and aW303ΔMSS51ts (ΔMSS51ts) were grown at 30°C in minimal 2% galactose medium. Mitochondrial protein synthesis was assayed as in Figure 1C either at 30 or 38°C for the indicated period of time. The incorporation of [35S]methionine into the mitochondrial translation products was essentially completed after 15 min of incubation at either temperature. (C) The wild-type W303-1A (W303) and the mtg1 null mutant aW303ΔMTG1 (ΔMTG1) were grown in 2% minimal galactose medium at 30°C, and the mtg1 ts mutant aW303ΔMSS51ts (ΔMSS51ts) was grown in the same medium at 30 and 38°C. The different strains were incubated for 15 min at 30 or 38°C before translation was uninitiated at the same temperature. Samples were taken 15 and 30 min after addition of [35S]methionine. The radioactively labeled products are identified in the margin as in Figure 1C.
Mitochondrial translation was assayed in whole cells by measuring [35S]methionine incorporation in the presence of cycloheximide. Cells grown at 30°C were incubated for 10 min either at 30 or 38°C before addition of [35S]methionine. Translation was also measured in the wild-type, the mtg1 null mutant, and an mss51 ts mutant. The results of this experiment indicated that labeling of mitochondrial translation products was not significantly different in the wild-type and the mtg1 ts mutant pretreated either at 30 or 38°C, even when incorporation of [35S]methionine was allowed to proceed for 1 h at 38°C (Figure 5B). A similar incubation of the mss51 ts mutant at 38°C resulted in significant reduction of labeling of Cox1p. Mss51p is a mitochondrial protein required for translation of subunit 1 (Cox1p) of cytochrome oxidase (Decoster et al., 1990; Siep et al., 2000).
When the mtg1 ts mutant was grown and assayed at the two different temperatures, incorporation of [35S]methionine was two or three times less in cells that had been grown at 38°C (Figure 5C). The reduced translation seen in cells grown at 38°C is consistent with their slow growth at this temperature.
Suppression of the mtg1 Null Mutant
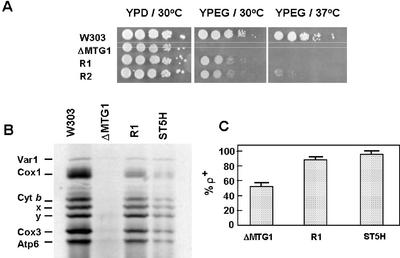
The mtg1 null mutant W303ΔMTG1 spontaneously reverts to respiratory competence on YPEG. Growth of the revertants on YPEG is slower than that of the wild-type parent at 30°C (Figure 6A). The difference in growth of the revertant and mutant is even more striking at 37°C (Figure 6A). The revertants are also cold sensitive for growth on YPEG at 24°C (unpublished data).
Figure 6.
Properties of mtg1 revertants. (A) Serial dilutions of the respiratory competent strain W303-1A (W303), the mtg1 null mutant aW303ΔMTG1 (ΔMTG1), and two revertants, aW303ΔMTG1/R1 (R1) and R2, were spotted on YPD and YPEG plates and incubated at 30 and 37°C for 2.5 d. (B) In vivo labeling of mitochondrial products with [35S]methionine in the presence of cycloheximide. The wild-type strain W303-1A (W303), the mtg1 null mutant aW303ΔMTG1 (ΔMTG1), the revertant aW303ΔMTG1/R1 (R1), and W303ΔMTG1/ST5H, the null mutant transformed with hMTG1 on a high-copy plasmid, were grown in 20 ml YPGal overnight. The cells were collected and transferred to 10 ml of fresh YPGal medium containing 2 mg/ml chloramphenicol. After a 2-h incubation at 30°C the cells were collected, washed two times in 50 ml of sterile water and transferred to 10 ml of minimal 2% galactose medium (Difco, nitrogen base without amino acids) supplemented with the prototrophic requirements and cycloheximide at a concentration of 0.5 mM. After 2 min, 0.2 mCi of [35S]methionine was added and incorporation allowed to proceed for 30 min. Protein synthesis was quenched by addition of 1.2 M sorbitol containing 1 mM cycloheximide. The cells were collected, washed several times in the same buffer, and digested with zymolyase for 5 min in 2 ml of 1.2 M sorbitol, 20 mM potassium phosphate, pH 7.5, 1 mM EDTA, 140 mM β-mercaptoethanol, and 20 mg/ml zymolyase. The spheroplasts were lysed in 0.5 M sorbitol and 0.2 mg/ml phenylmethylsulfonylfluoride, and mitochondria were isolated by differential centrifugation of the lysate. Total mitochondrial proteins (80 μg) were separated on a 12.5% SDS-PAGE gel containing 6 M urea and 6% glycerol (Claisse et al., 1980). The gel was dried before autoradiography. The mitochondrial translation products are identified in the left-hand margin as in Figure 1C. The two bands below Cyt b (labeled X and Y) have not been identified. They are consistently seen under these conditions of labeling and sample preparation. (C) Stability of mtDNA. The mtg1 null mutant (ΔMTG1), the revertant (R1), and the mutant transformed with the human hMTG1 gene (ST5H) were purified and 5 ρ+ colonies were inoculated into YPGal and grown to early stationary phase. The cultures were spread for single colonies on YPD and after several days of growth at 30°C they were replicated on a lawn of a ρo tester spread on minimal glucose medium. The diploid colonies formed on minimal glucose were replicated on YPEG and growth scored after overnight incubation at 30°C. The bars represent the average ± SD of the results corresponding to five independent colonies.
The partial rescue of respiration in the revertants correlates with a 25–30% restoration of mitochondrial protein synthesis (Figure 6B). The enhanced translational activity of mitochondria is paralleled by partial restoration of cytochromes a, a3, and b (Figure 1A) and increased stability of mtDNA. Although ∼50% of cells in a vegetatively grown culture of the mutant W303ΔMTG1 consist of ρo/- derivatives, this percentage is decreased to 10% in the revertant (Figure 6C).
Crosses of four independent revertants (aW303ΔMTG1/R1, R2, R3, and R8) to the mtg1 null mutant produced respiratory competent diploid cells, indicating that the mutations behave as dominant suppressors. The suppressors were ascertained to be mitochondrial by several criteria. Elimination of mtDNA from the revertants by ethidium bromide treatment abolished their ability to restore respiration in the mtg1 mutant N472. The suppressor did not segregate as a nuclear gene. Diploid cells obtained from a cross of aW303ΔMTG1/R1 to N472 were sporulated and tetrads dissected. Of 28 tetrads analyzed, only four had 4 viable spores. The spores in each of the 4 complete tetrads were respiratory competent, consistent with the presence of a dominant mitochondrial suppressor. Approximately 25% of the diploid cells formed from the cross of the mutant to the revertant were respiratory defective indicated that the suppressor segregated as a cytoplasmic factor.
The mitochondrial suppressors in two of the revertants were mapped by deletion analysis. aW303ΔMTG1/R1 and aW303ΔMTG1/R2 were converted to ρ- mutants by treatment with ethidium bromide, and panels of the ρ- clones were crossed to the null mutant, to different mit- and syn- mutants, and to a wild-type strain with the erythromycin resistance marker in the 21S rRNA. The resultant diploid cells were scored for growth on YPEG and for the presence of the erythromycin resistance allele. The suppressor was ascertained to be linked to a region of mtDNA with the erythromycin resistance marker and with the downstream cysteine tRNA. Attempts to further dissect this region by repeated mutagenesis with ethidium bromide were not successful, probably because of the inherent stability of the ρ- genome.
The suppressor mutations in the four revertants were further localized by sequence analysis of the mitochondrial 21S rRNA gene. The complete sequence of the gene in aW303ΔMTG1/R2 revealed a G to A transition at nucleotide 1956 based on the 21S rRNA sequence in GenBank (NC001224). This mutation was also found in 21S rRNA gene of aW303ΔMTG1/R1, whereas in aW303ΔMTG1/R3 there was G to T transversion at the neighboring nucleotide 1957 (Figure 7). The fourth revertant aW303ΔMTG1/R8 had a T to A transversion in the same region of the RNA at nucleotide 4020 of the gene. The three mutations are in a stem structure connecting domain V to the central peptidyl transferase domain of the RNA (Figure 7).
Figure 7.
Locations of the suppressor mutations in the 21S rRNA. Schematic representation of a portion of domain V of the yeast mitochondrial 21S RNA. The mutations of W303ΔMTG1/R1 and R2 (ΔMTG1/R1,R2), W303ΔMTG1/R3 (ΔMTG1/R3), and W303ΔMTG1/R8 (ΔMTG1/R8) are indicated. The nucleotide numbering is based on the DNA sequence of the gene reported in GenBank (NC 001224). The corresponding E. coli (Ec) nucleotides numbers are indicated in parenthesis. Two mutations in the stem structure, previously reported to produce erythromycin resistance (eryR); (Sor and Fukuhara, 1982; Cui and Mason, 1989), are also shown.
MTG1 Has a Human Homologue That Partially Complements the Yeast mtg1 Null Mutant
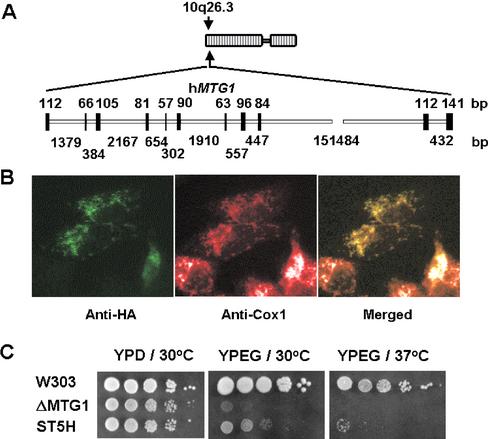
A human cDNA clone (IMAGE 3638994) codes for a putative homologue of Mtg1p. The cDNA sequence was mapped to locus 92170 on chromosome 10q26.3. The human gene, here referred to as hMTG1, consists of 11 exons and 10 introns (Figure 8A).
Figure 8.
Properties and expression of the human gene homologue of yeast MTG1. (A) Localization and genomic structure of human hMTG1 gene based on the result of a BLAST search of the human genome for homology to the hMTG1 cDNA sequence. The closed and open bars depict the exon and intron regions, respectively. Their lengths in nucleotides are indicated above and below the gene. (B) Subcellular localization of hMtg1p in human osteosarcoma 143B cells. Cells were transiently transfected with a cDNA coding for a hemagluttin-tagged hMtg1p (HA). The protein was visualized by indirect immunofluorescence using antibodies to human Cox1p and to the hemagluttin epitope (HA). A merged image is shown on the right. (C) Serial dilutions of the respiratory competent strain W303-1A (W303), the mtg1 null mutant aW303ΔMTG1 (ΔMTG1), and aW303ΔMTG1/ST5H (ST5H), the null mutant transformed with the hMTG1 cDNA in the high-copy shuttle plasmid YEp351. The plates were incubated at 30°C in YPD for 2.5 d and at 30 or 37°C for 4 d in YPEG (compare with the growth of revertants after 2.5 d shown in Figure 5A).
The subcellular location of hMtg1p was studied by transient expression of the protein tagged with a hemagluttin epitope (HA) in human osteosarcoma 143B cells. Immunohistochemical assays for the HA epitope in the transformed cells showed a typical punctate mitochondrial pattern that colocalized with subunit 1 of cytochrome c oxidase (Figure 8B).
Evidence for the functional equivalence of the two proteins was obtained by testing the ability of hMTG1 to complement a yeast mtg1 null mutant. The human cDNA was cloned in the yeast multicopy shuttle plasmid YEp351. Transformation of W303ΔMTG1 with this construct (pG132/ST5H) partially restored growth of the mutant on YPEG (Figure 8C). In vivo assays of mitochondrial translation indicated that the transformant had 5–10% of wild-type activity (Figure 6B). Human Mtg1p also alleviated the tendency of the yeast mutant to undergo deletions in mtDNA. The percentage of ρ+ cells in a vegetatively growing culture of the transformant was estimated to be greater than 95% compared with 50% in the null mutant (Figure 6C).
DISCUSSION
MTG1 codes for a peripheral protein of the mitochondrial inner membrane in yeast. Homologues exist in plants and animals, including humans. The functional equivalence of the yeast and human (hMTG1) homologues is supported by two lines of evidence. HA-tagged human Mtg1p colocalizes with a mitochondrial cytochrome oxidase marker in human osteosarcoma cells. Secondly, an hMTG1 cDNA partially rescues a yeast mtg1 null mutant strain as evidenced by growth of the transformant on a nonfermentable carbon source and increased mitochondrial translational activity.
The phenotype of mtg1 mutants indicates that the function of Mtg1p is related to mitochondrial translation. Mutations in MTG1 induce an instability of the mitochondrial genome resulting in a high rate of conversion to ρo/- mutants. Despite the tendency of mtDNA to undergo deletions, ∼50% of a vegetatively grown culture of the null mutant consists of ρ+ cells. This is significant in view of previous observations that mutations abolishing mitochondrial translational lead to a quantitative loss of the ρ+ genome (Myers et al., 1985). The presence of normal mtDNA in a sizeable fraction of mtg1 mutants indicates that they retain a low level of mitochondrial translation. This is consistent with the results of in vivo mitochondrial translation assays.
Three lines of evidence argue against a direct function of Mtg1p in mitochondrial translation. The absence of the protein in the large or small ribosomal subunits indicates that it is not a ribosomal protein. Second, Mtg1p is not homologous to any known prokaryotic or eukaryotic proteins with described functions in translation (e.g., elongation, initiation factors). Mitochondrial translation assay of the mtg1 ts mutant are also more consistent with a role of the protein in expression of the translational apparatus than in translation itself. Mitochondrial protein synthesis was inhibited when the ts mutant was grown at the nonpermissive temperature. Growth of the ts mutant at the permissive temperature, however, did not affect mitochondrial translation at either the permissive and restrictive temperature. Although these results are more compatible with a requirement of Mtg1p for some aspect of the biogenesis of the translational apparatus, they do not totally exclude the possibility of an involvement of Mtg1p in translation. For example, the ts allele could block maturation or localization of the protein in its proper compartment when cells are grown at the nonpermissive temperature.
The translational defect of mtg1 mutants is compensated by mutations in the 21S rRNA. This suggests that the target of Mtg1p action is likely to be the 21S rRNA or the large subunit of mitochondrial ribosomes. The presence of mature size 21S rRNA in yeast mtg1 null and point mutants excludes Mtg1p from having a function in processing of the primary rRNA transcripts. It is also unlikely that Mtg1p catalyzes rRNA modification because the enzymes involved in the three modifications of the 21S rRNA have been identified (Sirum-Connolly and Mason, 1995; Ansmant et al., 2000; Pintard et al., 2002).
Mtg1p is a member of a larger GTPase superfamily, although this activity has not been experimentally demonstrated in the yeast protein. The only member of this superfamily with known functions are the Nug proteins, which have been shown to be involved in export of preribosomal particles from the nucleolus to the nucleoplasm (Bassler et al., 2001). It is difficult to imagine how Mtg1p could be involved in transport of a preribosomal particle (Bassler et al., 2001) as the rRNAs are transcribed in the matrix compartment where ribosome assembly is presumed to take place.
Although the exact function Mtg1p is difficult to pinpoint, our data are most consistent with a role in ribosome assembly. Three different suppressor mutations have been identified in the 21S rRNA that partially compensate for the absence of Mtg1p. The three mutations are located in a stem structure in loop V of the peptidyl transferase domain. This region of the prokaryotic rRNAs has been shown to bind several different antibiotics, including erythromycin (Sor and Fukuhara, 1982; Ettayebi et al., 1985; Cui and Mason, 1989; Harris et al., 1989). The U to A change at nucleotide 2877 of one of the revertants confers resistance to erythromycin (unpublished data). In E. coli this region of the rRNA is close to the opening in the tunnel of the 50S subunit through which the polypeptide chain traverses during elongation (Ban et al., 2000). This region of the rRNA also interacts with the L4 and L22 subunits, both of which can acquire mutations causing erythromycin resistance (Chittum and Champney, 1994). Mtg1p may be required for assembly of this important structure in the large subunit by catalyzing a modification of a ribosomal protein that interacts with domain V or transiently stabilizes an RNA fold.
Acknowledgments
We thank Marina Gorbatyuk for technical assistance. This research was supported by National Institutes of Health Research Grant HL2274 and by a postdoctoral fellowship MDACU01991001 from the Muscular Dystrophy Association (to A.B.).
References
- Ansmant, I., Massenet, S., Grosjean, H., Motorin, Y., and Branlant, C. (2000). Identification of the Saccharomyces cerevisiae RNA: pseudouridine synthase responsible for formation of psi(2819) in 21S mitochondrial ribosomal RNA. Nucleic Acids Res. 28, 1941-1946. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ban, N., Nisson, P., Hansen, J., Moore, P.B., and Steitz, T.S. (2000). The complete atomic structure of the large ribosomal subunit at 2.4Å resolution. Science 289, 905-920. [DOI] [PubMed] [Google Scholar]
- Barrientos, A., Korr, D., and Tzagoloff, A. (2002). Shy1p is necessary for full expression of mitochondrial COX1 in the yeast model of Leigh's syndrome. EMBO J. 21, 43-52. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bassler, J., Grandi, P., Gadal, O., Lessman, T., Petfalski, E., Tollerway, D., Lechner, J., and Hurt, E. (2001). Identification of a 60S preribosomal particle that is closely linked to nuclear export. Mol. Cell 8, 517-529. [DOI] [PubMed] [Google Scholar]
- Beers, J., Glerum, D.M., and Tzagoloff, A. (1997). Purification, characterization, and localization of yeast Cox17p, a mitochondrial copper shuttle. J. Biol. Chem. 272, 33191-33196. [DOI] [PubMed] [Google Scholar]
- Botstein, D., and Davis, R.W. (1982). In: The molecular biology of the yeast Saccharomyces cerevisiae: metabolism and gene expression, ed. J.N. Strathern, E.W. Jones, and J.R. Broach, Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press, 607-636.
- Chittum, H.S., and Champney, W.S. (1994). Ribosomal protein gene sequence changes in erythromycin resistant mutants of Escherichia coli. J. Bacteriol. 176, 6192-6198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Claisse, M., Slonimski, P.P., Johnston, J., and Malher, H.R. (1980). Mutations within an intron and its flanking sites: patterns of novel polypeptides generated by mutants in one segment of the cob-box region of yeast mitochondrial DNA. Mol. Gen. Genet. 177, 375-387. [DOI] [PubMed] [Google Scholar]
- Cui, Z., and Mason, T.L. (1989). A single nucleotide substitution at the rib2 locus of the yeast mitochondrial gene for 21S rRNA confers resistance to erythromycin and cold-sensitive ribosome assembly. Curr. Genet. 16, 273-279. [DOI] [PubMed] [Google Scholar]
- Decoster, E., Simon, M., Hatat, D., and Faye, G. (1990). The MSS51 gene product is required for the translation of the COX1 mRNA in yeast mitochondria. Mol. Gen. Genet. 224, 111-118. [DOI] [PubMed] [Google Scholar]
- Ettayebi, M., Prasad, S.M., and Morgan, E.A. (1985). Chloramphenicol-erythromycin resistance mutations in a 23S rRNA gene of Escherichia coli. J. Bacteriol. 162, 551-557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Faye, G., Kujawa, C., and Fukuhara, H. (1974). Physical and genetic organization of petite and grande yeast mitochondrial DNA. IV. In vivo transcription products of mitochondrial DNA and localization of 23 S ribosomal RNA in petite mutants of Saccharomyces cerevisiae. J. Mol. Biol. 88, 185-203. [DOI] [PubMed] [Google Scholar]
- Glick, B., and Pon, L.A. (1995). Isolation of highly purified mitochondria from Saccharomyces cerevisiae. Methods Enzymol. 260, 213-223. [DOI] [PubMed] [Google Scholar]
- Graack, H.-R., and Wittmann-Liebold, B. (1998). Mitochondrial ribosomal proteins (MRPs) of yeast. Biochem. J. 329, 433-448. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harris, E.H., Burkhart, B.D., Gillham, N.W., and Boynton, J.E. (1989). Antibiotic resistance mutations in the chloroplast 16S and 23S rRNA genes of Chlamydomonas reinhardtii: correlation of genetic and physical maps of the chloroplast genome. Genetics 123, 281-292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hell, K., Herrmann, J.M., Pratje, E., Neupert, W., and Stuart, R.A. (1998). Oxa1p, an essential component of the N-tail protein export machinery in mitochondria. Proc. Natl. Acad. Sci. USA 95, 2250-2255. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hell, K., Tzagoloff, A., Neupert, W., and Stuart, R.A. (2000). Identification of Cox20p, a novel protein involved in the maturation and assembly of cytochrome oxidase subunit 2. J. Biol. Chem. 275, 4571-4578. [DOI] [PubMed] [Google Scholar]
- Hill, J.E., Myers, A.M., Koerner, T.J., and Tzagoloff, A. (1986). Yeast/E. coli shuttle vectors with multiple unique restriction sites. Yeast 2, 163-167. [DOI] [PubMed] [Google Scholar]
- Jin, C., Myers, A.M., and Tzagoloff, A. (1997). Cloning and characterization of MRP10, a yeast gene coding for a mitochondrial ribosomal protein. Curr. Genet. 31, 228-234. [DOI] [PubMed] [Google Scholar]
- Laemmli, U.K. (1970). Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227, 680-685. [DOI] [PubMed] [Google Scholar]
- Lowry, O.H., Rosebrough, N.J., Farr, A.L., and Randall, R.J. (1951). Protein measurement with the Folin phenol reagent. J. Biol. Chem. 193, 265-275. [PubMed] [Google Scholar]
- Maniatis, T., Fritsch, E.F., and Sambrook, J. (1982). Molecular cloning: a laboratory manual. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press.
- Maxam, A.M., and Gilbert, W. (1977). A new method for sequencing DNA. Proc. Natl. Acad. Sci. USA 74, 560-564. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morales, M.J., Dang, Y.L., Lou, Y.C., Sulo, P., and Martin, N.C. (1992). A 105-kDa protein is required for yeast mitochondrial RNase P activity. Proc. Natl. Acad. Sci. USA 89, 9875-9879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Myers, A.M., Crivellone, M.D., and Tzagoloff, A. (1987). MRP1 and MRP2, two yeast nuclear genes coding for mitochondrial ribosomal proteins. J. Biol. Chem. 262, 3388-97. [PubMed] [Google Scholar]
- Myers, A.M., Pape, L.K., and Tzagoloff, A. (1985). Mitochondrial protein synthesis is required for the maintenance of intact mitochondrial genomes in S. cerevisiae. EMBO J. 4, 2087-2092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pintard, L., Bujnicki, J.M., Lapeyre, B., and Bonnerot, C. (2002). MRM2 encodes a novel yeast mitochondrial 21S rRNA methyltransferase. EMBO J. 21, 1139-1147. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rothstein, R.J. (1983). One-step gene disruption in yeast. Methods Enzymol. 101, 202-211. [DOI] [PubMed] [Google Scholar]
- Schiestl, R.H., and Gietz, R.D. (1989). High efficiency transformation of intact yeast cells using single stranded nucleic acids as a carrier. Curr. Genet. 16, 339-346. [DOI] [PubMed] [Google Scholar]
- Schmidt, R.J., Myers, A.M., Gillham, N.W., and Boynton, J.E. (1984). Immunological similarities between specific chloroplast ribosomal proteins from Chlamydomonas reinhardtii and ribosomal proteins from Escherichia coli. Mol. Biol. Evol. 1, 317-334. [DOI] [PubMed] [Google Scholar]
- Sciacco, M., and Bonilla, E. (1996). Cytochemistry and immunocytochemistry of mitochondria in tissue sections. Methods Enzymol. 264, 509-521. [DOI] [PubMed] [Google Scholar]
- Siep, M., van Oosterum, K., Neufeglise, H., van der Spek, H., and Grivell, L.A. (2000). Mss51p, a putative translational activator of cytochrome c oxidase subunit-1 (COX1) mRNA, is required for synthesis of Cox1p in Saccharomyces cerevisiae. Curr. Genet. 37, 213-220. [DOI] [PubMed] [Google Scholar]
- Sikorski, R.S., and Hieter, P. (1989). A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122, 19-27. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sor, F., and Fukuhara, H. (1982). Identification of two erythromycin resistance mutations in the mitochondrial gene coding for the large ribosomal RNA in yeast. Nucleic Acids. Res. 10, 6571-6577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sirum-Connolly, K., and Mason, T.L. (1995). The role of nucleotide modifications in the yeast mitochondrial ribosome. Nucleic Acids Symp. Ser. 33, 73-75. [PubMed] [Google Scholar]
- Staples, R.R., and Dieckmann, C.L. (1993). Generation of temperature-sensitive cbp1 strains of Saccharomyces cerevisiae by PCR mutagenesis and in vivo recombination: characteristics of the mutant strains imply that CBP1 is involved in stabilization and processing of cytochrome b pre-mRNA. Genetics 135, 981-991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ten Berge, A.M., Zoutewelle, G., and Needleman, R.B. (1974). Regulation of maltose fermentation in Saccharomyces carlsbergensis. 3. Constitutive mutations at the MAL6-locus and suppressors changing a constitutive phenotype into a maltose negative phenotype. Mol. Gen. Genet. 131, 113-121. [DOI] [PubMed] [Google Scholar]
- Terpstra, P., Zanders, E., and Butow, R.A. (1979). The association of var1 with the 38S mitochondrial ribosomal subunit in yeast. J. Biol. Chem. 254, 12653-12661. [PubMed] [Google Scholar]
- Tzagoloff, A. (1971). Assembly of the mitochondrial membrane system. Role of mitochondrial and cytoplasmic protein synthesis in the biosynthesis of the rutamycin-sensitive adenosine triphosphatase. J. Biol. Chem. 246, 3050-3056. [PubMed] [Google Scholar]
- Tzagoloff, A., Akai, A., and Foury, F. (1976). Assembly of the mitochondrial membrane system XVI. Modified form of the ATPase proteolipid in oligomycin-resistant mutants of Saccharomyces cerevisiae. FEBS Lett. 65, 391-395. [DOI] [PubMed] [Google Scholar]
- Tzagoloff, A., Akai, A., and Needleman, R.B. (1975). Assembly of the mitochondrial membrane system. Characterization of nuclear mutants of Saccharomyces cerevisiae with defects in mitochondrial ATPase and respiratory enzymes. J. Biol. Chem. 250, 8228-8235. [PubMed] [Google Scholar]
- Tzagoloff, A., and Dieckmann, C.L. (1990). PET genes of Saccharomyces cerevisiae. Microbiol. Rev. 54, 211-225. [DOI] [PMC free article] [PubMed] [Google Scholar]