Abstract
The areR gene in Acinetobacter sp. strain ADP1 regulates the expression of the areCBA genes, which determine growth on benzyl alkanoates. AreR is a member of the NtrC/XylR family of regulatory proteins as determined by sequence homology. Seventy-nine bases upstream of the start of transcription is a region carrying two overlapping inverted repeat (IR) sequences that we predict to be the AreR binding site, also known as the upstream activator site (UAS). IR1 is a near-perfect (16 of 17 bp) repeat separated by 1 bp, and IR2 consists of 9- and 7-bp perfect repeats with a 3-bp gap, with the central bases of the two arms of the repeat separated by 44 and 22 bp. We report here a method for site-directed mutagenesis of chromosomal genes in ADP1 in which linear fragments generated by overlap extension PCR are used to transform ADP1 via its natural transformation system and recombinants are selected by a marker exchange-eviction strategy with a newly created sacB-Km cassette. This method was used to generate 38 strains with designed mutations in the putative UAS upstream of areCBA. The effects of the mutations on areCBA expression were measured by enzyme assays of benzyl alcohol dehydrogenase (AreB) and by reporter gene assays of lacZ inserted into areA. Substitutions or deletions in IR1 had more deleterious effects upon expression when they were in its central region, which overlaps the left arm of IR2, than when they were in its outer regions. By contrast, substitutions in the right arm of IR2 resulted in mutants with relatively high expression levels compared to that of the wild type. Effects of deletions in the right arm of IR2 were very dependent upon the length of the deletion, with 3- or 5-bp deletions reducing expression by >90% whereas an 11-bp deletion in the same area reduced the expression levels by only 50%, suggesting that alterations in the distance and the orientation of the UAS relative to the −24, −12 σ54 promoter are critical.
AreR regulates the expression of the areCBA operon, which encodes the enzymes for the sequential catabolism of (hydroxy)benzyl esters into (hydroxy)benzoates by Acinetobacter sp. strain ADP1 (11, 12). Expression of areCBA from a σ54-dependent promoter is induced only in the presence of pathway intermediates and is much reduced in a mutant with defective rpoN (encoding the σ54 subunit) (12). AreR has the domain structure distinctive of the NtrC-type family of σ54-type transcriptional regulators (12): it has the signal receptive A domain, the C domain responsible for ATP hydrolysis, and the DNA binding D domain (20, 26). The NtrC family of σ54-dependent regulators activate the holoenzyme RNA polymerase (E) by binding to DNA at upstream activator sites (UAS), which are located upstream of the associated −24, −12 promoter regions (18, 29). The regulators come into proximity with the Eσ54 via DNA looping mediated either by integration host factor (IHF) (10) or by a poly(A) region that forms an intrinsic bend in the DNA (4). The ATPase activity of the C domain of the regulator hydrolyzes ATP, which results in isomerization of the closed promoter complex, thus forming an open promoter complex that is capable of initiating transcription (1, 31).
There is no real consensus for the UAS of σ54-dependent systems except that they involve repeat sequences and are located between about bases −70 and −200 upstream of the transcriptional start (29). Two such well-studied systems for the catabolism of aromatic compounds in Pseudomonas strains are controlled by the regulators DmpR and XylR, where the UAS in each case is an inverted repeat (IR) sequence (22, 28).
Two overlapping IRs are present between bases −79 and −140 upstream of the areCBA transcriptional start (12) and were suggested as possible UAS (Fig. 1).
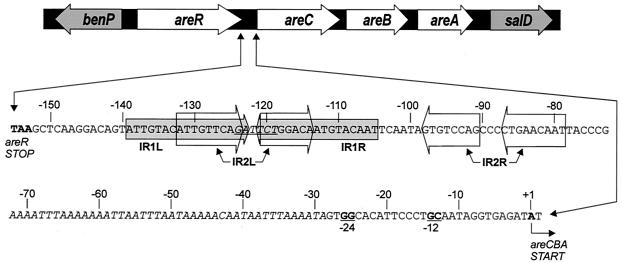
FIG. 1.
Physical map of the DNA in the areCBA promoter region showing the areCBA genes and their locations relative to one end of the supraoperonic ben-cat cluster. The nucleotide sequence of the UAS-promoter region between the stop codon for areR and the start of transcription of areCBA is expanded, with relevant elements highlighted. The start of transcription and the stop codon for the translation of AreR are labeled. The −24, −12 σ54-binding site is underlined and shown in bold. The AT-rich sequence containing a putative IHF binding site is italicized. The two IR regions (IR1 and IR2) are encased by arrows. The six bases between the two repeats of IR1 which were changed to a BamHI site (GGATCC) to insert the sacB-Km cassette are underlined and italicized.
The creation of site-directed mutations is very important for understanding gene function. Here we report the development of a reliable and efficient method for generating site-directed chromosomal mutations by allelic replacement (homologous recombination) based on the two-step marker exchange-eviction mutagenesis strategy (23). This has been developed for use in the naturally transformable bacterium Acinetobacter sp. strain ADP1, which has been used as a model for understanding the organization and integration of catabolism of carbon compounds, particularly arising from plants (21). We use a newly constructed cassette containing a gene conferring kanamycin resistance and the sacB gene from Bacillus subtilis that codes for levan sucrase and confers lethality when strains are grown in the presence of sucrose (7, 27). The sacB gene is used as a counterselectable marker when homologous mutated DNA fragments, created by overlap extension PCR (OEP) (9), are used to transform Acinetobacter sp. strain ADP1, which leads to the eviction of the sacB gene by homologous recombination. This mutational strategy was used in this study to create a variety of specific mutations within the putative UAS upstream of the areCBA promoter region as a means of confirming its position and better understanding the role of the two overlapping IR sequences.
MATERIALS AND METHODS
Strains and plasmids.
The plasmids and bacterial strains used in this study are listed in Table 1.
TABLE 1.
Bacterial strains and plasmids
| Strain or plasmida | Genotype and/or phenotypeb | Reference or source |
|---|---|---|
| Plasmids | ||
| pGEM-TEasy | TA cloning vector | Promega |
| pKOK6.1 | Apr Kmr; promoterless lacZ | 17 |
| pKOKD | pKOK6.1 with EcoRI deletion | This study |
| pKNG101 | Plasmid carrying sacB | 14 |
| pSacB | pGEM-TEasy with sacB cloned | This study |
| pRMJ1 | pKOKD with sacB from pSacB cloned into EcoRI site | This study |
| pADPW38 | pADPW36 with lacZ-Kmr cassette from pKOK6.1 cloned into NsiI site in areA | 12 |
| pADPW47 | 6.2-kbp HindIII fragment cloned from ADPW56, containing areR and areC | 12 |
| pADPW112 | 1.6-kbp amplified OEP mutagenesis fragment with BamHI site created between IR1L and IR2R cloned into pGEMeasy | This study |
| pADPW114 | pADPW112 with sacB-Km cassette from pRMJ1 cloned into BamHI site | This study |
| Acinetobacter strains | ||
| ADP1 (BD413) | Wild type | 13 |
| ADPW63 | areC::lacZ-Kmr; transformation of ADP1 with pADPW47 | 12 |
| ADPW112 | Carries sacB-Km cassette between IR1L and IR1R; transformation of ADP1 with pADPW114 | This study |
| ADPW114 | Designed BamHI site between IR1L and IR1R; ADPW112 transformed with pADPW112 | This study |
| ADPW151-153, 164-167, 171-201 | Designed mutations in and around UAS (Fig. 4) | This study |
All ADPW mutants with UAS mutations and lacZ insertions in areA are designated by the eponymous mutant number followed by the letter Z, e.g., ADPW151-Z for the mutant which has bases −112 to −130 of the ADP1 Parc UAS deleted and lacZ in areA.
Substitution mutations were always as follows: A→G, G→A, C→T, and T→C.
Bacterial strains and growth conditions.
Aromatic substrates were obtained from Sigma-Aldrich Co. Luria-Bertani medium (LB) (25) was used to cultivate bacteria unless noted otherwise. For growth on minimal medium, single carbon sources were added to the minimal salts medium (3) at the following concentrations: benzyl acetate at 2.5 mM and succinate at 10 mM. Where appropriate, ampicillin at 100 μg/ml and kanamycin at 50 μg/ml for Escherichia coli and at 10 μg/ml for Acinetobacter were used. Sucrose at 50 g liter−1 was added to LB when used for sacB counterselection.
PCR amplification.
PCR amplifications were performed in 50-μl reaction mixture volumes containing 10 ng of template DNA, 100 pmol of each primer, 2.5 nmol of each deoxynucleoside triphosphate, and 1 U of Pfu polymerase (Promega) in the reaction buffer supplied by the manufacturer. Reaction components were the same for OEP, except that the template DNA consisted of the two overlapping PCR-amplified fragments purified by excision from an 0.8% agarose gel. The PCR mixtures were subjected to a 2-min hot start at 94°C and then to 35 cycles, with one cycle consisting of 30 s at 94°C, 1 min at 56°C, and 2 min per kilobase at 72°C. PCR fragments to be cloned into pGEM-TEasy cloning vector (Promega Inc.) were incubated at 72°C for 10 min in the presence of Taq DNA polymerase, polymerase buffer, and 0.5 mM dATP prior to ligation.
Construction of pRMJ1 carrying the sacB-Km cassette.
EcoRI digestion and religation of pKOK6.1 (17) created pKOKD, a 6.85-kb ampicillin- and kanamycin-resistant derivative of pKOK6.1 but with the lacZ reporter gene and its adjacent BamHI, PstI, and SalI sites deleted (Fig. 2). The vector pKNG101 contains sacB and its promoter region (14), and primers were designed to amplify both on a single fragment. The forward primer sacB-1 (5′-AGATATCGGATCCGTCGACCTGCAGGCCCATGCAACAGAAAC-3′) contained adjacent engineered BamHI, PstI, and SalI sites (underlined), and the reverse primer sacB-2 had the sequence 5′-GGTTAGGAATGATATCAGCCATTTGCCTGCTTTTA-3′. The amplification product was A-tailed and cloned into the pGEM-TEasy cloning vector (Promega Inc.), creating pSacB. The inserted sacB PCR fragment in pSacB was flanked by the vector EcoRI sites. This enabled it to be excised as an EcoRI fragment after partial digestion to take account of the additional EcoRI site within sacB. This fragment was then cloned into the single EcoRI site within pKOKD, creating pRMJ1. Multiple restriction digests determined that sacB is in the orientation opposite that of the kanamycin resistance gene in pRMJ1 and that the two genes are transcribed divergently. Restriction digests also showed that the sacB-Km cassette thus created could be excised on a single BamHI, PstI, or SalI fragment.
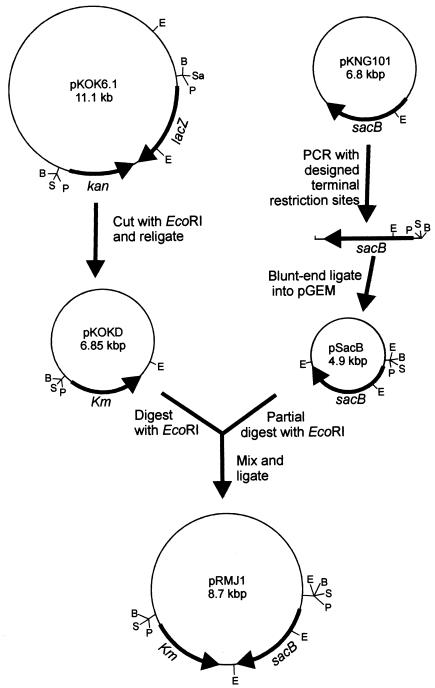
FIG. 2.
Construction of pRMJ1 containing the sacB-Km cassette. The plasmids and genes are not shown to scale. Relevant restriction sites are designated as follows: E, EcoRI; P, PstI, S, SalI; B, BamHI.
Creation of lacZ reporter gene fusions.
To generate the areA::lacZ fusion, the promoterless lacZ-Km cartridge of pKOK6.1 (17) was isolated as a PstI fragment and inserted into the compatible NsiI site in areA of plasmid pADPW36, creating pADPW38 (12). A linear DNA fragment carrying the areA::lacZ fusion was generated by digestion of pADPW38 with restriction endonucleases KpnI and XbaI and used to transform ADP1, thereby generating ADPW61. The same plasmid was linearized and transformed into each of the constructed ADP1 mutants, thereby creating a secondary set of mutants with mutations in the UAS and lacZ reporter genes in areA. Induction of the areCBA operons in these mutants was measured by β-galactosidase activities of whole cells (19) after growth to late log phase on succinate in the presence of benzyl alcohol as described above.
OEP.
PCR for OEP (9) was routinely performed by using Pfu DNA polymerase (Promega). The same two external primers, OEP-1 (5′-CACTTACTTGAATTCGAACTTTTCGGCTATG-3′) and OEP-2 (5′-TGGTACCTGCAGGCAGTAATCCCTTTGG-3′), were used for all the mutants and were complementary to regions of about 800 bp on either side of the UAS region. The mutations were designed on two overlapping internal primers (Table 2) which were routinely about 30 mers. At least 20 bases at the 5′ end of each were complementary to each other and constituted the region into which the designed mutations were incorporated. The ∼10 bases at the 3′ end of each were identical to the chromosomal sequence.
TABLE 2.
Primers used in this study
| Primer | Sequence (5′ → 3′) | Mutation created by OEP mutagenesis |
|---|---|---|
| 151/3 | GAATTGTACATGTACAATACTGTCCTTGAGC | Deletion of bases −112 to −130 |
| 151/4 | GTATTGTACATGTACAATTCAATAGTGTCC | Deletion of bases −112 to −130 |
| 152/3 | GAATTGTACAACTGTCCTTGAGCTTATTACTG | Deletion of bases −113 to −139 |
| 152/4 | CAAGGACAGTTGTACAATTCAATAGTGTCC | Deletion of bases −113 to −139 |
| 153/3 | ACACTATTGATGTACAATACTGTCCTTGAGC | Deletion of bases −105 to −131 |
| 153/4 | GTATTGTACATCAATAGTGTCCAGCCCCTG | Deletion of bases −105 to −131 |
| 164/3 | GTTCAGGGGCTCACTATTGAATTGTAC | Deletion of bases −93 to −95 |
| 164/4 | AATAGTGAGCCCCTGAACAATTACCCG | Deletion of bases −93 to −95 |
| 165/3 | GTTCAGGGGCCTATTGAATTGTAC | Deletion of bases −92 to −97 |
| 165/4 | TCAATAGGCCCCTGAACAATTACCCG | Deletion of bases −92 to −97 |
| 166/3 | AATTGTTCAAAAACTGGACACTATTG | Replacement of −87 to −90 |
| 166/4 | CCAGTTTTTGAACAATTACCCG | Replacement of −87 to −90 |
| 167/3 | TTTAAATAATTGTTCAGGGGCTGGAC | Replacement of −63 to −66 |
| 167/4 | CCCCTGAACAATTATTTAAAAATTTAAAA | Replacement of −63 to −66 |
| 171/3 | GAACAACTCGCAATACTGTCCTTGAGCT | Replacement of −132 to −135 |
| 171/4 | GACAGTATTGCGAGTTGTTCAGATTCTG | Replacement of −132 to −135 |
| 172/3 | ATCTGAGTGGTGTACAATACTGTCCTTGAG | Replacement of −128 to −131 |
| 172/4 | GTATTGTACACCACTCAGATTCTGGACAATG | Replacement of −128 to −131 |
| 173/3 | CAGAATTCAGACAATGTACAATACTGTCCTTG | Replacement of −124 to −127 |
| 173/4 | TGTACATTGTCTGAATTCTGGACAATGTAC | Replacement of −124 to −127 |
| 174/3 | TCTGAACAACTCGTGGCACTGTCCTTGAGCT | Replacement of −132 to −139 |
| 174/4 | GACAGTGCCACGAGTTGTTCAGATTCTGGAC | Replacement of −132 to −139 |
| 175/3 | ATCTGAGTGGCTCGTGGCACTGTCCTTGAGCT | Replacement of −128 to −139 |
| 175/4 | GACAGTGCCACGAGCCACTCAGATTCTGGACAATG | Replacement of −128 to −139 |
| 176/3 | CCAGAATTCAGGTGGCTCGTGGCACTGTCCTTGAGCT | Replacement of −124 to −139 |
| 176/4 | GACAGTGCCACGAGCCACCTGAATTCTGGACAATGTACAATTC | Replacement of −124 to −139 |
| 177/3 | ATCTGAGTGGCTCGCAATACTGTCCTTGAGCT | Replacement of −128 to −135 |
| 177/4 | GACAGTATTGCGAGCCACTCAGATTCTGGACAATG | Replacement of −128 to −135 |
| 178/3 | CCAGAATTACAGGTGGCTCGCAATACTGTCCTTGAGCT | Replacement of −124 to −135 |
| 178/4 | GACAGTATTGCGAGCCACCTGAATTCTGGACAATGTACAATTC | Replacement of −124 to −135 |
| 179/3 | CCAGAATTCAGGTGGTGTACAATACTGTCCTTGAGCT | Replacement of −124 to −131 |
| 179/4 | GACAGTATTGTACACCACCTGAATTCTGGACAATGTACAATTC | Replacement of −124 to −131 |
| 180/3 | TTGTACATTGTTTGAAATCTGAACAATGTACAATACTG | Replacement of −117 to −120 |
| 180/4 | TTCAGATTTCAAACAATGTACAATTCAATAGT | Replacement of −117 to −120 |
| 181/3 | TGTACACCACCCAGAATCTGAACAATGTACAATACTG | Replacement of −113 to −116 |
| 181/4 | TTCAGATTCTGGGTGGTGTACAATTCAATAGTGTCCA | Replacement of −113 to −116 |
| 182/3 | GAATTGCGTGTTGTCCAGAATCTGAACAATGTACA | Replacement of −109 to −112 |
| 182/4 | TTCAGATTCTGGACAACACGCAATTCAATAGTGTCCAGCC | Replacement of −109 to −112 |
| 183/3 | CTATTGAGCCATACATTGTCCAGAATCTGAAC | Replacement of −105 to −108 |
| 183/4 | CTGGACAATGTATGGCTCAATAGTGTCCAGCC | Replacement of −105 to −108 |
| 184/3 | TTGTACACCACTTGAAATCTGAACAATGTACAATACTG | Replacement of −113 to −120 |
| 184/4 | TTCAGATTTCAAGTGGTGTACAATTCAATAGT | Replacement of −113 to −120 |
| 185/3 | GAATTGCGTGCCACTTGAAATCTGAACAATGTACAATACTG | Replacement of −109 to −120 |
| 185/4 | TTCAGATTTCAAGTGGCACGCAATTCAATAGTGTCCA | Replacement of −109 to −120 |
| 186/3 | TGAGCCACGTGCCACTTGAAATCTGAACAATGTACAATACTG | Replacement of −105 to −120 |
| 186/4 | TTCAGATTTCAAGTGGCACGTGGCTCAATAGTGTCCAGCC | Replacement of −105 to −120 |
| 187/3 | GAATTGCGTGCCACCCAGAATCTGAACAATGTACAATACTG | Replacement of −109 to −116 |
| 187/4 | TTCAGATTCTGGGTGGCACGCAATTCAATAGTGTCCAGCC | Replacement of −109 to −116 |
| 188/3 | TTGAGCCACGTGCCACCCAGAATCTGAACAATGTACA | Replacement of −105 to −116 |
| 188/4 | TTCAGATTCTGGGTGGCACGTGGCTCAATAGTGTCCAGCC | Replacement of −105 to −116 |
| 189/3 | CTATTGAGCCACGTGTTGTCCAGAATCTGAACAATGTACA | Replacement of −105 to −112 |
| 189/4 | TTCAGATTCTGGACAACACGTGGCTCAATAGTGTCCAGCC | Replacement of −105 to −112 |
| 190/3 | TCAGGGGCTGGGTACTATTGAATTGTACATTGT | Replacement of −95 to −96 |
| 190/4 | CAATAGTACCCAGCCCCTGAACAATTACCCGA | Replacement of −95 to −96 |
| 191/3 | TCAGGGGCTAAACACTATTGAATTGTACATTGT | Replacement of −93 to −94 |
| 191/4 | CAATAGTGTTTAGCCCCTGAACAATTACCCGA | Replacement of −93 to −94 |
| 192/3 | TCAGGGGTCGGACATATTGAATTGTACATTGT | Replacement of −91 to −92 |
| 192/4 | CAATAGTGTCCGACCCCTGAACAATTACCGA | Replacement of −91 to −92 |
| 193/3 | TCAGGGGCTAAGTACTATTGAATTGTACATTGT | Replacement of −93 to −96 |
| 193/4 | CAATAGTACTTAGCCCCTGAACAATTACCCGA | Replacement of −93 to −96 |
| 194/3 | TCAGGGGTCAAGTACTATTGAATTGTACATTGT | Replacement of −91 to −96 |
| 194/4 | CAATAGTACTTGACCCCTGAACAATTACCCGA | Replacement of −91 to −96 |
| 195/3 | TCAGGGGTCAAACACTATTGAATTGTACATTGT | Replacement of −91 to −94 |
| 195/4 | CAATAGTGTTTGACCCCTGAACAATTACCCGA | Replacement of −91 to −94 |
| 197/3 | TTGTTCAGGTATTGAATTGTACATTGTCCAG | Deletion of bases −90 to −98 |
| 197/4 | GTACAATTCAATACCCTGAACAATTACCCGA | Deletion of bases −90 to −98 |
| 198/3 | TGTTCAGGGATTGAATTGTACATTGTCCAG | Deletion of bases −90 to −99 |
| 198/4 | GTACAATTCAATCCCTGAACAATTACCCGA | Deletion of bases −90 to −99 |
| 199/3 | TTGTTCAGGATTGAATTGTACATTGTCCAG | Deletion of bases −90 to −100 |
| 199/4 | GTACAATTCAATCCTGAACAATTACCCGA | Deletion of bases −90 to −100 |
| 200/3 | GTACAATACTGCTTTTGAGCTTATTACTG | Replacement of −144 to −146 |
| 200/4 | GCTCAAAAGCAGTATTGTACATTGTTCAG | Replacement of −144 to −146 |
| 201/3 | GTACAATACCACTTCTGAGCTTATTACTG | Replacement of −142 to −147 |
| 201/4 | GCTCAGAAGTGGTATTGTACATTGTTCAG | Replacement of −142 to −147 |
Natural transformation of ADP1 strains.
Recipient strains of ADP1 were transformed as described previously (6).
Cultures and preparation of cell extracts.
Overnight cultures of ADPW Pare mutants were prepared and inoculated into 1 liter of minimal medium containing 10 mM succinate and 1 mM benzyl alcohol. Following overnight growth on a shaking incubator at 30°C, additional succinate (to 10 mM) and benzyl acetate (to 1 mM) were added. The cells were grown for a further 4 h and harvested by centrifugation at 4°C, and the pellets were stored at −20°C. When required for assay, the pellets were resuspended in 50 mM phosphate buffer, pH 7.4, and the cells were disrupted ultrasonically. Disrupted cell suspension was clarified by centrifugation at 116,000 × g for 30 min. Samples of supernatants were stored at −20°C.
Enzyme assays.
Benzyl alcohol dehydrogenase (AreB) activity was measured as previously described (11). Assays were done in triplicate at two enzyme concentrations for two independently prepared cell extracts of each mutant.
RESULTS
Strategy for creation of chromosomal site-directed mutations. (i) Insertion of the sacB-Km cassette into the ADP1 Pare region.
The complete protocol used for construction of mutants is illustrated in Fig. 3. Because there was no appropriate restriction site (PstI, SalI, or BamHI) for insertion of the sacB-Km cassette into the region of the UAS, it was necessary to construct one. Consequently, OEP (9) was used to create a novel BamHI site in between the left arm of IR1 (IR1L) and the right arm of IR1 (IR1R) by replacing bases GATTCT at −124 to −119 with GGATCC (Fig. 1). Staggered overlapping complementary primers OEP-3 (5′-TTGTCCAGGATCCGAACAATGTACAATACT-3′) and OEP-4 (5′-ATTGTTCGGATCCTGGACAATGTACAATTCA-3′) were designed. The novel BamHI restriction endonuclease site is underlined, and the central complementary region is indicated in italics. OEP was carried out with these as the central primers and OEP-1 and OEP-2 (see above) as the external primers. The amplified 1.6-kbp OEP fragment was then A-tailed by incubation with dATP and Taq polymerase at 72°C for 10 min and cloned into pGEM-TEasy to create pADPW112. The sacB-Km cassette from pRMJ1 was then inserted into the BamHI restriction endonuclease site in pADPW112, creating pADPW114. pADPW114 was linearized and used to transform ADP1. Transformants with recombination in the sacB-Km cassette were selected on LB-kanamycin agar. The insertion of the sacB-Km cassette was confirmed by Southern blot analysis (data not shown). Transformants were then tested for sensitivity to sucrose by being patched onto LB-kanamycin agar plates containing sucrose. Whereas patches incubated at 37 or 30°C grew readily in the presence of sucrose, patches incubated at ambient temperatures (circa 16°C) did grow but very slowly and only after 5 days of incubation; by comparison, the wild type ADP1 readily grew to a large patch during the same period. Sucrose toxicity in ADP1 is thus temperature dependent, and ADP1::sacB-Km transformants were deemed sucrose sensitive only at ambient temperatures. A single typical transformant was maintained as ADPW112, and this transformant was used for the construction of all designed mutants.
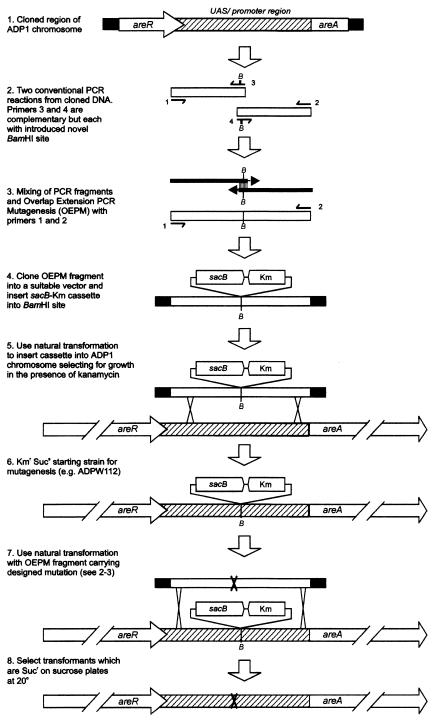
FIG. 3.
Generalized method of constructing site-directed mutations within the chromosome of Acinetobacter strain ADP1 by using a combination of OEP mutagenesis and marker exchange-eviction of the sacB-Km cassette originating from pRMJ1. The half arrows in panels 2 and 3 represent PCR primers 1 to 4. B represents an introduced BamHI site, which could also be a PstI or SalI site for the method to work (Fig. 2). In other situations, steps 1 to 3 would be unnecessary if there was a native BamHI, PstI, or SalI site in the vicinity where the mutations were to be introduced.
(ii) Marker exchange-eviction of the sacB-Km cassette from ADPW112.
The sacB-Km cassette from ADPW112 was evicted by marker exchange. The protocol was tested by transforming ADPW112 with pADPW112 (containing the novel BamHI site in the putative UAS region). Transformation mixtures of ADPW112 and linearized pADPW112 were grown overnight at ambient temperatures and then plated onto LB agar containing sucrose and incubated at ambient temperatures. Single colonies of transformants appeared after 48 h of incubation; a single transformant was designated ADPW114. No transformants were obtained when the same transformation was undertaken at 37 or 30°C. The transformants which were no longer sucrose sensitive were found to have lost kanamycin resistance. The adjacent DNA was amplified by PCR from genomic DNA from a representative transformant by using primers OEP-1 and OEP-2. A 1.6-kbp product was amplified that, on digestion with BamHI restriction endonuclease, showed the presence of two bands of 0.75 and 0.85 kbp, proving the presence of the created BamHI restriction site in ADPW114 by comparison with the product of PCR amplification of the same region in wild type ADP1, which failed to yield two fragments following BamHI digestion (data not shown).
(iii) Creation of ADP1 mutants with deletions or substitutions in the UAS.
ADP1 mutants with designed deletions or substitutions in the UAS were created essentially by the method described above but with different complementary primer pairs spanning the UAS region and carrying the designed mutation within each primer sequence (Table 2). The same flanking primers, OEP-1 and OEP-2, were used for all amplifications, and pADPW47 (cloned wild-type ADP1 DNA) (Table 1) was used as template DNA. The amplified OEP fragments were A-tailed and cloned into pGEMeasy, and each plasmid was linearized and used to transform ADPW112, where the eviction of the sacB marker gene was monitored by the selection on LB-sucrose plates of sucrose-resistant, kanamycin-sensitive transformants. The successful creation of the designed mutation was confirmed by sequencing a PCR fragment amplified from the genomic DNA of a transformant colony. Once the successful integration of the mutated fragment was confirmed, the mutant was transformed with a reporter lacZ gene. This was done by linearizing pADPW38 (areA::lacZ-Km), using it for transformation, and selecting for growth in the presence of kanamycin, thus creating a double mutant with a mutation in the UAS and a reporter lacZ gene in areA.
The effect of the mutation on are gene expression was tested (i) by using biochemical assays of AreB (benzyl alcohol dehydrogenase) activity of the primary mutant and (ii) by measuring β-galactosidase activity in the corresponding areA::lacZ derivative strain.
Structure of IR sequences at the UAS.
The UAS region has two IRs which overlap (Fig. 1). One is a near-perfect IR (16 of 17 bp) covering bases −105 to −139 from the +1 transcription start site and centered on base −122. This IR is referred to as IR1 (comprising IR1L and IR1R).
The other (IR2) stretches from −79 to −132 of the Pare UAS region. Each arm (IR2L and IR2R) consists of two stretches which are perfect 9- and 7-bp repeats and are separated by a 3-bp mismatch. The three mismatched bases in IR2L, ATT (bases −121 to −123), are centered between the two repeats of IR1 and would lie at the apex of any hairpin loop structure formed by IR1 (Fig. 1).
The central bases of the inner 7-bp repeats of IR2 (bases −94 and −117) are separated by 22 bp, and the central bases of the two 9-bp arms of IR2 (bases −83 and −128) are separated by 44 bp (Fig. 1). This is characteristic of regions shown to be involved in regulator binding in other σ54-dependent regulatory systems in which the IRs are separated by complete turns of the DNA helix, i.e., multiples of 11 bp (26).
Deletion mutations of the Pare UAS region.
Three mutants were created with major deletions in the UAS region. The deletions in mutants ADPW151 and ADPW152 were both 26 bp, that in mutant ADPW153 was 19 bp, and in all three cases the deletions were located in IR1 (Table 1; Fig. 3). Each mutation deleted nearly all of IR1 and the whole of IR2L (Fig. 1). Expression of the are genes was reduced by 85 to 90% by each mutation.
A further series of five mutants with deletions in IR2R was created. The mutants (ADPW164, ADPW165, ADPW197, ADPW198, and ADPW199) contained 3-, 6-, 9-, 10-, and 11-bp deletions, respectively, that were centered on base −94 at the center of the 7-bp arm of IR2R (Fig. 4A). Whereas the shorter 3- and 6-bp deletions caused major (90 to 98%) reductions in are gene expression, the longer the deletion the less was its effect: the 9- and 10-bp deletions caused ∼80% loss of expression, whereas the 11-bp deletion in ADPW199, which constituted one complete turn of the double helix, caused only a ∼50% loss of expression.
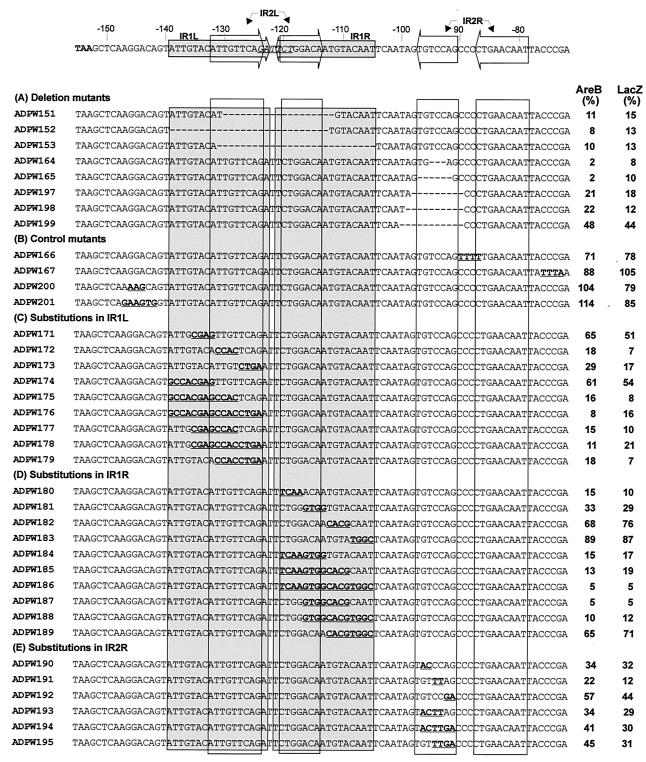
FIG. 4.
areCBA expression in mutants constructed with mutations in the putative UAS of the areCBA operon of Acinetobacter strain ADP1. The nucleotide sequence of the wild type and the positions of the two sets of IRs, IR1 and IR2, are shown at the top and the latter are boxed in the lower sequences. The corresponding sequence of each mutant is presented below: deletions are indicated by dashes, and substitutions are indicated by bold, underlined bases. The are gene expression of each mutant was determined by biochemical assays of AreB (benzyl alcohol dehydrogenase) activity, where the specific activity of the wild type ADP1 (0.33 μmol min−1 mg−1) was taken as 100% and the specific activities of mutants were recorded as percentages of the wild-type value. Expression was also measured by using β-galactosidase activities of double mutants carrying both the designed mutation in the UAS and the insertion of a promoterless lacZ in areA. Values were calculated in Miller units (19) and are presented as percentages of the wild-type activity (1,150 Miller U) measured in ADPW63, carrying only the areA::lacZ insert.
Control mutants.
Four control mutants were constructed (Fig. 4B): ADPW167, with a 4-bp substitution to the right of the entire IR region; ADPW200 (3 bp) and ADPW201 (6 bp), with substitutions to the left of IR1L; and ADPW166, with substitutions between the two repeats of IR2L. In none of these mutants was expression significantly lowered from that of the wild type.
Substitution mutations in IR1L.
Ten substitution mutants, designated ADPW171 to ADPW179, were created with substitutions in IR1L. The bases were systematically replaced in 4-, 8-, 12-, and 16-bp segments (Fig. 4C). In all cases, the substitutions were such that CG was replaced by AT and vice versa, thus destroying the degree of complementarity of the IRs. All of the mutations also affected the 9-bp arm of IR2L, except that in ADPW171 and ADPW174 only the left base of this arm was altered.
Replacing the inner bases of IR1, which includes the 9-bp arm of IR2L, reduced expression by up to 90%. However, replacing the outer bases of the left arm of this IR (in ADPW171 and ADPW174) had no such drastic effect on induction, reducing activity by only around 50%.
Substitution mutations in IR1R.
A similar set of mutants (ADPW180 to ADPW189) (Fig. 4D) was created with substitutions in IR1R. As with IR1L substitution mutants, replacing the inner eight bases of the right arm of IR1, which includes the 7-bp arm of IR2L, reduced expression severely by up to 95% (in ADPW180, ADPW181, ADPW184, ADPW185, ADPW186, ADPW187, and ADPW188). By contrast, replacing the outer bases of IR1R, which does not change the 7-bp arm of IR2L, had a much less drastic effect on induction, reducing activity by only around 10 to 30% (in ADPW182, ADPW183, and ADPW189).
Substitution mutations in the 7-bp arm of IR2R.
The final series of mutants (ADPW190 to ADPW195) was created with substitutions in the right arm of IR2 (Fig. 4E). The various mutations in this sequence of bases reduced gene expression by 50 to 80%. The greatest effect was from the replacement of 6 or 7 bp, which caused an 80% loss in enzyme induction, whereas replacing only 2 or 4 bp resulted in 50 to 70% loss of enzyme activity. In ADPW166 (Fig. 4B), the region between the two arms of IR2R was changed from CCCC to TTTT and this change had only a marginal effect upon expression (∼20%).
DISCUSSION
Acinetobacter strain ADP1 is an important model organism for studying the biochemistry, genetics, and molecular biology of degradation of natural products (8, 21). Its ability to undergo natural transformation has facilitated these investigations by enabling a broad range of experimentally simple techniques to be developed. Examples of these include the insertion of cassettes into specific genes (5, 24), gap repair to extract specific chromosomal segments into recombinant plasmids (2), random insertional mutagenesis (30), generation by PCR of random mutations in specific Acinetobacter genes (15) or in heterologous genes located in the ADP1 chromosome (16), and the identification of DNA complementing unknown mutations (32). To add to this arsenal, we have developed the first procedure for producing specific designed mutations within any ADP1 chromosomal gene. Starting from a single strain into which we inserted by natural transformation a sacB-Km cassette within the region to be mutated, we have generated 38 different deletion or substitution mutations within a span of about 50 bp on either side of the inserted cassette. All mutations were constructed on fragments generated by PCR by using two-step OEP with the same two external primers but with different pairs of overlapping internal primers each containing the designed mutation. In these studies, the chromosomal distance between the points to which the two external primers were homologous was chosen as 1.6 kb. Fragments thus generated were used to transform the strain carrying the sacB-Km cassette, and recombinants were selected in which the 3.7-kb sacB-Km cassette was lost. In this study, we aimed to ensure a high frequency of the desired transformants by using a fragment with long overlaps on either side of the site of the designed mutation as the OEP-generated amplicon. Further study will be necessary to ascertain how close the designed mutation can be to the end of the transforming DNA or how far from the inserted cassette to get an adequate frequency of transformants in which both the cassette has been replaced and the mutation has been incorporated. The only surprise we encountered in developing the method was that strains of ADP1 that contained the sacB-Km cassette grew perfectly adequately in the presence of the normally toxic 5% sucrose at both 38°C, the temperature at which ADP1 is normally cultured in the laboratory, and 30°C. In ADP1, sucrose toxicity is temperature sensitive and the lethality of sacB manifested itself only at room temperature; consequently, selection for eviction of the cassette had to be carried out at this lower temperature.
We have used this methodology to investigate putative regulatory DNA upstream of the catabolic are operon by a process of systematic mutagenesis. Whereas it could easily be used for mutations within structural genes, a particular advantage of this procedure for studying regulation is that the mutations are introduced into the single chromosomal copy of the gene and thereby overcome any problems due to copy number associated with using plasmids as a vector.
Transcription of the areCBA operon from the −24, −12 σ54-dependent promoter is strongly regulated, with areCBA expression measurable only in the presence of pathway substrates (11, 12). The sequence upstream of the −24, −12 promoter contains a 43-bp AT-rich region that is 98% AT (Fig. 1), abnormal even for an AT-rich bacterium such as Acinetobacter. This region separates the −24, −12 promoter from an upstream complex of IR sequences. This follows other σ54-dependent promoter systems (26) which have a UAS located 100 to 200 bp upstream of the −24, −12 region and separated from it by DNA which can form a loop to bring the UAS into physical proximity with the polymerase-σ54 complex. This DNA has been shown to contain IHF binding sites (28) which bring about the loop formation. The AT-rich region upstream of the are promoter consists of 42 AT pairs containing a single GC pair at position −42 from the start of transcription (Fig. 1). The sequence of bases containing the lone C (5′-AAAAACAATAATTT at −35 to −46) is consistent with the consensus sequence for IHF binding sites (http://www.bmb.psu.edu/seqscan)and is virtually identical to the identified binding region for IHF (IHF2; 5′-AAAAACAATACCTT) upstream of the σ54-dependent promoter system of the phenol-catabolic dmp operon (28).
The UAS of σ54-dependent promoters are normally arranged as IR sequences. In the putative are UAS there are two overlapping IR sequences, IR1 and IR2. Unlike most NtrC-like regulators, the areR gene is upstream of and transcribed in the same direction as its regulated genes (11), whereas the norm is divergent transcription. The experimental problem in attempting to assign separate roles for the two IR regions is that the left arm of IR2 (IR2L) sits centrally over both arms of IR1 and most mutations will affect both IRs simultaneously.
The results taken as a whole show that the 60-bp region is clearly involved in the regulation of areCBA expression. Both deletions and substitutions reduced are gene expression by up to 90%, whereas control substitutions just outside of the region had no effect upon expression. Deletions in IR1 that also disrupted IR2L sequences drastically reduced activity (Fig. 4A) but could not distinguish the relative importances of the two IRs. Deletions in IR2R (Fig. 4A) also caused loss in areCBA expression of up to 90%. Again, this does not assign the function of UAS to that particular region since it is known that deletions other than complete turns of the helix (multiples of 11 bp) between the UAS and the σ54-dependent promoter have a deleterious effect on gene expression (20). A 3- or 6-bp deletion effected the greatest reduction of expression, whereas deletions closer to one complete turn of the helix had less of an effect. The mutant ADPW199 with an 11-bp deletion retained ∼50% activity compared to the wild type, indicating that transcription is still activated despite the loss of one complete arm of IR2.
Substitutions in the region where IR1L and the 9-bp arm of IR2L overlap showed that only the bases which were also part of IR2L had a substantial effect on gene expression (Fig. 4C). Substitutions at the left end of IR1L, with little overlap with IR2L, although equally affecting the mismatch between IR1L and IR1R, decreased expression much less than similar substitutions that overlapped significantly with the 9-bp arm of IR2L.
Similarly, substitutions in IR1R (Fig. 4D) generally reduced expression more when they overlapped the 7-bp arm of IR2L than when they were at the outside end of IR1R. These results indicate that the central region of IR1, which also corresponds to IR2L, is more critical in determining expression, either when deleted or when replaced to reduce the complementarity of the two repeats, than are the two outer ends of IR1. By contrast, all the substitutions in the 7-bp arm of IR2R (Fig. 4E) have a relatively minor effect and it is apparent that transcriptional expression is much more tolerant of substitutions in this region.
The overall emphasis of the results indicate that the DNA of IR1 and IR2L carries the major role in determining the regulation of the are operon. It also implies that IR2 is probably a much less critical factor than IR1 in determining expression: all mutations in IR2L clearly have a severe effect, but the absence of reciprocal effects from similar mutations in IR2R would indicate that the mutations in IR2L are effective because they are also in IR1. If this implies that the repeat sequence of IR1 is the site where AreR binds, then it also follows from the data that there must be a graded reduction of critical contacts from the center to the periphery of the repeat.
It is unwise to conclude from the data presented that either of the IR elements has a direct role in terminating areR transcription. The results do not eliminate this as a possibility, but it would seem that a more convincing model would be that AreR, binding in the region of IR1, would act as a regulator of its own transcription.
Deletions within IR2R (Fig. 4A) do have a major effect on expression, but the correlation between the effect and the length of the deletion relative to the turn of the helix suggests that the explanation is due to a topological effect on the interaction of AreR bound to the UAS with the downstream −24, −12 promoter following loop formation rather than to the removal of an arm of the repeat.
A mutational study such as this cannot answer all the questions about the importance of specific residues in determining regulation. Analysis of DNA footprinting and gel shift assay studies of AreR on the DNA should lead to physical information on the protein-DNA contacts. This would then enable the design of mutants on a slightly more focused basis than we were able to use here, which should enable a tight correlation to be made between the physical binding data and the effect upon gene expression.
Acknowledgments
This research was funded by a BBSRC research grant (5/P12776).
We thank Victor de Lorenzo for advanced viewing of his review article.
REFERENCES
- 1.Austin, S., and R. Dixon. 1992. The prokaryotic enhancer binding protein NtrC has an ATPase activity which is phosphorylation and DNA dependent. EMBO J. 11:2219-2228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Averhoff, B., L. Gregg-Jolly, D. Elsemore, and L. N. Ornston. 1992. Genetic analysis of supraoperonic clustering by use of natural transformation in Acinetobacter calcoaceticus. J. Bacteriol. 174:200-204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bauchop, T., and S. R. Elsden. 1960. The growth of microorganisms in relation to energy supply. J. Gen. Microbiol. 23:457-469. [DOI] [PubMed] [Google Scholar]
- 4.Carmona, M., and B. Magasanik. 1996. Activation of transcription at σ54-dependent promoters on linear templates requires intrinsic or induced bending of the DNA. J. Mol. Biol. 261:348-356. [DOI] [PubMed] [Google Scholar]
- 5.DiMarco, A., and L. N. Ornston. 1994. Regulation of p-hydroxybenzoate hydroxylase synthesis by PobR bound to an operator in Acinetobacter calcoaceticus. J. Bacteriol. 176:4277-4284. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Elsmore, D. A., and L. N. Ornston. 1995. Unusual ancestry of dehydratases associated with quinate catabolism in Acinetobacter calcoaceticus. J. Bacteriol. 177:5971-5978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Gay, P., D. Le Coq, M. Steinmetz, T. Berkelman, and C. I. Kado. 1985. Positive selection procedure for entrapment of insertion sequence elements in gram-negative bacteria. J. Bacteriol. 164:918-921. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Harwood, C. S., and R. E. Parales. 1996. The β-ketoadipate pathway and the biology of self-identity. Annu. Rev. Microbiol. 50:553-590. [DOI] [PubMed] [Google Scholar]
- 9.Ho, S. N., H. D. Hunt, R. M. Horton, J. K. Pullen, and L. R. Pease. 1989. Site-directed mutagenesis by overlap extension using the polymerase chain reaction. Gene 77:51-59. [DOI] [PubMed] [Google Scholar]
- 10.Hoover, T. R., E. Santero, S. Porter, and S. Kustu. 1990. The integration host factor stimulates interaction of RNA polymerase with NifA, the transcriptional activator for nitrogen fixation operons. Cell 63:11-22. [DOI] [PubMed] [Google Scholar]
- 11.Jones, R. M., L. S. Collier, E. L. Neidle, and P. A. Williams. 1999. areABC genes determine the catabolism of aryl esters in Acinetobacter sp. strain ADP1. J. Bacteriol. 181:4568-4575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Jones, R. M., and P. A. Williams. 2001. areCBA is an operon in Acinetobacter sp. strain ADP1 and is controlled by AreR, a σ54-dependent regulator. J. Bacteriol. 183:405-409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Juni, E. 1972. Interspecies transformation of Acinetobacter: genetic evidence for a ubiquitous genus. J. Bacteriol. 112:917-931. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Kaniga, K., I. Delor, and G. R. Cornelis. 1991. A wide-host-range suicide vector for improving reverse genetics in Gram-negative bacteria: inactivation of the blaA gene of Yersinia enterocolitica. Gene 109:137-141. [DOI] [PubMed] [Google Scholar]
- 15.Kok, R. G., D. A. D'Argenio, and L. N. Ornston. 1997. Combining localized PCR mutagenesis and natural transformation in direct genetic analysis of a transcriptional regulator gene, pobR. J. Bacteriol. 179:4270-4276. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kok, R. G., D. A. D'Argenio, and L. N. Ornston. 1998. Mutation analysis of PobR and PcaU, closely related transcriptional activators in Acinetobacter. J. Bacteriol. 180:5058-5069. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Kokotek, W., and W. Lotz. 1989. Construction of a lacZ-kanamycin-resistance cassette, useful for site-directed mutagenesis and as a promoter probe. Gene 84:467-471. [DOI] [PubMed] [Google Scholar]
- 18.Kustu, S., E. Santero, J. Keener, D. Popham, and D. Weiss. 1989. Expression of σ54 (ntrA)-dependent genes is probably united by a common mechanism. Microbiol. Rev. 53:367-376. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Miller, J. H. 1972. Experiments in molecular genetics. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y.
- 20.Morett, E., and L. Segovia. 1993. The σ54 bacterial enhancer-binding protein family: mechanism of action and phylogenetic relationship of their functional domains. J. Bacteriol. 175:6067-6074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Parke, D., D. A. D'Argenio, and L. N. Ornston. 2000. Bacteria are not what they eat: that is why they are so diverse. J. Bacteriol. 182:257-263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Perez-Martin, J., and V. de Lorenzo. 1996. ATP binding to the sigma 54-dependent activator XylR triggers a protein multimerization cycle catalyzed by UAS DNA. Cell 86:331-339. [DOI] [PubMed] [Google Scholar]
- 23.Reid, J. L., and A. Collmer. 1987. An nptI-sacB-sacR cartridge for constructing directed, unmarked mutations in Gram-negative bacteria by marker exchange-eviction mutagenesis. Gene 57:239-246. [DOI] [PubMed] [Google Scholar]
- 24.Romero-Arroya, C. E., M. A. Schell, G. L. Gaines III, and E. L. Neidle. 1995. catM encodes a LysR-type transcriptional activator regulating catechol degradation in Acinetobacter calcoaceticus. J. Bacteriol. 177:5891-5898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Sambrook, J., E. F. Fritsch, and T. Maniatis. 1989. Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
- 26.Shingler, V. 1996. Signal sensing by sigma54-dependent regulators: derepression as a control mechanism. Mol. Microbiol. 19:409-416. [DOI] [PubMed] [Google Scholar]
- 27.Simon, R., B. Hötte, B. Klauke, and B. Kosier. 1991. Isolation and characterization of insertion sequence elements from gram-negative bacteria by using new broad-host-range, positive selection vectors. J. Bacteriol. 173:1502-1508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Sze, C. C., A. D. Laurie, and V. Shingler. 2001. In vivo and in vitro effects of integration host factor at the DmpR-regulated σ54-dependent Po promoter. J. Bacteriol. 183:2842-2851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Valls, M., and V. de Lorenzo. Transcription mediated by rpoN-dependent promoters: the σ54 sigmulon. In J.-M. Ramos (ed.), The pseudomonads, vol. II, in press. Kluwer, Amsterdam, The Netherlands.
- 30.Vosman, B., R. Kok, and K. J. Hellingwerth. 1991. Random insertional mutagenesis in Acinetobacter, p. 183-189. In K. J. Towner, E. Bergogne-Berezin, and C. A. Fewson (ed.), The biology of Acinetobacter. FEMS Symposia Series. Plenum Press, New York, N.Y.
- 31.Weiss, D. S., J. Batut, K. E. Klose, J. Keener, and S. Kustu. 1991. The phosphorylated form of the enhancer-binding protein NtrC has an ATPase activity that is essential for activation of transcription. Cell 67:155-167. [DOI] [PubMed] [Google Scholar]
- 32.Williams, P. A., and L. E. Shaw. 1997. mucK, a gene in Acinetobacter calcoaceticus ADP1 (BD413), encodes the ability to grow on exogenous cis,cis-muconate as the sole carbon source. J. Bacteriol. 179:5935-5942. [DOI] [PMC free article] [PubMed] [Google Scholar]