Abstract
Phenotypic characterization of the Arabidopsis thaliana transparent testa12 (tt12) mutant encoding a membrane protein of the multidrug and toxic efflux transporter family, suggested that TT12 is involved in the vacuolar accumulation of proanthocyanidin precursors in the seed. Metabolite analysis in tt12 seeds reveals an absence of flavan-3-ols and proanthocyanidins together with a reduction of the major flavonol quercetin-3-O-rhamnoside. The TT12 promoter is active in cells synthesizing proanthocyanidins. Using translational fusions between TT12 and green fluorescent protein, it is demonstrated that this transporter localizes to the tonoplast. Yeast vesicles expressing TT12 can transport the anthocyanin cyanidin-3-O-glucoside in the presence of MgATP but not the aglycones cyanidin and epicatechin. Inhibitor studies demonstrate that TT12 acts in vitro as a cyanidin-3-O-glucoside/H+-antiporter. TT12 does not transport glycosylated flavonols and procyanidin dimers, and a direct transport activity for catechin-3-O-glucoside, a glucosylated flavan-3-ol, was not detectable. However, catechin-3-O-glucoside inhibited TT12-mediated transport of cyanidin-3-O-glucoside in a dose-dependent manner, while flavan-3-ol aglycones and glycosylated flavonols had no effect on anthocyanin transport. It is proposed that TT12 transports glycosylated flavan-3-ols in vivo. Mutant banyuls (ban) seeds accumulate anthocyanins instead of proanthocyanidins, yet the ban tt12 double mutant exhibits reduced anthocyanin accumulation, which supports the transport data suggesting that TT12 mediates anthocyanin transport in vitro.
INTRODUCTION
Plants synthesize secondary metabolites for a multitude of biological functions, including UV protection or insect attraction via flower coloration. Since these functions are mostly dose dependent, secondary compounds accumulate constitutively or inducibly and often in a tissue- or cell-specific manner to concentrations starting in the millimolar range. At the same time, phenolic compounds are toxic to the cell due to their high chemical reactivity and protein denaturing properties (Matile, 1984). The major strategy of the plant to protect the cytosolic machinery against these toxic effects is the targeting of secondary metabolites in compartments with lower biosynthetic activity, notably the extracellular space and the large central vacuole. For many phenolic compounds, vacuolar accumulation is usually preceded by oxidation reactions mediated by cytochrome P450-dependent monoxygenases or dioxygenases and followed by conjugation to small hydrophilic (e.g., glycosyl) groups resulting in more water-soluble compounds (Coleman et al., 1997).
Flavonoids representing one class of phenolic compounds possess diverse chemical structures but share a common C6-C3-C6 body. Major classes are the monomeric red to purple anthocyanins in flowers and fruits that often form color-shifted copigments with UV-absorbing colorless to pale yellow flavonols or flavones (Harborne and Williams, 2000) and condensed tannins (proanthocyanidins [PAs]), which are colorless polymers of flavan-3-ols (e.g., epicatechin) that become brown after oxidation. Flavonoids appeared in the first land plants during evolution. In angiosperms, flavonoids are associated with sexual reproduction via pollination, seed dispersal, germination, and longevity (Debeaujon et al., 2000). In Arabidopsis thaliana, anthocyanins are found in sporophytic tissues but not in seeds, while the accumulation of PAs is restricted to PA-producing cells of the seed coat, and flavonols can be found in all plant parts (Shirley et al., 1995; Devic et al., 1999; Debeaujon et al., 2003; Routaboul et al., 2006). A histochemical analysis of developing Arabidopsis seeds suggested that PA accumulation is found in three different cell types: (1) in the innermost layer of the inner integument (ii1 layer or endothelium), (2) in a few cells of the ii2 layer at the micropyle, and (3) in cells forming the pigment strand at the chalaza (Debeaujon et al., 2003). Mutants deviating from the normal brown seed color arising from the oxidation of the colorless PAs at maturity have been termed transparent testa (tt), transparent testa glabra (ttg), tannin deficient seeds (tds), banyuls (ban), and aha10 (Lepiniec et al., 2006). Figure 1 displays a partial model of the flavonoid biosynthetic pathway in Arabidopsis. The BAN gene has recently been described to encode anthocyanidin reductase separating the anthocyanin and flavan-3-ol-derived (PA) side branches of the flavonoid pathway (Xie et al., 2003). It is essentially active in PA-producing cells of the seed coat (Debeaujon et al., 2003). Another gene, TT10, which encodes a laccase involved in oxidative flavonoid polymerization in late stages of seed development, is expressed mainly in PA-producing cells and the outer integument (Pourcel et al., 2005). By contrast, several mutants, such as tt4, tt5, tt7, and tt18/tds4, exhibit changes in overall flavonoid composition in vegetative tissues apart from affecting PA biosynthesis in the seed coat (Lepiniec et al., 2006).
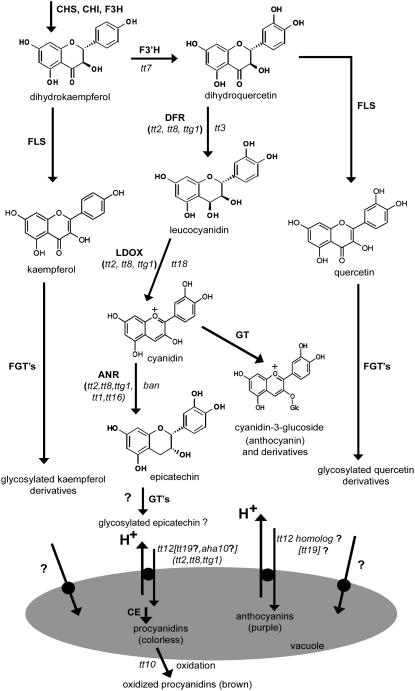
Figure 1.
Simplified Scheme of the Biosynthetic Pathway in Arabidopsis Leading to PAs, Flavonols, and Anthocyanins, Including Proposed Vacuolar Transport Steps.
Dihydrokaempferol is synthesized from 4-coumaroyl CoA and three molecules of malonyl CoA by the sequential action of chalcone synthase (CHS), chalcone isomerase (CHI), and flavanone 3-hydroxylase (F3H). Enzymes are represented in uppercase bold letters. Corresponding genetic mutations are in lowercase italic letters. Regulatory mutants are indicated by parentheses. AHA10, P-type H+-ATPase; ANR, anthocyanidin reductase; CE, condensing enzyme; DFR, dihydroflavonol reductase; F3′H, flavonol 3′-hydroxylase; FLS, flavonol synthase; (F)GT, (flavonol) glycosyltransferase; LDOX, leucoanthocyanidin dioxygenase.
With regard to secondary metabolites, biochemical analysis of vacuolar uptake mechanisms has led to the proposition of different transport and accumulation systems (Grotewold, 2004; Yazaki, 2005). Available data suggest that (1) vacuolar accumulation can involve trapping mechanisms, such as trapping of conformational isomers; (2) the conjugated moieties, such as sugar or acyl residues, are important determinants of transport specificity; and (3) plants possess specific transporters for their endogenously synthesized compounds as opposed to xenobiotic compounds possessing comparable conjugation patterns. Different transport mechanisms exist depending on the plant species chosen: while barley (Hordeum vulgare) uses a H+-antiport mechanism for the vacuolar uptake of endogenous glucosylated flavones, Arabidopsis, which does not synthesize flavones, uses an ATP binding cassette (ABC)–type directly energized vacuolar transport when supplied with barley flavones (Klein et al., 1996). In accordance with this latter observation, Zea mays plants with a reduced amount of transcript of the vacuolar ABC transporter Zm MRP3 exhibit reduced levels of anthocyanin accumulation in different plant parts (Goodman et al., 2004).
Vacuolar compartmentation and associated transport steps are important for proper flavonoid accumulation. Furthermore, in H. vulgare mesophyll cells, vacuolar flavonoid transport is directly controlled by the presence of the intact flavonoid biosynthetic pathway, suggesting that a biosynthetic intermediate posttranslationally activates the flavonoid transporter (Marinova et al., 2007). However, the molecular identity of the membrane proteins involved in flavonoid transport is just starting to be uncovered. A T-DNA insertion in the TT12 gene (At3G59030 or At DTX41) identified the first transporter-like membrane protein involved in PA biosynthesis as tt12 seeds lack accumulation of the flavan-3-ol epicatechin and PAs in vacuoles of the seed coat endothelium (Debeaujon et al., 2001). TT12 encodes a membrane protein with 12 predicted transmembrane helices and belongs to the multidrug and toxic compound extrusion (MATE) family (Brown et al., 1999). MATE proteins in prokaryotes, as exemplified by the NorM multidrug resistance protein from Vibrio parahaemolyticus and its homolog in Escherichia. coli, YdhE, mediate resistance against diverse drugs, including fluoroquinolone antibiotics (Morita et al., 1998). Functionally, NorM has been described as a Na+-driven multidrug efflux pump (Morita et al., 2000). MATE1, one of two orthologous genes in humans, encodes a transporter-mediating H+-coupled cellular efflux of toxic organic cations, such as tetraethylammonium, and thus represents the polyspecific organic cation exporter that transports toxic organic cations into urine and bile (Otsuka et al., 2005). In Arabidopsis, the MATE family consists of 56 genes (Li et al., 2002; Rogers and Guerinot, 2002). Apart from TT12, four MATE genes have been initially characterized by mutant analysis. They are involved in lateral root formation (ALF5/At DTX19; Diener et al., 2001), iron homeostasis (FRD3/At DTX43; Rogers and Guerinot, 2002), or disease resistance (EDS5/At DTX47; Nawrath et al., 2002). Complementation of the norfloxacin-hypersensitive E. coli strain KAM3, lacking the multidrug efflux pump AcrAB, resulted in functional cloning of At DTX1 (Li et al., 2002). Notably, recent evidence suggests that FRD3 is responsible for citrate secretion into the xylem and the rhizosphere, which extends the spectrum of potential substrates of MATE transporters from toxic substances or secondary compounds also to primary metabolites (Durrett et al., 2007).
Here, we identify the TT12 MATE transporter as a tonoplastic membrane protein with a clear biochemical function in the vacuolar deposition of flavonoids in the PA-synthesizing cells of the Arabidopsis seed coat. The substrate specificity of TT12 suggests that it mediates the transport of monomeric glycosylated flavan-3-ols as PA precursors in vivo, while it is able to transport anthocyanins in vitro as long as they are glycosylated. Mechanistically, TT12 acts as a proton antiporter.
RESULTS
Seeds of tt12 Have an Altered Flavonoid Metabolite Composition
It was previously demonstrated that tt12 seeds exhibit a defect in the vacuolar accumulation of PAs in immature seeds (Debeaujon et al., 2001). Apart from PAs, other flavonoid compounds, including flavonols, are synthesized during seed development in a cell type–specific manner (Pourcel et al., 2005; Kerhoas et al., 2006; Routaboul et al., 2006). To investigate the changes in the flavonoid composition in seeds caused by the absence of the TT12 MATE transporter, we performed a targeted metabolite analysis of mature wild-type and tt12 seeds by liquid chromatography–mass spectrometry (LC-MS).
In contrast with the wild type, neither the PA precursor epicatechin nor PA polymers (dimers to heptamers) were detected in tt12 seeds (Figure 2B). Acid-catalyzed hydrolysis of flavonoid extract and pellet remaining after extraction demonstrated that tt12 mutant seeds contained a small amount of PAs in the insoluble fraction (∼15% of the wild-type PAs) but no PAs in the soluble fraction (Figure 2C). Thus, absence of the transport activity mediated by TT12 ultimately leads to absence of epicatechin and its polymers in tt12 seeds.
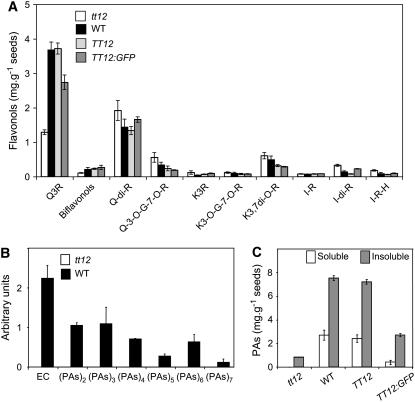
Figure 2.
Flavonoid Composition of Mature Seeds.
(A) Flavonol composition of seeds from the tt12 mutant, the wild type, TT12 overexpressor (TT12), and GFP5:TT12 overexpressor. Values represent the average and se of six independent measurements. Q-di-R, quercetin-3,7-di-O-rhamnoside; G, glucoside; H, hexoside; I, isorhamnetin; K, kaempferol; Q, quercetin; R, rhamnoside.
(B) Detection of epicatechin and soluble PAs (dimers to heptamers) by LC-MS. EC, epicatechin.
(C) Analysis of soluble and insoluble PAs measured after acid-catalyzed hydrolysis.
For (B) and (C), values represent the average and se of three independent measurements.
Wild-type seeds contained glycosylated quercetin and kaempferol derivatives, with quercetin-3-O-rhamnoside (Q3R) being quantitatively the most prominent compound, followed by quercetin-3,7-di-O-rhamnoside and kaempferol-3,7-di-O-rhamnoside (Figure 2A). In tt12, the amount of Q3R and biflavonols (dimers of Q3R) was reduced to ∼30 and 50% of wild-type levels, respectively, while the quantities of all other flavonols were not significantly affected by the tt12 mutation.
The TT12 Promoter Is Active Only in PA-Accumulating Cells
Previous in situ hybridization studies detected TT12 mRNA in the endothelium at ∼3 to 4 d after flowering (DAF) (Debeaujon et al., 2001). To specify the spatio-temporal expression pattern of TT12 during seed development, we analyzed the TT12 promoter activity using a 1.8-kb genomic fragment corresponding to the putative TT12 promoter region (ProTT12) together with the 5′ untranslated region fused to the uidA reporter gene. β-Glucuronidase (GUS) activity was analyzed in seeds at different stages of development. Faint GUS activity was restricted to the inner integument close to the micropyle on the day of flowering (Figure 3A). ProTT12:uidA expression progressed from the micropyle within the ii1 layer (2 DAF; Figure 3B) until GUS activity was observed in all cells of the ii1 layer and in cells of the chalaza from 3 DAF on (Figures 3C to 3E and 3G). Sectioning of immature transgenic ProTT12:uidA seeds 3 DAF confirmed TT12 expression in the ii1 layer of the seed coat, in chalazal cells, and in ii2 cells at the micropylar end (Figure 3G). At 6 DAF, GUS activity decreased in all cell types (Figure 3F). Taken together, TT12 is expressed specifically in cells actively synthesizing PAs in developing seeds, and TT12 expression correlates with the expression of BAN (Debeaujon et al., 2003).
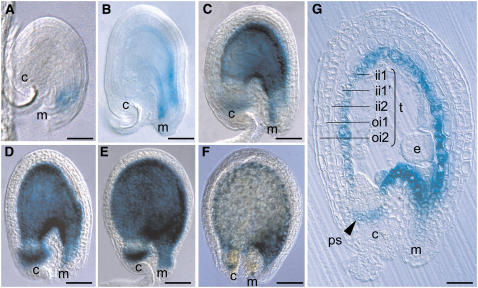
Figure 3.
Activity of the TT12 Promoter in Wild-Type Developing Seeds of Arabidopsis.
The expression of the ProTT12:uidA cassette was exclusively observed in developing seeds at 0 DAF (A), 2 DAF (B), 3 DAF ([C] and [G]), 4 DAF (D), 5 DAF (E), and 6 DAF (F). GUS activity was observed with Nomarski optics on whole mounts for (A) to (F) and on section for (G). The ii1 layer is also called endothelium. c, chalaza; e, embryo; ii, inner integument; m, micropyle; oi, outer integument; ps, pigment strand (chalazal tissue); t, testa. Bar = 25 μm in (A), 40 μm in (B), 65 μm in (C), 70 μm in (D), 80 μm in (E) and (F), and 33 μm in (G).
TT12 Is Localized to the Tonoplast
To examine the subcellular localization of TT12, the full-length TT12 cDNA was fused to the coding sequence of GFP5, resulting in N- and a C-terminal fusion of green fluorescent protein (GFP) to TT12. Expression of both fusions was driven by the dual cauliflower mosaic virus 35S RNA promoter (Pro35Sdual:GFP5:TT12:t35S and Pro35Sdual:TT12:GFP5:t35S simplified as GFP-cTT12 and cTT12-GFP in Figure 4, respectively) and analyzed after both stable and transient expression in plants. While expression of the unmodified TT12 cDNA fully complemented the tt12 mutant phenotype (including returning Q3R and insoluble and soluble PAs to wild-type levels; Figure 2), the Pro35Sdual:GFP5:TT12:t35S construct restored flavonoid levels at ∼75, 37, and 20% of the wild-type amount of Q3R and insoluble and soluble PAs, respectively (Figure 2), suggesting that a significant portion of GFP-cTT12 protein reached its proper destination within the cell and was functional.
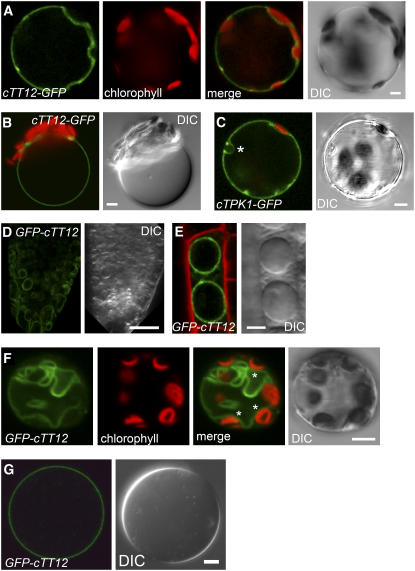
Figure 4.
Subcellular Localization of the TT12 Protein after Transient and Stable Expression.
(A) to (C) Mesophyll protoplasts isolated after transient expression of cTT12-GFP ([A] and [B]) or the tonoplast control cTPK1-GFP (C) by infiltration of N. benthamiana leaves with agrobacteria carrying the corresponding constructs. DIC, differential interference contrast. Bars = 5 μm.
(A) GFP and chlorophyll autofluorescence were false-colored in green and red, respectively. GFP fluorescence surrounds the plastids (see merged image).
(B) A vacuole released by gentle lysis exhibits GFP fluorescence on the tonoplast.
(C) Transient cTPK1-GFP expression is similar to cTT12-GFP, resulting in GFP fluorescence surrounding plastids and the nucleus (asterisk).
In (B) and (C), the green GFP and the red chlorophyll autofluorescence channels are merged.
(D) to (G) Confocal microscopy of Arabidopsis roots ([D] and [E]), mesophyll protoplasts (F), and isolated vacuoles (G) after stable transformation of GFP-cTT12 in the tt12 background.
(D) GFP fluorescence (green) of a root tip of a 7-d-old seedling. Bar = 20 μm.
(E) Merged image of GFP-cTT12 fluorescence (green) after propidium iodide counterstaining of cell walls (red). A single root cell possessing two large vacuoles exhibiting GFP-cTT12 fluorescence exclusively on the tonoplast membranes is shown. Bar = 5 μm.
(F) GFP-cTT12 fluorescence (green) is visible on an intracellular membrane surrounding the plastids exhibiting red chlorophyll autofluorescence (asterisks in the merged image) that delimits the vacuole(s) (DIC). Bar = 5 μm.
(G) Isolated leaf mesophyll vacuole prepared from stably transformed GFP-cTT12/tt12 plants exhibits GFP fluorescence on the tonoplast. Bar = 5 μm.
Transient expression by biolistic particle delivery of GFP-cTT12 together with the established tonoplast marker TPK1 (Czempinski et al., 2002) fused to DsRed2 in onion bulb epidermis cells demonstrated that both proteins coreside on membranes (see Supplemental Figure 1A online). Importantly, fluorescence of both fusion proteins is visible on strands traversing the vacuole. Transient expression of a translational GFP fusion of the Arabidopsis P31 protein (At3G01290), a protein containing prohibitin motives that was recently localized to the plasmalemma based on a plasma membrane–proteomic approach and verified by microscopy analysis of the corresponding GFP fusion (Marmagne et al., 2004), exhibited GFP fluorescence restricted to the cell periphery (see Supplemental Figure 1B online). In contrast with GFP-cTT12 and DsRed2-TPK1, P31-GFP-fluorescence was not detectable on transvacuolar strands, suggesting that TT12, in contrast with P31, does not reside on the plasmalemma.
The localization of TT12 was further investigated by transient expression of both TT12-GFP fusions in Nicothiana benthamiana leaves by infiltration of agrobacteria followed by isolation of mesophyll protoplasts (Figures 4A and 4B). The GFP signal obtained after expression of cTT12-GFP is restricted to a membrane that intracellularly surrounds the chloroplasts exhibiting red autofluorescence (Figure 4A). Gentle lysis of these mesophyll protoplasts demonstrates clearly that the GFP fluorescence surrounds the vacuole and thus localizes to the tonoplast (Figure 4B). This localization was identical when the N-terminal GFP fusion GFP-cTT12 was expressed, suggesting that the position of the GFP did not affect the targeting of the fusion protein (data not shown). As a control, TPK1 was separately expressed as a translational fusion with GFP fusion (Figure 4C). cTPK1-GFP fluorescence in N. benthamiana mesophyll protoplasts was restricted to the vacuolar membrane surrounding plastids and the nucleus, and its localization was therefore similar to TT12.
Confocal microscopy analysis of tt12 mutants ectopically expressing the GFP-cTT12 transgene exhibited membrane-bound GFP fluorescence in different tissues, such as roots or epidermis cells. Figure 4D illustrates GFP fluorescence of GFP-cTT12 on internal membranes of a root tip. Detailed inspection of single root cells in the apex of the primary root verified the presence of GFP-cTT12 on the tonoplast: GFP-fluorescence surrounded the vacuoles as seen after counterstaining of the cell walls with propidium iodide (red) (Figure 4E). When mesophyll protoplasts were isolated from these transgenic plants, GFP fluorescence of GFP-cTT12 was again visible on the intracellular membrane delimiting the vacuole and excluding the red autofluorescent chloroplasts (Figure 4F and Supplementary Figure 1C). Finally, tonoplast localization of GFP-cTT12 was confirmed by isolation of vacuoles starting from mesophyll protoplasts of transgenic Pro35Sdual:GFP5:TT12:t35S/tt12 plants. GFP fluorescence was detected on the membrane surrounding the vacuole (Figure 4G). Taken together, TT12 localizes to the tonoplast in plant cells.
TT12 Mediates Cyanidin-3-O-Glucoside Transport in Yeast Vesicles
To investigate the transport activity of TT12 biochemically, the full-length TT12 cDNA was expressed in Saccharomyces cerevisiae under the control of the strong constitutive PMA1 promoter. The presence of the TT12 protein and its subcellular localization was investigated in yeasts transformed with modified versions of TT12. As can be seen from the Supplemental Methods and Supplemental Figure 2 online, GFP fluorescence in yeasts cells transformed with GFP5-cTT12 was not restricted to the vacuolar membrane but was also present on other endomembranes and the plasma membrane. Based on this result, we decided to isolate total microsomal membrane vesicles rather than vacuolar membrane vesicles for transport experiments. Using N- and C-terminally His6-tagged versions of TT12, the presence of the TT12 proteins was verified in all independent yeast transformants tested (see Supplemental Figure 2B online). Finally, physiological intactness of the vesicle fractions isolated from yeasts that were either transformed with TT12 or with the empty vector control (NEV) was confirmed by the ability of the vesicle fractions to accumulate the weak base [14C]-methylamine in the presence of MgATP due to acidification of the vesicle lumen (see Supplemental Figure 2C and Supplemental Methods online). Since the uncharged methylamine base is membrane permeable but trapped inside the vesicles after protonation, this latter experiment also verifies the activity of the MgATP-dependent proton pumps regardless of the presence or absence of TT12.
Transport experiments with various structurally different flavonoids were performed with a standard substrate concentration of 1 mM using the rapid filtration technique (Tommasini et al., 1996). Since epicatechin represents the suggested immediate monomeric precursor of the PA polymers and since epicatechin is absent in tt12 seeds (Figure 2), we first tested the flavan-3-ols epicatechin and catechin. In the presence of MgATP, no time-dependent increase in the amounts of vesicle-associated flavan-3-ols was detectable either after photometric detection (see Supplemental Figure 3 online) or HPLC analysis (data not shown). These results suggested that under our experimental conditions, these substances are not transported by TT12.
Due to the absence of a leucoanthocyanidin reductase in Arabidopsis seeds, only epicatechin (cis-flavan-3-ol) and its derivatives are synthesized in this species (Abrahams et al., 2003; Routaboul et al., 2006). In our experiments, the PA precursor epicatechin was not modified by glycosylation. It has been shown in various experimental systems that sugar residues attached to the aglycone flavonoid core are important determinants of substrate recognition and transport activity. To test the influence of a sugar moiety, a related glucosylated flavonoid, the anthocyanin cyanidin-3-O-glucoside (C3G), was tested as a substrate for TT12-dependent transport and compared with its aglycone cyanidin.
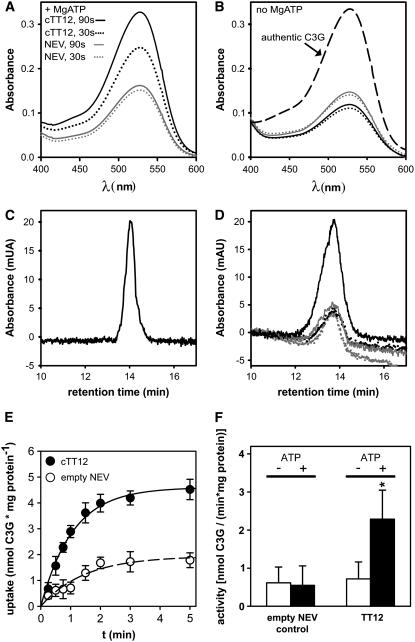
In the presence of MgATP and C3G, TT12-containing vesicles exhibited a time-dependent increase in the absorption spectrum maximum at 535 nm (corresponding to C3G) within 1 min of incubation, suggesting that uptake had occurred (Figure 5A). By contrast, absorption at 535 nm remained unchanged over time when vesicles isolated from empty vector–transformed yeasts were incubated with C3G and MgATP. In the absence of MgATP, no increase in absorption occurred when C3G was incubated with TT12-expressing vesicles (Figure 5B). The absorption spectra recorded after the vesicle transport experiment and filter elution were identical to authentic C3G standard (Figure 5B), which demonstrated that C3G was not subject to structural modification during TT12-mediated uptake. HPLC diode array analysis with detection at 520 nm of samples eluted from filters after transport confirmed the results of simple absorption measurements: single peaks, corresponding to C3G in retention time (Figure 5C) and absorption spectra (data not shown), increased over time only in the presence of both MgATP and TT12-containing vesicles (Figure 5D). We next investigated whether C3G transport by TT12 was time dependent. In the presence of MgATP, TT12-containing vesicles took up C3G linear with time for ∼90 s, while no further increase in vesicle-associated C3G was detectable between 2 and 5 min of transport (Figure 5E). By contrast, empty vector control vesicles exhibited only a marginal increase in the amount of vesicle-associated C3G. Taken together, C3G was transported into vesicles in a TT12-dependent manner and when energization by MgATP occurred (Figure 5F).
Figure 5.
TT12-Mediated Transport of C3G in Yeast Microsomal Vesicles.
(A) and (B) Absorption scans of C3G washed off from filters after the transport experiment representing the vesicle-associated amount of C3G.
(A) Result of a transport experiment in the presence of MgATP.
(B) Corresponding control in the absence of ATP. Transport was stopped by filtration after 30 (dotted lines) or 90 s of vesicular uptake (solid lines). Black and gray lines represent uptake experiments performed with vesicles isolated from cTT12- or empty vector control (NEV)–transformed yeasts, respectively. Dashed line in corresponds to authentic C3G standard.
(C) HPLC profile of authentic C3G standard.
(D) HPLC analysis of the uptake of C3G into microsomal vesicles prepared from yeasts transformed with TT12. Vesicles isolated from TT12-transformed yeasts have taken up C3G after 90 s (black solid line) compared with 30 s (black dotted line) of incubation. Corresponding control vesicles (empty vector; gray lines) do not display any time-dependent increase in C3G.
(E) Time-dependent uptake of C3G into vesicles isolated from yeasts transformed either with TT12 (closed symbols) or the empty vector (open symbols) in the presence of MgATP.
(F) Quantification of uptake activity. Averages ± se of 20 transport experiments performed with independent vesicle preparations are presented. Black and white bars stand for yeasts transformed with TT12 (pNEV-TT12) and empty vector (pNEV), respectively.
By contrast, when cyanidin was added as substrate, no time- and TT12-dependent changes in absorption or HPLC peak sizes were observed (see Supplemental Figure 3 online). Therefore, transport via TT12 is either specific for glucosylated flavonoids or low TT12-dependent transport rates of cyanidin and the flavan-3-ols are not detected because of the background caused by unspecific flavonoid binding to the membranes.
To confirm that the absorption differences observed for C3G were related to transport as well as to elucidate the type of energization mechanism, inhibitor experiments were performed (Table 1). None of the inhibitors tested altered the baseline level of C3G transport into vesicles isolated from empty vector–transformed yeasts. C3G uptake into TT12-containing vesicles, however, was almost completely inhibited by the sulfhydryl group blocker p-chloromercuribenzenesulfonic acid (PCMBS) (Bush, 1993). In the presence of NH4Cl-disrupting pH gradients or bafilomycin A1, a specific inhibitor of the vacuolar H+-ATPase (Dröse and Altendorf, 1997), C3G uptake was strongly reduced to ∼15% of the control or completely abolished, respectively. Therefore, TT12-mediated C3G uptake depends on a proton gradient established by the vacuolar proton pump and acts in vitro as a C3G/H+-antiporter.
Table 1.
Effect of Different Inhibitors on Vesicle Uptake of C3G Performed in the Presence of MgATP
| C3G Uptake (% of the Control)
|
||
|---|---|---|
| Condition | Empty Vector (pNEV) | TT12 |
| None | 24 ± 31 (n = 20) | 100 (n = 20) |
| 1 mM PCMBS | 32 ± 27 (n = 4) | 2 ± 4 (n = 4) |
| 5 mM NH4Cl | 17 ± 13 (n = 4) | 13 ± 19 (n = 4) |
| 0.1μM bafilomycin A1 | 20 ± 20 (n = 4) | 0 ± 0 (n = 4) |
The uptake activity measured in TT12 vesicles in the absence of inhibitors was set to 100% (control). n corresponds to the number of independent experiments.
To further investigate the substrate specificity of TT12-mediated transport, we next tested the glycosylated flavonols Q3R, quercetin-3-O-glucoside (Q3G), and the procyanidin dimers B1 and B2. In our vesicle uptake assay, all of these flavonoids were not transported (Table 2; see Supplemental Figure 3 online).
Table 2.
Synopsis of Transport Activities of TT12 Tested with Different Flavonoids as Substrates and Inhibition of TT12-Mediated C3G Transport by These Compounds
| Flavonoid Class | Compound | Transport by TT12 | Inhibition of C3G Transport by TT12 |
|---|---|---|---|
| Flavan-3-ol monomers | Epicatechin | − | − |
| Catechin | − | − | |
| Cat3G | − | + | |
| Flavan-3-ol dimers | Procyanidin B1 | − | ND |
| Procyanidin B2 | − | ND | |
| Anthocyani(di)ns | C3G | + | NA |
| Cyanidin | − | ND | |
| Flavonols | Q3R | − | − |
| Q3G | − | − |
Presence and absence of a transport activity with the corresponding substrate and of an inhibitory effect on C3G transport is indicated by + and −, respectively. ND, not determined; NA, not applicable.
Due to the specific expression of TT12 in the seed coat (Figure 3), a direct comparison of vacuolar transport activities using wild-type and mutant vacuoles is technically impossible. In an attempt to avoid heterologous expression, we isolated mesophyll vacuoles from the wild type and plants ectopically expressing ProCaMV35S:TT12 in the tt12 background to directly demonstrate the transport activity of TT12 in planta. However, anthocyanins and flavan-3-ols almost quantitatively formed precipitates in the presence of Percoll or Ficoll, which are osmotically inactive substances that are necessary to adjust the density of the vacuoles in the transport medium prior to silicone oil layer centrifugation.
A Glucosylated Flavan-3-ol Specifically Inhibits TT12-Mediated Cyanidin-3-O-Glucoside Transport
If TT12 mediates transport of C3G but not the aglycone cyanidin in vitro, it can be speculated that monomeric flavan-3-ols need to be glycosylated in order to be accepted by TT12 as a substrate. In support of this hypothesis, an epicatechin hexoside has been detected by LC-MS in mature seeds of Landsberg erecta and Cvi (J.-M. Routaboul, unpublished results). As a substantial step toward the identification of the in vivo substrate of TT12, we therefore wanted to investigate whether a glycosylated flavan-3-ol is transported by TT12. Unfortunately, glycosylated epicatechin was not available to us in preparative amounts. However, we were able to obtain chemically synthesized catechin-3-O-glucoside (cat3G).
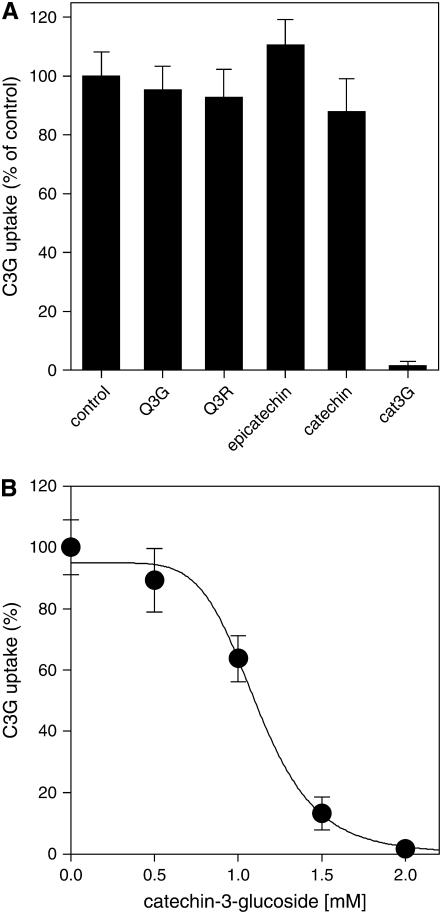
As for epicatechin and catechin, direct TT12-mediated transport of cat3G in yeast vesicles could not be detected (data not shown). However, cat3G inhibited C3G transport in a dose-dependent manner (Figure 6). With both C3G and cat3G at a concentration of 1 mM, MgATP-dependent C3G transport into TT12-containing yeast vesicles was reduced to ∼60% of the control activity in the absence of cat3G. Addition of 2 mM cat3G completely abolished C3G transport. To confirm that cat3G inhibition of TT12-mediated C3G transport is specific for the glucosylated flavan-3-ol, the unconjugated flavan-3ols epicatechin and catechin were added at 2 mM as competitors of C3G transport. In contrast with cat3G, neither catechin nor epicatechin inhibited C3G transport. Furthermore, the flavonols Q3R and Q3G had no effect on C3G transport (Figure 6). Based on these competition experiments, we suggest that a glucosylated flavan-3-ol is the in vivo TT12 substrate, while glycosylated flavonols are not transported by TT12.
Figure 6.
C3G Specifically Inhibits TT12-Mediated C3G Transport.
(A) Inhibition of the uptake of 1 mM C3G by the flavonols Q3G, Q3R, the nonglucosylated flavan-3-ols epicatechin and catechin, and by cat3G. All competitors were added at a final concentration of 2 mM. Experiments were performed in the presence of MgATP. The C3G transport activity with vesicles isolated from TT12-transformed yeasts in the absence of competitors was set to 100%. Transport rates were calculated after subtraction of unspecific binding.
(B) Dose-dependent inhibition of C3G uptake by cat3G. All experiments were performed in triplicate with two independent vesicle preparations.
TT12 Contributes to Vacuolar Anthocyanin Deposition in ban Seeds
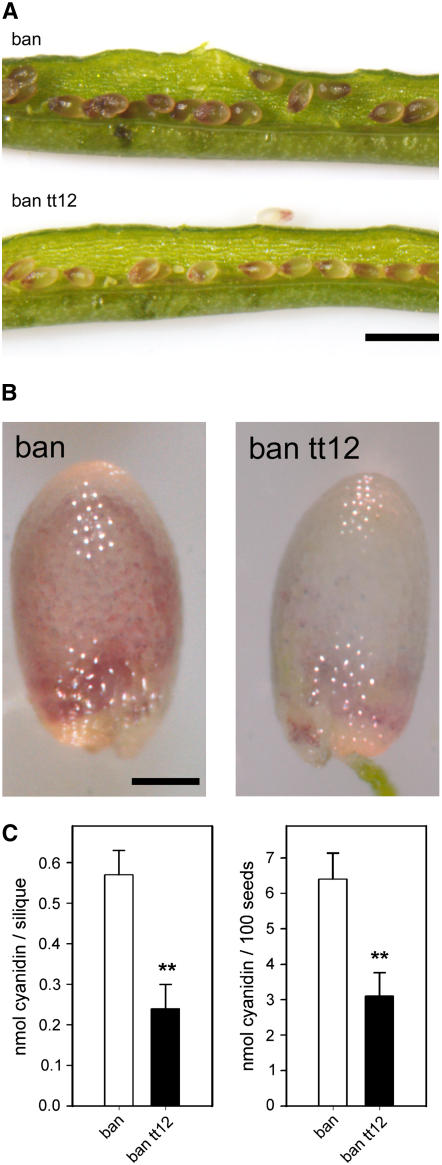
Our transport experiments suggest that TT12 mediates transport of glucosylated anthocyanidins and flavan-3-ol monomers. However, anthocyanins are not in vivo substrates in PA-accumulating cells of the Arabidopsis seed coat. To investigate whether anthocyanin accumulation is affected by TT12 in support of our in vitro data, we analyzed the seed phenotype of a ban tt12 double mutant (Figure 7). Immature seeds of the ban mutant contain anthocyanins of unknown conjugation pattern instead of PAs in PA-producing testa cells since they lack the anthocyanidin reductase activity (Albert et al., 1997; Devic et al., 1999; Xie et al., 2003), while wild-type and tt12 immature seeds do not exhibit any red coloration (data not shown). In contrast with ban, red coloration in immature ban tt12 seeds was strongly reduced. Furthermore, the amount of anthocyanins measured in extracts prepared in acidic methanol as absorption at 525 nm of either complete siliques or immature seeds was reduced to ∼50% in the ban tt12 double mutant compared with ban (Figure 7C).
Figure 7.
Anthocyanin Deposition in Immature Seeds of ban and the Double Mutant ban tt12.
(A) and (B) The characteristic red color observed in the endothelial layer of ban seed caused by anthocyanins is reduced and more diffuse in immature ban tt12 seeds as seen in whole siliques ([A]; bar = 1 mm) or individual immature seeds ([B]; bar = 250 μm).
(C) Entire siliques (left panel) or immature seeds (right panel) of identical developmental stages were extracted in acidic methanol, and the anthocyanin content was determined photometrically at 525 nm. ban tt12 siliques or seeds contain significantly less anthocyanins compared with ban (Student's t test, two-tailed distribution, P < 0.01).
DISCUSSION
Although flavonoid biosynthesis is one of the most thoroughly studied enzymatic pathways in plants, the mechanism(s) by which phenolic compounds reach their cellular destination, the vacuole or the apoplastic space, has not been investigated in detail (Grotewold, 2004). Biochemical investigation of vacuolar transport properties performed with flavonoids or simple phenolics, such as salicylic acid glucoside, as substrates indicated the presence of either directly energized ABC-type transporters or secondarily energized antiporters (for review, see Yazaki, 2005). The latter are usually driven by the H+ gradient across the tonoplast, which is ultimately established by the two vacuolar H+-pumps, the V-type H+-ATPase and H+-pyrophosphatase (Gaxiola et al., 2002). With regard to ABC transporters, the recent characterization of the maize multidrug resistance–associated protein (MRP) gene Zm MRP3 via an antisense approach suggests that MRP-type transporters are involved in vacuolar anthocyanin accumulation in this species since antisense plants lack accumulation of anthocyanins in leaves and tassels of pigmented maize plants (Goodman et al., 2004). However, the biochemical transport activity of tonoplast-localized Zm MRP3 has not been established.
Here, we demonstrate that the MATE transporter TT12 localizes to the tonoplast and mediates flavonoid transport by an H+-antiport mechanism. Therefore, and in accordance with genetic evidence, MATE transporters define a novel plant membrane protein family involved in vacuolar accumulation of phenolic compounds.
TT12 Is a H+-Driven Vacuolar Transporter for Flavonoids
The MATE superfamily has been identified recently as a membrane protein family (Brown et al., 1999) with 9 to 12 membrane-spanning domains (Schwacke et al., 2003). Biochemical data on transport properties and functional aspects of family members are scarce, as is the identification of functional conserved signatures or domains. MATE proteins in prokaryotes function as drug/Na+- or drug/H+-transporters (Morita et al., 2000; Chen et al., 2002; He et al., 2004), while extrusion of norfloxacin, the plant alkaloid berberine, and ethidium bromide mediated by the Arabidopsis MATE At DTX1 expressed in E. coli occurred in a H+-dependent fashion (Li et al., 2002). By contrast, the substrates transported by the MATE proteins encoded by ALF5 or EDS5 discovered via genetic mutant screens are unknown (Diener et al., 2001; Nawrath et al., 2002). Notably, FRD3 acts as a proton-coupled citrate exporter on the plasma membrane, extruding citrate into the xylem or the rhizosphere (Durrett et al., 2007).
Vesicle uptake experiments performed with transformed yeasts expressing TT12 demonstrated that TT12 exhibits time- and ATP-dependent transport of C3G. The strong inhibitory action of bafilomycin A1 and NH4Cl suggests that TT12-catalyzed C3G transport is H+ dependent. Thus, TT12 acts mechanistically like At DTX1 (Li et al., 2002), FRD3 (Durrett et al., 2007), and the human MATE1, which mediates H+-coupled extrusion of toxic organic cations (Otsuka et al., 2005). Moreover, GFP-tagged TT12 resides on the tonoplast. These data demonstrate that TT12 acts as a vacuolar anthocyanin/H+-antiporter under our heterologous expression conditions. In extension to these results, we hypothesize that plant MATE transporters represent the H+-dependent flavonoid transporters previously characterized in vacuolar uptake studies (Klein et al., 1996; Frangne et al., 2002; Marinova et al., 2007). Interestingly, a recent study provides circumstantial evidence for this hypothesis. The ant1 tomato (Solanum lycopersicum) mutant overexpressing a MYB transcription factor by activation tagging overproduces anthocyanins. Upregulated genes in the ant1 background include a MATE transporter gene exhibiting 36% amino acid sequence identity to TT12 and 61% to At4G25640 (Mathews et al., 2003). Therefore, it is tempting to speculate that TT12 homologs are ultimately responsible for vacuolar anthocyanin transport in vegetative plant parts.
What Are the in Vivo Substrates of TT12?
Three findings presented here extend the view on potential functions of TT12 in seeds. First, it was proposed that TT12 acted as a transporter for the PA precursors epicatechin and catechin. However, no detectable transport of nonglycosylated flavan-3-ol monomers and dimers was observed in TT12-expressing yeast vesicles (Table 2). Since TT12 mediated transport of C3G but not the aglycone cyanidin, we speculated that PA precursors need to be glycosylated to be transported by TT12. However, the activity of a glycosyltransferase providing the glycosylated PA precursor remains to be established. While modifications of flavan-3-ols by conjugation reactions have been reported for catechin (Porter, 1993), glucosylated epicatechin is presently not available in sufficient amounts for transport experiments with TT12. Although direct transport of chemically synthesized cat3G was undetectable, cat3G, but not the flavan-3-ol aglycones, was an efficient inhibitor of TT12-mediated C3G transport. Furthermore, cat3G inhibition was dose dependent (Figure 6, Table 2). Based on these findings, we suggest that TT12 is a transporter for glycosylated flavan-3-ol monomers in vivo, while TT12 exhibits an extended substrate specificity in vitro, accepting glycosylated anthocyanidins. Reduction of anthocyanins in ban tt12 double mutant seeds (Figure 7) suggests that absence of TT12 produced a novel phenotype in the ban background, which may be interpreted as a partial defect in vacuolar deposition of anthocyanins. However, since anthocyanins were not completely absent in the double mutant seeds, additional transporters that are able to transport the anthocyanins present in ban seeds have to be suggested.
Second, LC-MS analysis of tt12 seeds established that no soluble flavan-3-ols, PAs, and only low levels of insoluble PAs are present in the absence of the transporter. Furthermore, TT12 promoter activity (this work) and the TT12 mRNA (Debeaujon et al., 2001) were shown to colocalize with cells involved in PA biosynthesis. This situation is identical to the one encountered with the BAN gene and is relevant with the fact that the spatio-temporal patterns of expression for both genes are regulated by the same MYB, bHLH, and WD40 factors, respectively: TT2, TT8, and TTG1 (Nesi et al., 2002; Debeaujon et al., 2003). In view of the fact that TT12 is a member of a large gene family, it can be speculated that due to this specific pattern of expression and the strong tt12 mutant phenotype, TT12 is functionally unique in Arabidopsis in spite of potential redundancy.
Third, LC-MS analysis of tt12 seeds demonstrated a specific reduction in Q3R (the major flavonol component in dry seeds; Routaboul et al., 2006), while all other flavonol derivatives remained unchanged (Figure 2). Pourcel et al. (2005) showed with histochemical staining on immature seed sections that flavonols are essentially present in the oi1 layer of the outer integument in the Arabidopsis wild-type seed coat. TT12 is not expressed in this cell layer, according to the pattern of TT12 promoter activity during seed development (Figure 3). Moreover, neither Q3R nor the corresponding glucoside Q3G was a substrate for TT12-mediated transport into yeast vesicles (Table 2), and these flavonol glycosides were also unable to inhibit C3G transport (Figure 6). Thus, in extension to our results demonstrating that cat3G acts as an efficient competitor of C3G transport, it has to be suggested that flavonol glycosides are unlikely to be substrates of TT12. With regard to our expression and transport results, the finding that levels of Q3R accumulating in the outer integument are reduced in tt12 seeds is puzzling, as is the recent finding that in tt12 mutants, cytosolic flavonoids (presumably mainly flavonols) are not loaded from the cytosol into the tapetosomes of tapetum cells during pollen formation in anthers (Hsieh and Huang, 2007). Additional experiments will be necessary to understand the relationship between TT12 function and flavonol composition. It is possible that histochemical detection of Q3R is masked by tannins in endothelial cells, which may be investigated by looking for flavonol presence in Arabidopsis mutants deprived of tannins but having flavonols (e.g., tt3). It will be particularly important to know whether the flavonol phenotype results from a direct effect of TT12 absence or is an indirect consequence of PA metabolism modification.
If TT12 acts as a flavan-3-ol glycoside/H+-antiporter, the vacuolar lumen must be acidic to support epicatechin accumulation. Seeds of a knockout mutation in the Arabidopsis plasma membrane H+-pump gene AHA10 display features of a transparent testa phenotype, suggesting that AHA10 (though not predicted to localize to the tonoplast) could be involved in establishing a transmembrane H+-gradient driving TT12-mediated vacuolar flavonoid accumulation in PA-accumulating cells (Baxter et al., 2005). In tt18/tds4, aha10, and tt12 mutants, a fluorescent dye, which is targeted to the vacuole in wild-type endothelial cells, accumulates in multiple vesicular structures, suggesting that PA biosynthesis involves vesicle fusion to form the large vacuole (Abrahams et al., 2003; Baxter et al., 2005). Although it remains to be shown how PA biosynthesis affects vacuolar biogenesis in endothelial cells, a prospective model to flavonoid transport proposes vesicle-mediated trafficking of these compounds after their synthesis at large multienzyme complexes located at the cytoplasmic face of the endoplasmic reticulum, involved in channeled production of end products (Winkel, 2004). Vesicles implicated in the transport of flavonoid-derived compounds have been previously described (Grotewold, 2004). Following the model of Baxter et al. (2005) and including our results on the transport function of TT12, the rather static biosynthetic scheme presented in Figure 1 will have to be revised: PAs and other flavonoids are either directly loaded into vesicles or are guided through the cytosol by binding to glutathione S-transferases, such as TT19, toward vesicles destined to form the central vacuole. However, our localization of TT12 to the vacuolar membrane suggests that the transport step itself takes place on the level of the vacuole and not at the endoplasmic reticulum where the precursors are synthesized. TT12-mediated proton-dependent transport of PA precursors into vacuoles could be energized by the proton gradient established by AHA10. The subcellular localization of AHA10, predicted to be the plasma membrane, has not been determined experimentally. In extension of our results, we speculate that AHA10 also localizes to the tonoplast and is therefore a P-type H+-ATPase with an unusual localization. However, due to the effects of AHA10 absence on the vacuolar ultrastructure in the endothelium, it is also possible that AHA10 travels on vesicles (e.g., from the endoplasmic reticulum toward the vacuole or between the plasma membrane and prevacuolar compartments, as proposed by Baxter et al. [2005]). In the latter case, it remains to be determined whether the formation of the pH gradient takes place on the level of the vacuole or in vesicles destined to fuse with the vacuole.
The transport specificities of TT12 suggest a second extension of the late steps of PA biosynthesis in Arabidopsis seeds. According to our transport results, it has to be hypothesized that flavan-3-ols need to be glycosylated prior to vacuolar transport by TT12. However, a glycosylation step acting on the level of epicatechin is biochemically and molecularly unknown. Furthermore, since epicatechin and the derived PA polymers are not glycosylated according to the metabolite analysis of Arabidopsis seeds (Figure 2; Routaboul et al., 2006), an efficient glycosidase must be active in the vacuoles. Both steps have not yet been described on the level of mutants exhibiting a deviation in the seed color, which may reflect gene redundancy. Using the Botany Array Resource Expression Angler tool (Toufighi et al., 2005) with the AtGenExpress Plus and the Botany Array Resource data sets, we searched in silico for genes that are coexpressed with TT12. Three glycosylhydrolases (At2G39640, At4G13600, and At4G33850) and one glucosyltransferase (At3G01620) were found to be coexpressed with TT12 with r-values higher than 0.77. Hierarchical clustering analysis of these four genes together with TT12, BAN, and AHA10, all of which are closely related to PA metabolism in the seed coat, using the clustering tool integrated into Genevestigator V3 (Zimmermann et al., 2004) and the anatomy profile, reveals that the transferase and one of the hydrolases (At4G33850) that cluster together are highly expressed in pollen and stamen (see Supplemental Figure 4 online for details). Thus, it is rather unlikely that these genes are involved in glycosylation of flavan-3-ols and the subsequent hydrolysis of the vacuolar glycoside. By contrast, the hydrolase-encoding gene At2G39640, like AHA10, TT12, and BAN, is preferentially expressed in siliques, and its expression profile clusters closely with TT12 and AHA10. As a consequence, this glycosylhydrolase could be a putative candidate that processes glycosylated flavan-3-ol monomers once these enter the vacuole via TT12-mediated transport.
Our results support the hypothesis that different transporters are responsible for the vacuolar transfer of flavonoids, depending on their modification pattern. In Arabidopsis and tomato, glycosylated flavonoids are handled by MATE transporters, such as TT12 (Debeaujon et al., 2001; Mathews et al., 2003). On the other hand, the major maize anthocyanins, which are malonylated derivatives of C3G and therefore negatively charged at cytosolic pH, have been implicated to be substrates of typical MRP-type ABC transporters, such as Zm MRP3 (Goodman et al., 2004). Therefore, the molecular identification of MATE and ABC transporters as important but independent elements of vacuolar flavonoid accumulation confirms biochemical data suggesting the presence of secondary versus directly energized vacuolar uptake systems when glucosylated versus glucuronidated flavones were compared in rye (Secale cereale) and barley (Klein et al., 1996, 2000; Frangne et al., 2002).
In conclusion, we demonstrate that the Arabidopsis MATE transporter TT12 acts as a flavonoid/H+-antiporter on the vacuolar membrane of PA-synthesizing cells of the seed coat. Its substrate specificity, the ability of cat3G to reduce the transport of C3G, and the phenotype of the ban tt12 double mutant seeds suggests that TT12 can mediate the transport of anthocyanins and flavan-3-ols as long as they are glycosylated, while aglycones, flavan-3-ol dimers, and flavonol glycosides are not TT12 substrates. It therefore has to be postulated that PA precursors undergo a glycosylation step prior to TT12-mediated transfer into the seed coat vacuoles.
METHODS
Chemicals, Strains, Plant Material, and Cloning Procedures
C3G was isolated from blackberries by countercurrent chromatography, and its identity was confirmed by HPLC diode array detection, HPLC coupled to electron spray ionization–mass spectrometry, and nuclear magnetic resonance spectroscopy as described previously (Schwarz et al., 2003). cat3G was from the Nestlé Research Center collection of standard compounds. All other flavonoid compounds were from Extrasynthèse. For uptake experiments, C3G and cyanidin were dissolved as a 0.1 M stock in 50% methanol/0.1% HCl. Stocks of catechin, epicatechin (0.5 M), cat3G (0.2 M), PA dimers (0.1 M), and quercetin glycosides (0.1 M) were prepared in DMSO. Bafilomycin A1 was a kind gift of K. Altendorf (University Osnabrück, Germany). Cellulase YC and pectolyase Y23 were from Kyowa Chemical. PCMBS (Toronto Research Chemicals) was directly dissolved in transport buffer. All other chemicals were from Sigma-Aldrich.
The yeast strain YPH 499 (MATa ura3-52 lys2-801 ade2-101 trp1-Δ63 his3-Δ200 leu2-1) was used for heterologous expression of the TT12 cDNA and its derivatives and for the isolation of microsomal membrane vesicles. The Arabidopsis thaliana tt12 mutant isolated from a collection of T-DNA transformants in the Ws-1 background (Errampalli et al., 1991) and the ban tt12 double mutant have been described elsewhere (Debeaujon et al., 2000, 2001). For our analysis, ban tt12 F2 plants were grown and the F3 seeds were analyzed.
Plasmid Constructions and Transformation Procedures
Construction of a 2μ yeast expression vector pNEV-Ura (Sauer and Stolz, 1994) containing the TT12 cDNA (cTT12) or GFP5:TT12 was performed by amplification of cTT12 and GFP5:TT12 from Pro35Sdual:cTT12/pBIB-HYG and Pro35Sdual:GFP5:TT12:t35S/pAVA393 with the primers TT12-up (5′-ATAAGAATGCGGCCGCATGAGCTCCACAGAGAC-3′) or GFP-TT12-up (5′-ATAAGAATGCGGCCGCATGAGTAAAGGAGAA-3′), respectively, and TT12-low (5′-ATAAGAATGCGGCCGCTTAAACACCTGCGTTAG-3′), where NotI restriction sites are underlined. PCR products were cloned into the NotI site of pNEV-Ura, resulting in pN-TT12 and pN-GFPTT12, respectively. Likewise, N- and C-terminally RGSH6-tagged versions of TT12 cDNA in pNEV-Ura were obtained using the following primers: H6N-TT12up (5′-ATAAGAATGCGGCCGCATGAGAGGATCGCATCACCATCACCATCACTCCTCGATGAGCTCCACAGAGACATACG-3′) and TT12-low for pN-H6NTT12 (N-terminal RGSH6 tag) and TT12-up in combination with H6C-TT12low (5′-ATAAGAATGCGGCCGCTTAGTGATGGTGATGGTGATGCGATCCTCTCGAGGAAACACCTGCGTTAGCCATC-3′) for pN-TT12H6C (C-terminal RGSH6 tag) (NotI sites underlined; RGSH6 motif in italics). Yeast transformation followed standard procedures (Gietz and Woods, 2002), and transformants were selected on minimal synthetic dropout medium lacking uracil (SD-Ura).
To construct the ProTT12:uidA:t35S cassette, a 1.8-kb TT12 promoter was amplified by PCR from the pEB-junction plasmid (Debeaujon et al., 2001) using primers pTT12-5′SalI (5′-GTAGTCGACGAATTCACAATCGGAAAGTCAC-3′) and pTT12-3′NcoI (5′-TCTGTGGAGCCCATGGTCCGTTTATTAGTTC-3′; SalI and NcoI sites underlined, respectively). The SalI-NcoI–digested fragment was cloned into the XhoI and NcoI sites of the pBS-GUS vector described previously (Debeaujon et al., 2003). The ProTT12:uidA:t35S cassette was cloned as a SmaI-KpnI fragment into the pBIB-HYG binary vector (Becker, 1990). The Pro35Sdual:GFP5:TT12:t35S construct was generated as follows. Using primers TT12-ATG-BclI (5′-GGAACTAATAAATGATCAATGAGCTCCACAGAG-3′) and TT12-STOP-BclI (5′-GTATGATCATGCTGTTATCTTAAACACCTGC-3′; BclI sites underlined), the TT12 coding sequence was amplified and cloned at the BglII site of the pAVA393 plasmid. The resulting construct was KpnI-SmaI digested and transferred into pBIB-HYG at the KpnI-SmaI sites to give the Pro35Sdual:GFP5:TT12/pBIB-HYG plasmid. The Pro35Sdual:TT12:GFP/pMDC83 plasmid was obtained by Gateway cloning (Invitrogen). The TT12 cDNA was amplified omitting the stop codon by PCR using the primers 5′-GGGGACAAGTTTGTACAAAAAAGCAGGCTCCATGAGCTCCACAGAGACATACG-3′ and 5′-GGGGACCACTTTGTACAAGAAAGCTGGGTCAACACCTGCGTTAGCCATC-3′, where attB1 and attB2 sites are underlined, respectively. The resulting PCR product was transferred into pDONR Zeo (Invitrogen) and subsequently into pMDC83 (Curtis and Grossniklaus, 2003) by BP and LR clonase reactions according to the manufacturer's instructions.
Plasmids carrying the Pro35S:TPK1:GFP (Czempinski et al., 2002) or the Pro35S:P31:GFP cassette (Marmagne et al., 2004) were kindly obtained from K. Czempinski (University of Potsdam, Germany) and G. Ephritikhine (Centre National de la Recherche Scientifique, Gif-sur-Yvette Cedex, France). The DsRed2:TPK1 marker was constructed by replacing the GFP5 cDNA from pAVA393 by the DsRed2 cDNA (Clontech) to build the DsRed2/pAVA393 vector. Afterwards, the TPK1 cDNA (Czempinski et al., 2002) was amplified with primers cKCO1-5BamHI (5′-GTAGGATCCATGTCGAGTGATGCAGCTCGTAC-3′) and cKCO1-3BamHI (5′-GTAGGATCCTTACCTTTGAATCTGAGACGTG-3′), BamHI-digested, and cloned into the BglII site of DsRed2/pAVA393.
Complementation of the tt12 mutant was performed with cTT12, placed under the control of the dual 35S promoter. The cDNA was amplified from the λ clone isolated by Debeaujon et al. (2001) using the primers TT12-ATG-BspMI (5′-GTAACCTGCGGACCATGGGCTCCACAGAGAC-3′) and TT12-Stop-Bcl(-) (5′-GTATGATCATGCTGTTATCTTAAACGCCTGC-3′; BspMI and BclI sites underlined). Finally, the cDNA was cloned at the SmaI site of the pMagic plasmid (Nesi et al., 2002) to generate the Pro35Sdual:cTT12/pBIB-HYG construction.
For all constructs, PCR amplifications were performed using the proofreading Pfu Ultra DNA Polymerase (Stratagene) or the Expand High Fidelity PCR system (Roche). Correct integration and absence of errors was ultimately verified by sequencing. Stable transformation of Arabidopsis plants was realized as performed previously (Nesi et al., 2000).
Analysis of Seeds
GUS staining, sample mounting in a chloral hydrate solution, and inclusion in resin for the realization of sections and observations were performed according to Debeaujon et al. (2003). Seed flavonoids were extracted and analyzed as described previously (Routaboul et al., 2006).
Preparation of Yeast Membrane Vesicles and in Vitro Transport Studies
Yeast membrane vesicles for in vitro transport studies were isolated essentially as described by Tommasini et al. (1996) with the modifications reported by Klein et al. (2002). Uptake experiments to study transport of flavonoid substrates into membrane vesicles were performed using the rapid filtration technique with nitrocellulose filters (0.45-μm pore size; Millipore). One part of thawed vesicles was mixed with three parts of transport buffer (0.4 M glycerol, 0.1 M KCl, 20 mM Tris-MES, pH 7.4, and 1 mM DTT) in the presence of 1 mM flavonoids and in the absence or presence of 5 mM MgATP in a total reaction volume of 0.8 mL. At time points indicated, 0.35 mL were removed and added to 3 mL of ice-cold transport buffer, and the mixture was loaded on a prewet filter that was rapidly washed with 3× 2 mL of ice-cold transport buffer. Immediately after the experiment, filter-bound flavonoids were dissolved by adding 50% methanol/0.1% HCl. Flavonoid content eluted from the filters was analyzed in two ways: (1) absorption spectra scans between 220 and 600 nm with a Beckman Coulter DU 800 spectrophotometer, and (2) HPLC separation was performed on a CC 250/4 Nucleosil 100-5 C18HD column (Macherey-Nagel) with water/0.6% perchloric acid (A) and methanol (B) at a total flow rate of 1 mL/min and the following gradient: 0 to 20 min from 5 to 60% B over A. Parameters were controlled by a Gynkotek liquid chromatograph (Dionex) equipped with a UVD340S diode array detector set at 520, 330, and 280 nm for identification of anthocyanins, flavonols, and flavan-3-ols, respectively.
Subcellular Localization and Confocal Laser Scanning Microscopy Analysis
Subcellular localization of TT12-GFP fusions was investigated in transient assays and after stable transformation of Arabidopsis plants. Transient expression in onion blub epidermal cells was performed by biolistic particle delivery of Pro35Sdual:GFP5:TT12:t35S/pAVA393 alone or in the presence of equimolar amounts of Pro35Sdual:DsRed2:TPK1 using DNA-coated 0.6 μm gold particles as microcarriers and the BioRad He Biolistic PDS-1000/He system following the manufacturer's instructions. Transient expression in Nicotiana benthamiana leaves using infiltration of Agrobacterium tumefaciens GV31 01 transformed with the corresponding plasmids by electroporation was performed according to published procedures (Voinnet et al., 2003; Walter et al., 2004). Microscopic observations were made after 48 h at 25°C. Mesophyll protoplasts from infiltrated N. benthamiana leaves or stably transformed Pro35Sdual:GFP5:TT12/tt12 Arabidopsis plants were prepared as described (Frangne et al., 2002), and vacuoles were released during confocal analysis by gentle addition of water. Roots of sterile-grown transgenic tt12 plants ectopically expressing GFP5-cTT12 were analyzed after 7 d and were counterstained with 1 mM propidium iodide (Molecular Probes). Single optical sections were captured by confocal laser scanning microscopy using a TCS SP2-x1 full-spectrum confocal microsope attached to a Leica DM IRE2 inverted fluorescence microscope. GFP and DsRed or GFP and propidium iodide were simultaneously excited with a 488-nm Ar and a 543-nm HeNe laser, and fluorescence emission images averaged over eight frames were captured in independent channels (GFP, 500 to 520 nm; DsRed, 580 to 615 nm; propidium iodide and chlorophyll autofluorescence, 620 to 700nm; DD488/543 beam splitter). Images were false-colored in green (GFP) or red (DsRed, propidium iodide, and chlorophyll autofluorescence) using Adobe Photoshop 7.0.
Accession Numbers
The Arabidopsis Genome Initiative identifiers for the genes/proteins described in this article are as follows: TT12/At DTX41 (At3G59030), BAN/ANR (At1G61720), TT10 (At5G48100), ALF5/At DTX19 (At3G23560), FRD3/At DTX43 (At3G08040), EDS5/At DTX47 (At4g39030), At DTX1 (At2G04070), Arabidopsis P31 protein (At3G01290), AHA10 (At1G17260), TT19 (At5G17220), chalcone synthase (At5G13930), chalcone isomerase (At3G55120), flavanone 3-hydroxylase (At3G51240), flavonol 3′-hydroxylase (At5G07790), flavonol synthase (At5G08640), and leucoanthocyanidin dioxygenase (At4G22880).
Supplemental Data
The following materials are available in the online version of this article.
Supplemental Figure 1. Localization of GFP-cTT12 after Transient and Stable Expression in Plant Cells.
Supplemental Figure 2. Expression and Localization of TT12 in Saccharomyces cerevisiae and ATP-Dependent Formation of a pH Gradient.
Supplemental Figure 3. TT12 Transport Activity toward Various Flavonoid Substrates.
Supplemental Figure 4. Hierarchical Clustering Analysis of the Expression of TT12, BANYULS, AHA10, and Putative Coexpressed Glycosyltransferases or Glycosylhydrolases.
Supplemental Methods. Heterologous Expression of cTT12 in Saccharomyces cerevisiae and MgATP-Dependent pH Gradient Formation in Yeast Membrane Vesicles.
Supplementary Material
Acknowledgments
We thank E. Martinoia, S.E. Schauer (both of the University of Zurich), and L. Lepiniec (Institut National de la Recherche Agronomique Versailles) for comments on the manuscript, K. Czempinski (University of Potsdam) for the TPK1 clone, G. Ephritikhine (Centre National de la Recherche Scientifique, Gif-sur-Yvette Cedex, France) for P31-GFP, A. von Arnim (University of Tennessee, Knoxville) for pAVA393, and J.-D. Faure and C. Smyczynski (both of the Institut National de la Recherche Agronomique Versailles) for the DsRed2 cDNA. We also thank the reviewers for constructive and interesting suggestions. This work was supported by the Forschungskredit of the University of Zurich (M.K.), by the Swiss National Foundation (3100A0-116051; M.K.), by the Roche Research Foundation (Basel) (K.M. and M.K.), and by the Integrated Action Program “Germaine de Staël” (I.D. and M.K.).
The author responsible for distribution of materials integral to the findings presented in this article in accordance with the policy described in the Instructions for Authors (www.plantcell.org) is: Markus Klein (mklein@botinst.uzh.ch).
Online version contains Web-only data.
References
- Abrahams, S., Lee, E., Walker, A.R., Tanner, G.J., Larkin, P.J., and Ashton, A.R. (2003). The Arabidopsis TDS4 gene encodes leucoanthocyanidin dioxygenase (LDOX) and is essential for proanthocyanidin synthesis and vacuole development. Plant J. 35 624–636. [DOI] [PubMed] [Google Scholar]
- Albert, S., Delseny, M., and Devic, M. (1997). BANYULS, a novel negative regulator of flavonoid biosynthesis in the Arabidopsis seed coat. Plant J. 11 289–299. [DOI] [PubMed] [Google Scholar]
- Baxter, I.R., Young, J.C., Armstrong, G., Foster, N., Bogenschutz, N., Cordova, T., Peer, W.A., Hazen, S.P., Murphy, A.S., and Harper, J.F. (2005). A plasma membrane H+-ATPase is required for the formation of proanthocyanidins in the seed coat endothelium of Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 102 2649–2654. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Becker, D. (1990). Binary vectors which allow the exchange of plant selectable markers and reporter genes. Nucleic Acids Res. 18 203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown, M.H., Paulsen, I.T., and Skurray, R.A. (1999). The multidrug efflux protein NorM is a prototype of a new family of transporters. Mol. Microbiol. 31 394–395. [DOI] [PubMed] [Google Scholar]
- Bush, D.R. (1993). Inhibitors of the proton-sucrose symport. Arch. Biochem. Biophys. 307 355–360. [DOI] [PubMed] [Google Scholar]
- Chen, J., Morita, Y., Huda, M.N., Kuroda, T., Mizushima, T., and Tsuchiya, T. (2002). VmrA, a member of a novel class of Na+-coupled multidrug efflux pumps from Vibrio parahaemolyticus. J. Bacteriol. 184 572–576. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Coleman, J.O.D., Randall, R., and Blake-Kalff, M.M.A. (1997). Detoxification of xenobiotics in plant cells by glutathione conjugation and vacuolar compartmentalization: A fluorescent assay using monochlorobimane. Plant Cell Environ. 20 449–460. [Google Scholar]
- Curtis, M.D., and Grossniklaus, U. (2003). A gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant Physiol. 133 462–469. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Czempinski, K., Frachisse, J.M., Maurel, C., Barbier-Brygoo, H., and Mueller-Roeber, B. (2002). Vacuolar membrane localization of the Arabidopsis ‘two-pore’ K+ channel KCO1. Plant J. 29 809–820. [DOI] [PubMed] [Google Scholar]
- Debeaujon, I., Leon-Kloosterziel, K.M., and Koornneef, M. (2000). Influence of the testa on seed dormancy, germination, and longevity in Arabidopsis. Plant Physiol. 122 403–413. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Debeaujon, I., Nesi, N., Perez, P., Devic, M., Grandjean, O., Caboche, M., and Lepiniec, L. (2003). Proanthocyanidin-accumulating cells in Arabidopsis testa: Regulation of differentiation and role in seed development. Plant Cell 15 2514–2531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Debeaujon, I., Peeters, A.J.M., Leon-Kloosterziel, K.M., and Koornneef, M. (2001). The TRANSPARENT TESTA12 gene of Arabidopsis encodes a multidrug secondary transporter-like protein required for flavonoid sequestration in vacuoles of the seed coat endothelium. Plant Cell 13 853–871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Devic, M., Guilleminot, J., Debeaujon, I., Bechtold, N., Bensaude, E., Koornneef, M., Pelletier, G., and Delseny, M. (1999). The BANYULS gene encodes a DFR-like protein and is a marker of early seed coat development. Plant J. 19 387–398. [DOI] [PubMed] [Google Scholar]
- Diener, A.C., Gaxiola, R.A., and Fink, G.R. (2001). Arabidopsis ALF5, a multidrug efflux transporter gene family member, confers resistance to toxins. Plant Cell 13 1625–1637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dröse, S., and Altendorf, K. (1997). Bafilomycins and concanamycins as inhibitors of V-ATPases and P-ATPases. J. Exp. Biol. 200 1–8. [DOI] [PubMed] [Google Scholar]
- Durrett, T.P., Gassmann, W., and Rogers, E.E. (2007). The FRD3-mediated efflux of citrate into the root vasculature is necessary for efficient iron translocation. Plant Physiol. 144 197–205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Errampalli, D., Patton, D., Castle, L., Mickelson, L., Hansen, K., Schnall, J., Feldmann, K., and Meinke, D. (1991). Embryonic lethals and T-DNA insertional mutagenesis in Arabidopsis. Plant Cell 3 149–157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frangne, N., Eggmann, T., Koblischke, C., Weissenböck, G., Martinoia, E., and Klein, M. (2002). Flavone glucoside uptake into barley mesophyll and Arabidopsis cell culture vacuoles. Energization occurs by H+-antiport and ATP-binding cassette-type mechanisms. Plant Physiol. 128 726–733. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gaxiola, R.A., Fink, G.R., and Hirschi, K.D. (2002). Genetic manipulation of vacuolar proton pumps and transporters. Plant Physiol. 129 967–973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gietz, R.D., and Woods, R.A. (2002). Transformation of yeast by lithium acetate/single-stranded carrier DNA/polyethylene glycol method. Methods Enzymol. 350 87–96. [DOI] [PubMed] [Google Scholar]
- Goodman, C.D., Casati, P., and Walbot, V. (2004). A multidrug resistance-associated protein involved in anthocyanin transport in Zea mays. Plant Cell 16 1812–1826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grotewold, E. (2004). The challenges of moving chemicals within and out of cells: Insights into the transport of plant natural products. Planta 219 906–909. [DOI] [PubMed] [Google Scholar]
- Harborne, J.B., and Williams, C.A. (2000). Advances in flavonoid research since 1992. Phytochemistry 55 481–504. [DOI] [PubMed] [Google Scholar]
- He, G.X., Kuroda, T., Mima, T., Morita, Y., Mizushima, T., and Tsuchiya, T. (2004). An H+-coupled multidrug efflux pump, PmpM, a member of the MATE family of transporters, from Pseudomonas aeruginosa. J. Bacteriol. 186 262–265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsieh, K., and Huang, A.H.C. (2007). Tapetosomes in Brassica tapetum accumulate endoplasmic reticulum-derived flavonoids and alkanes for delivery to the pollen surface. Plant Cell 19 582–596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kerhoas, L., Aouak, D., Cingoz, A., Routaboul, J.M., Lepiniec, L., Einhorn, J., and Birlirakis, N. (2006). Structural characterization of the major flavonoid glycosides from Arabidopsis thaliana seeds. J. Agric. Food Chem. 54 6603–6612. [DOI] [PubMed] [Google Scholar]
- Klein, M., Mamnun, Y.M., Eggmann, T., Schuller, C., Wolfger, H., Martinoia, E., and Kuchler, K. (2002). The ATP-binding cassette (ABC) transporter Bpt1p mediates vacuolar sequestration of glutathione conjugates in yeast. FEBS Lett. 520 63–67. [DOI] [PubMed] [Google Scholar]
- Klein, M., Martinoia, E., Hoffmann-Thoma, G., and Weissenböck, G. (2000). A membrane-potential dependent ABC-like transporter mediates the vacuolar uptake of rye flavone glucuronides: Regulation of glucuronide uptake by glutathione and its conjugates. Plant J. 21 289–304. [DOI] [PubMed] [Google Scholar]
- Klein, M., Weissenböck, G., Dufaud, A., Gaillard, C., Kreuz, K., and Martinoia, E. (1996). Different energization mechanisms drive the vacuolar uptake of a flavonoid glucoside and a herbicide glucoside. J. Biol. Chem. 271 29666–29671. [DOI] [PubMed] [Google Scholar]
- Lepiniec, L., Debeaujon, I., Routaboul, J.M., Baudry, A., Pourcel, L., Nesi, N., and Caboche, M. (2006). Genetics and biochemistry of seed flavonoids. Annu. Rev. Plant Biol. 57 405–430. [DOI] [PubMed] [Google Scholar]
- Li, L., He, Z., Pandey, G.K., Tsuchiya, T., and Luan, S. (2002). Functional cloning and characterization of a plant efflux carrier for multidrug and heavy metal detoxification. J. Biol. Chem. 277 5360–5368. [DOI] [PubMed] [Google Scholar]
- Marinova, K., Kleinschmidt, K., Weissenböck, G., and Klein, M. (2007). Flavonoid biosynthesis in barley (Hordeum vulgare L.) primary leaves requires the presence of the vacuole and controls the activity of vacuolar flavonoid transport. Plant Physiol. 144 432–444. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marmagne, A., Rouet, M.-A., Ferro, M., Rolland, N., Alcon, C., Joyard, J., Garin, J., Barbier-Brygoo, H., and Ephritikhine, G. (2004). Identification of new intrinsic proteins in Arabidopsis plasma membrane proteome. Mol. Cell. Proteomics 3 675–691. [DOI] [PubMed] [Google Scholar]
- Mathews, H., Clendennen, S.K., Caldwell, C.G., Liu, X.L., Connors, K., Matheis, N., Schuster, D.K., Menasco, D.J., Wagoner, W., Lightner, J., and Wagner, D.R. (2003). Activation tagging in tomato identifies a transcriptional regulator of anthocyanin biosynthesis, modification, and transport. Plant Cell 15 1689–1703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matile, P. (1984). The toxic compartment of plant cells. Naturwissenschaften 71 18–24. [Google Scholar]
- Morita, Y., Kataoka, A., Shiota, S., Mizushima, T., and Tsuchiya, T. (2000). NorM of Vibrio parahaemolyticus is an Na+-driven multidrug efflux pump. J. Bacteriol. 182 6694–6697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morita, Y., Kodama, K., Shiota, S., Mine, T., Kataoka, A., Mizushima, T., and Tsuchiya, T. (1998). NorM, a putative multidrug efflux protein, of Vibrio parahaemolyticus and its homolog in Escherichia coli. Antimicrob. Agents Chemother. 42 1778–1782. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nawrath, C., Heck, S., Parinthawong, N., and Metraux, J.P. (2002). EDS5, an essential component of salicylic acid-dependent signaling for disease resistance in Arabidopsis, is a member of the MATE transporter family. Plant Cell 14 275–286. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nesi, N., Debeaujon, I., Jond, C., Pelletier, G., Caboche, M., and Lepiniec, L. (2000). The TT8 gene encodes a basic helix-loop-helix domain protein required for expression of DFR and BAN genes in Arabidopsis siliques. Plant Cell 12 1863–1878. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nesi, N., Debeaujon, I., Jond, C., Stewart, A.J., Jenkins, G.I., Caboche, M., and Lepiniec, L. (2002). The TRANSPARENT TESTA16 locus encodes the Arabidopsis BSISTER MADS domain protein and is required for proper development and pigmentation of the seed coat. Plant Cell 14 2463–2479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Otsuka, M., Matsumoto, T., Morimoto, R., Arioka, S., Omote, H., and Moriyama, Y. (2005). A human transporter protein that mediates the final excretion step for toxic organic cations. Proc. Natl. Acad. Sci. USA 102 17923–17928. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Porter, L.J. (1993). Flavans and proanthocyanidins. In The Flavonoids: Advances in Research Since 1986, J.B. Harborne, ed (London: Chapman and Hall), pp. 23–55.
- Pourcel, L., Routaboul, J.M., Kerhoas, L., Caboche, M., Lepiniec, L., and Debeaujon, I. (2005). TRANSPARENT TESTA10 encodes a laccase-like enzyme involved in oxidative polymerization of flavonoids in Arabidopsis seed coat. Plant Cell 17 2966–2980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rogers, E.E., and Guerinot, M.L. (2002). FRD3, a member of the multidrug and toxin efflux family, controls iron deficiency responses in Arabidopsis. Plant Cell 14 1787–1799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Routaboul, J.M., Kerhoas, L., Debeaujon, I., Pourcel, L., Caboche, M., Einhorn, J., and Lepiniec, L. (2006). Flavonoid diversity and biosynthesis in seed of Arabidopsis thaliana. Planta 224 96–107. [DOI] [PubMed] [Google Scholar]
- Sauer, N., and Stolz, J. (1994). SUC1 and SUC2: Two sucrose transporters from Arabidopsis thaliana; expression and characterization in baker's yeast and identification of the histidine-tagged protein. Plant J. 6 67–77. [DOI] [PubMed] [Google Scholar]
- Schwacke, R., Schneider, A., van der Graaff, E., Fischer, K., Catoni, E., Desimone, M., Frommer, W.B., Flügge, U.I., and Kunze, R. (2003). ARAMEMNON, a novel database for Arabidopsis integral membrane proteins. Plant Physiol. 131 16–26. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwarz, M., Hillebrand, S., Habben, S., Degenhardt, A., and Winterhalter, P. (2003). Application of high-speed countercurrent chromatography to the large-scale isolation of anthocyanins. Biochem. Eng. J. 14 179–189. [Google Scholar]
- Shirley, B.W., Kubasek, W.L., Storz, G., Bruggemann, E., Koornneef, M., Ausubel, F.M., and Goodman, H.M. (1995). Analysis of Arabidopsis mutants deficient in flavonoid biosynthesis. Plant J. 8 659–671. [DOI] [PubMed] [Google Scholar]
- Tommasini, R., Evers, R., Vogt, E., Mornet, C., Zaman, G.J.R., Schinkel, A.H., Borst, P., and Martinoia, E. (1996). The human multidrug resistance-associated protein functionally complements the yeast cadmium resistance factor 1. Proc. Natl. Acad. Sci. USA 93 6743–6748. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Toufighi, K., Brady, S.M., Austin, R., Ly, E., and Provart, N.J. (2005). The botany array resource: e-Northerns, expression angling, and promoter analyses. Plant J. 43 153–163. [DOI] [PubMed] [Google Scholar]
- Voinnet, O., Rivas, S., Mestre, P., and Baulcombe, D. (2003). An enhanced transient expression system in plants based on suppression of gene silencing by the p19 protein of tomato bushy stunt virus. Plant J. 33 949–956. [DOI] [PubMed] [Google Scholar]
- Walter, M., Chaban, C., Schutze, K., Batistic, O., Weckermann, K., Nake, C., Blazevic, D., Grefen, C., Schumacher, K., Oecking, C., Harter, K., and Kudla, J. (2004). Visualization of protein interactions in living plant cells using bimolecular fluorescence complementation. Plant J. 40 428–438. [DOI] [PubMed] [Google Scholar]
- Winkel, B.S.J. (2004). Metabolic channeling in plants. Annu. Rev. Plant Biol. 55 85–107. [DOI] [PubMed] [Google Scholar]
- Xie, D.Y., Sharma, S.B., Paiva, N.L., Ferreira, D., and Dixon, R.A. (2003). Role of anthocyanidin reductase, encoded by BANYULS in plant flavonoid biosynthesis. Science 299 396–399. [DOI] [PubMed] [Google Scholar]
- Yazaki, K. (2005). Transporters of secondary metabolites. Curr. Opin. Plant Biol. 8 301–307. [DOI] [PubMed] [Google Scholar]
- Zimmermann, P., Hirsch-Hoffmann, M., Hennig, L., and Gruissem, W. (2004). GENEVESTIGATOR. Arabidopsis microarray database and analysis toolbox. Plant Physiol. 136 2621–2632. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.