Abstract
ComFA is a membrane protein required for the uptake of transforming DNA following its binding to the Bacillus subtilis competent-cell surface. ComFA, which resembles members of the DEAD family of ATP-driven helicases, contains sequences similar to those found in many ATP-binding proteins and thought to represent the ATP-binding sites of these proteins. We have suggested that ComFA may function as a DNA translocase and/or helicase, using the energy of ATP hydrolysis to mediate the uptake of DNA. As a partial test of this hypothesis, we have introduced mutations into highly conserved glycyl and lysyl residues of the putative ATP-binding site, located, respectively, at positions 151 and 152, and determined the effects of these alterations on in vivo function. A substitution of the conserved lysyl by a glutamyl residue (K152E) and a double G151R-K152N mutation each resulted in a nearly 1,000-fold decrease in transformability, equivalent to that observed in a ComFA null mutant. A K152N mutation caused a partial loss-of-function phenotype. These effects were manifested at the level of DNA uptake; no marked effects on the final levels of DNA binding were noted. When either the K152E mutant allele or the G151R-K152N double mutant allele was combined in single copy with wild-type comFA, a dominant negative phenotype expressed on the level of DNA uptake was observed, suggesting that ComFA acts in a complex with other proteins, with additional molecules of ComFA, or with both.
Full text
PDF

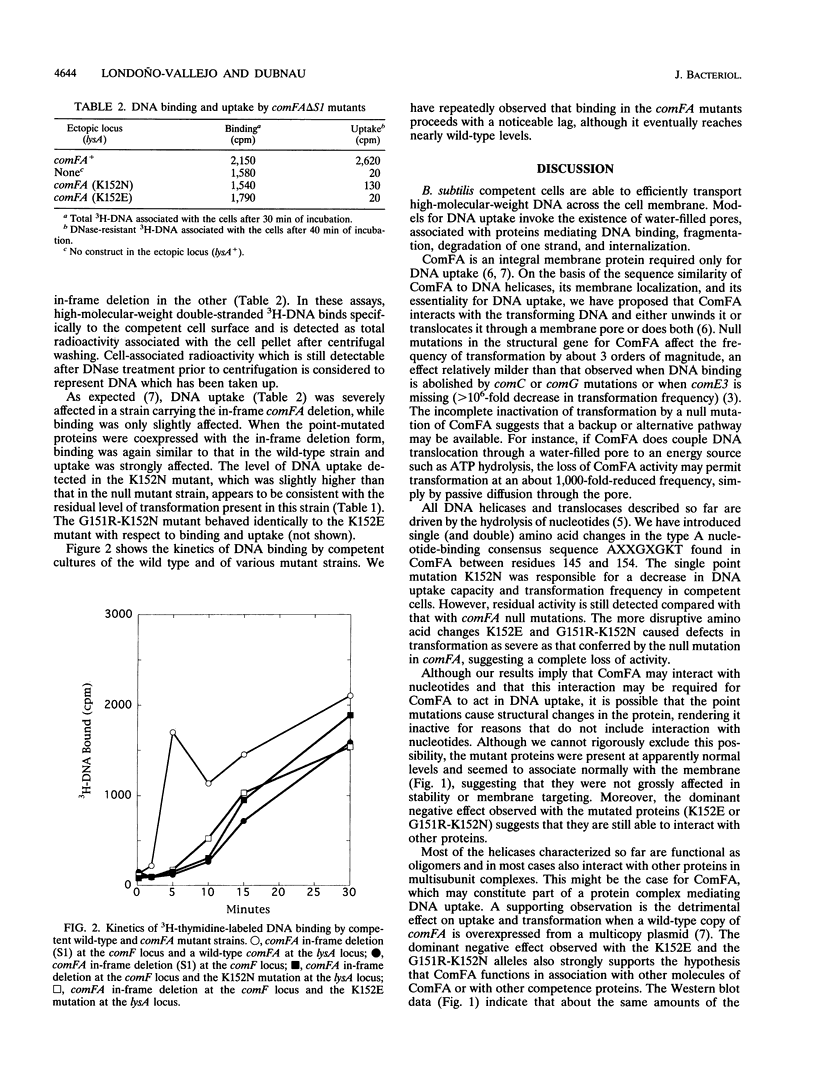

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Albano M., Hahn J., Dubnau D. Expression of competence genes in Bacillus subtilis. J Bacteriol. 1987 Jul;169(7):3110–3117. doi: 10.1128/jb.169.7.3110-3117.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dubnau D. Genetic competence in Bacillus subtilis. Microbiol Rev. 1991 Sep;55(3):395–424. doi: 10.1128/mr.55.3.395-424.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hahn J., Albano M., Dubnau D. Isolation and characterization of Tn917lac-generated competence mutants of Bacillus subtilis. J Bacteriol. 1987 Jul;169(7):3104–3109. doi: 10.1128/jb.169.7.3104-3109.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee M. S., Marians K. J. Differential ATP requirements distinguish the DNA translocation and DNA unwinding activities of the Escherichia coli PRI A protein. J Biol Chem. 1990 Oct 5;265(28):17078–17083. [PubMed] [Google Scholar]
- Londoño-Vallejo J. A., Dubnau D. comF, a Bacillus subtilis late competence locus, encodes a protein similar to ATP-dependent RNA/DNA helicases. Mol Microbiol. 1993 Jul;9(1):119–131. doi: 10.1111/j.1365-2958.1993.tb01674.x. [DOI] [PubMed] [Google Scholar]
- Nurse P., DiGate R. J., Zavitz K. H., Marians K. J. Molecular cloning and DNA sequence analysis of Escherichia coli priA, the gene encoding the primosomal protein replication factor Y. Proc Natl Acad Sci U S A. 1990 Jun;87(12):4615–4619. doi: 10.1073/pnas.87.12.4615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walker J. E., Saraste M., Runswick M. J., Gay N. J. Distantly related sequences in the alpha- and beta-subunits of ATP synthase, myosin, kinases and other ATP-requiring enzymes and a common nucleotide binding fold. EMBO J. 1982;1(8):945–951. doi: 10.1002/j.1460-2075.1982.tb01276.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weinrauch Y., Msadek T., Kunst F., Dubnau D. Sequence and properties of comQ, a new competence regulatory gene of Bacillus subtilis. J Bacteriol. 1991 Sep;173(18):5685–5693. doi: 10.1128/jb.173.18.5685-5693.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]




