Abstract
Cell differentiation is an inherent component of the Caulobacter crescentus cell cycle. The transition of a swarmer cell, with a single polar flagellum, into a sessile stalked cell includes several morphogenetic events. These include the release of the flagellum and pili, the proteolysis of chemotaxis proteins, the biogenesis of the polar stalk, and the initiation of DNA replication. We have isolated a group of temperature-sensitive mutants that are unable to complete this process at the restrictive temperature. We show here that one of these strains has a mutation in a homolog of the Escherichia coli secA gene, whose product is involved in protein translocation at the cell membrane. This C. crescentus secA mutant has allowed the identification of morphogenetic events in the swarmer-to-stalked cell transition that require SecA-dependent protein translocation. Upon shift to the nonpermissive temperature, the mutant secA swarmer cell is able to release the polar flagellum, degrade chemoreceptors, and initiate DNA replication, but it is unable to form a stalk, complete DNA replication, or carry out cell division. At the nonpermissive temperature, the cell cycle blocks prior to the de novo synthesis of flagella and chemotaxis proteins that normally occurs in the predivisional cell. Although interactions between the chromosome and the cytoplasmic membrane are believed to be a functional component of the temporal regulation of DNA replication, the ability of this secA mutant to initiate replication at the nonpermissive temperature suggests that SecA-dependent events are not involved in this process. However, both cell division and stalk formation, which is analogous to a polar division event, require SecA function.
Full text
PDF




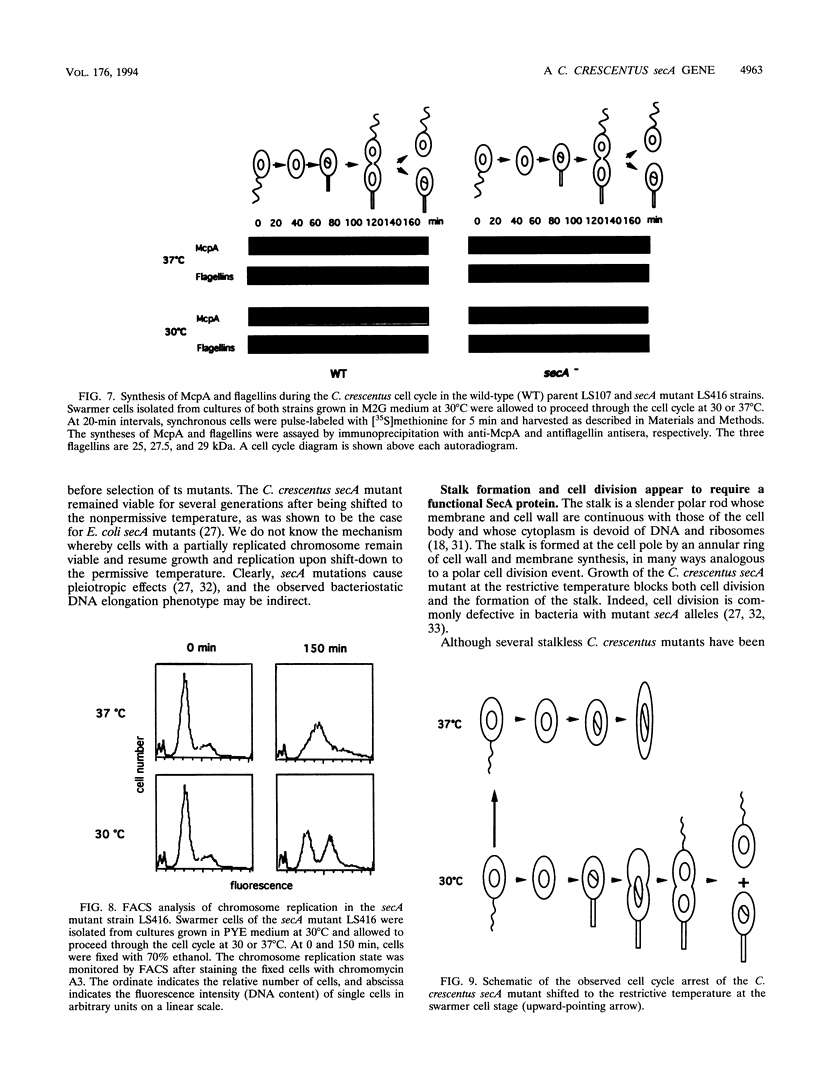


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alley M. R., Gomes S. L., Alexander W., Shapiro L. Genetic analysis of a temporally transcribed chemotaxis gene cluster in Caulobacter crescentus. Genetics. 1991 Oct;129(2):333–341. doi: 10.1093/genetics/129.2.333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alley M. R., Maddock J. R., Shapiro L. Requirement of the carboxyl terminus of a bacterial chemoreceptor for its targeted proteolysis. Science. 1993 Mar 19;259(5102):1754–1757. doi: 10.1126/science.8456303. [DOI] [PubMed] [Google Scholar]
- Degnen S. T., Newton A. Chromosome replication during development in Caulobacter crescentus. J Mol Biol. 1972 Mar 14;64(3):671–680. doi: 10.1016/0022-2836(72)90090-3. [DOI] [PubMed] [Google Scholar]
- Devereux J., Haeberli P., Smithies O. A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 1):387–395. doi: 10.1093/nar/12.1part1.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dingwall A., Shapiro L. Rate, origin, and bidirectionality of Caulobacter chromosome replication as determined by pulsed-field gel electrophoresis. Proc Natl Acad Sci U S A. 1989 Jan;86(1):119–123. doi: 10.1073/pnas.86.1.119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ditta G., Schmidhauser T., Yakobson E., Lu P., Liang X. W., Finlay D. R., Guiney D., Helinski D. R. Plasmids related to the broad host range vector, pRK290, useful for gene cloning and for monitoring gene expression. Plasmid. 1985 Mar;13(2):149–153. doi: 10.1016/0147-619x(85)90068-x. [DOI] [PubMed] [Google Scholar]
- Ely B., Ely T. W. Use of pulsed field gel electrophoresis and transposon mutagenesis to estimate the minimal number of genes required for motility in Caulobacter crescentus. Genetics. 1989 Dec;123(4):649–654. doi: 10.1093/genetics/123.4.649. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ely B. Transfer of drug resistance factors to the dimorphic bacterium Caulobacter crescentus. Genetics. 1979 Mar;91(3):371–380. doi: 10.1093/genetics/91.3.371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Evinger M., Agabian N. Envelope-associated nucleoid from Caulobacter crescentus stalked and swarmer cells. J Bacteriol. 1977 Oct;132(1):294–301. doi: 10.1128/jb.132.1.294-301.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fukuda A., Iba H., Okada Y. Stalkless mutants of Caulobacter crescentus. J Bacteriol. 1977 Jul;131(1):280–287. doi: 10.1128/jb.131.1.280-287.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fukuda A., Miyakawa K., Iida H., Okada Y. Regulation of polar surface structures in Caulobacter crescentus: pleiotropic mutations affect the coordinate morphogenesis of flagella, pili and phage receptors. Mol Gen Genet. 1976 Dec 8;149(2):167–173. doi: 10.1007/BF00332885. [DOI] [PubMed] [Google Scholar]
- Gober J. W., Shapiro L. Integration host factor is required for the activation of developmentally regulated genes in Caulobacter. Genes Dev. 1990 Sep;4(9):1494–1504. doi: 10.1101/gad.4.9.1494. [DOI] [PubMed] [Google Scholar]
- Gomes S. L., Shapiro L. Differential expression and positioning of chemotaxis methylation proteins in Caulobacter. J Mol Biol. 1984 Sep 25;178(3):551–568. doi: 10.1016/0022-2836(84)90238-9. [DOI] [PubMed] [Google Scholar]
- Hodgson D., Shaw P., Letts V., Henry S., Shapiro L. Genetic analysis and characterization of a Caulobacter crescentus mutant defective in membrane biogenesis. J Bacteriol. 1984 May;158(2):430–440. doi: 10.1128/jb.158.2.430-440.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hodgson D., Shaw P., O'Connell M., Henry S., Shapiro L. Caulobacter crescentus fatty acid-dependent cell cycle mutant. J Bacteriol. 1984 Apr;158(1):156–162. doi: 10.1128/jb.158.1.156-162.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson R. C., Ely B. Isolation of spontaneously derived mutants of Caulobacter crescentus. Genetics. 1977 May;86(1):25–32. doi: 10.1093/genetics/86.1.25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jordan T. L., Porter J. S., Pate J. L. Isolation and characterization of prosthecae of Asticcacaulis biprosthecum. Arch Mikrobiol. 1974 Mar 1;96(1):1–16. doi: 10.1007/BF00590158. [DOI] [PubMed] [Google Scholar]
- Josefsson L. G., Randall L. L. Different exported proteins in E. coli show differences in the temporal mode of processing in vivo. Cell. 1981 Jul;25(1):151–157. doi: 10.1016/0092-8674(81)90239-7. [DOI] [PubMed] [Google Scholar]
- Marczynski G. T., Dingwall A., Shapiro L. Plasmid and chromosomal DNA replication and partitioning during the Caulobacter crescentus cell cycle. J Mol Biol. 1990 Apr 20;212(4):709–722. doi: 10.1016/0022-2836(90)90232-B. [DOI] [PubMed] [Google Scholar]
- Marino W., Ammer S., Shapiro L. Conditional surface structure mutants of Caulobacter crescentus temperature-sensitive flagella formation due to an altered flagellin monomer. J Mol Biol. 1976 Oct 25;107(2):115–130. doi: 10.1016/s0022-2836(76)80021-6. [DOI] [PubMed] [Google Scholar]
- Mizusawa S., Nishimura S., Seela F. Improvement of the dideoxy chain termination method of DNA sequencing by use of deoxy-7-deazaguanosine triphosphate in place of dGTP. Nucleic Acids Res. 1986 Feb 11;14(3):1319–1324. doi: 10.1093/nar/14.3.1319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nathan P., Newton A. Identification of two new cell division genes that affect a high-molecular-weight penicillin-binding protein in Caulobacter crescentus. J Bacteriol. 1988 May;170(5):2319–2327. doi: 10.1128/jb.170.5.2319-2327.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Newton A., Ohta N. Regulation of the cell division cycle and differentiation in bacteria. Annu Rev Microbiol. 1990;44:689–719. doi: 10.1146/annurev.mi.44.100190.003353. [DOI] [PubMed] [Google Scholar]
- Ohta N., Lane T., Ninfa E. G., Sommer J. M., Newton A. A histidine protein kinase homologue required for regulation of bacterial cell division and differentiation. Proc Natl Acad Sci U S A. 1992 Nov 1;89(21):10297–10301. doi: 10.1073/pnas.89.21.10297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oliver D. B., Beckwith J. E. coli mutant pleiotropically defective in the export of secreted proteins. Cell. 1981 Sep;25(3):765–772. doi: 10.1016/0092-8674(81)90184-7. [DOI] [PubMed] [Google Scholar]
- Oliver D. B. SecA protein: autoregulated ATPase catalysing preprotein insertion and translocation across the Escherichia coli inner membrane. Mol Microbiol. 1993 Jan;7(2):159–165. doi: 10.1111/j.1365-2958.1993.tb01107.x. [DOI] [PubMed] [Google Scholar]
- Osley M. A., Newton A. Mutational analysis of developmental control in Caulobacter crescentus. Proc Natl Acad Sci U S A. 1977 Jan;74(1):124–128. doi: 10.1073/pnas.74.1.124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Osley M. A., Newton A. Temporal control of the cell cycle in Caulobacter crescentus: roles of DNA chain elongation and completion. J Mol Biol. 1980 Mar 25;138(1):109–128. doi: 10.1016/s0022-2836(80)80007-6. [DOI] [PubMed] [Google Scholar]
- POINDEXTER J. S. BIOLOGICAL PROPERTIES AND CLASSIFICATION OF THE CAULOBACTER GROUP. Bacteriol Rev. 1964 Sep;28:231–295. doi: 10.1128/br.28.3.231-295.1964. [DOI] [PMC free article] [PubMed] [Google Scholar]
- STOVEPOINDEXTER J. L., COHEN-BAZIRE G. THE FINE STRUCTURE OF STALKED BACTERIA BELONGING TO THE FAMILY CAULOBACTERACEAE. J Cell Biol. 1964 Dec;23:587–607. doi: 10.1083/jcb.23.3.587. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sadaie Y., Kada T. Bacillus subtilis gene involved in cell division, sporulation, and exoenzyme secretion. J Bacteriol. 1985 Aug;163(2):648–653. doi: 10.1128/jb.163.2.648-653.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sadaie Y., Takamatsu H., Nakamura K., Yamane K. Sequencing reveals similarity of the wild-type div+ gene of Bacillus subtilis to the Escherichia coli secA gene. Gene. 1991 Feb 1;98(1):101–105. doi: 10.1016/0378-1119(91)90110-w. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmidt J. M., Stanier R. Y. The development of cellular stalks in bacteria. J Cell Biol. 1966 Mar;28(3):423–436. doi: 10.1083/jcb.28.3.423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shapiro L., Maizel J. V., Jr Synthesis and structure of Caulobacter crescentus flagella. J Bacteriol. 1973 Jan;113(1):478–485. doi: 10.1128/jb.113.1.478-485.1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sommer J. M., Newton A. Turning off flagellum rotation requires the pleiotropic gene pleD: pleA, pleC, and pleD define two morphogenic pathways in Caulobacter crescentus. J Bacteriol. 1989 Jan;171(1):392–401. doi: 10.1128/jb.171.1.392-401.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stephens C. M., Shapiro L. An unusual promoter controls cell-cycle regulation and dependence on DNA replication of the Caulobacter fliLM early flagellar operon. Mol Microbiol. 1993 Sep;9(6):1169–1179. doi: 10.1111/j.1365-2958.1993.tb01246.x. [DOI] [PubMed] [Google Scholar]
- Stoker N. G., Broome-Smith J. K., Edelman A., Spratt B. G. Organization and subcloning of the dacA-rodA-pbpA cluster of cell shape genes in Escherichia coli. J Bacteriol. 1983 Aug;155(2):847–853. doi: 10.1128/jb.155.2.847-853.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang S. P., Sharma P. L., Schoenlein P. V., Ely B. A histidine protein kinase is involved in polar organelle development in Caulobacter crescentus. Proc Natl Acad Sci U S A. 1993 Jan 15;90(2):630–634. doi: 10.1073/pnas.90.2.630. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waxman D. J., Strominger J. L. Penicillin-binding proteins and the mechanism of action of beta-lactam antibiotics. Annu Rev Biochem. 1983;52:825–869. doi: 10.1146/annurev.bi.52.070183.004141. [DOI] [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]