Abstract
The kil-kor regulon was first identified on the broad-host-range IncP alpha plasmid RK2 by the presence of multiple kil loci (kilA, kilB, kilC, and recently kilE) that are lethal to Escherichia coli host cells in the absence of regulation by kor functions in various combinations. Whereas the kilB operon is required for mating-pair formation during conjugation, the functions encoded by the other kil loci are not known. They are not essential for replication or conjugal transfer, but their coregulation with replication and transfer genes indicates that they are likely to be important for RK2. In this report, we describe molecular and genetic studies on kilC. We determined the nucleotide sequence of the kilC region, which is located between the origin of vegetative replication (oriV) and transposon Tn1 on RK2. Primer extension analysis identified the transcriptional start site and showed that a sequence corresponding to a strong sigma 70 promoter is functional. The abundance of RNA initiated from the kilC promoter is reduced in the presence of korA and korC, as predicted from genetic analysis of kilC regulation. The first gene of the kilC operon (klcA) is sufficient to express the host-lethal phenotype of the kilC determinant in the absence of korA and korC. By comparing RK2 to the related IncP alpha plasmids pUZ8 and R995, we determined that the Tn1 transposon in RK2 interrupts a gene (klcB) immediately downstream of klcA. Thus, the kilC determinant is normally part of an autoregulated operon of three genes: klcA, klcB, and korC. klcA is predicted to encode a 15,856-Da polypeptide that is related to the ArdB antirestriction protein of the IncN plasmid pKM101, suggesting a role for klcA in the broad host ranges of IncP alpha plasmids. The predicted product of the uninterrupted klcB gene is a polypeptide of 51,133 Da that contains a segment with significant similarity to the RK2 regulatory proteins KorA and TrbA. Located 145 bp upstream of the kilC promoter is a 10th copy of the 17-bp oriV iteron sequence in inverted orientation relative to that of the other nine iterons of oriV. Iteron 10 is identical to the "orphan" iteron 1, and both have identical 6-bp flanking sequences that make them likely to be strong binding sites for the TrfA replication initiator protein. The locations and relative orientation of orphan iterons 10 and 1 raise the possibility that these iterons promote the formation of a DNA loop via protein-protein interactions by bound TrfA and lead us to propose that they demarcate the functional origin of replication. This analysis of the kilC region and our previous studies on the other kil loci of RK2 have revealed that the region between oriV and the korABF operon in wild-type IncP alpha plasmids is saturated by the kilC, kilE, and kilA loci arranged in four kor-regulated operons encoding a total of 12 genes.
Full text
PDF







Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ayres E. K., Saadi S., Schreiner H. C., Thomson V. J., Figurski D. H. Differentiation of lethal and nonlethal, kor-regulated functions in the kilB region of broad host-range plasmid RK2. Plasmid. 1991 Jan;25(1):53–63. doi: 10.1016/0147-619x(91)90006-i. [DOI] [PubMed] [Google Scholar]
- Ayres E. K., Thomson V. J., Merino G., Balderes D., Figurski D. H. Precise deletions in large bacterial genomes by vector-mediated excision (VEX). The trfA gene of promiscuous plasmid RK2 is essential for replication in several gram-negative hosts. J Mol Biol. 1993 Mar 5;230(1):174–185. doi: 10.1006/jmbi.1993.1134. [DOI] [PubMed] [Google Scholar]
- Bechhofer D. H., Figurski D. H. Map location and nucleotide sequence of korA, a key regulatory gene of promiscuous plasmid RK2. Nucleic Acids Res. 1983 Nov 11;11(21):7453–7469. doi: 10.1093/nar/11.21.7453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belogurov A. A., Delver E. P., Rodzevich O. V. IncN plasmid pKM101 and IncI1 plasmid ColIb-P9 encode homologous antirestriction proteins in their leading regions. J Bacteriol. 1992 Aug;174(15):5079–5085. doi: 10.1128/jb.174.15.5079-5085.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belogurov A. A., Delver E. P., Rodzevich O. V. Plasmid pKM101 encodes two nonhomologous antirestriction proteins (ArdA and ArdB) whose expression is controlled by homologous regulatory sequences. J Bacteriol. 1993 Aug;175(15):4843–4850. doi: 10.1128/jb.175.15.4843-4850.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chattoraj D. K., Mason R. J., Wickner S. H. Mini-P1 plasmid replication: the autoregulation-sequestration paradox. Cell. 1988 Feb 26;52(4):551–557. doi: 10.1016/0092-8674(88)90468-0. [DOI] [PubMed] [Google Scholar]
- Cohen S. N., Chang A. C., Hsu L. Nonchromosomal antibiotic resistance in bacteria: genetic transformation of Escherichia coli by R-factor DNA. Proc Natl Acad Sci U S A. 1972 Aug;69(8):2110–2114. doi: 10.1073/pnas.69.8.2110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Datta N., Hedges R. W., Shaw E. J., Sykes R. B., Richmond M. H. Properties of an R factor from Pseudomonas aeruginosa. J Bacteriol. 1971 Dec;108(3):1244–1249. doi: 10.1128/jb.108.3.1244-1249.1971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Delver E. P., Kotova V. U., Zavilgelsky G. B., Belogurov A. A. Nucleotide sequence of the gene (ard) encoding the antirestriction protein of plasmid colIb-P9. J Bacteriol. 1991 Sep;173(18):5887–5892. doi: 10.1128/jb.173.18.5887-5892.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Devereux J., Haeberli P., Smithies O. A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 1):387–395. doi: 10.1093/nar/12.1part1.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dunn J. J., Elzinga M., Mark K. K., Studier F. W. Amino acid sequence of the gene 0.3 protein of bacteriophage T7 and nucleotide sequence of its mRNA. J Biol Chem. 1981 Mar 10;256(5):2579–2585. [PubMed] [Google Scholar]
- Eckdahl T. T., Anderson J. N. Conserved DNA structures in origins of replication. Nucleic Acids Res. 1990 Mar 25;18(6):1609–1612. doi: 10.1093/nar/18.6.1609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Figurski D. H., Pohlman R. F., Bechhofer D. H., Prince A. S., Kelton C. A. Broad host range plasmid RK2 encodes multiple kil genes potentially lethal to Escherichia coli host cells. Proc Natl Acad Sci U S A. 1982 Mar;79(6):1935–1939. doi: 10.1073/pnas.79.6.1935. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Figurski D. H., Young C., Schreiner H. C., Pohlman R. F., Bechhofer D. H., Prince A. S., D'Amico T. F. Genetic interactions of broad host-range plasmid RK2: evidence for a complex replication regulon. Basic Life Sci. 1985;30:227–241. doi: 10.1007/978-1-4613-2447-8_19. [DOI] [PubMed] [Google Scholar]
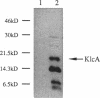
- Goncharoff P., Saadi S., Chang C. H., Saltman L. H., Figurski D. H. Structural, molecular, and genetic analysis of the kilA operon of broad-host-range plasmid RK2. J Bacteriol. 1991 Jun;173(11):3463–3477. doi: 10.1128/jb.173.11.3463-3477.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hanahan D. Studies on transformation of Escherichia coli with plasmids. J Mol Biol. 1983 Jun 5;166(4):557–580. doi: 10.1016/s0022-2836(83)80284-8. [DOI] [PubMed] [Google Scholar]
- Harley C. B., Reynolds R. P. Analysis of E. coli promoter sequences. Nucleic Acids Res. 1987 Mar 11;15(5):2343–2361. doi: 10.1093/nar/15.5.2343. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harrington R. E. DNA curving and bending in protein-DNA recognition. Mol Microbiol. 1992 Sep;6(18):2549–2555. doi: 10.1111/j.1365-2958.1992.tb01431.x. [DOI] [PubMed] [Google Scholar]
- Hawley D. K., McClure W. R. Compilation and analysis of Escherichia coli promoter DNA sequences. Nucleic Acids Res. 1983 Apr 25;11(8):2237–2255. doi: 10.1093/nar/11.8.2237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hedges R. W., Matthew M. Acquisition by Escherichia coli of plasmid-borne beta-lactamases normally confined to Pseudomonas spp. Plasmid. 1979 Apr;2(2):269–278. doi: 10.1016/0147-619x(79)90045-3. [DOI] [PubMed] [Google Scholar]
- Hershfield V., Boyer H. W., Yanofsky C., Lovett M. A., Helinski D. R. Plasmid ColEl as a molecular vehicle for cloning and amplification of DNA. Proc Natl Acad Sci U S A. 1974 Sep;71(9):3455–3459. doi: 10.1073/pnas.71.9.3455. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ingram L. C., Richmond M. H., Sykes R. B. Molecular characterization of the R factors implicated in the carbenicillin resistance of a sequence of Pseudomonas aeruginosa strains isolated from burns. Antimicrob Agents Chemother. 1973 Feb;3(2):279–288. doi: 10.1128/aac.3.2.279. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jagura-Burdzy G., Khanim F., Smith C. A., Thomas C. M. Crosstalk between plasmid vegetative replication and conjugative transfer: repression of the trfA operon by trbA of broad host range plasmid RK2. Nucleic Acids Res. 1992 Aug 11;20(15):3939–3944. doi: 10.1093/nar/20.15.3939. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kornacki J. A., Burlage R. S., Figurski D. H. The kil-kor regulon of broad-host-range plasmid RK2: nucleotide sequence, polypeptide product, and expression of regulatory gene korC. J Bacteriol. 1990 Jun;172(6):3040–3050. doi: 10.1128/jb.172.6.3040-3050.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kornacki J. A., Chang C. H., Figurski D. H. kil-kor regulon of promiscuous plasmid RK2: structure, products, and regulation of two operons that constitute the kilE locus. J Bacteriol. 1993 Aug;175(16):5078–5090. doi: 10.1128/jb.175.16.5078-5090.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lessl M., Balzer D., Lurz R., Waters V. L., Guiney D. G., Lanka E. Dissection of IncP conjugative plasmid transfer: definition of the transfer region Tra2 by mobilization of the Tra1 region in trans. J Bacteriol. 1992 Apr;174(8):2493–2500. doi: 10.1128/jb.174.8.2493-2500.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- Miron A., Mukherjee S., Bastia D. Activation of distant replication origins in vivo by DNA looping as revealed by a novel mutant form of an initiator protein defective in cooperativity at a distance. EMBO J. 1992 Mar;11(3):1205–1216. doi: 10.1002/j.1460-2075.1992.tb05161.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Motallebi-Veshareh M., Balzer D., Lanka E., Jagura-Burdzy G., Thomas C. M. Conjugative transfer functions of broad-host-range plasmid RK2 are coregulated with vegetative replication. Mol Microbiol. 1992 Apr;6(7):907–920. doi: 10.1111/j.1365-2958.1992.tb01541.x. [DOI] [PubMed] [Google Scholar]
- Mukherjee S., Erickson H., Bastia D. Enhancer-origin interaction in plasmid R6K involves a DNA loop mediated by initiator protein. Cell. 1988 Feb 12;52(3):375–383. doi: 10.1016/s0092-8674(88)80030-8. [DOI] [PubMed] [Google Scholar]
- Mulligan M. E., Hawley D. K., Entriken R., McClure W. R. Escherichia coli promoter sequences predict in vitro RNA polymerase selectivity. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 2):789–800. doi: 10.1093/nar/12.1part2.789. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olsen R. H., Shipley P. Host range and properties of the Pseudomonas aeruginosa R factor R1822. J Bacteriol. 1973 Feb;113(2):772–780. doi: 10.1128/jb.113.2.772-780.1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pal S. K., Chattoraj D. K. P1 plasmid replication: initiator sequestration is inadequate to explain control by initiator-binding sites. J Bacteriol. 1988 Aug;170(8):3554–3560. doi: 10.1128/jb.170.8.3554-3560.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pansegrau W., Lanka E., Barth P. T., Figurski D. H., Guiney D. G., Haas D., Helinski D. R., Schwab H., Stanisich V. A., Thomas C. M. Complete nucleotide sequence of Birmingham IncP alpha plasmids. Compilation and comparative analysis. J Mol Biol. 1994 Jun 24;239(5):623–663. doi: 10.1006/jmbi.1994.1404. [DOI] [PubMed] [Google Scholar]
- Perri S., Helinski D. R. DNA sequence requirements for interaction of the RK2 replication initiation protein with plasmid origin repeats. J Biol Chem. 1993 Feb 15;268(5):3662–3669. [PubMed] [Google Scholar]
- Pinkney M., Diaz R., Lanka E., Thomas C. M. Replication of mini RK2 plasmid in extracts of Escherichia coli requires plasmid-encoded protein TrfA and host-encoded proteins DnaA, B, G DNA gyrase and DNA polymerase III. J Mol Biol. 1988 Oct 20;203(4):927–938. doi: 10.1016/0022-2836(88)90118-0. [DOI] [PubMed] [Google Scholar]
- Prentki P., Krisch H. M. In vitro insertional mutagenesis with a selectable DNA fragment. Gene. 1984 Sep;29(3):303–313. doi: 10.1016/0378-1119(84)90059-3. [DOI] [PubMed] [Google Scholar]
- Read T. D., Thomas A. T., Wilkins B. M. Evasion of type I and type II DNA restriction systems by IncI1 plasmid CoIIb-P9 during transfer by bacterial conjugation. Mol Microbiol. 1992 Jul;6(14):1933–1941. doi: 10.1111/j.1365-2958.1992.tb01366.x. [DOI] [PubMed] [Google Scholar]
- Saiki R. K., Gelfand D. H., Stoffel S., Scharf S. J., Higuchi R., Horn G. T., Mullis K. B., Erlich H. A. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science. 1988 Jan 29;239(4839):487–491. doi: 10.1126/science.2448875. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shine J., Dalgarno L. Determinant of cistron specificity in bacterial ribosomes. Nature. 1975 Mar 6;254(5495):34–38. doi: 10.1038/254034a0. [DOI] [PubMed] [Google Scholar]
- Shine J., Dalgarno L. The 3'-terminal sequence of Escherichia coli 16S ribosomal RNA: complementarity to nonsense triplets and ribosome binding sites. Proc Natl Acad Sci U S A. 1974 Apr;71(4):1342–1346. doi: 10.1073/pnas.71.4.1342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stalker D. M., Thomas C. M., Helinski D. R. Nucleotide sequence of the region of the origin of replication of the broad host range plasmid RK2. Mol Gen Genet. 1981;181(1):8–12. doi: 10.1007/BF00338997. [DOI] [PubMed] [Google Scholar]
- Studier F. W., Moffatt B. A. Use of bacteriophage T7 RNA polymerase to direct selective high-level expression of cloned genes. J Mol Biol. 1986 May 5;189(1):113–130. doi: 10.1016/0022-2836(86)90385-2. [DOI] [PubMed] [Google Scholar]
- Thomas C. M., Ibbotson J. P., Wang N. Y., Smith C. A., Tipping R., Loader N. M. Gene regulation on broad host range plasmid RK2: identification of three novel operons whose transcription is repressed by both KorA and KorC. Nucleic Acids Res. 1988 Jun 24;16(12):5345–5359. doi: 10.1093/nar/16.12.5345. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas C. M., Stalker D. M., Helinski D. R. Replication and incompatibility properties of segments of the origin region of replication of the broad host range plasmid RK2. Mol Gen Genet. 1981;181(1):1–7. doi: 10.1007/BF00338996. [DOI] [PubMed] [Google Scholar]
- Thomson V. J., Jovanovic O. S., Pohlman R. F., Chang C. H., Figurski D. H. Structure, function, and regulation of the kilB locus of promiscuous plasmid RK2. J Bacteriol. 1993 Apr;175(8):2423–2435. doi: 10.1128/jb.175.8.2423-2435.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- VanWye J. D., Bronson E. C., Anderson J. N. Species-specific patterns of DNA bending and sequence. Nucleic Acids Res. 1991 Oct 11;19(19):5253–5261. doi: 10.1093/nar/19.19.5253. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Villarroel R., Hedges R. W., Maenhaut R., Leemans J., Engler G., Van Montagu M., Schell J. Heteroduplex analysis of P-plasmid evolution: the role of insertion and deletion of transposable elements. Mol Gen Genet. 1983;189(3):390–399. doi: 10.1007/BF00325900. [DOI] [PubMed] [Google Scholar]
- Waters S. H., Rogowsky P., Grinsted J., Altenbuchner J., Schmitt R. The tetracycline resistance determinants of RP1 and Tn1721: nucleotide sequence analysis. Nucleic Acids Res. 1983 Sep 10;11(17):6089–6105. doi: 10.1093/nar/11.17.6089. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]
- York D., Filutowicz M. Autoregulation-deficient mutant of the plasmid R6K-encoded pi protein distinguishes between palindromic and nonpalindromic binding sites. J Biol Chem. 1993 Oct 15;268(29):21854–21861. [PubMed] [Google Scholar]
- Young C., Bechhofer D. H., Figurski D. H. Gene regulation in plasmid RK2: positive control by korA in the expression of korC. J Bacteriol. 1984 Jan;157(1):247–252. doi: 10.1128/jb.157.1.247-252.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young C., Burlage R. S., Figurski D. H. Control of the kilA gene of the broad-host-range plasmid RK2: involvement of korA, korB, and a new gene, korE. J Bacteriol. 1987 Mar;169(3):1315–1320. doi: 10.1128/jb.169.3.1315-1320.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]