Abstract
Background
The endemic seagrass Posidonia oceanica (L.) Delile colonizes soft bottoms producing highly productive meadows that play a crucial role in coastal ecosystems dynamics. Human activities and natural events are responsible for a widespread meadows regression; to date the identification of "diagnostic" tools to monitor conservation status is a critical issue. In this study the feasibility of a novel tool to evaluate ecological impacts on Posidonia meadows has been tested. Quantification of a putative stress indicator, i.e. phenols content, has been coupled to 2-D electrophoretic protein analysis of rhizome samples.
Results
The overall expression pattern from Posidonia rhizome was determined using a preliminary proteomic approach, 437 protein spots were characterized by pI and molecular weight. We found that protein expression differs in samples belonging to sites with high or low phenols: 22 unique protein spots are peculiar of "low phenols" and 27 other spots characterize "high phenols" samples.
Conclusion
Posidonia showed phenols variations within the meadow, that probably reflect the heterogeneity of environmental pressures. In addition, comparison of the 2-D electrophoresis patterns allowed to highlight qualitative protein expression differences in response to these pressures. These differences may account for changes in metabolic/physiological pathways as adaptation to stress. A combined approach, based on phenols content determination and 2-D electrophoresis protein pattern, seems a promising tool to monitor Posidonia meadows health state.
Background
Posidonia oceanica (L.) Delile (fig. 1) is a Mediterranean endemism. Plants colonize soft bottoms producing large meadows that span from the sea surface to 35–40 m depth. Meadows are highly productive ecosystems, as they produce high amount of oxygen and organic compounds, sustain complex food nets, act as a nursery/refuge for several species. They also play a crucial role in coastal preservation, by stabilizing sediments and reducing hydrodynamics effects (see fig. 1) [1,2].
Figure 1.
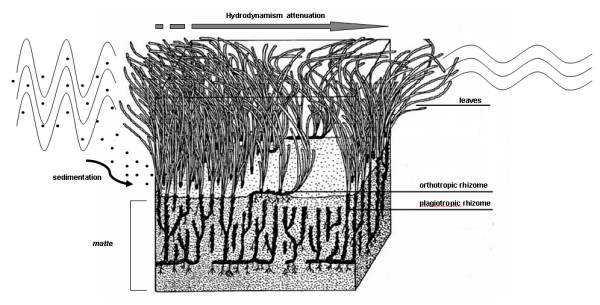
Schematic representation of Posidonia meadow (matte, rhizomes and leaves) and its effect on sediment stabilization and reduction of hydrodynamism (modified from Boudouresque and Meinesz [55]).
Many human activities and natural events are responsible for the widespread meadows regression, such as modified hydrogeological regime and littoral transport [3-6], pollution [7-11], aquaculture [12-15], trawling [16,17], anchorages [18-21], placing of cable/pipes or damping [22,23], in addition to grazing, sea storms, climatic changes, etc. [24-26]. All lead to alterations of Posidonia ecosystems.
Phenolic compounds are widespread secondary metabolites in plants. They play a role in herbivore/pathogen protection [27,28] and are considered stress indicators in terrestrial plants [29-38]. Phenols compounds have been identified in marine phanerogames [39-41] and high concentrations of phenolic compounds in Posidonia leaves were found in a few cases a) under competition with Caulerpa taxifolia [42,43], b) under mercury contamination [44], c) nearby offshore aquaculture cages [45]. An attempt to identify specific phenolic compounds in Posidonia leaves in response to different environmental pressures did not give clear-cut results [46]. This may be due to the fact that Posidonia leaves are temporary structures with a relatively short lifespan (about 7 months) at the Mediterranean mid-latitudes. Therefore they show marked seasonal fluctuations in common physiological processes – including synthesis and accumulation of phenolic compounds [47]. Rhizomes have a lifespan much longer than leaves: consequently, they undergo less marked fluctuations and may carry the memory of experienced environmental pressures.
The 2-D electrophoresis protein analysis produces maps of all the expressed proteins, in a given time and under a specific environmental condition. The protein pattern is a dynamic entity varying from cell to cell in the same organism, it is constantly modulated by external and internal signalling and reflects changes in the physiological state. The proteomic approach, based on the simultaneous separation of hundreds of proteins in the same 2D-electrophoretic gel, represents a powerful tool to monitor the "health state" of ecosystems, by comparing quantitative/qualitative pattern differences of protein expression in organisms living in polluted/non-polluted areas.
In this work we choose the rhizome, in particular the basal section, as the most reliable plant portion to evaluate possible alterations of both phenols content and protein expression.
The aim of this study was to verify the feasibility of phenols quantification coupled to 2-D electrophoretic protein analysis in rhizomes, as a novel "diagnostic" tool to monitor Posidonia meadows conservation status.
Results and discussion
Phenols content
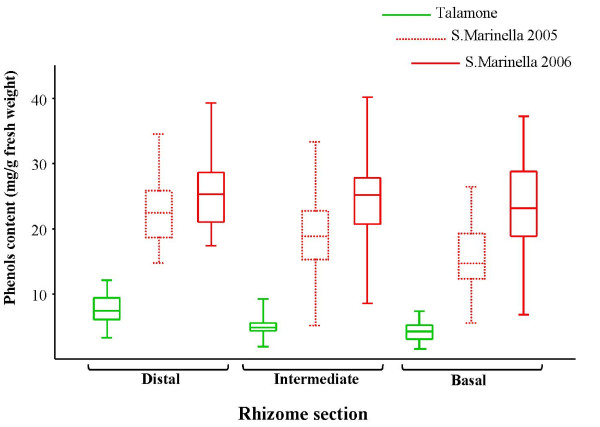
Total phenols were measured in distal, intermediate and basal sections of sampled rhizomes. Mean value (mg/g fresh weight) of the distal sections was 25.6 (n = 60; SE 1.1) ranging from 18.9 to 35, intermediate sections mean was 25.1 (n = 60; SE 1.1) ranging from 15.3 to 36.7 and basal sections mean was 23.7 (n = 60; SE 1.4) from 7.1 to 35.2.
Data of total phenols content in samples collected in 2006 from the S. Marinella meadow were compared to data previously obtained with identical experimental procedure from rhizomes of the same meadow collected in 2005 and from rhizomes of the Talamone meadow (Grosseto, Italy, 2002). Samples from the well preserved meadow of Talamone [54] showed the lowest and less scattered phenols values, samples from the S. Marinella meadow (year 2006) showed highly scattered values (Fig. 2). We found that the overall phenols content in S. Marinella-2005 samples was lower than the content detected in 2006 samples, however in both cases values were higher than the ones obtained from the Talamone samples (compare box-plots in Fig.2).
Figure 2.
Distribution of phenols content in rhizome distal, intermediate and basal sections. Phenols values (mg/g fresh weight) are represented as box-plots: the box contains 50% data (the extremes of that box are the Q1 and Q3, 1st and 3rd quartiles), the internal horizontal segments represent median of the distributions (Q2 value, 2nd quartile), 'whiskers' range from the lowest to the highest value. The box plots from S. Marinella meadow samples collected in 2006 (red, solid line) and 2005 (red, dotted line), and from Talamone meadow samples (green) are reported.
Phenols content differences between Talamone and S. Marinella-2006 samples are statistically significant for all the sections (Mann-Whitney, p << 0.001), differences between S. Marinella-2005 and S. Marinella-2006 samples are highly significant for basal and intermediate sections (Mann-Whitney, p << 0.001), significant for the distal section (Mann-Whitney, p = 0.00146). The lowest phenols content may be consistently associated with the good health state of the Talamone meadow [53].
In this study on the S. Marinella meadow the three lowest phenols contents found in rhizome basal sections were: 7.1 mg/g ± 0.5, 14.4 mg/g ± 0.2 and 14.5 mg/g ± 0.8; the three highest were: 28.9 mg/g ± 0.2, 33.6 mg/g ± 2.0 and 35.2 mg/g ± 2.8. Posidonia shoots from these six sampling sites were chosen for protein analyses.
Protein analysis
In order to assess a possible match between different phenols content and variations in protein expression, at first we determined the overall expression pattern of Posidonia rhizome by 2-D electrophoresis. The polypeptides falling within the experimental window of pI 3–9 and 12–200 kDa and sufficiently abundant to be detected by the silver staining procedure were taken into account. Protein patterns from low and high phenols samples were combined and the experimental values of pI and molecular weight for each isoelectric spot were calculated by a dedicated computer software using reference proteins with known pI and molecular weight, commonly called "anchors" (see Materials and Methods section). Proteins, accounting for a total of 437 spots, ranged from pI 5.11 to pI 8.66 with an apparent molecular weight ranging from 13306 Da to 95563 Da (see additional file 1).
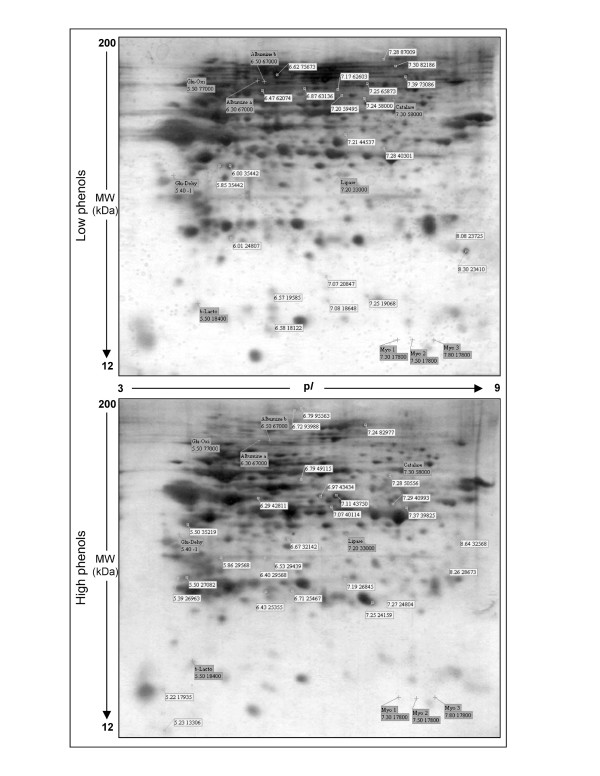
Two representative 2-D gels from low and high phenols are shown in Fig. 3. Computer-assisted cross-comparison revealed qualitative differences that accounted for differentially expressed proteins: 22 spots are peculiar of low phenols whereas 27 spots characterize high phenols samples, accounting for 5.0 and 6.4% of the entire map, respectively. These differences were consistently found in all the examined samples.
Figure 3.
Representative 2-DE patterns of Posidonia rhizome proteins. Upper panel, proteins isolated from low phenols samples. Lower panel, proteins isolated from high phenols samples. About 15 μg of proteins from dry powder of Posidonia rhizome were separated on IPG gel strip (7 cm, 3–10 NL) followed by SDS-PAGE on a vertical mini-gel (12%T). Peculiar protein spots are labelled, pI and molecular weight values are indicated (white background). Positions of protein markers are indicated and labelled with name and pI/molecular weight values (gray background). Protein markers (in alphabetic order) are: Albumine a; Albumine b; Catalase; Glucose-1-Dehydronase (Glu-Dehy); Glucose oxidase (Glu-Oxi); β-Lactoglobuline (b-Lacto); Myoglobine subunits (Myo 1, Myo 2, Myo 3). Numbers on the left refer to the position of the molecular weight standards and numbers in the middle indicate the pI range.
At low phenols content, differentially expressed proteins ranged from pI 5.85 to pI 8.30, with an apparent molecular weight ranging from 18122 Da to 87009 Da. In high phenols samples, isoelectric point of differentially expressed proteins was comprised between pI 5.22 and pI 8.64, with an apparent molecular weight from 13306 Da to 95563 Da (Table 1).
Table 1.
Peculiar protein spots identified in the 2-D electrophoretic maps of low phenols (22 spots) and high phenols (27 spots) Posidonia rhizome samples
| Low phenols | High phenols | ||
| pI | MW (Da) | pI | MW (Da) |
| 5.85 | 35442 | 5.22 | 17935 |
| 6.00 | 35442 | 5.23 | 13306 |
| 6.01 | 24807 | 5.39 | 26963 |
| 6.47 | 62074 | 5.50 | 35219 |
| 6.57 | 19585 | 5.50 | 27082 |
| 6.58 | 18122 | 5.50 | 28926 |
| 6.62 | 75673 | 5.86 | 29568 |
| 6.87 | 63136 | 6.29 | 42811 |
| 7.07 | 20847 | 6.40 | 29568 |
| 7.08 | 18648 | 6.43 | 25355 |
| 7.17 | 62603 | 6.53 | 29439 |
| 7.20 | 59495 | 6.67 | 32142 |
| 7.21 | 44537 | 6.71 | 25467 |
| 7.24 | 58000 | 6.72 | 93988 |
| 7.25 | 65873 | 6.79 | 95563 |
| 7.25 | 19068 | 6.79 | 49115 |
| 7.28 | 87009 | 6.97 | 43434 |
| 7.28 | 40301 | 7.07 | 40114 |
| 7.30 | 82186 | 7.11 | 43750 |
| 7.39 | 73086 | 7.19 | 26845 |
| 8.08 | 23725 | 7.24 | 82977 |
| 8.30 | 23410 | 7.25 | 24159 |
| 7.27 | 24804 | ||
| 7.28 | 50556 | ||
| 7.29 | 40993 | ||
| 7.37 | 39825 | ||
| 8.26 | 28673 | ||
| 8.64 | 32568 | ||
The experimental values of pI and molecular weight (MW) for every isoelectric spot were calculated with ImageMaster 2D Platinum System.
Although an identity was not assigned to the differentially expressed polypeptides, they have been firstly characterized by assigning a molecular mass and a total charge.
Conclusion
It has been suggested that phenols content is an indicative trait of environmental stress in Posidonia oceanica [42-47,53]. In this work we choose the rhizome basal section as the most reliable material to measure this putative marker of ecosystem imbalance. The Posidonia shoots utilized in this study showed that variations in the phenols content exists within the examined meadow (S. Marinella, Italy), probably reflecting environmental pressures heterogeneity. Moreover, comparison of phenols content in the same meadow at one year time distance showed an increase in the overall values.
We have constructed the first 2-D electrophoretic map of Posidonia oceanica rhizome, made of 437 protein spots, characterized by pI and molecular weight.
Usually, the bi-dimensional protein pattern is typical of the physiological state and varies under different environmental conditions. Thus, the patterns comparison allows the highlighting of protein expression differences in response to stresses. Here we showed, by comparison of samples belonging to sites with low or high phenols content, that certain protein spots present in "low phenols" are absent in "high phenols" and vice versa. This may account for changes in metabolic/physiological pathways as adaptation to stress, including activation/repression of coordinate sets of genes. Differences cover the 5–6% of the entire protein map and match the plants phenols response. To the best of our knowledge, this is the first attempt to correlate the pattern of expressed proteins to a putative stress indicator, such as phenols content, in Posidonia rhizome.
Although the identification of each protein spot needs further investigation and might be hampered by the lack of extensive database information, the combined approach, based on phenols content determination and 2-D electrophoresis protein pattern, seems a promising tool to monitor Posidonia meadows health state.
Methods
Sampling, conservation and sample selection
Posidonia oceanica was sampled from the Santa Marinella meadow (Rome, Italy), Site of Community Importance (according to Habitat Directive 92/43/EEC), spanning from Capo Linaro to Santa Severa, for a 13.5 km coastline and covering a surface of 1,800 ha.
Shoots were sampled in May 2006 in 20 randomly chosen sites (depths from 7.5 to 13.5 m). At least 4 orthotropic shoots per sampling site were collected and maintained at 4°C in the dark until the arrival to the laboratory. Plants were first rinsed in 0.1 Triton-X (Sigma) and then in distilled water to remove epiphytes and contaminants.
At least 3 shoots per site were stored at -20°C until processing for phenol analysis, and at least 1 at -80°C for protein analyses. Total phenols concentration was determined in duplicate on three different rhizomes for sampling site according to Folin-Ciocalteau procedure [48], each shoot was dissected into three sections (about 1/3 of the total length): basal, intermediate and distal; about 125 mg fresh weight of each section were separately processed. Once established the sampling sites in which the three highest and lowest phenols values were found, shoots from these sampling sites were selected for protein analysis.
Protein extraction and electrophoresis
Protein were extracted according to the Wang et al. [49] protocol originally developed for recalcitrant plant tissues (leaves and flesh). This protocol, previously utilized for Posidonia leaves [50], was modified for Posidonia rhizomes.
Briefly, frozen rhizomes were peeled off to remove all cortical tissues, and a piece of the basal section (250 mg) was ground in liquid N2 using a mortar with pestle. The powdered tissue was subjected to phenols extraction in the presence of SDS, as described [49].
The protein pellet was dried and dissolved in 2-DE rehydration solution [8 M urea, 2 M thiourea, 2% CHAPS, 50 mM dithiothreitol (DTT), 0.2% carrier ampholytes (3–10 Bio-Lyte Ampholyte, Bio-Rad Laboratories)], supplemented with proteases and phosphatases inhibitors (Sigma). Protein content was measured by Bradford protein assay (Bio-Rad Laboratories) using Bovine Serum Albumine as standard.
Protein samples (15 μg) were applied in 155 μl of 2-DE rehydration solution to 7 cm Readystrip IPG (Immobilized pH Gradient, pH 3–10 NL, Bio-Rad Laboratories), by incubating overnight. The isoelectrofocusing (IEF) was performed at room temperature using the ZOOM IPG Runner™ Mini-cell (Invitrogen), applying 500 V for 4 hrs (1 mA/strip; 0.5 W/strip).
Focused strips were equilibrated using DTT and iodoacetamide solutions, positioned on a 12% acrylamide SDS-PAGE minigel 1 mm thick [51], according to standard procedures. After electrophoresis, resolved proteins were visualized by acidic silver staining that allows to detect as low as 0.5–1 ng of protein per spot [52]. Proteome pI markers were from SERVA Electrophoresis (Heidelberg). Each protein sample was subjected at least to 2 parallel runs of isolectrofocusing and second dimension electrophoretic separation to assess proteomic pattern reproducibility.
Image processing and data analysis
Silver-stained gels were digitised using a Trust EasyConnect 19200 scanner, generating 2.4 Mb images. The images were saved as Tiff format and imported into the ImageMaster 2-D Platinum software (Amersham, version 6.0). Spot selection was performed using default selection parameters [53]. For the attribution of isoelectric points and relative molecular masses we utilized as internal standard a mixture of 10 protein with known identities (see Fig. 3 and related legend).
Authors' contributions
LM and AG conceived and designed the study. AR sampled plants. AR and DR carried out the laboratory analyses, and analysed the data. NNA processed 2-DE images. LM and AG wrote the manuscript with help from AR and DR. All authors read and approved the final manuscript.
Supplementary Material
List of the 437 protein spots identified from Posidonia rhizome. The experimental values of pI and molecular weight (MW) for every isoelectric spot were calculated with ImageMaster 2D Platinum System. The percent of each spot with respect to the total spots volume is also reported.
Acknowledgments
Acknowledgements
The authors are grateful to E Fresi for the introduction to the fantastic world of Posidonia. Thanks also go to M Scardi e L Valiante for samples collection. We are also grateful to I Pucci-Minafra for her insightful comments. This work was supported by CNR/MiUR funds to AG; ECON grant by commitment of ENEL Produzione S.p.A. (n. 3000043814/2004) and 60% grant from University of Tor Vergata to LM.
Contributor Information
Luciana Migliore, Email: luciana.migliore@uniroma2.it.
Alice Rotini, Email: alice.rotini@virgilio.it.
Davide Randazzo, Email: bio_davide@libero.it.
Nadia N Albanese, Email: nadiaalbanese@yahoo.it.
Agata Giallongo, Email: giallongo@ibim.cnr.it.
References
- Mazzella L, Scipione B, Gambi MC, Fresi E, Buia MC, Russo F, De Maio R, Lorenti M, Rando A. Le praterie sommerse del Mediterraneo. Stazione Zoologica, Napoli, Pubblicazione a cura del laboratorio di Ecologia del Benthos; 1986. [Google Scholar]
- Arata P, Diviacco G. Importanza delle praterie di Posidonia oceanica nel sistema marino costiero edegli interventi per la loro salvaguardia. Acqua Aria. 1989;5:555–571. [Google Scholar]
- Astier JM. Impact des aménagements littoraux de la rade de Toulon, liés aux techniques d'endigage, sur les herbiers à Posidonia oceanica. In: Boudouresque CF, Jeudy De Grissac, A, Olivier J, editor. International Workshop on Posidonia oceanica beds. Vol. 1. France: GIS Posidonie publications; 1984. pp. 255–259. [Google Scholar]
- Blanc JJ, Jeudy De Grissac A. Réflexion géologique sur la régression des herbiers à Posidonies (départements du Var et des Bouches-du-Rhône) In: Boudouresque CF, Meinesz A, Fresi E, Gravez V, editor. Second international Workshop on Posidonia beds. Vol. 2. France: GIS Posidonie publications; 1989. pp. 273–285. [Google Scholar]
- Guidetti P, Fabiano M. The use of lepidochronology to asses the impact of terrigenous discharges on the primary leaf production of the Mediterranean seagrass Posidonia oceanica. Mar Poll Bull. 2000;40:449–453. doi: 10.1016/S0025-326X(99)00229-5. [DOI] [Google Scholar]
- Ruiz JM, Romero J. Effects of disturbances caused by coastal constructions on spatial structure, growth dynamics and Photosynthesis of the seagrass Posidonia oceanica. Mar Poll Bull. 2003;46:1523–1533. doi: 10.1016/j.marpolbul.2003.08.021. [DOI] [PubMed] [Google Scholar]
- Pérès JM, Picard J. Causes de la raréfaction et de la disparition des herbiers de Posidonia oceanica sur les côtes françaises de la Méditerranée. Aquat Bot. 1975;1:133–139. doi: 10.1016/0304-3770(75)90018-2. [DOI] [Google Scholar]
- Pérès JM. La régression des herbiers à Posidonia oceanica. In: Boudouresque CF, Meinesz A, Fresi E, Gravez V, editor. International Workshop on Posidonia oceanica beds. Vol. 1. France: GIS Posidonie publications; 1984. pp. 445–454. [Google Scholar]
- Bourcier M. Régression des herbiers à Posidonia oceanica (L.) Delile, à l'Est de Marseille, sous l'action conjuguée des activités humaines et des modifications climatiques. In: Boudouresque CF, Meinesz A, Fresi E, Gravez V, editor. International Workshop on Posidonia oceanica beds. Vol. 2. France: GIS Posidonie publications; 1989. pp. 287–293. [Google Scholar]
- Pergent-Martini C, Pergent G. Proceedings of the Second International conference on the Mediterranean coastal environment: 24–27 October 1995. Vol. 95. Terragona, Spain. MEDCOAST; 1995. Impact of a sewage treatment plant on the Posidonia oceanica meadow: Assessment criteria; pp. 1389–1399. [Google Scholar]
- Balestri E, Benedetti-Cecchi L, Lardicci C. Variability in patterns of growth and morphology of Posidonia oceanica exposed to urban and industrial wastes: contrasts with two reference locations. J Exp Mar Biol Ecol. 2004;308:1–21. doi: 10.1016/j.jembe.2004.01.015. [DOI] [Google Scholar]
- Pergent G, Mendez S, Pergent-Martini C, Pasqualini V. Preliminary data on the impact of fish farming facilities on Posidonia oceanica meadows in the Mediterranean. Oceanol Acta. 1999;22:95–107. doi: 10.1016/S0399-1784(99)80036-X. [DOI] [Google Scholar]
- Delgado O, Ruiz J, Perez M, Romero J, Ballesteros E. Effects of fish farming on seagrass (Posidonia oceanica) in a Mediterranean bay: seagrass decline after organic loading cessation. Oceanol Acta. 1999;22:109–117. doi: 10.1016/S0399-1784(99)80037-1. [DOI] [Google Scholar]
- Cancemi G, De Falco G, Pergent G. Impact of a fish farming facility on a Posidonia oceanica meadow. Biol Mar Medit. 2000;7:341–344. [Google Scholar]
- Ruiz JM, Perez M, Romero J. Effects of fish farm loadings on seagrass (Posidonia oceanica) distribution, growth and photosynthesis. Mar Poll Bull. 2001;42:749–760. doi: 10.1016/S0025-326X(00)00215-0. [DOI] [PubMed] [Google Scholar]
- Sànchez-Lizaso JL, Guillen-Nieto JE, Ramos-Espla AA. Theregression of Posidonia oceanica meadows in El Campello (SE Spain) Rapports et Procès-Verbaux des Réunion de la Commission internationationale pour l'Exploration scientifique de la Mèditerranèe. 1990;32:7. [Google Scholar]
- Ruiz JM, Gutièrrez Ortega JM, Garcìa Charton JA, Pèrez Ruzafa A. Spatial characterization of environmental impact by bottom trawling on meadows in artificial reef areas of the southeastern cost of Spain. Proceedings of the Seventh International Conference on Artificial Reefs 7th CARAH. 1999. pp. 664–674.
- Porcher M. Impact de mouillages forains sur le herbiers à Posidonia oceanica. In: Boudouresque, CF, De Grissac AJ, Oliver J Marseille, editor. International Workshop on Posidonia oceanica beds. France: GIS Posidonie publications; 1984. pp. 45–148. [Google Scholar]
- Garcia-Charton JA, Bayle JT, Sànchez-Lizaso JL, Chiesa P, Llaurdò F, Pérez C, Djian H. Respuesta de la pradera de Posidonia oceanica y su ictiofauna asociada al anclaje de embarcaciones en el parque Nacional de Port-Cros Francia. Publ Espec – Inst Esp Oceanogr. 1993;11:423–430. [Google Scholar]
- Francour P, Ganteaume A, Poulain M. Effects of boat anchoring in Posidonia oceanica seagrass beds in the Port-Cros National Park (north-western Mediterranean Sea) Aquat Cons. 1999;9:391–400. doi: 10.1002/(SICI)1099-0755(199907/08)9:4<391::AID-AQC356>3.0.CO;2-8. [DOI] [Google Scholar]
- Milazzo M, Badalamenti F, Seccherelli G, Chemello R. Boat anchoring on Posidonia oceanica beds in a marine protected area (Italy, western Mediterranean): effect of anchor types in different anchoring stages. J Exp Mar Biol Ecol. 2004;299:51–62. doi: 10.1016/j.jembe.2003.09.003. [DOI] [Google Scholar]
- Peirano A, Bianchi NC. Decline of the seagrass Posidonia oceanica in response to environmental disturbance: a simulation like approach off Liguria (NW Mediterranean Sea) Proceedings of 30th European marine biological Symposium; Southampton. 1995. pp. 87–95.
- Pasqualini V, Pergent-Martini C, Pergent G. Environmental impacts identification along corsican coasts (Mediterranean Sea) using image processing. Aquat Bot. 1999;65:311–320. doi: 10.1016/S0304-3770(99)00048-0. [DOI] [Google Scholar]
- Marbà N, Duarte CM, Cebrian J, Gallegos ME, Olesen B, Sand-Jensen K. Growth and population dynamics of Posidonia oceanica on the Spanish Mediterranean coast: elucidating seagrass decline. Mar Ecol Prog Ser. 1996;137:203–213. doi: 10.3354/meps137203. [DOI] [Google Scholar]
- Boudouresque CF, Giraud G, Thommeret J, Thommeret Y. First attempt at dating by 14C the undersea beds of dead Posidonia oceanica in the bay of Port-Man (Port-Cros, Var, France) Trav Sci Parc nation Port-Cros, France. 1980;6:239–242. [Google Scholar]
- Boudouresque CF, Bernard G, Bonhomme P, Charbonnel E, Diviacco G, Meinesz A, Pergent G, Pergent-Martini C, Ruitton S, Tunesi L. Préservation et conservation des herbiers à Posidonia oceanica. Accord RAMOGE publications: Marseille, France; 2006. [Google Scholar]
- Swain T. Secondary compounds as protective agents. Ann Rev Plant Physiol Plant Mol Biol. 1977;28:479–501. doi: 10.1146/annurev.pp.28.060177.002403. [DOI] [Google Scholar]
- Pisani JM, Distel RA. Inter and intraspecific variations in production of spines and phenols in Prosopis caldemia and Prosopis flexuosa. J Chem Ecol. 1998;24:23–36. doi: 10.1023/A:1022380627261. [DOI] [Google Scholar]
- Contour-Ansel D, Louguet P. Variations du taux de polyphenols dans les aiguilles d'épicéa (Picea abies), présentant différents degrés de dépérissement. Pollut Atmos. 1986;28:270–274. [Google Scholar]
- Dixon RA, Paiva NL. Stress-induced phenylpropanoid metabolism. Plant Cell. 1995;7:1085–1097. doi: 10.1105/tpc.7.7.1085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muzika RM. Terpenes and phenolics in response to nitrogen fertilization: a test of carbon/nutrient balance hypothesis. Chemoecology. 1993;4:3–7. doi: 10.1007/BF01245890. [DOI] [Google Scholar]
- Karolewski P, Giertych MJ. Influence of toxic metal ions on phenols in needles and roots and on root respiration of scots pine seedlings. Acta Soc Bot Pol. 1994;63:29–35. [Google Scholar]
- Peñuelas J, Estiarte M, Kimball BA, Idso SB, Pinter PJ, Wall GW, Garcia RL, Hansaker DJ, LaMorte RL, Hendrix DL. Variety of responses of plant phenolic concentration to CO2 enrichment. J Exp Bot. 1996;47:1463–1467. doi: 10.1093/jxb/47.9.1463. [DOI] [Google Scholar]
- Giertych MJ, Karolewski P. Phenolic compounds distribution along the length of Scots pine needles in a polluted and control environment and its connection with necroses formation. Acta Soc Bot Pol. 2000;69:127–130. [Google Scholar]
- Castells E, Roumet C, Peñuelas J, Roy J. Intraspecific variability of phenolic concentrations and their responses to elevated CO2 in two Mediterranean perennial grasses. Environ Exp Bot. 2002;47:205–216. doi: 10.1016/S0098-8472(01)00123-X. [DOI] [Google Scholar]
- Loponen J, Ossipov V, Lempa K, Haukioja E, Pihlaja K. Concentrations and among-compound correlations of individual phenolics in white birch leaves under air pollution stress. Chemosphere. 1998;37:1445–1456. doi: 10.1016/S0045-6535(98)00135-0. [DOI] [Google Scholar]
- Robles C, Greff S, Pasqualini V, Garzino S, Bousquet-Mélou A, Fernandez C, et al. Phenols and flavonoids in Aleppo pine needles as bioindicators of air pollution. J Environ Qual. 2003;32:2265–2271. doi: 10.2134/jeq2003.2265. [DOI] [PubMed] [Google Scholar]
- Rivero RM, Ruiz JM, Garcia PC, Lopez-Lefebre LR, Sanchez E, Romero L. Resistance to cold and heat stress: accumulation of phenolic compounds in tomato and watermelon plants. Plant Science. 2001;160:315–321. doi: 10.1016/S0168-9452(00)00395-2. [DOI] [PubMed] [Google Scholar]
- Carriello L, Zanetti L, De Stefano S. Posidonia ecosystem – V. Phenolic compounds from marine phanerogames,Cymodocea nodosa and Posidonia oceanica. Comp Biochem Physiol B: Biochem Molec Biol. 1979;62:159–161. doi: 10.1016/0305-0491(79)90304-3. [DOI] [Google Scholar]
- Zapata O, Mc Millan C. Phenolic acids in seagrasses. Aq Bot. 1979;7:307–317. doi: 10.1016/0304-3770(79)90032-9. [DOI] [Google Scholar]
- Cozza R, Chiappetta A, Petrarulo M, Salimonti A, Rende F, Bitonti M B, Innocenti M. Cytophysiological features of Posidonia oceanica as putative markers of environmental conditions. Chem Ecol. 2004;20:215–223. doi: 10.1080/02757540410001689777. [DOI] [Google Scholar]
- Cuny P, Serve L, Jupin H, Boudouresque CF. Water soluble phenolic compounds of the marine phanerogam Posidonia oceanica in a Mediterranean area colonised by the introduced chlorophyte Caulerpa taxifolia. Aq Bot. 1995;52:237–242. doi: 10.1016/0304-3770(95)00504-8. [DOI] [Google Scholar]
- Dumay O, Costa J, Desjobert JM, Pergent G. Variations in the concentration of phenolic compounds in the seagrass Posidoniaoceanica under conditions of competition. Phytochemistry. 2004;65:3211–3220. doi: 10.1016/j.phytochem.2004.09.003. [DOI] [PubMed] [Google Scholar]
- Ferrat L, Pergent-Martini C, Romeo M, Pergent G. Hydrosoluble phenolic compounds production in a Mediterranean seagrass according to mercury contamination. Gul Mex Sci. 2003;21:108. [Google Scholar]
- Cannac M, Ferrat L, Pergent-Martini C, Pergent G, Pasqualini V. Effects of fish farming on flavonoids in Posidonia oceanica. Sci Tot Environ. 2006;370:91–98. doi: 10.1016/j.scitotenv.2006.07.016. [DOI] [PubMed] [Google Scholar]
- Agostini S, Desjobert JM, Pergent G. Distribution of phenolic compounds in the seagrass Posidonia oceanica. Phytochemistry. 1998;48:611–617. doi: 10.1016/S0031-9422(97)01118-7. [DOI] [PubMed] [Google Scholar]
- Dumay O, Costa J, Desjobert JM, Pergent G. Variations in the concentration of phenolic compounds in the seagrass Posidonia oceanica under conditions of competition. Phytochemistry. 2004;65:3211–3220. doi: 10.1016/j.phytochem.2004.09.003. [DOI] [PubMed] [Google Scholar]
- Booker FL, Miller JE. Phenylpropanoid metabolim and phenolic composition of soybean (Glycine max L.) leaves following exposure to ozone. J Exp Bot. 1998;49:1191–1202. doi: 10.1093/jexbot/49.324.1191. [DOI] [Google Scholar]
- Wang W, Vignani R, Cresti M, Scali M. A universal and rapid protocol for protein extraction from recalcitrant plant tissues for proteomic analysis. Short communication Electrophoresis. 2006;27:2782–2786. doi: 10.1002/elps.200500722. [DOI] [PubMed] [Google Scholar]
- Bucalossi D, Leonzio C, Casini S, Fossi MC, Marsili L, Ancora S, Wang W, Scali M. Application of a suite of biomarkers in Posidonia oceanica (L.) delile to assess the ecotoxicological impact on the coastal environment. Short communication Mar Environ Res. 2006;62:s327–s331. doi: 10.1016/j.marenvres.2006.04.014. [DOI] [PubMed] [Google Scholar]
- Laemmli UK. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970;227:680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Switzer RCIII, Merrril CR, Shifrin S. A highly sensitive silver stain for detecting proteins and peptides in polyacrylamide gels. Anal Biochem. 1979;98:231–237. doi: 10.1016/0003-2697(79)90732-2. [DOI] [PubMed] [Google Scholar]
- Expasy http://www.expasy.org/melanie/
- Fresi E, Dolce T, Forni C, Lorenzi C, Migliore L, Rizzelli D, Scardi M. Proceedings of the Conference Le scienze naturali, economiche e giuridiche nello studio e per la gestione degli ambienti acquatici. Terrasini, Palermo. CONISMA-AIOL; 2004. La prateria di Posidonia oceanica (L.) Delile di Talamone (Grosseto, Italia): struttura e stato di salute. 18–22 October 2004. [Google Scholar]
- Boudouresque CF, Mainesz A. Découverte de l'herbier de Posidonie. Cahier Parc Nat Port-Cros, France. 1982;4:1–81. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
List of the 437 protein spots identified from Posidonia rhizome. The experimental values of pI and molecular weight (MW) for every isoelectric spot were calculated with ImageMaster 2D Platinum System. The percent of each spot with respect to the total spots volume is also reported.