Abstract
HLA-E-restricted T cell receptor αβ+ CD8+ cytolytic T lymphocytes (CTLs) exist as monoclonal expansions in the peripheral blood of some individuals. Here, we show that they recognize, with high avidity, peptides derived from the UL40 protein of different human cytomegalovirus (CMV) strains. Recognition results in the induction of cytotoxicity, IFN-γ production and cell proliferation. Autologous cells pulsed with CMV-derived peptides become susceptible to lysis by HLA-E-restricted CTLs and induce their proliferation. The high avidity for CMV-derived peptides may explain how these cells are generated in vivo and suggest their possible role in the host defenses against CMV, a virus that evolved various mechanisms to down-regulate classical HLA class I molecules, thus escaping detection by conventional CTLs.
CD8+ cytolytic T lymphocytes (CTLs) expressing T cell receptor (TCR) αβ generally recognize antigen peptides in association with classical MHC class I molecules (HLA-A, -B and -C, in humans) (1, 2). Limited information exists on the capability of nonclassical MHC class I molecules (HLA-E, -F, -G, and -H) to function as molecular target of T cell-mediated recognition (3). However, recent reports indicated that HLA-E may represent a novel restriction element for the human CTL (4–7). HLA-E is a class Ib molecule characterized by a limited polymorphism and a broad expression on different cell types (8). Under physiological conditions, HLA-E associates with peptides derived from the leader sequence of various HLA class I molecules (9, 10). HLA-E-binding peptides that derived either from viruses, including cytomegalovirus (CMV), Epstein–Barr virus (EBV), and influenza virus, or from heat shock proteins (hsp); e.g., hsp60 have also been identified (11–13). HLA-E-dependent presentation of a Mycobacterium tuberculosis-derived antigen to human CD8+ T cells has also been reported; however, the epitopes recognized have not been identified so far (7). In addition, it has been proposed that TCRVβ-derived peptides may be associated with HLA-E molecules expressed by CD4+ T cells, rendering these cells susceptible to CTL-mediated lysis (4). Various reports in mice suggest that Qa-1, i.e., the murine HLA-E homologue, may play a major role in the host response to infections. Thus, Qa-1-restricted CTL specific for Salmonella typhimurium or Lysteria monocytogenes have been identified (14–16). Importantly, the Qa-1-bound nonameric peptide from Salmonella typhimurium GroEL is highly conserved in various Gram bacteria, including Salmonella typhi, Escherichia coli, Yersinia enterocolitica, Klebsiella pnenumoniae, and Helicobacter pylori (14).
A CD8+ CTL subset, originally described in our laboratory (17–19), has recently been shown to recognize HLA-E (5, 20, 21). These cells display a broad cytolytic activity against various allogeneic natural killer (NK)-susceptible tumor cell lines (a function referred to as NK-like activity). Because of this function they were named NK cytotolytic T lymphocytes (NK-CTLs). NK-CTLs are characterized by a memory phenotype and by the surface expression of HLA class I-specific inhibitory NK receptors (iNKRs) (18, 19). A remarkable property of NK-CTL is the capability to undergo extensive proliferation in response to allogeneic cells in mixed lymphocytes culture (MLC) (20). Interestingly, both freshly isolated NK-CTL and MLC-derived NK-CTL populations are characterized by monoclonal TCR rearrangements. NK-CTLs have also been shown to lyse phytohemagglutinin (PHA)-induced blasts (or B-EBV cell lines) derived from most (but not all) HLA-unmatched donors. Remarkably, NK-CTLs failed to lyse autologous cells. This broad cytolytic activity could be, at least in part, explained by the finding that NK-CTL can recognize HLA-E bound to different peptides derived from the leader sequences of many HLA class I alleles. This concept was supported by the capability of NK-CTL of lysing the transporter associated with antigen processing (TAP)-2-deficient RMA-S murine cell line cotransfected with HLA-E*01033 and human β2-microglobulin (RMA-S/HLA-E) that had been loaded with saturating amounts of different HLA-E-binding peptides. These data suggest a degenerated peptide recognition by the TCR of NK-CTL (20).
In this article, we show that NK-CTLs recognize, with high avidity, peptides derived from the UL40 protein isolated from different human CMV strains. CMV is a β-herpesvirus that infects most of the human population, resulting in lifelong persistent infection, which may cause severe morbidity in immunocompromised individuals. Both T and NK cells appear to play a critical role in controlling CMV infections (22–24). Our present data suggest that HLA-E-restricted CTLs may be induced in vivo as a consequence of CMV infection. Because CMV is known to escape detection by conventional CTL due to the down-regulation of HLA class I molecules (25), it is conceivable that NK-CTLs may play a role in T cell defenses against this virus.
Materials and Methods
mAbs, Tetramers, and Flow Cytofluorimetric Analysis. The following mAbs were used in this study: GL183 (IgG1, anti-KIR2DL2/S2/L3), Y249 (IgM, anti-KIR2DL2/S2/L3), EB6 (IgG1, anti-KIR2DL1/S1), XA141 (IgM, anti-KIR2DL1/S1), F278 (IgG1, anti-ILT2/LIR1), Y9 (IgM, anti-CD94), Z270 (IgG1, anti-NKG2A), KD1 (IgG2a, anti-CD16), BAB281 (IgG1, anti-NKp46), A13 (IgG1, anti-Vδ1), and BB3 (IgG1, anti-Vδ2) were produced in our laboratory. Leu-2a (IgG1, anti-CD8) and Leu-3a (IgG1, anti-CD4) mAbs were purchased from Becton Dickinson. mAbs anti-TCRVβ16 (IgG1), anti-TCRVβ22 (IgG1), anti-TCRVβ9 (IgG2a), and anti-TCRVβ5.1 (IgG1) were purchased from Immunotech (Marseilles, France). FITC- and phycoerythrin (PE)-conjugated anti-isotype goat anti-mouse were purchased from Southern Biotechnology Associates. The reactivity of mAbs with cell populations was assessed by indirect immunofluorescence and cytofluorimetric analysis as described (26). Briefly, 105 cells were stained with the corresponding mAb followed appropriate FITC- or PE-conjugated anti-isotype-specific goat anti-mouse second reagents (Southern Biotechnology Associates). Soluble HLA-E tetramers were supplied by the National Institute of Allergy and Infectious Diseases Tetramer Facility, and the National Institutes of Health AIDS Research and Reference Reagent Program. HLA-E*0101 and β2-microglobulin proteins were refolded with the following synthetic peptides: VMAPRTLIL and VMAPRTLVL. To enumerate VMAPRTLIL– or VMAPRTLVL-specific T cells, samples were stained with tetramersPE (at ≈15 μg/ml) for 30 min at room temperature. Five micrograms per ml XA185 (anti-CD94) mAb was used in blocking experiments. All samples were analyzed on a FACScan flow cytometer (Becton Dickinson).
Isolation, Culture, and Cloning of CD8+ T Lymphocytes. Peripheral blood mononuclear cells (PBMCs) were isolated from blood of normal donors by a Ficoll–Hypaque density gradient, as described (26). PBMCs were depleted of CD4+, CD16+, NKp46+, Vδ1+, and Vδ2+ cells by negative selection by using appropriate mAbs and magnetic beads coated with anti-mouse IgG (Dynal, Oslo). The culture medium used was RPMI medium 1640 (Seromed, Berlin) supplemented with 10% FCS (Sigma-Aldrich), 1% antibiotic mixture (5 mg/ml penicillin and 5 mg/ml streptomycin stock solution) and human recombinant IL-2 was kindly provided by Chiron. Ficoll–Hypaque (Histopaque 1077) was purchased from Sigma-Aldrich. NK-CTL populations were obtained in MLC. MLCs were set up as described (26) by culturing CD8+ T cells in round-bottom microwells (105 cells per well) in the presence of irradiated (5,000 rads) allogeneic PBMCs (2 × 105 cells). At day 3, 25 units/ml IL-2 was added to cultures. At day 8, cells were either restimulated or cloned under limiting dilution conditions (27). Priming of CD8+ T cells has been performed as follows: B-EBV were pulsed overnight with 104 nM VMAPRTLVL, VMAPRTLIL, and VMAPRTLLL peptides, irradiated, and added in the culture at a ratio variable from 5:1 to 10:1 (CD8+/APC) in the presence of irradiated autologous PBMCs (2 × 105 cells). At day 3, 25 units/ml IL-2 was added to cultures. At day 8, the specificity of the resulting of NK-CTL expanded after MLCs or specific priming was evaluated in a cytolytic assay and immunofluorescence analysis was performed.
UL40 Construct and Transfection. The set of primers hCMVUL40 up (5′-CAG GAC TTT TTA ATG GC) and hCMVUL40 down (5′-AAA TCG CAC CTG TGA CCT C) was used to amplify the UL40 ORF from hCMV strain AD169 (kindly provided by A. Puccetti, University of Genoa, Genoa, Italy) by using an appropriate subgenomic DNA clone as template. Thirty cycles of PCR (30 s at 95°C, 30 s at 55°C, and 30 s at 72°C) were performed by using TAQ-GOLD (Perkin–Elmer/Applied Biosystems) after a preactivation of 12 min at 95°C. The amplification products were subcloned into pcDNA3.1/V5/His TOPO vector by using the Eukaryotic TOPO TA cloning kit (Invitrogen) and sequenced. DNA sequencing was performed by using d-Rhodamine terminator cycle sequencing kit and a 3100 ABI automatic sequencer (Perkin–Elmer/Applied Biosystems).
HEK-293T cells (4 × 105 per plate) were cotransfected with UL40-pcDNA3.1/V5/His and HLA-E*01033-RSV5 neo (kindly provided by J. E. Coligan, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda) constructs using FuGene6 reagent (Roche Applied Science). After 48 h, transfected cells were selected in medium containing 1 mg/ml G418. G418-resistent cells were cloned by limiting dilution and clones were tested by RT-PCR (using the above described protocol). To exclude that PCR amplifications were due to DNA contaminations, RNA of positive samples was further amplified without retrotranscription. The clones in which PCR fragments were obtained only in the RT+ condition were scored as UL40 positive.
Amplifications and Sequencing of TCRVβ22 Rearrangement. RNA samples were extracted by using RNAzol (Cinna/Biotecx Laboratories, Houston) and oligo(dT)-primed cDNA were prepared by using standard techniques. The primers Vβ22 (5′-TGG GAC AGG AAG TGA TCT TGC) and Vβ cost (5′-GTG GGA GAT CTC TGC TTC TGA TCG CTC AAA) were used to specifically amplify the Vβ22 transcripts by PCRs (30 cycles: 30 s at 95°C, 45 s at 55°C, and 1 min at 72°C). The amplification products were cloned into pcDNA3.1/V5/His TOPO vector, and at least 10 independent clones were sequenced.
Cytolytic Activity and IFN-γ Assays. NK-CTLs derived from four different donors were tested for cytolytic activity in a 4-h 51Cr-release assay as described (28) in the presence or in the absence of mAbs. The concentration of the mAbs used for the masking experiments was 10 μg/ml. The E:T ratio used in all experiments was 10:1, unless otherwise indicated. Target cells used in these experiments were represented by a panel of PHA blasts, and B-EBV was derived from a large panel of donors. The murine TAP2-deficient T cell lymphoma RMA-S cell line cotransfected with human β2-microglobulin and HLA-E*01033 allele (kindly provided by J. E. Coligan) was also tested. The RMA-S/HLA-E cell line was previously incubated overnight at 37°C, either alone or in the presence of serial dilutions of synthetic peptides. The peptides used in these experiments were as follows: VMAPRTLVL, VMAPRTLIL, VMAPRTLLL, VMAPRALLL, and SQAPLPCVL. The human fibroblast cell line HEK-293T, either untransfected or cotransfected with AD169-derived UL40 and HLA-E*01033 allele was also tested. B-EBV and untransfected HEK-293T cell lines were also tested after overnight incubation at 37°C with the aforementioned peptides.
NK-CTLs derived from four different donors were analyzed for their IFN-γ release by using the following procedure: responding cells (75 × 104 cells per ml) were incubated overnight with RMA-S/HLA-E (25 × 104 cells per ml) pulsed with serial dilutions of VMAPRTLVL and VMAPRTLIL synthetic peptides in 96-well culture plates. After 18 h, culture supernatants were collected for measurements of IFN-γ release by standard ELISA (R & D Systems).
Results
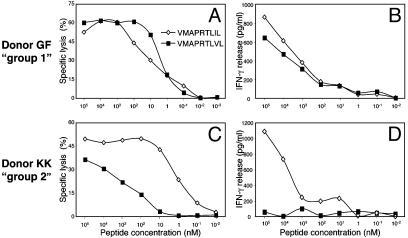
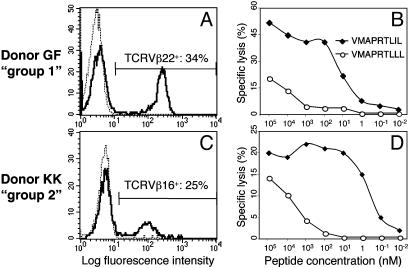
Analysis of TCR Avidity for Different HLA-E-Binding Peptides. We have recently shown that NK-CTLs derived from different donors can recognize HLA-E in association with various peptides loaded at saturating amounts (105 nM) on the murine TAP2-deficient RMA-S/HLA-E cell line. The HLA-E-binding peptides tested were derived from the leader sequence of HLA class I alleles and from viral proteins (20). In view of this apparent broad specificity of NK-CTLs, it was important to analyze the TCR avidity for different HLA-E-bound peptides. To this end, NK-CTL clones derived from four donors (KK, GG, GF, and DP) were tested for their ability to lyse the TAP2-deficient RMA-S/HLA-E cell line incubated with serial dilutions of the VMAPRTLIL, VMAPRTLVL, VMAPRTLLL, and VMAPRALLL peptides. It is of note that the VMAPRTLIL, VMAPRTLVL, and VMAPRTLLL peptides are present in the leader sequence of the UL40 protein derived from different CMV strains (11, 29, 30). Importantly, however, they are also part of the leader sequence of various HLA class I alleles (see Table 1). As to the VMAPRALLL HLA-Cw*07-derived peptide, no viral counterpart has been identified so far. The SQAPLPCVL peptide derived from the BZLF1 protein of EBV (12) (that has no HLA class I counterpart) was also tested. NK-CTLs derived from one or another of the four donors analyzed could recognize VMAPRTLLL, VMAPRALLL, and SQAPLPCVL peptides only when RMA-S/HLA-E target cells were pulsed with high concentrations (105 nM) of these peptides (data not shown). On the other hand, the CMV-derived VMAPRTLIL and VMAPRTLVL peptides could be recognized on RMA-S/HLA-E also when added at very low concentrations. Two groups of donors could be distinguished for their ability to recognize low dilutions of these CMV-derived peptides. Thus, the NK-CTL clone GF2.1, isolated from donor GF (representative of group 1), and characterized by the TCRVβ22+ iNKR+ phenotype, recognized, with high avidity, both the VMAPRTLIL and VMAPRTLVL peptides, because 50% of maximal lysis (IC50) of pulsed RMAS/HLA-E was detected between 1 and 10 nM concentrations of both peptides (Fig. 1A). Although not shown, similar results were obtained with NK-CTLs derived from group 1 donor DP expressing a different TCR rearrangement (TCRVβ5.1). In contrast, clone KK3.2 (TCRVβ16+, iNKR+) derived from donor KK (representative of group 2) recognized with high avidity only the VMAPRTLIL peptide. Thus, IC50 was detected at 1–10 nM concentrations of the VMAPRTLIL peptide, but only at concentrations as high as 102 to 103 nM of the VMAPRTLVL peptide (Fig. 1C). Similar results were obtained with clones derived from donor GG also belonging to group 2 but characterized by the TCRVβ9 rearrangement (data not shown). Thus, NK-CTLs from group 1 donors display a high avidity for both peptides, whereas those from group 2 are characterized by high avidity for VMAPRTLIL peptide and low avidity for VMAPRTLVL peptide. NK-CTLs derived from group 1 and group 2 donors were also analyzed for the production of IFN-γ in a similar experimental setting. As shown in Fig. 1B, clone GF2.1 (group 1) produced IFN-γ, in response to RMA-S/HLA-E pulsed with serial dilutions of both VMAPRTLIL and VMAPRTLVL peptides. On the other hand, IFN-γ production by clone KK3.2 (group 2) only occurred in response to RMA-S/HLA-E pulsed with VMAPRTLIL, but not VMAPRTLVL peptide.
Table 1. Leader sequence peptides from HLA class I and their relationship to UL40-derived peptides.
| HLA class I | Leader sequence peptide3-11 | HLA class I allotypes† |
|---|---|---|
| HLA-A | VMAPRTLLL‡ | HLA-A*01, -A*03, -A*11, -A*29, -A*30, -A*31, -A*32, -A*33, -A*36, and -A*74 |
| VMAPRTLVL‡ | HLA-A*02, -A*23, -A*24, -A*25, -A*26, -A*3402, -A*43, -A*66, and -A*69 | |
| IMAPRTLVL | HLA-A*3401 | |
| VMPPRTLLL | HLA-A*80 | |
| HLA-B | VMAPRTVLL | HLA-B*07, -B*08, -B*14, -B*38, -B*39, -B*42, -B*67, -B*73, and -B*81 |
| VTAPRTLLL | HLA-B*13, -B*18, -B*27, -B*35§, -B*37, -B*40§, -B*44§, -B*47, -B*54, -B*56, -B*58, -B*59, -B*82, and -B*83 | |
| VTAPRTVLL | HLA-B*15, -B*35§, -B*40§, -B*41, -B*44§, -B*45, -B*49, -B*50, -B*51, -B*52, -B*57, and -B*78 | |
| HLA-C | VMAPRTLIL‡ | HLA-Cw*01, -Cw*03, -Cw*04, -Cw*05, -Cw*06, -Cw*0801-03, -Cw*12, -Cw*14, -Cw*16, and -Cw*1702 |
| VMAPRTLLL‡ | HLA-Cw*2 and -Cw*15 | |
| VMAPRTLTL | HLA-Cw*0809 | |
| VMAPRALLL | HLA-Cw*07 and -Cw*18 | |
| VMAPQALLL | HLA-Cw*1701 and -Cw*1703 | |
| HLA-G | VMAPRTLFL | HLA-G*01 |
The amino acid sequences of the various HLA class I alleles can be accessed at: www.anthonynolan.com/HIG/data.html.
These peptides are identical to peptides contained in the UL40 ORF from various human CMV strains.
HLA-*B35, -*B40, and -B*44 are characterized by different leader sequence peptides according to subtypes.
Fig. 1.
Different avidity in peptide recognition by NK-CTLs from different donors. Two representative NK-CTL clones derived from group 1 (GF2.1) and group 2 (KK3.2) donors were tested for cytolytic activity (in a 4-h 51Cr-release assay) (A and C) or for IFN-γ production (in an 18-h assay) (B and D) against RMA-S/HLA-E cells pulsed with graded amounts of the indicated HLA-E-binding peptides. Similar results were obtained in nine independent experiments by using clones derived from the same or different donors.
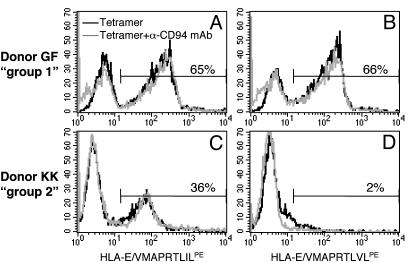
To further substantiate data on the differential avidity of the TCR of NK-CTLs for HLA-E-bound peptides, we used HLA-E tetramers. HLA-E*0101 tetramers refolded with either VMAPRTLIL or VMAPRTLVL peptide were used to stain NK-CTL populations from both groups of donors. A correlation existed between the TCR avidity of NK-CTLs belonging to group 1 or group 2 donors and the ability to bind HLA-E tetramers refolded with different peptides. Fig. 2 shows that the NK-CTL-enriched population derived from donor GF (containing 65% of TCRVβ22+ iNKR+ cells) bound HLA-E tetramers refolded with either VMAPRTLIL (Fig. 2A) or VMAPRTLVL peptide (Fig. 2B). On the other hand, the NK-CTL-enriched population derived from donor KK (containing 36% of TCRVβ16+ iNKR+ cells) only bound HLA-E tetramers refolded with VMAPRTLIL (Fig. 2C), but not with VMAPRTLVL peptide (Fig. 2D). Notably, NK-CTLs from both groups of donors were characterized by the expression of low-density HLA-E-specific CD94/NKG2 receptors. To analyze whether the expression of these receptors could interfere with binding of HLA-E tetramers to TCR (8, 31), control experiments were performed by preincubating cells with anti-CD94 mAb. As shown in Fig. 2, mAb-mediated masking of CD94 did not substantially interfere with tetramer binding to TCR.
Fig. 2.
Binding of HLA-E tetramers to the TCR of NK-CTLs. NK-CTL populations from donor GF (group 1) or from donor KK (group 2), characterized by the expression of TCRVβ22 and TCRVβ16, 65% and 35%, respectively, were stained with HLA-E tetramersPE refolded with either VMAPRTLIL (A and C) or VMAPRTLVL (B and D) peptides. Experiments were performed in the absence or presence of anti-CD94 mAb. Similar results were obtained in five independent experiments.
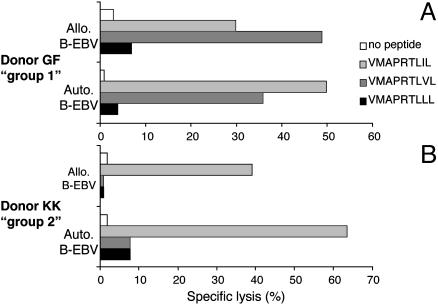
Target Cells Resistant to NK-CTL-Mediated Lysis Become Susceptible in the Presence of CMV-Derived, HLA-E-Binding Peptides. In a previous report (20), we showed that NK-CTLs are characterized by a broad alloreactivity; i.e., they are capable of lysing target cells (PHA-blasts, B-EBV, or tumor cell lines) derived from most (but not all) HLA class I unmatched donors. Remarkably, autologous cells were resistant to lysis. We investigated whether incubation with peptides that were recognized with high avidity by NKCTLs could render susceptible to lysis target cells that were otherwise resistant. Toward this end, resistant autologous or allogeneic B-EBV cells were pulsed with saturating amounts (105 nM) of the VMAPRTLIL or VMAPRTLVL peptides or with the VMAPRTLLL peptide as control. Fig. 3A shows that the NK-CTL population derived from donor GF (group 1) acquired the ability to lyse both autologous and allogeneic B-EBV cells that had been pulsed with either VMAPRTLIL or VMAPRTLVL peptide. On the other hand, only the VMAPRTLIL peptide rendered the same target cells susceptible to lysis by the NK-CTL population derived from donor KK (group 2) (Fig. 3B). Note that when resistant target cells were pulsed with the control VMAPRTLLL peptide they did not become susceptible to lysis by NK-CTLs of both groups (Fig. 3 A and B). Hence, the same peptides that are recognized with high avidity on RMA-S/ HLA-E cell transfectants could render susceptible to lysis autologous as well as (resistant) allogeneic B-EBV cell lines.
Fig. 3.
Peptide-induced lysis of resistant target cells by NK-CTLs. NK-CTL populations were derived from group 1 (A) and group 2 (B) donors and analyzed for their ability to lyse either autologous or resistant allogeneic B-EBV target cells (in a 4-h 51Cr-release assay). Target cells were either unpulsed or pulsed overnight at 37°C with the indicated HLA-E-binding peptides. Similar results were obtained in five independent experiments.
NK-CTLs Have Been Identified in Donors Displaying a Particular HLA-C Haplotype. The above experiments provide insight in our understanding the pattern of NK-CTL specificity. Along this line, differences in the avidity displayed by NK-CTLs for certain HLA-E-bound peptides could explain why allogeneic target cells are either susceptible or resistant to NK-CTL-mediated lysis. Interestingly, the VMAPRTLIL peptide that is recognized with high avidity on RMA-S/HLA-E is present not only in the CMV UL40 protein but also in the leader sequence of the majority of HLA-Cw alleles (with few exceptions, including HLA-Cw*02 and HLA-Cw*07; see Table 1). Accordingly, NK-CTLs derived both from group 1 and group 2 donors could lyse allogeneic target cells expressing at least one VMAPRTLIL-bearing HLA-C allele (Table 2). However, NKCTLs derived from group 1 could also lyse certain allogeneic target cells expressing HLA-Cw*02 and/or HLA-Cw*07 alleles (that lack the VMAPRTLIL peptide; see Table 2). The explanation of these data were provided by the analysis of the HLA-A alleles expressed by these target cells. Indeed, they expressed at least one HLA-A allele bearing the VMAPRTLVL peptide, also recognized with high avidity by group 1 (but not group 2) NK-CTLs (see Tables 1 and 2). It is of note that, in agreement with the above data, the NK-CTLs analyzed in this article were derived only from HLACw*02+ and/or HLA-Cw*07+ donors. In addition, donors belonging to group 2 expressed at least one VMAPRTLV-bearing HLA-A allele (i.e., HLA-A*02), whereas group 1 donors did not (Table 2). In summary, VMAPRTLIL represents a non-self peptide for group 1 and group 2 donors, and is recognized with high avidity by NK-CTLs from both groups. On the other hand, VMAPRTLVL represents a self peptide for group 2 donors, and is therefore recognized only by group 1 donors.
Table 2. Cytolytic activity of NK-CTLs against allogeneic cells.
| Donor | Haplotype | Group 1 GF: A1;3, B27;44, Cw2 DP: A3, B18;27, Cw2;7 | Group 2 KK: A2;32, B44, Cw7 GG: A2;25, B7;18, Cw7 |
|---|---|---|---|
| LM | A3;29 B35;44 Cw4;5 | Susceptible | Susceptible |
| CB | A26;68 B27;62 Cw2;4 | Susceptible | Susceptible |
| RB | A11;25 B18;44 Cw5 | Susceptible | Susceptible |
| GB1 | A2;11 B44;51 Cw1;4 | Susceptible | Susceptible |
| GB2 | A3;32 B7;35 Cw5;7 | Susceptible | Susceptible |
| AC | A2 B8;35 Cw2;7 | Susceptible | Resistant |
| LN | A24;26 B8;35 Cw7 | Susceptible | Resistant |
| BM15 | A1 B49 Cw7 | Resistant | Resistant |
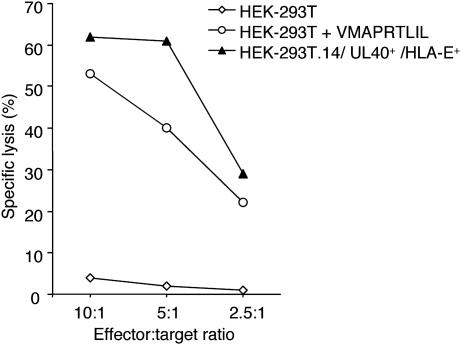
NK-CTL-Mediated, HLA-E-Restricted Recognition of a Naturally Processed CMV-Derived Peptide. The above data clearly demonstrate that NK-CTLs recognize with high avidity, in an HLA-E-restricted fashion, the VMAPRTLIL peptide common to the leader sequences of UL40 protein of different CMV strains (including AD169, Towne, and various CMV clinical isolates; ref. 11), and to those of most HLA-Cw alleles. In addition, the VMAPRTLVL peptide recognized by group 1 NK-CTLs is present in various HLA-A alleles, as well as in the UL40 protein of another CMV strain (Toledo strain; ref. 11). During CMV infection VMAPRTLIL peptide is processed and presented at the cell surface in association with HLA-E (11, 29, 32). These data offer a clue to explain how NK-CTLs might be generated and suggest a possible role for NK-CTLs in the immune response against CMV. In this context, it is important to stress that NK-CTLs have been detected, among donors with an appropriate HLA class I haplotype (see above), only in those who were seropositive for anti-CMV antibodies. To investigate whether naturally processed, UL40-derived endogenous peptides could be recognized by NK-CTLs, the human cell line HEK-293T was cotransfected with HLA-E*01033 RSV5 neo and AD169-UL40 pcDNA3.1 constructs. Cell clones derived from HEK-293T/UL40+/HLA-E+ cell transfectants were tested in RT-PCR for the expression of UL40 mRNA (data not shown). Notably, HEK-293T cell line (haplotype: HLA-A*02; -*07; and -Cw*07) lacks VMAPRTLIL-bearing HLA-Cw alleles. Accordingly, it is not recognized by group 2 NK-CTLs (Fig. 4). On the other hand, because it does express VMAPRTLVL-bearing HLA-A alleles, it is susceptible to lysis by group 1 NK-CTLs. Thus, NK-CTLs derived from group 2 donors were tested for their ability to lyse the UL40+/HLA-E+/HEK-293T.14 cell clone. As shown in Fig. 4, the UL40+/HLA-E+/HEK-293T.14 clone was lysed by the TCRVβ16+ NK-CTL population from donor KK (group 2). Importantly, lysis was inhibited by an ant-TCRVβ16 mAb added to the cytolytic assay (data not shown). In addition, in a control experiment, untransfected HEK-293T cells became susceptible to lysis when pulsed with saturating concentrations (105 nM) of exogenously provided AD169-UL40-derived peptide (VMAPRTLIL).
Fig. 4.
NK-CTLs recognize a naturally processed CMV peptide in UL40+ cell transfectants. NK-CTL population derived from group 2 donor KK (TCRVβ16+) was tested in a cytolytic assay against HEK-293T tumor cell line, either untransfected (⋄) or cotransfected (▴) with HLA-E*01033 and AD169-derived UL40 in a 4-h 51Cr-release assay. Untransfected HEK-293T cell line was also analyzed after pulsing with exogenous AD169-UL40-derived peptide (VMAPRTLIL) (○). Similar results were obtained in eight independent experiments.
UL40-Derived Peptides Can Induce NK-CTL Proliferation in an Autologous Setting. The above data clearly demonstrate that an UL40-derived peptide, that has been naturally processed in UL40 cell transfectants, can be recognized by NK-CTLs in the context of HLA-E. Next, we asked whether the same CMV peptide could also induce specific proliferation of NK-CTLs when presented by autologous antigen-presenting cells (APCs). To directly verify this possibility, CD8+ T lymphocytes derived from group 1 or group 2 donors were cultured with autologous irradiated B-EBV cells. These cells, used as a source of APCs, were pulsed with the UL40-derived VMAPRTLIL peptide or VMAPRTLLL self peptide as control. After 7 days, CD8+ T lymphocytes derived from both donor GF (group 1) and KK (group 2) displayed a marked expansions of TCRVβ22+ or TCRVβ16+ cells, respectively, in cultures pulsed with the CMV-derived VMAPRTLIL peptide (Fig. 5 A and C). This display did not occur when cultures were pulsed with the VMAPRTLLL self peptide (data not shown). Both the TCRVβ22+ and the TCRVβ16+ T cell expansions specifically lysed RMA-S/HLA-E pulsed with the VMAPRTLIL peptide (IC50: 1–10 nM peptide concentration). On the other hand, no specific cytolytic activity was detected when RMA-S/HLA-E were pulsed with the VMAPRTLLL self peptide as control (Fig. 5 B and D). A similar TCRVβ22+ cell expansion was obtained in donor GF (group1) when CD8+ T cells were cultured with autologous B-EBV cells pulsed also with the other non-self CMV-derived peptide (VMAPRTLVL) (data not shown). Notably, the NK-CTL expansions obtained from this donor in response to both CMV-derived peptides expressed the same TCRVβ rearrangement and the same iNKR phenotype as the original NK-CTLs present in small percentages in the peripheral blood of the same donor. These data provide direct evidence that NK-CTL can undergo specific expansion in response to an HLA-E-binding CMV peptide in an autologous setting, and offer an interesting clue to explain how NK-CTL populations may be generated in vivo.
Fig. 5.
NK-CTL proliferation in response to autologous APC pulsed with UL40-derived peptide (VMAPRTLIL). NK-CTL-specific expansions from donor GF (group 1; A) or from donor KK (group 2; C), obtained after priming CD8+ T cells with autologous APC pulsed with VMAPRTLIL peptide, were characterized by the expression of TCRVβ22 and TCRVβ16, 34% and 25%, respectively. These cell populations were tested for cytolytic activity (in a 4-h 51Cr-release assay) against RMA-S/HLA-E cells pulsed with graded amounts of the indicated peptides (B and D). Similar results were obtained in four independent experiments.
Discussion
Our present study provides, to our knowledge, the first evidence that human CD8+ TCRαβ+ T cells can recognize CMV-derived antigen peptides in the context of the nonclassical HLA-E molecule. This mechanism may have particular relevance in the immune defenses against CMV, because this virus evolved a number of strategies to down-regulate the surface expression of MHC class Ia molecules, thus escaping the control of classical CD8+ CTL (25). In addition, it provides an important clue to understand how NK-CTL expressing high avidity to TCR may be generated in CMV-seropositive donors in response to CMV infections and how they may crossreact with HLA class I-derived peptides expressed by various allogeneic cells.
Recently, we and other groups have shown that the function of the nonclassical HLA-E molecules is not simply confined to the modulation of NK cell responses as it actually represents a restriction element for the TCRαβ-mediated recognition (4–7). The surface expression of HLA-E requires (and it is therefore controlled by) the availability of a set of highly conserved nonameric peptides derived from the leader sequence of various HLA class I molecules including HLA-A, -B, -C, and -G, but not HLA-F (9, 10, 33). When an appropriate peptide is available, the resultant molecular complex can interact with CD94/NKG2A or CD94/NKG2C heterodimers expressed at the cell surface of NK cells resulting, respectively, either in inhibition or in triggering of the NK cell-mediated cytotoxicity and cytokine production (8, 31). Previous reports showed that HLA-E-binding peptides are also derived from the leader sequence of the human CMV UL40 protein, and may increase the levels of HLA-E expression on the surface of infected cells (11, 29, 32). Therefore, the CMV-induced upregulation of HLA-E may provide a mechanism by which CMV-infected cells can escape lysis by NK cells expressing the CD94/NKG2A inhibitory receptor (11, 29, 31, 32). Association of the UL40-derived peptide to HLA-E is apparently TAP-independent, because it is not affected by the CMV US6 glycoprotein, known to inhibit the TAP transporter and thus to down-regulate the surface expression of classical HLA class I molecules (11, 30). Down-regulation of classical HLA class I molecules represents an important mechanism of immunoevasion of the CMV-infected target cells from conventional CTL (25), although it may render infected cells susceptible to lysis by CD94/NKG2A-negative NK cells. Indeed, NK cells have been shown to play an important role in defenses against herpesvirus infections (24). Our data indicate that also HLA-E-restricted CTLs may actually play a relevant defensive role during CMV infection. They also suggest how NK-CTLs could be generated in vivo and what their physiological role could be. It is conceivable that NK-CTLs, recognizing, with high avidity, a UL40-derived peptide, may represent monoclonal expansions generated in response to CMV infection. The fact that, in various individuals, NK-CTLs represent a sizeable fraction of CD8+ T cells and that they persist for many years (ref. 19 and our unpublished observation) may reflect the continuous stimulation by CMV, and may as well be consequent to the antiapoptotic effect induced by the engagement of iNKRs (34, 35), which are typically expressed at the NK-CTL surface.
Interestingly, donors from which NK-CTLs specific for the VMAPRTLIL peptide could be derived were characterized by the HLA-Cw*02 and/or HLA-Cw*07 haplotype. It is of note that VMAPRTLIL represents, for these donors, a non-self peptide because it is not present in their HLA-Cw alleles. On the other hand, donors who express HLA-Cw alleles carrying the VMAPRTLIL peptide fail to generate NK-CTL specific for this self peptide (unpublished data). It is conceivable that in these donors NK-CTL precursors may have been negatively selected by HLAE/VMAPRTLIL complexes. Along this line, previous reports (36) provided evidence that MHC class Ib molecules can mediate thymic selection in mice. If this finding holds true, these donors would be tolerant to CMV strains that bear the VMAPRTLIL peptide. It is of note that different CMV strains, as well as various CMV clinical isolates, have a mutated form of UL40 from which other HLA-E-binding peptides can be obtained (11, 29). For example, a peptide identical to that of the HLA-A*02 allele (VMAPRTLVL) is present in the Toledo strain (11), whereas the 3C CMV clinical isolate contains a peptide (VMAPRTLLL) identical to that of HLA-Cw*02 allele (29). This result may suggest that the HLA class I host haplotype, as well as the infecting CMV strain, may greatly influence the ability of different individuals to exploit NK-CTL-mediated defenses against CMV infections.
Because CMV UL40-derived peptides are identical to those derived from the leader sequences of various HLA class I alleles, the recognition of allogeneic cells by NK-CTL may be considered as a side effect of the specific response against CMV. It is of note that NK-CTL express iNKRs, which can prevent their TCR-induced activation. We have previously shown (20) that inhibition of NK-CTL cytotoxicity can be mediated by KIR2DL receptors, but not by CD94/NKG2A or LIR1/ILT2 (possibly due to their low surface density), provided that allogeneic (non-CMV infected) cells express the relevant HLA class I alleles recognized by KIR2DL. It should be stressed, however, that in the case of CMV-infected target cells characterized by down-regulation of classical HLA class I molecules, even the KIR2DL/HLA class I interaction may be insufficient to inhibit NK–CTL function.
The ability of NK-CTLs to recognize and lyse allogeneic cells should be carefully taken into consideration in transplantation using mismatched donors, because they could play a relevant role in the pathogenesis of graft-versus-host disease and graft rejection. In this context, it is noteworthy that CMV has repeatedly been associated with rejection of solid organs, or with graft-versus-host disease, in the case of bone marrow transplantation (37). Nonetheless, the exact mechanism of any interaction between the virus and the disease remains unclear (37). It is tempting to speculate that at least in mismatched transplantation HLA-E-restricted NK-CTLs might play a role in the pathogenesis of these phenomena due to the cross-reactivity between HLA-E-bound CMV peptides and those derived from the host HLA class I antigens.
Acknowledgments
This work was supported by grants awarded by Associazione Italiana per la Ricerca sul Cancro; Istituto Superiore di Sanità; Ministero della Salute; and Ministero dell'Università e della Ricerca Scientifica e Tecnologica 5%-Consiglio Nazionale delle Ricerche Biotechnology program 95/95; Consiglio Nazionale delle Ricerche; Progetto Finalizzato Biotecnologie; and Compagnia di San Paolo.
Abbreviations: CTL, cytolytic T lymphocyte; TCR, T cell receptor; CMV, cytomegalovirus; EBV, Epstein–Barr virus; NK, natural killer; iNKR, NK inhibitory receptors; MLC, mixed lymphocytes culture; TAP, transporter associated with antigen processing.
References
- 1.Rock, K. L. & Goldberg, A. L. (1999) Annu. Rev. Immunol. 17, 739–779. [DOI] [PubMed] [Google Scholar]
- 2.Townsend, A. & Bodmer, H. (1989) Annu. Rev. Immunol. 7, 601–624. [DOI] [PubMed] [Google Scholar]
- 3.Braud, V. M., Allan, D. S. & McMichael, A. J. (1999) Curr. Opin. Immunol. 11, 100–108. [DOI] [PubMed] [Google Scholar]
- 4.Li, J., Goldstein, I., Glickman-Nir, E., Jiang, H. & Chess, L. (2001) J. Immunol. 167, 3800–3808. [DOI] [PubMed] [Google Scholar]
- 5.Pietra, G., Romagnani, C., Falco, M., Vitale, M., Castriconi, R., Pende, D., Millo, E., Anfossi, S., Biassoni, R., Moretta, L. & Mingari, M. C. (2001) Eur. J. Immunol. 31, 3687–3693. [DOI] [PubMed] [Google Scholar]
- 6.Garcia, P., Llano, M., de Heredia, A. B., Willberg, C. B., Caparros, E., Aparicio, P., Braud, V. M. & Lopez-Botet, M. (2002) Eur. J. Immunol. 32, 936–944. [DOI] [PubMed] [Google Scholar]
- 7.Heinzel, A. S., Grotzke, J. E., Lines, R. A., Lewinsohn, D. A., McNabb, A. L., Streblow, D. N., Braud, V. M., Grieser, H. J., Belisle, J. T. & Lewinsohn, D. M. (2002) J. Exp. Med. 196, 1473–1481. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Lee, N., Llano, M., Carretero, M., Ishitani, A., Navarro, F., Lopez-Botet, M. & Geraghty, D. E. (1998) Proc. Natl. Acad. Sci. USA 95, 5199–5204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Lee, N., Goodlett, D. R., Ishitani, A., Marquardt, H. & Geraghty, D. E. (1998) J. Immunol. 160, 4951–4960. [PubMed] [Google Scholar]
- 10.Braud, V., Jones, E. Y. & McMichael, A. J. (1997) Eur. J. Immunol. 27, 1164–1169. [DOI] [PubMed] [Google Scholar]
- 11.Tomasec, P., Braud, V. M., Rickards, C., Powell, M. B., McSharry, B. P., Gadola, S., Cerundolo, V., Borysiewicz, L. K., McMichael, A. J. & Wilkinson, G. W. G. (2000) Science 287, 1031–1033. [DOI] [PubMed] [Google Scholar]
- 12.Ulbrecht, M., Modrow, S., Srivastava, R., Peterson, P. E. & Weiss, E. H. (1998) J. Immunol. 160, 4375–4385. [PubMed] [Google Scholar]
- 13.Michaelsson, J., Teixeira de Matos, C., Achour, A., Lanier, L. L., Karre, K. & Soderstrom, K. (2002) J. Exp. Med. 196, 1403–1414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Lo, W. F., Woods, A. S., DeCloux, A., Cotter, R. J., Metcalf, E. S. & Soloski, M. J. (2000) Nat. Med. 6, 215–218. [DOI] [PubMed] [Google Scholar]
- 15.Lo, W. F., Ong, H., Metcalf, E. S. & Soloski, M. J. (1999) J. Immunol. 162, 5398–5406. [PubMed] [Google Scholar]
- 16.Seaman, M. S., Wang, C. R. & Forman, J. (2000) J. Immunol. 165, 5192–5201. [DOI] [PubMed] [Google Scholar]
- 17.Mingari, M. C., Vitale, C., Cambiaggi, A., Schiavetti, F., Melioli, G., Ferrini, S. & Poggi, A. (1995) Int. Immunol. 7, 697–703. [DOI] [PubMed] [Google Scholar]
- 18.Mingari, M. C., Ponte, M., Vitale, C., Maggi, E., Romagnani, S., Demarest, J., Pantaleo, G., Fauci, A. S. & Moretta, L. (1996) Proc. Natl. Acad. Sci. USA 93, 12433–12438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Mingari, M. C., Moretta, A. & Moretta, L. (1998) Immunol. Today 19, 153–157. [DOI] [PubMed] [Google Scholar]
- 20.Romagnani, C., Pietra, G., Falco, M., Millo, E., Mazzarino, P., Biassoni, R., Moretta, A., Moretta, L. & Mingari, M. C. (2002) Proc. Natl. Acad. Sci. USA 99, 11328–11333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Moretta, L., Romagnani, C., Pietra, G., Moretta, A. & Mingari, M. C. (2003) Trends Immunol. 24, 136–143. [DOI] [PubMed] [Google Scholar]
- 22.Reddehase, M. J. (2000) Curr. Opin. Immunol. 12, 390–396. [DOI] [PubMed] [Google Scholar]
- 23.Walter, E. A., Greenberg, P. D., Gilbert, M. J., Finch, R. J., Watanabe, K. S., Thomas, E. D. & Riddell, S. R. (1995) N. Engl. J. Med. 333, 1038–1044. [DOI] [PubMed] [Google Scholar]
- 24.Biron, C. A., Byron, K. S. & Sullivan J. L. (1989) N. Engl. J. Med. 320, 1731–1735. [DOI] [PubMed] [Google Scholar]
- 25.Reddehase, M. J. (2002) Nat. Rev. Immunol. 2, 831–844. [DOI] [PubMed] [Google Scholar]
- 26.Mingari, M. C., Cambiaggi, A., Vitale, C., Schiavetti, F., Bellomo, R. & Poggi, A. (1996) Int. Immunol. 8, 203–209. [DOI] [PubMed] [Google Scholar]
- 27.Moretta, A., Pantaleo, G., Moretta, L., Mingari, M. C. & Cerottini, J. C. (1983) J. Exp. Med. 158, 571–585. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Poggi, A, Biassoni, R., Pella, N., Paolieri, F., Bellomo, R., Bertolini, A., Moretta, L. & Mingari, M. C. (1990) J. Exp. Med. 172, 1409–1418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Cerboni, C., Mousavi-Jazi, M., Wakiguchi, H., Carbone, E., Karre, K. & Soderstrom, K. (2001) Eur. J. Immunol. 31, 2926–2935. [DOI] [PubMed] [Google Scholar]
- 30.Ulbrecht, M., Martinozzi, S., Grzeschik, M., Hengel, H., Ellwart, J. W., Pla, M. & Weiss, E. H. (2000) J. Immunol. 164, 5019–5022. [DOI] [PubMed] [Google Scholar]
- 31.Braud, V. M., Allan, D. S., O'Callaghan, C. A., Soderstrom, K., D'Andrea, A., Ogg, G. S., Lazetic, S., Young, N. T., Bell, J. I., Phillips, J. H., et al. (1998) Nature 391, 795–799. [DOI] [PubMed] [Google Scholar]
- 32.Wang, E. C., McSharry, B., Retiere, C., Tomasec, P., Williams, S., Borysiewicz, L. K., Braud, V. M. & Wilkinson, G. W. (2002) Proc. Natl. Acad. Sci. USA 99, 7570–7575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.O'Callaghan, C. A. & Bell, J. I. (1998) Immunol. Rev. 163, 129–138. [DOI] [PubMed] [Google Scholar]
- 34.Ugolini, S., Arpin, C., Anfossi, N., Walzer, T., Cambiaggi, A., Forster, R., Lipp, M., Toes, R. E., Melief, C. J., Marvel, J. & Vivier, E. (2001) Nat. Immunol. 2, 430–435. [DOI] [PubMed] [Google Scholar]
- 35.Young, N. T. & Uhrberg, M. (2002) Trends Immunol. 23, 71–75. [DOI] [PubMed] [Google Scholar]
- 36.Maurice, M. M., Gould, D. S., Carroll, J., Vugmeyster, Y. & Ploegh, H. L. (2001) Proc. Natl. Acad. Sci. USA 98, 7437–7442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Griffiths P. D., Clark D. A. & Emery V. C. (2000) J. Antimicrob. Chemother. 45, Suppl. 3, 29–34. [DOI] [PubMed] [Google Scholar]