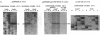Abstract
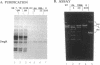
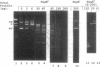
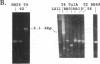
Although not encoded by an intron, the bacteriophage T4 SegA protein shares common amino acid motifs with a family of proteins found within mobile group I introns present in fungi and phage. Each of these intron-encoded proteins is thought to initiate the homing of its own intron by cleaving the intronless DNA at or near the site of insertion. Previously, we have found that SegA also cleaves DNA. In this report, we have purified the SegA protein and characterized this endonuclease activity extensively. SegA protein cleaved circular and linear plasmids, DNA containing unmodified cytosines, and wild-type T4 DNA containing hydroxymethylated, glucosylated cytosines. In all cases, certain sites on the DNA were highly preferred for cleavage, but with increasing protein concentration or time of incubation, cleavage occurred at many sites. SegA cleaving activity was stimulated by the presence of ATP or ATP gamma S. Sequence analysis of three highly preferred cleavage sites did not reveal a simple consensus sequence, suggesting that even among highly preferred sites, SegA tolerates many different sequences. A T4 segA amber mutant that we constructed had no phenotype, and PCR analyses indicated that several T-even-related phages lack the segA gene. Taken together, our results show that SegA is an endonuclease with a hierarchy of site specificity, and these results are consistent with the insertion of segA DNA into the T4 genome some time after the divergence of the closely consistent with the insertion of segA DNA into the T4 genome some time after the divergence of the closely related T-even phages.
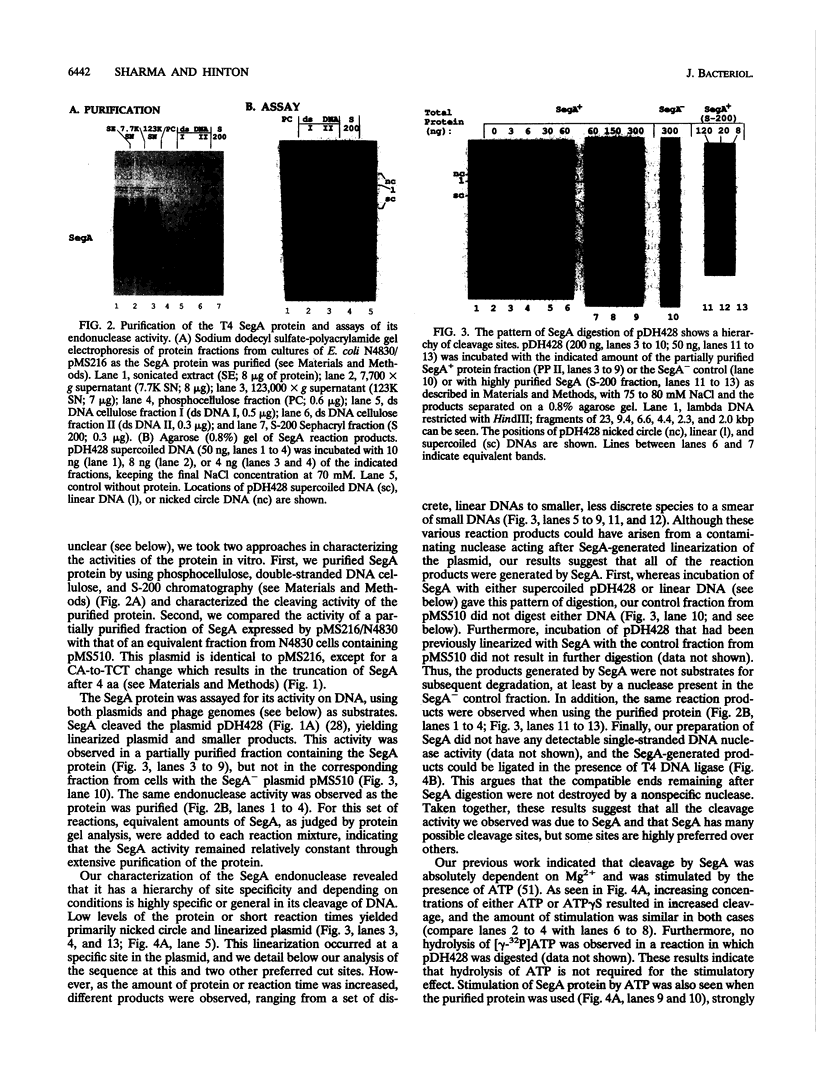
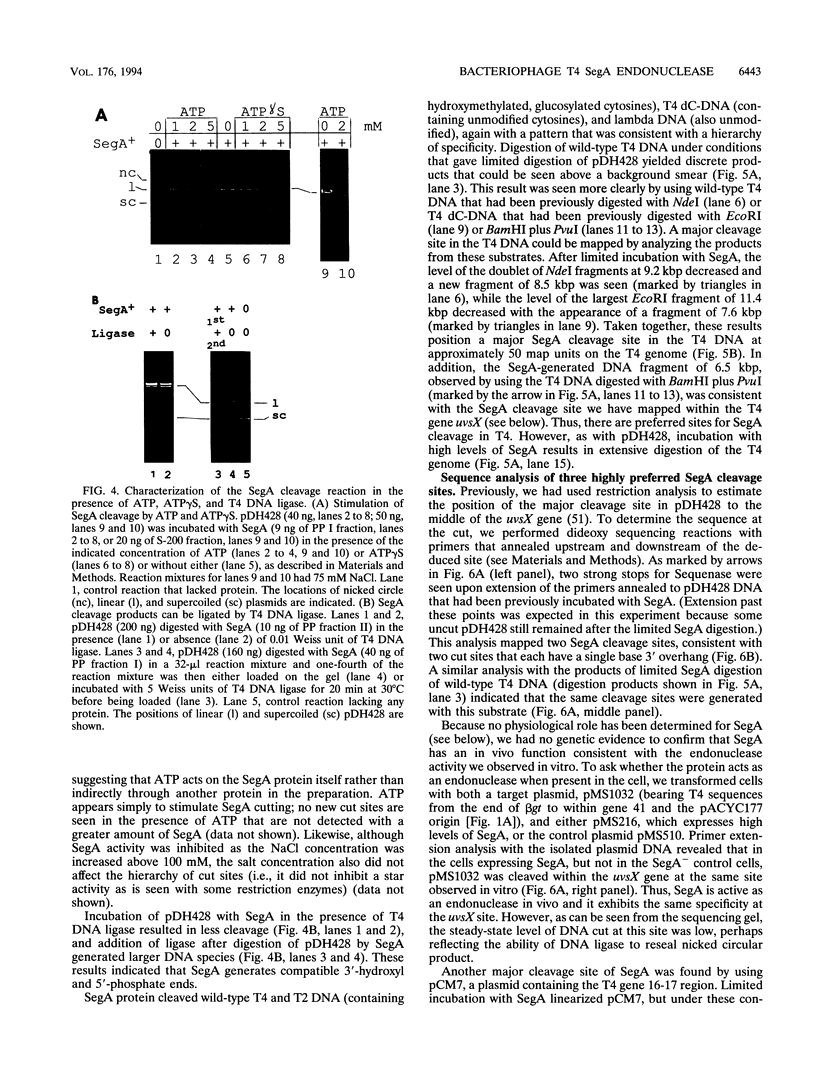
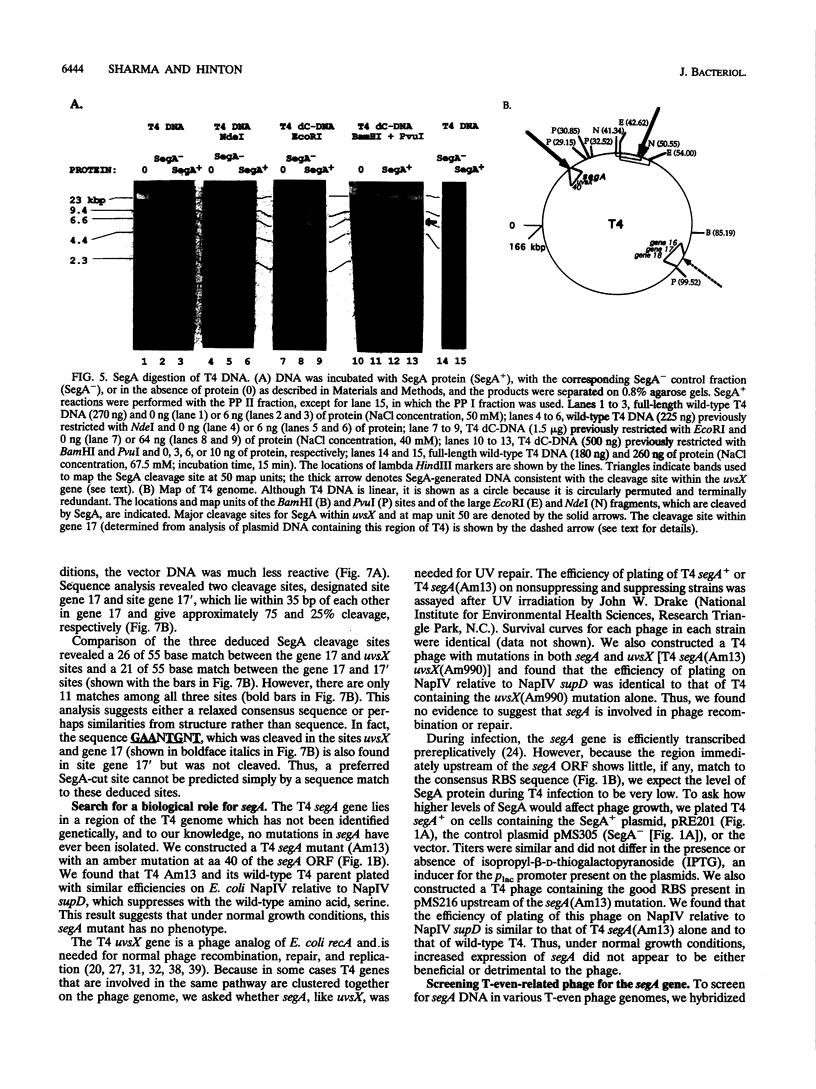
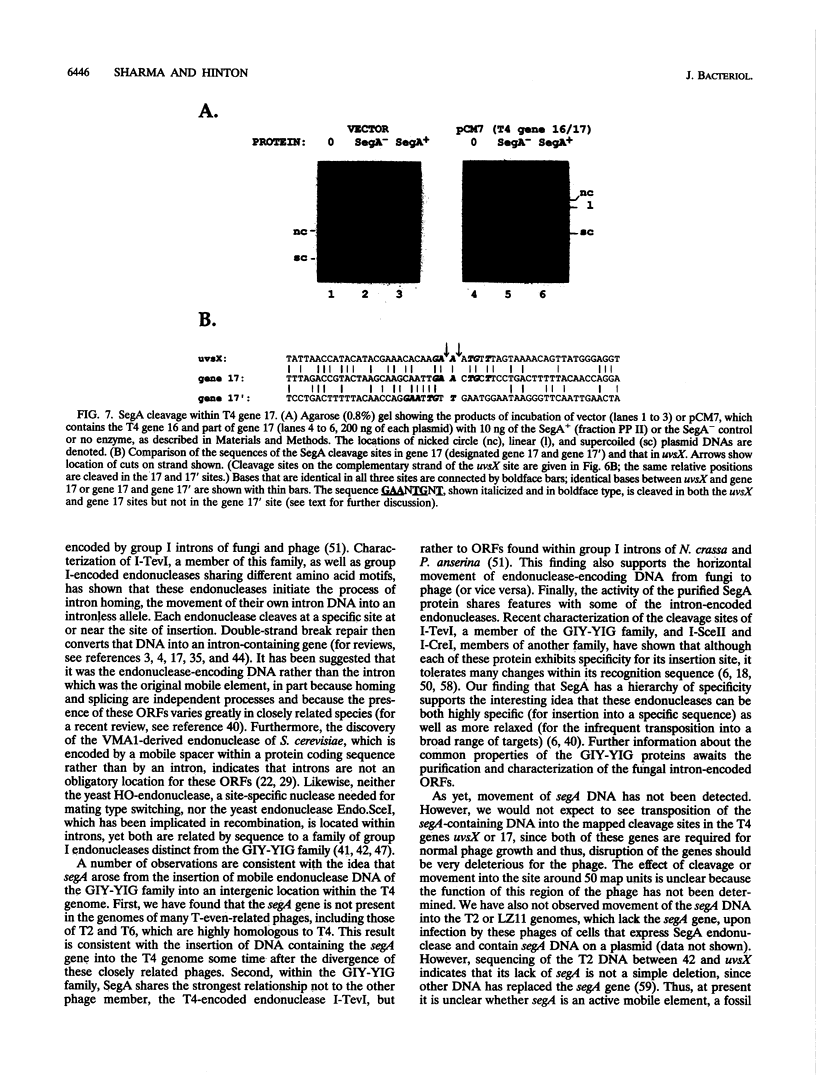
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Adhya S., Gottesman M. Promoter occlusion: transcription through a promoter may inhibit its activity. Cell. 1982 Jul;29(3):939–944. doi: 10.1016/0092-8674(82)90456-1. [DOI] [PubMed] [Google Scholar]
- Belfort M. Phage T4 introns: self-splicing and mobility. Annu Rev Genet. 1990;24:363–385. doi: 10.1146/annurev.ge.24.120190.002051. [DOI] [PubMed] [Google Scholar]
- Belfort M. Self-splicing introns in prokaryotes: migrant fossils? Cell. 1991 Jan 11;64(1):9–11. doi: 10.1016/0092-8674(91)90201-9. [DOI] [PubMed] [Google Scholar]
- Bell-Pedersen D., Quirk S., Clyman J., Belfort M. Intron mobility in phage T4 is dependent upon a distinctive class of endonucleases and independent of DNA sequences encoding the intron core: mechanistic and evolutionary implications. Nucleic Acids Res. 1990 Jul 11;18(13):3763–3770. doi: 10.1093/nar/18.13.3763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bryk M., Quirk S. M., Mueller J. E., Loizos N., Lawrence C., Belfort M. The td intron endonuclease I-TevI makes extensive sequence-tolerant contacts across the minor groove of its DNA target. EMBO J. 1993 May;12(5):2141–2149. doi: 10.1002/j.1460-2075.1993.tb05862.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Burger G., Werner S. The mitochondrial URF1 gene in Neurospora crassa has an intron that contains a novel type of URF. J Mol Biol. 1985 Nov 20;186(2):231–242. doi: 10.1016/0022-2836(85)90100-7. [DOI] [PubMed] [Google Scholar]
- Chu F. K., Maley G. F., Maley F., Belfort M. Intervening sequence in the thymidylate synthase gene of bacteriophage T4. Proc Natl Acad Sci U S A. 1984 May;81(10):3049–3053. doi: 10.1073/pnas.81.10.3049. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chu F. K., Maley G. F., West D. K., Belfort M., Maley F. Characterization of the intron in the phage T4 thymidylate synthase gene and evidence for its self-excision from the primary transcript. Cell. 1986 Apr 25;45(2):157–166. doi: 10.1016/0092-8674(86)90379-x. [DOI] [PubMed] [Google Scholar]
- Colleaux L., Michel-Wolwertz M. R., Matagne R. F., Dujon B. The apocytochrome b gene of Chlamydomonas smithii contains a mobile intron related to both Saccharomyces and Neurospora introns. Mol Gen Genet. 1990 Sep;223(2):288–296. doi: 10.1007/BF00265065. [DOI] [PubMed] [Google Scholar]
- Collins R. A., Reynolds C. A., Olive J. The self-splicing intron in the Neurospora apocytochrome b gene contains a long reading frame in frame with the upstream exon. Nucleic Acids Res. 1988 Feb 11;16(3):1125–1134. doi: 10.1093/nar/16.3.1125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crouse G. F., Frischauf A., Lehrach H. An integrated and simplified approach to cloning into plasmids and single-stranded phages. Methods Enzymol. 1983;101:78–89. doi: 10.1016/0076-6879(83)01006-x. [DOI] [PubMed] [Google Scholar]
- Cummings D. J., Domenico J. M., Michel F. DNA sequence and organization of the mitochondrial ND1 gene from Podospora anserina: analysis of alternate splice sites. Curr Genet. 1988 Sep;14(3):253–264. doi: 10.1007/BF00376746. [DOI] [PubMed] [Google Scholar]
- Datta D. B., Arden B., Henning U. Major proteins of the Escherichia coli outer cell envelope membrane as bacteriophage receptors. J Bacteriol. 1977 Sep;131(3):821–829. doi: 10.1128/jb.131.3.821-829.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Delahodde A., Goguel V., Becam A. M., Creusot F., Perea J., Banroques J., Jacq C. Site-specific DNA endonuclease and RNA maturase activities of two homologous intron-encoded proteins from yeast mitochondria. Cell. 1989 Feb 10;56(3):431–441. doi: 10.1016/0092-8674(89)90246-8. [DOI] [PubMed] [Google Scholar]
- Dujon B. Group I introns as mobile genetic elements: facts and mechanistic speculations--a review. Gene. 1989 Oct 15;82(1):91–114. doi: 10.1016/0378-1119(89)90034-6. [DOI] [PubMed] [Google Scholar]
- Dujon B. Sequence of the intron and flanking exons of the mitochondrial 21S rRNA gene of yeast strains having different alleles at the omega and rib-1 loci. Cell. 1980 May;20(1):185–197. doi: 10.1016/0092-8674(80)90246-9. [DOI] [PubMed] [Google Scholar]
- Dürrenberger F., Rochaix J. D. Characterization of the cleavage site and the recognition sequence of the I-CreI DNA endonuclease encoded by the chloroplast ribosomal intron of Chlamydomonas reinhardtii. Mol Gen Genet. 1993 Jan;236(2-3):409–414. doi: 10.1007/BF00277141. [DOI] [PubMed] [Google Scholar]
- Formosa T., Alberts B. M. Purification and characterization of the T4 bacteriophage uvsX protein. J Biol Chem. 1986 May 5;261(13):6107–6118. [PubMed] [Google Scholar]
- Gimble F. S., Thorner J. Homing of a DNA endonuclease gene by meiotic gene conversion in Saccharomyces cerevisiae. Nature. 1992 May 28;357(6376):301–306. doi: 10.1038/357301a0. [DOI] [PubMed] [Google Scholar]
- Hensgens L. A., Bonen L., de Haan M., van der Horst G., Grivell L. A. Two intron sequences in yeast mitochondrial COX1 gene: homology among URF-containing introns and strain-dependent variation in flanking exons. Cell. 1983 Feb;32(2):379–389. doi: 10.1016/0092-8674(83)90457-9. [DOI] [PubMed] [Google Scholar]
- Hinton D. M., Nossal N. G. Bacteriophage T4 DNA replication protein 61. Cloning of the gene and purification of the expressed protein. J Biol Chem. 1985 Oct 15;260(23):12858–12865. [PubMed] [Google Scholar]
- Hinton D. M., Nossal N. G. Cloning of the bacteriophage T4 uvsX gene and purification and characterization of the T4 uvsX recombination protein. J Biol Chem. 1986 Apr 25;261(12):5663–5673. [PubMed] [Google Scholar]
- Hinton D. M. Transcript analyses of the uvsX-40-41 region of bacteriophage T4. Changes in the RNA as infection proceeds. J Biol Chem. 1989 Aug 25;264(24):14432–14439. [PubMed] [Google Scholar]
- Hinton D. M. Transcription from a bacteriophage T4 middle promoter using T4 motA protein and phage-modified RNA polymerase. J Biol Chem. 1991 Sep 25;266(27):18034–18044. [PubMed] [Google Scholar]
- Hirata R., Ohsumk Y., Nakano A., Kawasaki H., Suzuki K., Anraku Y. Molecular structure of a gene, VMA1, encoding the catalytic subunit of H(+)-translocating adenosine triphosphatase from vacuolar membranes of Saccharomyces cerevisiae. J Biol Chem. 1990 Apr 25;265(12):6726–6733. [PubMed] [Google Scholar]
- Jacquier A., Dujon B. An intron-encoded protein is active in a gene conversion process that spreads an intron into a mitochondrial gene. Cell. 1985 Jun;41(2):383–394. doi: 10.1016/s0092-8674(85)80011-8. [DOI] [PubMed] [Google Scholar]
- Kunkel T. A., Roberts J. D., Zakour R. A. Rapid and efficient site-specific mutagenesis without phenotypic selection. Methods Enzymol. 1987;154:367–382. doi: 10.1016/0076-6879(87)54085-x. [DOI] [PubMed] [Google Scholar]
- Lambowitz A. M. Infectious introns. Cell. 1989 Feb 10;56(3):323–326. doi: 10.1016/0092-8674(89)90232-8. [DOI] [PubMed] [Google Scholar]
- Macreadie I. G., Scott R. M., Zinn A. R., Butow R. A. Transposition of an intron in yeast mitochondria requires a protein encoded by that intron. Cell. 1985 Jun;41(2):395–402. doi: 10.1016/s0092-8674(85)80012-x. [DOI] [PubMed] [Google Scholar]
- Michel F., Dujon B. Genetic exchanges between bacteriophage T4 and filamentous fungi? Cell. 1986 Aug 1;46(3):323–323. doi: 10.1016/0092-8674(86)90651-3. [DOI] [PubMed] [Google Scholar]
- Mosig G. The essential role of recombination in phage T4 growth. Annu Rev Genet. 1987;21:347–371. doi: 10.1146/annurev.ge.21.120187.002023. [DOI] [PubMed] [Google Scholar]
- Nakagawa K., Morishima N., Shibata T. A maturase-like subunit of the sequence-specific endonuclease endo.SceI from yeast mitochondria. J Biol Chem. 1991 Jan 25;266(3):1977–1984. [PubMed] [Google Scholar]
- Nakagawa K., Morishima N., Shibata T. An endonuclease with multiple cutting sites, Endo.SceI, initiates genetic recombination at its cutting site in yeast mitochondria. EMBO J. 1992 Jul;11(7):2707–2715. doi: 10.1002/j.1460-2075.1992.tb05336.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Perlman P. S., Butow R. A. Mobile introns and intron-encoded proteins. Science. 1989 Dec 1;246(4934):1106–1109. doi: 10.1126/science.2479980. [DOI] [PubMed] [Google Scholar]
- Quirk S. M., Bell-Pedersen D., Tomaschewski J., Rüger W., Belfort M. The inconsistent distribution of introns in the T-even phages indicates recent genetic exchanges. Nucleic Acids Res. 1989 Jan 11;17(1):301–315. doi: 10.1093/nar/17.1.301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ripley L. S., Dubins J. S., deBoer J. G., DeMarini D. M., Bogerd A. M., Kreuzer K. N. Hotspot sites for acridine-induced frameshift mutations in bacteriophage T4 correspond to sites of action of the T4 type II topoisomerase. J Mol Biol. 1988 Apr 20;200(4):665–680. doi: 10.1016/0022-2836(88)90479-2. [DOI] [PubMed] [Google Scholar]
- Russell D. W., Jensen R., Zoller M. J., Burke J., Errede B., Smith M., Herskowitz I. Structure of the Saccharomyces cerevisiae HO gene and analysis of its upstream regulatory region. Mol Cell Biol. 1986 Dec;6(12):4281–4294. doi: 10.1128/mcb.6.12.4281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Russell R. L., Huskey R. J. Partial exclusion between T-even bacteriophages: an incipient genetic isolation mechanism. Genetics. 1974 Dec;78(4):989–1014. doi: 10.1093/genetics/78.4.989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sargueil B., Hatat D., Delahodde A., Jacq C. In vivo and in vitro analyses of an intron-encoded DNA endonuclease from yeast mitochondria. Recognition site by site-directed mutagenesis. Nucleic Acids Res. 1990 Oct 11;18(19):5659–5665. doi: 10.1093/nar/18.19.5659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharma M., Ellis R. L., Hinton D. M. Identification of a family of bacteriophage T4 genes encoding proteins similar to those present in group I introns of fungi and phage. Proc Natl Acad Sci U S A. 1992 Jul 15;89(14):6658–6662. doi: 10.1073/pnas.89.14.6658. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Szostak J. W., Orr-Weaver T. L., Rothstein R. J., Stahl F. W. The double-strand-break repair model for recombination. Cell. 1983 May;33(1):25–35. doi: 10.1016/0092-8674(83)90331-8. [DOI] [PubMed] [Google Scholar]
- Tabor S., Richardson C. C. A bacteriophage T7 RNA polymerase/promoter system for controlled exclusive expression of specific genes. Proc Natl Acad Sci U S A. 1985 Feb;82(4):1074–1078. doi: 10.1073/pnas.82.4.1074. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tian G. L., Michel F., Macadre C., Slonimski P. P., Lazowska J. Incipient mitochondrial evolution in yeasts. II. The complete sequence of the gene coding for cytochrome b in Saccharomyces douglasii reveals the presence of both new and conserved introns and discloses major differences in the fixation of mutations in evolution. J Mol Biol. 1991 Apr 20;218(4):747–760. doi: 10.1016/0022-2836(91)90263-6. [DOI] [PubMed] [Google Scholar]
- Turmel M., Boulanger J., Schnare M. N., Gray M. W., Lemieux C. Six group I introns and three internal transcribed spacers in the chloroplast large subunit ribosomal RNA gene of the green alga Chlamydomonas eugametos. J Mol Biol. 1991 Mar 20;218(2):293–311. doi: 10.1016/0022-2836(91)90713-g. [DOI] [PubMed] [Google Scholar]
- Wernette C., Saldanha R., Smith D., Ming D., Perlman P. S., Butow R. A. Complex recognition site for the group I intron-encoded endonuclease I-SceII. Mol Cell Biol. 1992 Feb;12(2):716–723. doi: 10.1128/mcb.12.2.716. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Winkler M., Rüger W. Cloning and sequencing of the genes of beta-glucosyl-HMC-alpha-glucosyl-transferases of bacteriophages T2 and T6. Nucleic Acids Res. 1993 Mar 25;21(6):1500–1500. doi: 10.1093/nar/21.6.1500. [DOI] [PMC free article] [PubMed] [Google Scholar]