Abstract
Unraveling the molecular basis of the life cycle of hepatitis C virus (HCV), a prevalent agent of human liver disease, entails the identification of cell-encoded factors that participate in the replication of the viral RNA genome. This study provides evidence that the so-called NF/NFAR proteins, namely, NF90/NFAR-1, NF110/NFAR-2, NF45, and RNA helicase A (RHA), which mostly belong to the dsRBM protein family, are involved in the HCV RNA replication process. NF/NFAR proteins were shown to specifically bind to replication signals in the HCV genomic 5′ and 3′ termini and to promote the formation of a looplike structure of the viral RNA. In cells containing replicating HCV RNA, the generally nuclear NF/NFAR proteins accumulate in the cytoplasmic viral replication complexes, and the prototype NFAR protein, NF90/NFAR-1, stably interacts with a viral protein. HCV replication was inhibited in cells where RNAi depleted RHA from the cytoplasm. Likewise, HCV replication was hindered in cells that contained another NF/NFAR protein recruiting virus. The recruitment of NF/NFAR proteins by HCV is assumed to serve two major purposes: to support 5′–3′ interactions of the viral RNA for the coordination of viral protein and RNA synthesis and to weaken host–defense mechanisms.
Keywords: dsRBM, hepatitis C virus, NF90, NFAR, ILF3, RHA
INTRODUCTION
Worldwide, approximately 200 million people are infected with hepatitis C virus (HCV) and at serious risk of developing chronic hepatitis and hepatocellular carcinoma. No vaccination procedures are available, and current therapies fail to eliminate the virus from a large number of patients (Pawlotsky 2006). A central issue for the development of alternative antiviral strategies entails the characterization of molecular determinants that govern the HCV life cycle.
HCV belongs to the Flaviviridae family, which includes other important human and animal pathogens as the flavivirus West Nile and the pestivirus, bovine viral diarrhea virus (BVDV). The HCV genome is a single-stranded RNA of positive polarity that consists of a long open reading frame (ORF) and 5′ and 3′ nontranslated regions (NTRs). As with all (+)-strand RNA viruses, the HCV genome has a double function in the host cell's cytoplasm. Following infection, the genome first operates as an mRNA. An internal ribosomal entry site (IRES) in the 5′ NTR mediates translation of the ORF by the cell's translation apparatus. The resultant polyprotein is processed by viral and cellular proteases to yield the viral structural and nonstructural (NS) proteins (Fig. 1). At a certain stage, the viral genome changes roles to become part of a membrane-associated replication complex (Gosert et al. 2003; Miyanari et al. 2003; Shi et al. 2003). Starting from the genomic 3′ end, this complex synthesizes (−)-strand intermediates, which, in a second step, act as templates for the transcription of progeny (+)-strand RNA molecules.
FIGURE 1.
HCV and BVDV RNAs. Organization of HCV and BVDV genomes and replicons. The depicted mono- and bicistronic HCV replicon constructs were applied throughout this work; the bicistronic replicons encoded an additional G418 resistance (NEO) gene (for details, see Grassmann et al. (2005). NTRs are shown as lines, genetic units as shaded boxes, known functions of the viral proteins are indicated (Lindenbach et al. 2007). Proteolytic cleavage sites in the polyprotein are marked as follows: arrow, cleavage by NS3/NS4A; circle, cleavage by signal peptidases; A, autoprotease activity; ubi, cleavage by ubiquitin carboxy hydrolase.
The molecular features that regulate viral translation and RNA replication are unknown. Studies with poliovirus, the prototype (+)-strand RNA virus, showed that translation and RNA replication are mutually exclusive processes (Barton et al. 1999). Other data obtained with poliovirus and with the flaviviruses Kunjin/West Nile and Dengue suggest that the viral RNAs form a circular structure that supports the coordination of viral protein and RNA synthesis (Herold and Andino 2001; Khromykh et al. 2001; Alvarez et al. 2005; Filomatori et al. 2006).
The analysis of the intracellular translation and replication process of the Flaviviridae RNA genome was considerably facilitated by cDNA constructs permitting the transcription and mutagenesis of functional viral RNA molecules and by the finding that subgenomic RNA molecules (“RNA replicons”) replicate independently from virion formation in transfected cells (Khromykh and Westaway 1997; Behrens et al. 1998; Lohmann et al. 1999). HCV replicons and replicons of the narrowly HCV-related BVDV show the same genomic organization; they essentially consist of the NTRs and the coding regions of the viral NS proteins NS3, NS4A, NS4B, NS5A, and NS5B (Fig. 1). Genetic studies of HCV and BVDV defined distinct RNA motifs in the conserved 5′ and 3′ ends of the viral genomes that are essential for the replication process (Frolov et al. 1998; Yanagi et al. 1999; Yu et al. 1999; Becher et al. 2000; Friebe et al. 2001; Friebe and Bartenschlager 2002; Yi and Lemon 2003; Lee et al. 2004; Pankraz et al. 2005). Experimental data obtained with BVDV replicons moreover showed that formation of the initial replication complex requires both genomic termini, and RNA motifs in the BVDV 5′- and 3′NTR were indicated to be active components of the functional switch between translation and replication. Thus, some signals in the 5′-terminal domain I of the BVDV 5′NTR determine the efficiency of the IRES while other elements are mandatory for RNA replication (Yu et al. 2000; Grassmann et al. 2005). The BVDV 3′NTR displays a bipartite organization: the 3′-terminal conserved segment (3′C) operates as a promoter element of the first replication step, while the upstream variable portion of the 3′NTR (3′V) appears acting as a decisive modulator of the translation and replication process. Jointly, these data implied that the coordination of translation and RNA replication involves dynamic 5′–3′ interactions of the BVDV RNA (Isken et al. 2003, 2004).
The characterization of RNA replicons established that, on the virus' side, the proteins encoded by genetic units NS3 to NS5B are necessary and sufficient to catalyze both RNA replication steps (Fig. 1; Grassmann et al. 2001). Thus, a vital question concerns the identity of host proteins, which, besides the viral proteins, are involved in RNA replication (for recent reviews on the status of recently identified host factors of Flaviviridae, see Behrens and Isken 2006; Lindenbach et al. 2007). Recently, we described a set of cellular proteins binding at high affinity to the 5′ and to the 3′NTR of the BVDV genome. The protein interaction sites were shown to involve RNA elements in the 3′V region of the BVDV 3′NTR and domains I and III of the BVDV 5′NTR, which suggested that the factors participate in the regulation of translation and RNA replication (see also Fig. 3B; Isken et al. 2003, 2004). We identified these proteins as a group of primarily nuclear factors including NF90/NFAR-1 (termed also DRBP76 or ILF3), NF110/NFAR-2 (an isoform of NF90/NFAR-1, also known as ILF3α), NF45, and RNA helicase A (RHA; NDHII). Two yet unidentified BVDV RNA-binding proteins, p64 and p130, were indicated to be related forms of NF90/NFAR-1 (p64) and RHA (p130), and NF45, NF90/NFAR1, and RHA were confirmed to be subunits of a cytoplasmic protein complex (termed here as the [NF90/NFAR-1, NF45, RHA] complex) (Liao et al. 1998). Thus, using the idiom “NF/NFAR proteins,” we emphasized that the proteins are either isoforms or interaction partners of NF90/NFAR-1. NF90/NFAR-1, NF110/NFAR-2, and RHA contain double-strand RNA-binding motifs (dsRBM) and hence belong to a protein family with widespread functional diversity, including, for example, the RNA regulated protein kinase PKR and the RNase III-like proteins Dicer and Drosha. Several reports suggest that NF/NFAR proteins are implicated in the regulation of gene expression on the transcriptional and post-transcriptional level. Other data imply that NF/NFAR proteins are occupied in RNA transport and that these factors are in a yet undefined manner part of antiviral measures of the cell (for review, see Saunders and Barber 2003; see also Discussion).
FIGURE 3.
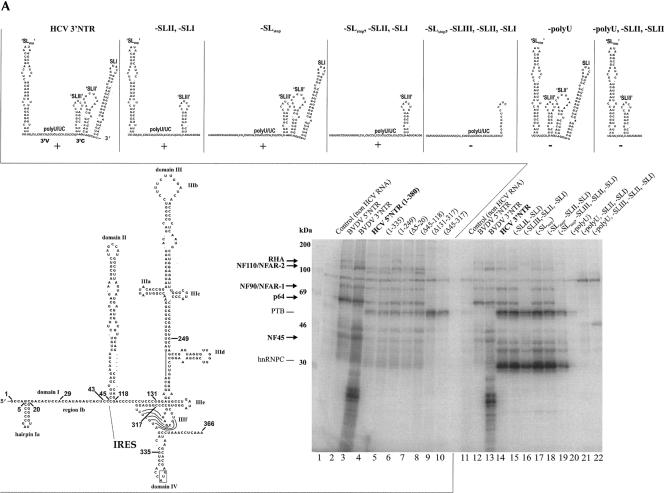
Mapping of the NFAR protein binding sites in the HCV NTRs. (A) UV cross-linking/label-transfer experiments with mutant transcripts of the HCV 5′- and HCV 3′NTR. (Bottom, left) Scheme of the structure of the HCV (strain 1b) 5′NTR; sites that were used to introduce deletions are indicated (SDS-PAGE); lanes 1–10, cross-linking experiments that were performed with HCV 5′NTR wild-type and mutant (Δ) transcripts (BVDV 5′- and 3′NTR applied as positive controls). The protein bands corresponding to the NFAR proteins, PTB, and hnRNPC are indicated (see also Fig. 2). (Top) Schematic representations of the structures of the wild-type and applied mutant HCV (strain 1b) 3′NTR transcripts (calculated by mfold). (SDS-PAGE) Lanes 11–22, cross-linking experiments performed with the HCV 3′NTR wild-type and mutant transcripts (BVDV 5′- and 3′NTR applied as positive controls). (B) Scheme of the NFAR protein-binding sites in the HCV and BVDV NTRs. The depicted RNA structures (HCV strain 1B; BVDV strain CP7) were experimentally proven or predicted by mfold (quotation marks indicate unverified motifs). The translational start and stop-codons are boxed; arrows indicate the considered 5′-boundary of the IRES and frames of the 3′V and 3′C regions. The known functions of the different RNA domains during translation and replication are indicated (see also text; question marks indicate preliminary data). Shadowed regions designate the defined NFAR protein-binding sites.
Here we provide several lines of experimental evidence indicating that NF/NFAR proteins are actively involved in the HCV replication process. Most interestingly, the [NF90/NFAR-1, NF45, RHA] complex was shown to trigger the formation of a loop structure of the HCV RNA that appears to be crucial for viral replication. Hence, this study revealed new important insights into molecular and possibly pathogenic features of the HCV life cycle.
RESULTS
NF/NFAR proteins specifically bind to the HCV 5′- and 3′NTR
The initial experiment of this study aimed at identifying cellular proteins that specifically interact with the HCV NTRs. For this purpose, we used the tobramycin-tag affinity purification method, which proved to be highly efficient and specific during analysis of the protein composition of prespliceosomal complexes (Hartmuth et al. 2002). For the purification procedure, we applied an RNA transcript that consisted of a tobramycin RNA aptamer (TobA), the HCV 5′NTR, a short portion of the open reading frame (sORF), and the HCV 3′NTR (TobA-5′NTR-sORF-3′NTR) (Fig. 2A). The sORF component was included because it was reported to permit correct functional folding of the complex RNA structure of the HCV 5′NTR (Honda et al. 1996a,b; Kieft et al. 2001). The RNA was incubated with cytoplasmic extract of Huh-7 hepatoma cells, which effectively support HCV replication (Lohmann et al. 1999; see also below), and protein complexes assembled on the TobA-5′NTR-sORF-3′NTR were affinity purified by binding to tobramycin–Sepharose (Fig. 2B, lane 3). Analogous procedures that applied no RNA or RNA lacking the TobA (5′NTR-sORF-3′NTR) served as negative controls (Fig. 2B, lanes 1,2). Proteins that remained binding to the RNA under stringent conditions were subsequently analyzed by SDS-PAGE and silver staining and, following extraction and digestion with trypsin, identified by mass spectrometry analysis (see Materials and Methods). Thus, among others, we identified the proteins hnRNP C, polypyrimidine tract binding (PTB) protein, and nucleolin, which were earlier reported to bind to the HCV NTRs (Gontarek et al. 1999; Ito and Lai 1999; Lu et al. 2004). Most interestingly, we also found the NF/NFAR proteins NF90/NFAR-1, NF45, and RNA helicase A (RHA) specifically binding to the HCV NTRs (Fig. 2B).
FIGURE 2.
NFAR proteins associate specifically with the HCV 5′- and 3′NTR. (A) Schematic representation of HCV RNA transcripts that were applied in the described affinity purification and UV cross-linking procedures, respectively. (B) Silver-stained SDS-PAGE of affinity-purified proteins: lane 1, control purification without RNA; lane 2, control with 5′NTR-sORF-3′NTR; lane 3, purification performed with TobA-5′NTR-sORF-3′NTR. The bands that contained RHA, NF90/NFAR-1, nucleolin, PTB, NF45, and hnRNP C (all proteins were not present in the controls) are marked. (C) Side-by-side UV cross-linking/competition and Western blot experiments. Cross-linking was performed with BVDV as well as HCV 5′- and 3′NTR transcripts and with non-HCV and HCV non-NTR control transcripts (for details, see Materials and Methods). Left panel: UV cross-linking/competition experiments that were carried out with the HCV 5′NTR transcript and cytoplasmic extracts of Huh-7 and HeLa cells, respectively. For comparison, cross-linking/label transfer experiments with the BVDV 5′- and 3′NTR were separated on the same gel; the supposed protein bands of p130, RHA, NF110/NFAR-2, NF90/NFAR-1, p64, NF45, PTB, and hnRNPC are indicated. Competition was carried out with the indicated molar excess (10× = 10 fold; 100× = 100 fold) of nonlabeled competitor RNA transcript. Right panel: Analogous UV cross-linking/competition experiments carried out with the HCV 3′NTR transcript. Right: for orientation, a Western blot was performed with total protein of HeLa S10 extract and a combination of anti NF45 and anti NF90/NFAR-1 antisera. The Western blot was performed on the very same SDS gel as the cross-linking experiments shown on the left. Extreme right: UV cross-linking/label-transfer experiment performed with HeLa S10 extract and non-NTR and non-HCV (Isken et al. 2003) control transcripts. In the case of the non-NTR control, an RNA transcript corresponding to nucleotides 7891–8162 of Con1 RNA was used (note that experiments with other non-NTR HCV probes led to identical results, not shown).
To confirm and further specify these findings, we next tested [32P]-labeled RNA transcripts that encompassed either the individual HCV 5′NTR or the HCV 3′NTR (Fig. 2A) in an established UV cross-linking/label transfer assay using again cytoplasmic extracts of Huh-7 cells as a protein source (for details, see Materials and Methods; Isken et al. 2003). Unrelated RNA transcripts (“non-HCV” RNAs) and various transcripts that encoded portions of the HCV replicon ORF (“non-NTR” HCV RNAs) were applied as negative controls. To focus the assay on the detection of RNA/protein interactions that occur at high affinity/specificity, it was performed at stringent conditions (e.g., in the presence of a 2000-fold molar excess of tRNA).
UV cross-linking/label transfer assays with transcripts of the BVDV 5′- and 3′NTR, which were carried out as positive controls, showed that under the applied conditions the NF/NFAR proteins and a few, yet undefined cellular proteins associate with the viral RNAs. Binding of NF/NFAR proteins was undetectable with “non-HCV” RNAs as well as with “non-NTR” HCV transcripts (one of five experiments with different “non-NTR” HCV transcripts is shown in Figs. 2C, 3A). The equivalent experiments with transcripts of the HCV NTRs gave rise to an RNA-charged protein pattern that showed distinct differences but also noticeable similarities to the BVDV situation. As expected, cross-linking of the HCV 3′NTR revealed the binding of PTB (57 kDa) and hnRNP C (35 kDa) (Gontarek et al. 1999) that were not observed with BVDV (Fig. 2C, lanes 21,28; Fig. 3A, lane 14). In accord with the report of Ito and Lai (1999), PTB was found interacting also with the HCV 5′NTR (see, for example, Fig. 2C, lanes 4,11; Fig. 3A, lane 4). However, in striking homology with BVDV, cross-linking/label transfer assays with the HCV 5′NTR or the HCV 3′NTR detected in each case a prominent RNA-charged protein that migrated at the same position as NF90/NFAR-1 (Fig. 2C, cf. lanes 4,11 and 21,28; Fig. 3A, cf. lanes 4 and 14; Fig. 2C, NF90/NFAR1 and NF45 were also stained by Western blotting). Moreover, we observed evident signals of proteins that were cross-linked to the HCV 5′NTR and 3′NTR, which corresponded closely to the RNA-labeled bands of NF110/NFAR-2, p64, NF45 (often migrating as a double band), and RHA in the BVDV experiments (Fig. 2C, cf. lanes 4,11 and 21,28; best seen in Fig. 3A, cf. lanes 4 and 14). These results were obtained irrespective of whether extracts of Huh-7 or HeLa cells were applied (Fig. 2C). Importantly, the cellular protein/HCV NTR interactions remained unaffected by the presence of an excess of unlabeled nonspecific RNA but were chased away by added unlabeled HCV 5′NTR or HCV 3′NTR (Fig. 2C, cf. lanes 5–8,12–15 and 22–25,29–32): i.e., while a 10- to 20-fold molar excess of HCV competitor yielded only little competition, we observed an evident competition with a 100-fold molar excess with respect to the nonspecific control. Consistent with the idea that we were dealing with the same set of proteins, also the BVDV 5′- and 3′NTRs were effective as competitors of protein binding to the HCV NTRs. Competition turned out to be even more active or prominent with the BVDV NTRs, i.e., already with a 20-fold excess of BVDV NTR transcript, we observed effective competition (Fig. 2C, cf. lanes 9,10,16,17 and 26,27,33, 34). This was congruent with the observation that the NF/NFAR proteins generally associated at higher efficiency with the BVDV NTRs (compare, for example, Fig. 2C, cf. lanes 2,3 and 4,21). Finally, in concurrence with the fact that in comparison to the situation with HCV RNA PTB and hnRNP C were binding only poorly to the BVDV NTRs, the PTB and hnRNP signals detected in the cross-linking assays with HCV 3′NTR remained mainly unaffected in competition assays with BVDV RNA but were chased away by competition with HCV RNA (Fig. 2C, cf. lanes 24–26 and 32–34).
We took these data as significant clues that NF/NFAR proteins were binding in a specific manner to each of the HCV NTRs.
Binding of NF/NFAR proteins to the HCV 5′- and 3′NTRs involves defined RNA motifs
To disclose the viral RNA motifs engaged in the interaction with the cellular proteins, we performed a systematic deletion mutagenesis of the HCV 5′- and 3′NTRs, where we deleted domains of defined RNA secondary structure. The mutant RNAs were analyzed with the above cross-linking assay for protein binding. Performing time-course RT-PCR experiments, all listed deletion mutants were tested in the context of total replicon RNA as well as in the transcripts used for UV cross-linking and shown to have no effect on the stability of the RNAs under the conditions of replication and cross-linking experiments (data not shown; Grassmann et al. 2005).
Thus, with the 5′NTR, we examined transcripts with deletions of the hairpin Ia (Δ5–20) or the Ib region (Δ29–43) in domain I. Moreover, we tested 5′NTR transcripts that lacked domain II (Δ45–118) and/or domain III (Δ45–317; Δ131–317), the entire 3′ portion (1–249), or the domain IV segment (1–335) (see Fig. 3A). We found that binding of the cellular proteins was still detectable with transcripts that lacked parts of domain I (data not shown; Δ5–20; Δ29–43; Fig. 3A, lane 7), the entire domain II (Δ45–118; Fig. 3A, lane 8), domain IV (1–335; Fig. 3A, lane 5), or the downstream portion of domain III (1–249; Fig. 3A, lane 6). In contrast, protein binding did not occur with transcripts that lacked the entire domain III (Δ45–317; Δ131–317; Fig. 3A, lanes 9,10). Although the exact definition of the protein interaction site in the 5′NTR requires further thorough mapping analysis that will be addressed in a separate study, the available data clearly suggest that binding of the cellular proteins to the 5′NTR involves specifically the apical portion of domain III (summarized in Fig. 3B).
With the HCV 3′NTR, we performed a similar mutagenesis approach deleting stepwise individual RNA motifs (schematized in Fig. 3A). While testing these mutants by cross-linking/label transfer analysis, we observed that association of the cellular proteins with the 3′NTR occurred as well in the absence of a stem–loop structure named SLstop (owing to its vicinity to the ORF stop-codon), which according to mfold estimates may be formed by the variable 3′V region. Protein binding did also not require the presence of the stem–loop structures SLI and SLII that are folded by the conserved 3′C region near the genomic 3′ end (Fig. 3A, lanes 15,17,18). Conversely, we observed that association of the cellular proteins with the viral RNA was strictly reliant on the presence of the poly-U/UC stretch, a specific feature of the HCV 3′NTR, as well as on the presence of a predicted stem–loop motif, SLIII, in the upstream portion of the 3′C region (Fig. 3A, cf. lanes 16 and 19–21). Since protein binding was only detectable with transcripts that contained both intact elements, we took this as an indication that poly-U/UC and SLIII are parts of a common RNA motif that is crucial for the formation of the cellular protein/3′NTR complex (summarized in Fig. 3B).
To conclusively confirm the data shown in Figures 2 and 3, we decided to perform RNA-protein coimmunoprecipitation (IP) experiments applying a rabbit antiserum against NF45 and a monoclonal antibody (mAb) against NF90/NFAR-1. Both antibodies were previously shown to precipitate the individual proteins NF45 and NF90/NFAR-1, the NF45, NF90/NFAR-1 heterodimer (Kao et al. 1994), as well as the [NF90/NFAR-1, NF45, RHA] complex from cytoplasmic extracts (see Fig. 4; O. Isken, unpubl. data). As a source for precipitation, we used S10 extracts of Huh-7 cells that were supplemented with [32P]-labeled HCV 5′- or HCV 3′NTR transcripts, respectively. As controls, we employed extracts that contained equal or higher molar amounts of nonrelated RNAs (not shown) or 5′- or 3′NTR transcripts where the previously defined NFAR binding sites were defective caused by the deletion of the IRES domains II and III or the polyU/UC stretch, respectively (Fig. 4, lanes 1–4; see also Fig. 3A).
FIGURE 4.
Coimmunoprecipitation of HCV NTR transcripts by anti-NF90/NFAR-1 and anti-NF45 antibodies. The IPs were carried out with a non-immune (control) rabbit antiserum, a rabbit anti NF45 antiserum, and a monoclonal anti NF90/NFAR-1 antibody (see also Materials and Methods). S10 extract of Huh-7 cells was applied as a protein source; the extract was supplemented with equal masses (10 ng) of (32P)-labeled transcripts of the wild-type or mutant HCV 5′ and 3′NTR, respectively (the organization of the applied mutant transcripts is depicted in Fig. 3A). (Top) RNA analysis on a 7 M urea, Tris/borate acrylamide gel; lanes 1–4, transcript input (10% of total input shown; due to their lower molecular weights, the mutant transcripts were applied at a slight molar excess); lanes 5–16, immunoprecipitated RNAs. (Bottom) Protein analysis by SDS-PAGE and Western blot: lanes 5–16, the protein samples that correspond to the above RNA precipitation were probed with a combination of rabbit antisera against NF45 and NF90/NFAR-1. Note that these antisera were different from the originally applied precipitating antibodies (see Materials and Methods). The precipitated NF45 and NF90/NFAR-1 and the costained IgG antibody bands are indicated; lane 17, for orientation, a fraction of nuclear extract was loaded and probed by Western blot and anti-NF90 and anti-NF45 antibody.
While control antibodies precipitated neither proteins nor RNA transcripts (Fig. 4, lanes 5–8), both the anti-NF45 as well as the anti-NF90/NFAR-1 antibody precipitated specifically NF45 and NF90/NFAR-1 and also coprecipitated the indicated HCV NTR transcripts (Fig. 4, cf. lanes 9,11 and 13,15). In contrast, RNA coprecipitations were untraceable with non-HCV or non-NTR transcripts (not shown). Most importantly, neither the anti-NF45 nor the anti-NF90/NFAR-1 antibody coprecipitated HCV 5′- or 3′NTR RNAs that lacked the NF/NFAR protein interaction sites (Fig. 4, cf. lanes 10,12 and 14,16). Matching results were obtained when we performed the experiments with equimolar mixtures of wild-type and mutant transcripts: i.e., also in this case, the NF/NFAR proteins coprecipitated exclusively with the wild-type 5′- and 3′NTR transcripts (not shown).
Thus, in sum, the situation with HCV resembled that of BVDV, i.e., with both viral RNAs, NF/NFAR proteins were found binding at high specificity to defined RNA motifs in each of the nontranslated regions of the viral genome (see Fig. 3B and Discussion).
NF/NFAR proteins promote 5′–3′ interactions of the HCV RNA
The fact that NF/NFAR proteins were observed associating with the 5′NTR as well as with the 3′NTR of the HCV genome suggested that the proteins were involved in mediating physical interactions between both ends of the viral genome. To test this notion, we pursued two experimental approaches. First, we performed a pull-down assay that applied streptavidin beads loaded with biotinylated transcripts of the viral 5′NTR (or 3′NTR) to test for an interaction/coprecipitation of radioactively labeled transcripts of the 3′NTR (or 5′NTR) (see Fig. 5A and Materials and Methods). Thus, RNA–RNA coprecipitation was attempted in the absence of protein, in the presence of an unrelated protein (BSA), an unrelated RNA binding protein (hsp90), an RNA-binding protein involved in HCV replication (HCV NS3), and in the presence of the purified [NF90/NFAR-1, NF45, RHA] protein complex (abbreviated [NFAR] in Fig. 5A). As also shown in Figure 5A, we detected only weak RNA–RNA coprecipitations in the experiments that applied either biotinylated nonrelated control RNA (Fig. 5A, cf. lanes 4–7 and 19–22) no protein (Fig. 5A, cf. lanes 4,8,12 and 19,23,27), BSA (not shown), hsp90, or HCV NS3 (Fig. 5A, cf. lanes 1,2,5,6,9,10,13,14 and 16,17,20,21,24,25,28,29). In contrast, we observed a significant level of 5′–3′ RNA coprecipitation when we performed the experiment with the purified [NF90/NFAR-1, NF45, RHA] complex (Fig. 5A, lanes 15,30; see Materials and Methods for details). Importantly, the (NFAR)-mediated RNA–RNA contacts were only detectable in the pull-down experiments where we applied the HCV 5′NTR and 3′NTR while this was not the case with nonspecific RNAs or when we applied RNAs of equal polarity in the assay (e.g., labeled and biotinylated 3′NTR transcripts; Fig. 5A, lanes 7,11,22,26). Hence, these data supported the idea that the [NF90/NFAR-1, NF45, RHA] complex promotes specific interactions between the HCV 5′- and 3′NTR.
FIGURE 5.
NFAR proteins mediate RNA 5′–3′ interactions. (A) 5′–3′ coprecipitation assay. (Left) Silver-stained SDS-PAGE of a chromatographic fraction of the purified [NF90/NFAR-1, NF45, RHA] complex (see also Isken et al. [2003]). (Middle) Schematic sketch of the experimental assay. (Right) Autoradiographs of coprecipitation experiments that were supplemented with the indicated proteins (see Materials and Methods; [NF90/NFAR-1, NF45, RHA] complex abbreviated as [NFAR]). The RNA–RNA coprecipitation experiments were performed in all conceivable combinations, i.e., with the (32P)-labeled NTRs alone (lanes 1–4 and 16–18); with the (32P)-labeled NTRs and biotinylated control RNA (lanes 4–7 and 19–22); with (32P)-labeled 5′NTR and biotinylated 5′NTR (lanes 8–11); (32P)-labeled 5′NTR and biotinylated 3′NTR (lanes 12–15); (32P)-labeled 3′NTR and biotinylated 3′NTR (lanes 23–26); (32P)-labeled 3′NTR and biotinylated 5′NTR (lanes 27–30). The percentile of precipitated labeled RNA is indicated. (B) RNA electron micrographs. (Top) Scheme of the applied RNA molecule (see also Materials and Methods) and analytical agarose gel stained with ethidium bromide indicating the formation of RNA duplex. (Middle) Representative EM images of RNA molecules with wild-type NTRs that were supplemented with BSA or with the purified [NF90/NFAR-1, NF45, RHA] complex. The BSA-supplemented RNAs appeared nearly exclusively (∼98%) as ∼300-nm elongated structures, while ∼14% formed looplike structures in the presence of the NFAR complex (as shown). (Bottom) Statistics of images with wild-type and mutant RNAs; the applied mutant RNAs had a deficient NFAR-binding site in the 3′NTR (−poly-U; see Fig. 3A).
To further substantiate this concept, we decided to directly picture protein-mediated changes of the HCV RNA conformation by electron microscopy (EM). Considering previous studies on the conformation of mRNA molecules (Wells et al. 1998), we conducted these experiments with RNA molecules that consisted of the single-stranded NTRs and a double-stranded portion of the ORF. Fixed with uranyl acetate, these RNAs were clearly distinguishable by EM (Fig. 5B). Examining approximately 100 images of RNAs that contained wild-type NTRs or NTRs that lacked parts of the NF/NFAR protein-binding site in the 5′- or in the 3′NTR (e.g., the poly-UC/U stretch in the 3′NTR, see Fig. 3A), we found that irrespective of the presence of unrelated protein (BSA) most molecules existed primarily in a stretched conformation. However, with the added, purified [NF90/NFAR-1, NF45, RHA] complex, we observed a significant number of wild-type RNAs displaying a “looplike” configuration (i.e., ∼14%), while this was not the case with the mutant RNAs (Fig. 5B).
These data corroborated the notion that NF/NFAR proteins associating with the viral NTRs support the formation of a looplike conformation of the HCV RNA.
Experimental evidence that NF/NFAR proteins are components of the HCV replication complex
The above in vitro data implied that NF/NFAR proteins, while binding to the viral RNA, participate in the functional assembly of the HCV replication complex. The following experiments were aimed at further testing this notion.
First, we wanted to find out whether interactions of NF/NFAR proteins, such as that of NF90/NFAR-1 with the viral RNA, occurred also in cells that contained active viral replication complexes. Accordingly, we performed an IP experiment that applied the aforementioned anti-NF90/NFAR-1 mAb and, as a protein source, S10 extracts of Huh-7 cells that were persistently transfected with HCV replicon. The precipitate was tested by Western blot for the NF90/NFAR-1 antigen and by semiquantitative RT-PCR for coprecipitated HCV RNA. Thus, in contrast to experiments with unrelated antibody, we found that IPs with anti NF90/NFAR-1 precipitated, as expected, NF90/NFAR-1 (shown in Fig. 6B; see also Fig. 4) and also coprecipitated a significant amount of HCV RNA in the replicon-transfected cells (Fig. 6A). We took this result as indication for a specific interaction between NF90/NFAR-1 and HCV RNA in HCV replicon-harboring cells.
FIGURE 6.
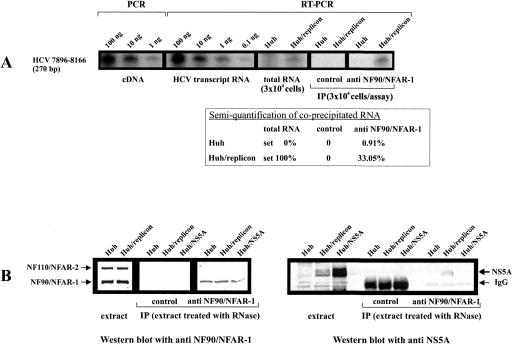
NF90/NFAR-1 interacts with the viral RNA and NS5A in HCV-transfected cells. (A) Coimmunoprecipitation (IP) of NF90/NFAR-1 and HCV RNA. The IP was carried out with anti NF90/NFAR-1 mAb, a control mAb and S10 extracts of 3 × 106 naive or HCV replicon-transfected Huh-7 cells. Coprecipitated viral RNA was detected by RT-PCR, which generated a 270-bp product. As controls and for semiquantification, we performed PCRs or RT-PCRs with the indicated amounts of cDNA, replicon transcript, and total RNA extracted from 3 × 104 naive or HCV replicon-transfected cells. The amount of HCV RNA that was coprecipitated by the anti NFAR-1 antibody and by the control mAb was estimated with respect to these controls (considering that these controls applied a 100-fold lower amount of cells): the table shows average values of four experiments. Note that precipitations that were carried out in the presence of micrococcal nuclease (75 U/extract of 106 cells) resulted negative results (not shown). (B) Co-IP of NF90/NFAR-1 and NS5A. The IP was performed with anti NF90/NFAR-1 mAb and a control mAb using RNase-treated S10 extracts of naive Huh-7 (Huh), Huh-7 that carried HCV replicon (Huh/replicon), and Huh-7 that had been transfected with a plasmid expressing the HCV NS5A (Huh/NS5A). Precipitated NF90/NFAR-1 was detected by Western blots with an antiserum that also detects NF110/NFAR-2 (see control with total, untreated extract; note that the anti-NF90/NFAR-1 mAb applied for IP did not coprecipitate NF110). Expression and coprecipitation of NS5A was detected by Western blots with a rabbit serum against the viral protein (staining NS5A in each case as a double band). Note that the control antibody yielded a stronger IgG band than the NF90/NFAR-1 mAb. Co-IP of NF90/NFAR-1 and NS5A was detected irrespective of whether the extract was treated with RNase A (20 μg/extract of 106 cells) or not.
Next, we performed the analogous IP experiment with anti-NF90/NFAR-1 antibody but analyzed the precipitate for coprecipitating viral proteins. While the detection of potentially coprecipitating NS3, NS4A/B, and NS5B remained uncertain for technical reasons (not shown), we observed an evident and specific coprecipitation of NF90/NFAR-1 and the HCV NS5A protein (Fig. 6B). Importantly, a specific co-IP of NF90/NFAR-1 and NS5A was observed irrespective of whether the applied extract was treated with RNase. In contrast, a co-IP was undetectable with extracts of cells that contained heterologously expressed NS5A (Fig. 6B). These findings were confirmed by inverse IPs with a precipitating anti-NS5A antiserum; i.e., also in this case we observed a specific coprecipitation of NS5A and NF90/NFAR-1, but only with lysates of HCV replicon-transfected cells (not shown). We interpreted these experiments such that the cellular NF90/NFAR-1 protein gets into stable contact with the viral NS5A protein, but only in those cells that form functional HCV replication complexes. Apparently, this contact involves neither RNA-mediated nor direct protein–protein interactions but stable though indirect interactions between NF90/NFAR1 and NS5A (see Discussion).
To directly explore interactions of NF/NFAR proteins with the HCV replication complex, we next investigated persistently HCV replicon-transfected Huh-7 cells with a double-staining immunofluorescence (IF) procedure. To this end, we stained the cells with specific rabbit antisera against NF90/NFAR-1, NF110/NFAR-2, RHA, or NF45 and with monoclonal antibodies (mAbs) against the HCV NS3 or NS5A proteins and examined the cells by confocal microscopy (note that IF experiments with anti-NS3 mAb and anti-NS5A mAb yielded identical results). Employing the same set of anti-NFAR antisera and an anti-BVDV NS3 mAb, we performed analogous IF experiments with transiently BVDV replicon-transfected Huh-7 cells and with BVDV-infected MDBK cells.
It is important to note that the applied IF procedures detected HCV and BVDV proteins solely in the case of viral replication. In other words, viral protein synthesis was imperceptible in cells that were transfected with translation-competent but replication-defective RNAs (see also Fig. 9; Behrens et al. 1998). We found that with persistently HCV-transfected Huh-7 cells, ∼40%–60% showed a measurable level of expression of NS3 and NS5A. Characteristic of functional HCV replication complexes, the viral proteins accumulated around the perinuclear region with the endoplasmic reticulum (Fig. 7; Gosert et al. 2003). The residual cells were not stained by the anti-NS3 or anti-NS5A mAb, because the viral replication cycle was most probably in an early or late stage (Miyanari et al. 2003). When we counterstained these replication-negative cells with the various anti-NFAR antisera, the NF/NFAR proteins were found nearly completely in the nucleus (Fig. 7), which was indistinguishable from the situation observed with naive (nontransfected) Huh-7 cells (not shown). For that reason, we considered the replication-negative cells as internal negative controls. In remarkable contrast, in the cells that were positive for HCV replication the entire set of NFAR proteins (RHA; NF110/NFAR-2, NF90/NFAR-1, and NF45) was also detectable in the cellular cytoplasm where the proteins largely colocalized with the viral replication complexes (Fig. 7). Conversely, when we investigated cells that expressed the HCV NS3 or NS5A protein individually, NF/NFAR proteins were undetectable in the cytoplasm (not shown). Hence, in agreement with the above coprecipitation data, only in those cells that showed active HCV replication, the mainly nuclear NF/NFAR proteins assembled at significant amounts in the cellular cytoplasm. Identical results were obtained with the BVDV experiments showing that also under the conditions of transient transfection (not shown) and infection experiments, the NF/NFAR proteins were specifically accumulating in the cytoplasmic viral replication complexes (Fig. 7; see also Discussion).
FIGURE 9.
Interference of HCV and BVDV replication. Naive and HCV replicon carrying Huh-7 cells were transfected with replicative or non-replicative (RdRp inactivated) monocistronic BVDV replicon. Replication of HCV and BVDV was monitored by complementary IF staining of the cells with anti-HCV NS5A rabbit antiserum and anti-BVDV NS3 mAb, respectively. The bar diagram reflects a representative experiment that determined the fractions of cells of a tissue culture plate (the total number of cells was calculated by DAPI staining of the nuclei) that support HCV or BVDV replication in the absence (set 100%) or presence of the respective other RNA. (Right panel) Transfection experiments of naïve and HCV replicon carrying Huh-7 cells with fluorescent-marked (Cy3) siRNAs indicating identical receptiveness of both cell types for RNA transfections.
FIGURE 7.
NF/NFAR proteins accumulate at the sites of viral replication in the cytoplasm. Representative confocal images (600× enlarged) of MDBK cells infected with BVDV and of persistently HCV replicon-transfected Huh-7 cells. (Upper panel) MDBK cells were infected at 0.1 MOI with BVDV strain CP7, 24 h post-infection, the cells were stained with an anti BVDV NS3 mAb and counterstained with a rabbit antiserum against NF45. (Lower panels) Huh-7 cells carrying the HCV (Con1, strain 1B) replicon were stained with a mAb against NS5A and counterstained with antisera against the different NFAR proteins. Both fields were independently analyzed, and a colocalization of viral and cellular proteins is shown by merging. (Bottom) HCV replicon-carrying cells stained with anti-NS5A and anti-NF90/NFAR-1 antibody (enlarged).
In sum, these experiments provided consistent evidence that the cellular NF90/NFAR-1, NF110/NFAR-2, NF45, and RHA proteins are recruited components of the HCV replication complex.
Significance of NF/NFAR proteins for HCV replication
To address the question whether NF/NFAR proteins actively participate in the HCV replication process, we performed three sets of experiments.
First, we wanted to test HCV replication in cells where the levels of NFAR proteins had been reduced by RNAi (Elbashir et al. 2001). Pilot experiments established that naive Huh-7 cells and Huh-7 cells carrying the HCV replicon were equally transfectable with siRNAs or other replicon RNAs (i.e., at 80%–90% efficiency; see Fig. 9). However, attempts to deplete the cells by RNAi from NF90/NFAR-1 or NF45 failed as considerable changes of the protein quantities caused instant cell death (not shown). Nevertheless, by transfecting a mixture of two anti-RHA siRNAs it was possible to reduce the level of cytoplasmic RHA by ∼40%–60% during the following 2 days without affecting cell viability over at least 5 days (Fig. 8; Isken et al. 2003). Most importantly, in contrast to Huh-7 replicon cells that were transfected with control siRNAs, we observed an immediate reduction of HCV replication by 60%–80% in cells that had been treated with the anti-RHA siRNAs (Fig. 8).
FIGURE 8.
HCV replication is inhibited in cells with lower levels of RHA. The experiment was carried out with RNAi depleted Huh-7 cells that were transiently transfected with monocistronic HCV replicon RNA (see Materials and Methods). (Upper panel) Western blot indicating the level of RHA at day 4 post-transfection of anti-RHA or control siRNAs (error bars indicate mean deviations of three different experiments. (Lower panel) Representative RNase protection experiment measuring HCV RNA replication at days 2–5 post-transfection in cells that had been transfected with anti RHA or control siRNAs. The same cell preparations were also applied in previous experiments with BVDV; GAPDH was used as a loading control. Note that the cells showed no signs of apoptosis over this time and that replication was equally inhibited when we transfected varying amounts of HCV RNA (Isken et al. [2003] and data not shown).
Detailed mutagenesis studies of the HCV 5′NTR and of the HCV 3′NTR were reported by several laboratories, and these studies strongly suggested that the 5′NTR domain III region and the 3′NTR poly-U/UC and SLIII motifs, which turned out here to be involved in NF/NFAR protein binding (Fig. 3A), are essential for RNA replication (for examples, see Yanagi et al. 1999; Friebe et al. 2001; Friebe and Bartenschlager 2002; Yi and Lemon 2003; Grassmann et al. 2005). Nevertheless, since most of the earlier examined mutants differed from the mutants that were applied in the protein interaction experiments shown in Figure 3A, we considered it important to understand whether a precise removal of these domains prohibiting NF/NFAR protein binding (see Fig. 4) would interfere with RNA replication. Monitoring RNA replication via selection experiments, RNase protection or RT-PCR in comparison with wild-type controls, we investigated the replication of selectable bicistronic HCV replicons as well as of monocistronic HCV replicons that contained deletions of the 5′NTR domain III (Δ131–317), of the 3′NTR poly-U/UC stretch (−poly-U/UC), and/or of the 3′NTR SLIII (−SLIII), respectively (see Fig. 3A). In broad congruence with the previously published data, we found that each of these mutations completely blocked HCV replication (data not shown).
The last-performed experiment of this series followed the idea that the NFAR proteins were essential and perhaps limiting host factors of HCV replication. Such a scenario would imply that the presence of another NF/NFAR protein recruiting virus as BVDV would interfere with HCV replication and vice versa. To address this, we first compared the rate of BVDV replication in naive and HCV replicon-transfected Huh-7 cells (as already stated, both cell types were evenly transfectable with RNA) (Fig. 9). HCV and BVDV replication were determined simultaneously by complementary IF-staining that measured the replication-associated synthesis of the HCV NS5A and BVDV NS3 proteins. Thus, evaluating approximately 200 cells per experiment, we found that the fraction of Huh-7 cells that supported BVDV replication was five times lower if the cells contained replicating HCV RNA. Conversely, with the HCV replicon containing Huh-7 cells, a cotransfection of replicative BVDV RNA reduced the number of cells that were positive for HCV replication by almost 70%, while the level of HCV replication remained unaffected if the cells were cotransfected with a translation-competent but nonreplicative BVDV RNA (Fig. 9). Most importantly, the total number of cells that evidently supported viral replication (either BVDV or HCV) was found to be the same in each of the different experiments, and side-by-side replication of BVDV and HCV was never detected in the same cell (Fig. 9).
In sum, these experiments supported the notion that NF/NFAR proteins actively participate in the replication of HCV (see Discussion).
DISCUSSION
In this study, we examined the cellular “NF/NFAR proteins,” a group of predominantly nuclear proteins including NF90/NFAR-1, NF110/NFAR-2, NF45, and RHA, for a role in the replication of the important human pathogen HCV. The NF/NFAR proteins mostly belong to the dsRBM family, many members of which are in the current focus of interest due to their emerging roles in RNA metabolism (Saunders and Barber 2003).
Initial evidence pointing to a function of NF/NFAR proteins in HCV replication came from affinity purification, cross-linking, and immunoprecipitation assays that revealed specific interactions of the whole set of proteins with distinct RNA motifs in the HCV 5′NTR and HCV 3′NTR (Figs. 2, 3). These data were considerably strengthened by immunofluorescence studies showing that in cells that contain replicating HCV RNA, NF90/NFAR-1 and NF45, and also NF110/NFAR-2 and RHA, are specifically recruited to the viral replication complexes in the cytoplasm (Fig. 7). In addition, we detected stable contacts between NF90/NFAR-1 and the viral RNA and between NF90/NFAR-1 and NS5A in HCV-transfected cells (Fig. 6).
The idea that NF/NFAR proteins are functional components of the HCV replication process was sustained by a series of findings. (1) Accumulation of NF/NFAR proteins in the cytoplasm and interactions between NF90/NFAR-1 and NS5A were exclusively detectable in cells that formed productive HCV replication complexes (Figs. 6, 7). (2) Interactions of the NF/NFAR proteins to the HCV 5′- and 3′NTRs involve RNA motifs that are indispensable for viral replication (Fig. 3). (3) HCV replication was inhibited in cells with lower levels of RHA in the cytoplasm (Fig. 8). (4) HCV replication was inhibited by coreplicating BVDV RNA (Fig. 9). The interpretation of the RNAi, mutagenesis, and BVDV-HCV competition experiments is certainly limited. For example, RNAi-mediated depletion of RHA may cause unrelated changes in the cell's physiology, which, in turn, block viral replication. Along the same line, the here-determined NF/NFAR protein binding sites are also important for the binding of other protein factors; in other words, the inhibitory effect observed upon deletion of these regions of the viral RNA is not necessarily linked to NF/NFAR protein binding. Finally, the observed incompatibility of BVDV and HCV replication may certainly be due to other reasons than a restricted availability of NF/NFAR proteins. However, the striking consistency of data that was obtained with rather different assay systems clearly support the concept that the NF/NFAR proteins are representing HCV host factors.
Our experiments suggest that the cytoplasmic NF45, NF90/NFAR-1, and RHA proteins are largely active as a protein complex and that a main function of this complex in HCV replication may be to interact with the 5′- and 3′NTRs of the viral RNA and to bring the RNA's termini into close proximity. This notion was supported by pull-down assays, which showed that HCV 5′–3′ RNA/RNA interactions are specifically augmented by the purified [NF90/NFAR-1, NF45, RHA] complex. Moreover, the added NFAR protein complex significantly stimulated the formation of looplike RNA structures as demonstrated by EM studies (Fig. 5). While the mechanism by which [NF90/NFAR-1, NF45, RHA] promotes HCV RNA 5′–3′ interactions has yet to be defined, the fact that the NF/NFAR proteins were found to interact specifically and exclusively with essential replication signals in the HCV NTRs suggests that genome circularization is a decisive precondition of the HCV life cycle. Hence, these studies support the paradigm of 5′–3′ RNA–RNA interactions in the replication of (+)-strand RNA viruses. Most interestingly, our data revealed that the circularization of HCV RNA apparently demands the formation of a unique viral/cellular ribonucleoprotein complex.
The 5′–3′ interactions of the viral RNA are an assumed critical determinant of the coordination of viral translation and RNA replication (see Introduction). Accordingly, our observations raise the question of the role of the NF/NFAR proteins in this important stage of the viral life cycle. Suggestions may come from a recent study of the BVDV 3′NTR showing that formation of the viral RNA/NFAR protein complex with the 3′NTR 3′V region (see Fig. 3B) is important for optimal activity of the BVDV IRES. In addition, we found that the 3′V/NFAR protein complex is crucial for accurate termination of translation at the ORF stop-codon, which, in turn, was demonstrated to be an essential prerequisite for the initiation of the viral replication cycle (see Isken et al. [2004] and model herein). The BVDV data therefore imply that during the early mRNA phase of the viral genome, the NFAR-mediated RNA looplike structure favors translation, perhaps by enabling ribosomes to reinitiate after an accomplished round of translation. Concurrently, the 3′V/NFAR protein complex prohibits the translation machinery from interfering with the assembly of the replication complex, which, in the case of BVDV involves the 3′NTR 3′C region and the conserved 5′-terminus of the viral genome (Yu et al. 1999, 2000; see also Fig. 3B). In view of this scenario, it is intriguing that with the HCV 3′NTR the NFAR proteins were found to bind to the HCV poly-UC/U, SLIII segment at the border between 3′V and 3′C (Fig. 3B). Preliminary data indeed suggest that the HCV poly-UC/U, SLIII may modulate translation and RNA replication in a similar way as the BVDV 3′V region (O. Isken, unpubl.). Concerning the HCV 5′NTR, our data indicate a potential overlap of the NFAR and eIF3 binding sites (Fig. 3B; Sizova et al. 1998). This again has interesting implications, since, at a later stage of the infection process and, perhaps, in a different context of factors, NF/NFAR proteins could also be part of the inhibition of translation by interfering with the interaction of eIF3 with the IRES. Experiments testing this possibility are underway.
Two related concepts are currently discussed of how (+)-strand viral RNAs may switch from translation to replication. One model posits that the emerging replication complex causes conformational changes of the viral RNA, which eventually block the IRES (Isken et al. 2004). The second scenario favors direct feedback inhibition of translation by a viral protein (Gamarnik and Andino 1998). To this end, it is remarkable that we observed stable interactions between NF90/NFAR-1 and NS5A, the only viral protein that operates in trans during formation of the replication complex and which may, in fact, operate as a feedback inhibitor (Grassmann et al. 2001; Appel et al. 2005; Kalliampakou et al. 2005). NS5A has gained much attention since the protein, besides being part of the viral replicase, is thought to modulate the interferon response and physiology of infected cells (Macdonald and Harris 2004). Moreover, NS5A is known to interact with a whole set of cellular factors, one of which is the SNARE-like protein hVAP-33 that is suggested to play a fundamental role in intracellular vesicle biogenesis and trafficking. There is strong evidence that hVAP-33, via its contact with NS5A, essentially participates in HCV replication (Evans et al. 2004; Gao et al. 2004). Thus, it will be exciting to understand whether hVAP-33 is involved in the stable, although apparently indirect, contact between NF90/NFAR-1 and NS5A (Fig. 6).
Besides our reports on HCV and BVDV, there are hints that other flaviviruses and also picornaviruses recruit NF/NFAR proteins during translation and RNA replication (O. Isken, unpubl.; Merrill and Gromeier 2006). This leads to the question of why different viruses engage this group of cellular factors particularly in their reproduction process. As already mentioned, NF90/NFAR-1, NF110/NFAR-2, and RHA attracted notice as potential regulators of gene expression on the transcriptional level (Reichman et al. 2002; Reichman and Mathews 2003; Zhang and Grosse 2004). Thus, it is remarkable that in interferon-stimulated cells, RHA translocates to promyelocytic leukemia nuclear bodies (PML NBs) suggesting that the enzyme acts as a potential regulator of interferon-inducible genes attached to PML NBs (Fuchsova et al. 2002). This, in turn, fuels the idea that NF/NFAR protein recruiting viruses act in two ways to their own needs as the exploitation of NF/NFAR proteins as components of the viral replicase in the cytoplasm may remove the proteins from sites in the nucleus involved in the stimulation of an antiviral state. In accord with this idea, it was reported that overexpression of NF90/NFAR-1 activates the cellular antiviral expression cascade (Krasnoselskaya-Riz et al. 2002), and NF90/NFAR-1 was shown to modulate the activity of PKR, another key factor of the cell's interferon response (Parker et al. 2001; Saunders et al. 2001). Hence, as an intriguing scenario, various viruses may subvert NF/NFAR proteins from potential opponents to supporters of their infection route. Future work needs to address how the interplay of the NF/NFAR proteins with the viral RNA contributes to HCV pathogenesis and to assess the [NF90/NFAR-1, NF45, RHA] complex as a potential target for antivirals.
MATERIALS AND METHODS
Plasmid constructs
All plasmid constructs used for in vitro transcription of mono- or bicistronic HCV replicon RNAs derived from the Con1 isolate of Lohmann et al. (1999) and contained adaptive mutations to allow efficient replication in Huh-7 cells (described in detail by Grassmann et al. [2005]). The plasmid pHCV-bi (Grassmann et al. 2005) was used to generate the following replicon NTR mutants by Qickchange (Stratagene): 3′NTR, pHCV-polyU (Con1; Δ9425–9499); pHCV-SLstop (Δ9375–9417); pHCV-SLIII (Δ9506–9529); pHCV-polyU-SLIII (Δ9425–9526); and 5′NTR, pHCVΔ131–317. For transcription by T7 polymerase, the plasmids were linearized with ScaI. Corresponding cDNA constructs that allowed the transcription of mutant monocistronic replicon molecules were generated in a similar way using the plasmid construct pHCV-mono (Grassmann et al. 2005). To generate transcripts encoding the HCV 3′NTR for UV cross-linking, the 3′NTR coding regions of pHCV-bi, pHCV-poly-U, or pHCV-SLstop were amplified by PCR and cloned KpnI–SpeI into pBluescript. T3 transcription of the wild-type and mutant 3′NTRs was performed after linearization of the resulting plasmids with ScaI. Further truncated transcripts were obtained by linearization with NheI (Con1 position 9536; -SLII-SLI) or DdeI (9519; -SLIII-SLII-SLI). To generate transcripts encoding the HCV 5′NTR for UV cross-linking, pHCV-bi or mutant pHCV-bi plasmids Δ5–20, Δ29–43, Δ45–317, Δ44–118, and Δ131–317 (Grassmann et al. 2005) were linearized with AscI, ApaLI, or NheI and transcribed with T7 RNA polymerase. The HCV 5′NTR transcripts included also 36 nucleotides of the ORF. To generate plasmids that encode non-NTR portions of the HCV genome, we inserted Con1 residues 3420–3641, 4724–5030, or 7891–8162 into the SmaI site of pBluescript. The resulting plasmids were linearized with XbaI and transcribed with T3 polymerase.
The plasmid 5′NTR-sORF-3′NTR contains the HCV 5′NTR (nt 1–341), the first 24 nucleotides of the HCV ORF, followed by the sequence of a FLAG-tag and the HCV 3′NTR (cloning details will be provided on request). The TobA-5′NTR-sORF-3′NTR plasmid was generated by cloning the J6f1 tobramycin aptamer (TobA) (Hartmuth et al. 2002; Jiang and Patel 2004) upstream of the HCV 5′NTR. For T7 in vitro transcription, the plasmids were linearized with NotI.
The plasmids to generate the BVDV transcripts and the probes for the RNase protection assay (RPA) were described previously (Isken et al. 2003; Grassmann et al. 2005). The plasmid to generate the non-HCV transcript used in all shown experiments was described by Isken et al. (2003). Alternatively, we applied the mRNA of the homeodomain cofactor DCOH (data not shown; Mendel et al. 1991).
To generate the RNA molecules for the EM studies, the ApaI fragment encoding most of the HCV ORF (Con1 positions 3627–8392) was removed from pHCV-mono to result pHRT. To produce (+)-strand RNA, pHRT was linearized with ScaI and transcribed with T7 RNA polymerase. To generate the complementary (−)-strand RNA, the shortened HCV ORF/NTR fragment was cloned HindIII–KpnI from pHRT into pBluescript. The resulting plasmid was cleaved with AscI and transcribed by T3 RNA polymerase. The pcDNA construct that expressed the HCV NS5A protein was generated by cloning the cDNA of the Con1 NS5A gene (generated by PCR) with an artificial translation initiation codon BamHI/XbaI into pcDNA3 (Invitrogen).
RNA transfection
Transfection of HCV RNAs into Huh-7 cells was performed by electroporation as previously described (Grassmann et al. 2005). Transfection efficiencies were estimated by electroporation of 200 pmol of Cy3-labeled siRNA per 3 × 106 cells (Fig. 9).
Cells and cytoplasmic extracts
Huh-7, HeLa, and MDBK cells were grown in Dulbecco's modified Eagle's medium supplemented with 5% fetal calf serum. Cytoplasmic extracts were prepared as described by Behrens et al. (1998) and Grassmann et al. (2005). The final ion concentration of these extracts was 50 mM Tris/Cl, pH 8, 100 mM NaCl, 5 mM MgCl2, and 0.5% NP40. To generate cytoplasmic extract that was free of RNA (Fig. 6), the NP40 concentration was increased to 1% and the extract digested for 1 h at 37°C with RNase A (20 μg/extract of 106 cells). To control the digestion of RNA, the nucleic acids were extracted by phenol/chloroform and, after ethanol precipitation, 5′ and 3′ [32P] end-labeled (standard procedures) and analyzed by electrophoresis on a denaturing (7 M urea) 20% Tris/borate acrylamide gel.
Antibodies
The rabbit antisera against the HCV NS proteins were from Dr. De Francesco (IRBM, Rome), the anti-BVDV NS3 mAb was from Dr. Dubovi (Cornell University), and the anti-HCV NS3 and NS5A mAbs were from Dr. Moradpour (University of Lausanne). The rabbit antisera against the NF/NFAR proteins were raised against peptides of these proteins (Affinity Bioreagents). One antiserum against NF45 was shown to precipitate NF45, the heterodimer consisting of NF45 and NF90 and the [NF90/NFAR-1, NF45, RHA] complex, respectively (Figs. 4, 5; O. Isken, unpubl. data). The precipitating anti NF90/NFAR-1 mAb (Figs. 4, 5) was purchased from Pharmingen.
UV cross-linking
The assay was described previously (Isken et al. 2003). Briefly, 15 μL of Huh-7 cytoplasmic extract (20 μg of protein/assay) were incubated with gel-purified UTP [32P]-labeled RNA probe (500,000 cpm/∼10 ng) in the presence of 20 μg of tRNA, followed by exposure to UV light and treatment with 7 μg of RNaseA and 20 U of RNase T1. RNA-linked proteins were detected by SDS-PAGE and autoradiography.
Determining HCV RNA replication
Replication of transfected RNAs was determined either on the level of the viral RNA by probing total cytoplasmic RNA at the indicated time points with an established RNase protection (RPA) protocol detecting progeny (+)-strand RNA (Grassmann et al. 2005) or by selecting G418-resistant cells at 400 μg of G418/mL (Lohmann et al. 1999). G418-resistant colonies were fixed with methanol and stained with Coomassie.
RNAi
RNAi to reduce the amount of cytoplasmic RHA was performed as described (Isken et al. 2003). Briefly, 3 × 106 HCV replicon containing Huh-7 cells were transfected twice within 48 h with 600 pmol of anti-RHA or control siRNAs. Cells were harvested at the indicated time points and analyzed for the level of RHA and (+)-strand viral RNA by Western blot and RPA.
5′–3′ Coprecipitation assay
Two micrograms of biotinylated RNA, prepared by in vitro transcription in the presence of 0.16 mM biotinylated UTP (Roche), were bound to 15 μL streptavidin-coated Sepharose beads. The [32P]-labeled RNA probe (∼10 ng; concentration optimized) was incubated with 100 ng (concentration optimized) of the purified [NF90/NFAR-1, NF45, RHA] complex, BSA, purified HCV NS3 (Grassmann et al. 1999), or purified hsp90 (Stressgene) in the presence of 10 μg of tRNA. The [NF90/NFAR-1, NF45, RHA] complex was purified following the protocol of Liao et al. (1998). The RNA–protein complexes were then incubated for 2 h with the RNA-coated streptavidin beads. The beads were collected and washed, and coprecipitated labeled RNA was analyzed on a denaturing 5% Tris/borate acrylamide gel.
Tobramycin affinity purification of RNP complexes
The RNAs were synthesized using a T7 RNA polymerase kit (Megascript, Ambion) according to the manufacturer's recommendations. The tobramycin affinity purification technique (Hartmuth et al. 2002) was applied with modifications. Binding buffer (BP) was prepared as a 4× concentrated stock solution (80 mM Tris-HCl, pH 8.1, 4 mM CaCl2, 4 mM MgCl2, 0.8 mM DTT). N-Hydroxysuccinimide-activated (NHA)-Sepharose 4 Fast Flow was derivatized with 5 mM tobramycin; 20-μL aliquots of that tobramycin matrix were blocked overnight with 250 μL of blocking buffer (BP, supplemented with 300 mM KCl, 0.1 mg/mL Escherichia coli tRNA, 0.5 mg/mL BSA, 0.01% NP-40) by rotation at 4°C; 28 pmol TobA-5′NTR-sORF-3′NTR or 5′NTR-sORF-3′NTR RNA was incubated with the tobramycin matrix in 400 μL of BP for 1.5 h by rotation at 4°C. Afterward, the matrix was washed three times in BP, supplemented with 145 mM KCl, 5 mM MgCl2. 400 μL of 40% cytoplasmic extract containing 3.2 mg of total protein buffered with 160 mM HEPES, pH 7.6, were loaded onto the matrix and incubated for 30 min by rotation at 30°C. The matrix was washed three times in BP, supplemented with 75 mM KCl and 5 mM MgCl2. Bound complexes were eluted with 250 μL of BP, supplemented with 5 mM tobramycin, 145 mM KCl, and 2 mM MgCl2 for 30 min by rotation at RT.
Proteins were recovered by methanol–chloroform precipitation (Jiang et al. 2004) and analyzed by 5%–15% SDS-PAGE, and silver staining (Shevchenko et al. 1996).
Mass spectrometry
Protein bands were cut out of the gel, washed, and digested with trypsin (Promega) in 30 μL of 10 mM ammonium bicarbonate, pH 8.0, overnight at 37°C (Shevchenko et al. 1996). Tryptic peptides were extracted and liquid chromatography–tandem mass spectrometry (LC/MS/MS) analyses were performed using a Micromass CapLC column 75 μm × 15 cm, 3 μm dC18 Atlantis NanoEase (Waters) coupled to a quadrupole time-of-flight mass spectrometer (Q-TOF2, Micromass). Peptide fragment ion spectra were searched against the nonredundant protein database for exact matches using the MASCOT program (http://www.matrixscience.com).
Immunoprecipitation
Twenty microliters of rabbit anti-serum or 1.5 μg of mAb were bound to 30 μL of protein G-Sepharose (PGS) under standard conditions using buffer A (20 mM Tris/Cl, pH 8, 150 mM KCl, 1% NP40, 1 mM PMSF, 1 mM EDTA, pH 8, 1 mM DTT). Depending on the type of experiment, the antibody-Sepharose suspension was incubated for 3 h at 4°C with cytoplasmic extract (3 × 106 cells/assay) of naive or HCV-replicon containing Huh-7 cells or with extract of naive cells that had been preincubated (1 h, 4°C) with [32P]-labeled RNA transcript (∼10 ng). Following the incubation with extract, the PGS was washed with buffer A containing 150, 300, and 150 mM KCl, respectively, and analyzed for bound cellular or viral protein or for bound viral RNA. Precipitated proteins were detected by Western blots with antisera against the NFAR or HCV NS proteins; coprecipitated RNAs were detected by gel electrophoresis and autoradiography or by RT-PCR. In the latter case, the beads were treated with proteinase K and the extracted RNA precipitated and resuspended in RNase-free water. cDNA synthesis was carried out with the SuperScriptII kit (Invitrogen) for 1 h at 45°C using a DNA oligomer that encompassed Con1 positions 8146–8166. One-fifth of the cDNA reaction was used for PCR with primers Con1 7896–7916 and 8146–8166 in the presence of 0.5 μL of [32P]-dATP; the PCR products were analyzed on a denaturing 5% Tris-borate acrylamide gel.
Purification of the [NF90/NFAR-1, NF45, RHA] complex
The procedure was carried out essentially as described by Liao et al. (1998). Briefly, the S10 fraction of HeLa cells was loaded onto a poly(I)·poly(C)-agarose column equilibrated with buffer B (20 mM HEPES/NaOH, pH 7.6, 3 mM MgCl2, 200 mM KCl). The column was washed with buffer B containing 0.01% Triton X-100 and the proteins eluted with a linear gradient of 0.25–2 M KCl in the same buffer.
Electron microscopy (EM)
Partially double-stranded RNA molecules were prepared by mixing equal amounts of (+)- and (−)-strand RNA in buffer A (10 mM HEPES, pH 7.4, 100 mM KCl, 0.2 mM EDTA). Protein–RNA complexes were formed incubating 100 ng of protein (BSA or [NFAR]) with 100 ng of RNA in buffer A for 20 min at 30°C and stabilized by cross-linking with 1% glutaraldehyde at 4°C for 12 h. To prepare specimens, the samples were diluted 1:250 in solution B (5.4 M urea, 3 mM triethanolamine-HCl, pH 7.9) and mixed 1:3 with benzyldimethylalkylammonium chloride. From 100 μL droplets, an RNA film was picked up on carbon-coated grids; the grids were washed with water, dehydrated in ethanol, and dried. After they were stained with uranyl acetate (2% in 90% ethanol), the grids were destained and dried prior to rotary shadowing with carbon-platinum (150–200 Hz of Pt/C at 7°C). Micrographs were taken at 68,000× magnification on a FEI Technai 12 TEM and an AMT 2k × 2k wide-angle CCD camera.
Immunofluorescence
For confocal microscopy (Fig. 7), the cells were grown on coverslips for 24 h. The cells were washed with PBS, fixed with ice-cold ethanol, blocked, and permeabilized in PBS containing 5% FBS and 0.05% Tween20 and incubated with the indicated antibodies. Primary antibodies were detected with RedoxRed-conjugated anti-mouse antibody (Jackson Laboratory) and Alexa-488 conjugated anti-rabbit antibody (Molecular Probes), respectively. Photomicrographs were taken with a Bio-Rad radiant 2000 LSM with a Plan APO 60× water-immersion objective (Nikon). To exclude bleaching effects, the fields were analyzed independently by digital confocal fluorescence microscopy using LaserSharp 2000 software (Zeiss). In the competition experiments (Fig. 9), the replicon-transfected cells were fixed with 4% paraformaldehyde and permeabilized with 1% N-octyl glucoside. After exposure to the indicated antibodies, the cells were counterstained with appropriate secondary antibodies; the nuclei were stained with DAPI (4′,6′-diamidino-2-phenylindole). The cells were counted and examined for RNA replication with an inverted Nikon TE2000 microscope using a 40× Plan Fluor objective (Nikon) and MetaVue software (Molecular Devices).
Competition assay
Naive or HCV replicon-containing Huh-7 cells (106 cells/assay) were transfected with 2 μg of wild-type or mutant (RdRp-defective) BVDV replicon RNA (Behrens et al. 1998); 20 h post-transfection, RNA replication was determined by immunofluorescent staining of the HCV NS5A and BVDV NS3 protein, respectively.
ACKNOWLEDGMENTS
This work was supported by the NIH (RO1DK062847), the BMBF Kompetenznetz Hepatitis (Projekt 13.3), the Exzellenzinitiative of the State of Saxony-Anhalt, and the DFG (BE 1885/6-1). The EM, cell culture, and imaging facilities at FCCC and the MS facility at the Institute of Biochemistry and Biotechnology in Halle (Dr. Angelika Schierhorn) are gratefully acknowledged. Thanks to the Viral Pathogenesis group at FCCC for helpful discussions and to Wolfram Gerlich (University of Giessen) for support.
Footnotes
Article published online ahead of print. Article and publication date are at http://www.rnajournal.org/cgi/doi/10.1261/rna.594207.
REFERENCES
- Alvarez, D.E., Lodeiro, M.F., Luduena, S.J., Pietrasanta, L.I., Gamarnik, A.V. Long-range RNA–RNA interactions circularize the dengue virus genome. J. Virol. 2005;79:6631–6643. doi: 10.1128/JVI.79.11.6631-6643.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Appel, N., Herian, U., Bartenschlager, R. Efficient rescue of hepatitis C virus RNA replication by trans-complementation with nonstructural protein 5A. J. Virol. 2005;79:896–909. doi: 10.1128/JVI.79.2.896-909.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barton, D.J., Morasco, B.J., Flanegan, J.B. Translating ribosomes inhibit poliovirus negative-strand RNA synthesis. J. Virol. 1999;73:10104–10112. doi: 10.1128/jvi.73.12.10104-10112.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Becher, P., Orlich, M., Thile, H.J. Mutations in the 5′ nontranslated region of bovine viral diarrhea virus result in altered growth characteristics. J. Virol. 2000;74:7884–7894. doi: 10.1128/jvi.74.17.7884-7894.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Behrens, S.E., Isken, O. Cis and trans-acting factors in Flaviviridae replication. In: Kalitzky M., Borowski P., editors. Molecular Biology of the Flavivirus. Horizon Bioscience; Norwich, UK: 2006. pp. 101–134. [Google Scholar]
- Behrens, S.E., Grassmann, C.W., Thiel, H.J., Meyers, G., Tautz, N. Characterization of an autonomous subgenomic pestivirus RNA replicon. J. Virol. 1998;72:2364–2372. doi: 10.1128/jvi.72.3.2364-2372.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elbashir, S.M., Harborth, J., Lendeckel, W., Yalcin, A., Weber, K., Tuschl, T. Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature. 2001;411:494–498. doi: 10.1038/35078107. [DOI] [PubMed] [Google Scholar]
- Evans, M.J., Rice, C.M., Goff, S.P. Phosphorylation of hepatitis C virus nonstructural protein 5A modulates its protein interactions and viral RNA replication. Proc. Natl. Acad. Sci. 2004;101:13038–13043. doi: 10.1073/pnas.0405152101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Filomatori, C.V., Lodeiro, M.F., Alvarez, D.E., Samsa, M.M., Pietrasanta, L., Gamarnik, A.V. A 5′ RNA element promotes dengue virus RNA synthesis on a circular genome. Genes & Dev. 2006;20:2238–2249. doi: 10.1101/gad.1444206. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Friebe, P., Bartenschlager, R. Genetic analysis of sequences in the 3′ nontranslated region of hepatitis C virus that are important for RNA replication. J. Virol. 2002;76:5326–5338. doi: 10.1128/JVI.76.11.5326-5338.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Friebe, P., Lohmann, V., Krieger, N., Bartenschlager, R. Sequences in the 5′ nontranslated region of hepatitis C virus required for RNA replication. J. Virol. 2001;75:12047–12057. doi: 10.1128/JVI.75.24.12047-12057.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frolov, I., McBride, M.S., Rice, C.M. cis-acting RNA elements required for replication of bovine viral diarrhea virus–hepatitis C virus 5′ nontranslated region chimeras. RNA. 1998;4:1418–1435. doi: 10.1017/s1355838298981031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fuchsova, B., Novak, P., Kafkova, J., Hozak, P. Nuclear DNA helicase II is recruited to IFN-α–activated transcription sites at PML nuclear bodies. J. Cell Biol. 2002;158:463–473. doi: 10.1083/jcb.200202035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gamarnik, A.V., Andino, R. Switch from translation to RNA replication in a positive-stranded RNA virus. Genes & Dev. 1998;12:2293–2304. doi: 10.1101/gad.12.15.2293. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gao, L., Aizaki, H., He, J.W., Lai, M.M.C. Interactions between viral nonstructural proteins and host protein hVAP-33 mediate the formation of hepatitis C virus RNA replication complex on lipid raft. J. Virol. 2004;78:3480–3488. doi: 10.1128/JVI.78.7.3480-3488.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gontarek, R.R., Gutshall, L.L., Herold, K.M., Tsai, J., Sathe, G.M., Mao, J., Prescott, C., Del Vecchio, A.M. hnRNP C and polypyrimidine tract-binding protein specifically interact with the pyrimidine-rich region within the 3′NTR of the HCV RNA genome. Nucl. Acids Res. 1999;27:1457–1463. doi: 10.1093/nar/27.6.1457. http://nar.oxfordjournals.org/cgi/content/full/27/6/1457. [DOI] [PMC free article] [PubMed]
- Gosert, R., Egger, D., Lohmann, V., Bartenschlager, R., Blum, H.E., Bienz, K., Moradpour, D. Identification of the hepatitis C virus RNA replication complex in Huh-7 cells harboring subgenomic replicons. J. Virol. 2003;77:5487–5492. doi: 10.1128/JVI.77.9.5487-5492.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grassmann, C.W., Isken, O., Behrens, S.E. Assignment of the multifunctional NS3 protein of bovine viral diarrhea virus during RNA replication: An in vivo and in vitro study. J. Virol. 1999;73:9196–9205. doi: 10.1128/jvi.73.11.9196-9205.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grassmann, C.W., Isken, O., Tautz, N., Behrens, S.E. Genetic analysis of the pestivirus nonstructural coding region: Defects in the NS5A unit can be complemented in trans. J. Virol. 2001;75:7791–7802. doi: 10.1128/JVI.75.17.7791-7802.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grassmann, C.W., Yu, H., Isken, O., Behrens, S.E. Hepatitis C virus and the related bovine viral diarrhea virus considerably differ in the functional organization of the 5′ nontranslated region: Implications for the viral life cycle. Virology. 2005;333:349–366. doi: 10.1016/j.virol.2005.01.007. [DOI] [PubMed] [Google Scholar]
- Hartmuth, K., Urlaub, H., Vornlocher, H.P., Will, C.L., Gentzel, M., Wilm, M., Lührmann, R. Protein composition of human prespliceosomes isolated by a tobramycin affinity selection method. Proc. Natl. Acad. Sci. 2002;99:16719–16724. doi: 10.1073/pnas.262483899. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Herold, J., Andino, R. Poliovirus RNA replication requires genome circularization through a protein–protein bridge. Mol. Cell. 2001;7:581–591. doi: 10.1016/S1097-2765(01)00205-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Honda, M., Brown, E.A., Lemon, S.M. Stability of a stem–loop involving the initiator AUG controls the efficiency of internal initiation of translation on hepatitis C virus RNA. RNA. 1996a;2:955–968. [PMC free article] [PubMed] [Google Scholar]
- Honda, M., Ping, L.H., Rijnbrand, R.C., Amphlett, E., Clarke, B., Rowlands, D., Lemon, S.M. Structural requirements for initiation of translation by internal ribosome entry within genome-length hepatitis C virus RNA. Virology. 1996b;222:31–42. doi: 10.1006/viro.1996.0395. [DOI] [PubMed] [Google Scholar]
- Isken, O., Grassmann, C.W., Sarisky, R.T., Kann, M., Zhang, S., Grosse, F., Kao, P.N., Behrens, S.E. Members of the NF90/NFAR protein group are involved in the life cycle of a positive-strand RNA virus. EMBO J. 2003;22:5655–5665. doi: 10.1093/emboj/cdg562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Isken, O., Grassmann, C.W., Yu, H., Behrens, S.E. Complex signals in the genomic 3′ nontranslated region of bovine viral diarrhea virus coordinate translation and replication of the viral RNA. RNA. 2004;10:1637–1652. doi: 10.1261/rna.7290904. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ito, T., Lai, M.M.C. An internal polypyrimidine-tract-binding protein-binding site in the hepatitis C virus RNA attenuates translation, which is relieved by the 3′-untranslated sequence. Virology. 1999;254:288–296. doi: 10.1006/viro.1998.9541. [DOI] [PubMed] [Google Scholar]
- Jiang, L., Patel, D.J. Solution structure of the tobramycin–RNA aptamer complex. Nat. Struct. Biol. 2004;5:769–774. doi: 10.1038/1804. [DOI] [PubMed] [Google Scholar]
- Jiang, L., He, L., Fountoulakis, M. Comparison of protein precipitation methods for sample preparation prior to proteomic analysis. J. Chromatogr. A. 2004;1023:317–320. doi: 10.1016/j.chroma.2003.10.029. [DOI] [PubMed] [Google Scholar]
- Kalliampakou, K.I., Kalamvoki, M., Mavromara, P. Hepatitis C virus (HCV) NS5A protein downregulates HCV IRES-dependent translation. J. Gen. Virol. 2005;86:1015–1025. doi: 10.1099/vir.0.80728-0. [DOI] [PubMed] [Google Scholar]
- Kao, P.N., Chen, L., Brock, G., Ng, J., Kenny, J., Smith, A.J., Corthesy, B. Cloning and expression of cyclosporin A- and FK506-sensitive nuclear factor of activated T-cells: NF45 and NF90. J. Biol. Chem. 1994;269:20691–20699. [PubMed] [Google Scholar]
- Kieft, J.S., Zhou, K., Jubin, R., Doudna, J.A. Mechanism of ribosome recruitment by hepatitis C IRES RNA. RNA. 2001;7:194–206. doi: 10.1017/s1355838201001790. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khromykh, A.A., Westaway, E.G. Subgenomic replicons of the flavivirus Kunjin: construction and applications. J. Virol. 1997;71:1497–1505. doi: 10.1128/jvi.71.2.1497-1505.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khromykh, A.A., Meka, H., Guyatt, K.J., Westaway, E.G. Essential role of cyclization sequences in flavivirus RNA replication. J. Virol. 2001;75:6719–6728. doi: 10.1128/JVI.75.14.6719-6728.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krasnoselskaya-Riz, I., Spruill, A., Chen, Y.W., Schuster, D., Teslovich, T., Baker, C., Kumar, A., Stephan, D.A. Nuclear factor 90 mediates activation of the cellular antiviral expression cascade. AIDS Res. Hum. Retroviruses. 2002;18:591–604. doi: 10.1089/088922202753747941. [DOI] [PubMed] [Google Scholar]
- Lee, H., Shin, H., Wimmer, E., Paul, A.V. cis-acting RNA signals in the NS5B C-terminal coding sequence of the hepatitis C virus genome. J. Virol. 2004;78:10865–10877. doi: 10.1128/JVI.78.20.10865-10877.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liao, H.J., Kobayashi, R., Mathews, M.B. Activities of adenovirus virus-associated RNAs: Purification and characterization of RNA binding proteins. Proc. Natl. Acad. Sci. 1998;95:8514–8519. doi: 10.1073/pnas.95.15.8514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lindenbach, B.D., Thiel, H.J., Rice, C.M. Flaviviridae: The viruses and their replication. In: Fields B.N., editor. Virology. 5th ed. Lippincott-Raven Press; Philadelphia: 2007. pp. 1101–1151. [Google Scholar]
- Lohmann, V., Korner, F., Koch, J., Herian, U., Theilmann, L., Bartenschlager, R. Replication of subgenomic hepatitis C virus RNAs in a hepatoma cell line. Science. 1999;285:110–113. doi: 10.1126/science.285.5424.110. [DOI] [PubMed] [Google Scholar]
- Lu, H., Li, W., Noble, W.S., Payan, D., Anderson, D.C. Riboproteomics of the hepatitis C virus internal ribosome entry site. J. Proteome Res. 2004;3:949–957. doi: 10.1021/pr0499592. [DOI] [PubMed] [Google Scholar]
- Macdonald, A., Harris, M. Hepatitis C virus NS5A: Tales of a promiscuous protein. J. Gen. Virol. 2004;85:2485–2502. doi: 10.1099/vir.0.80204-0. [DOI] [PubMed] [Google Scholar]
- Mendel, D.B., Khavari, P.A., Conley, P.B., Graves, M.K., Hansen, L.P., Admon, A., Crabtree, G.R. Characterization of a cofactor that regulates dimerization of a mammalian homeodomain protein. Science. 1991;254:1762–1767. doi: 10.1126/science.1763325. [DOI] [PubMed] [Google Scholar]
- Merrill, M.K., Gromeier, M. The double-stranded RNA binding protein 76:NF45 heterodimer inhibits translation initiation at the rhinovirus type 2 internal ribosome entry site. J. Virol. 2006;80:6936–6942. doi: 10.1128/JVI.00243-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miyanari, Y., Hijikata, M., Yamaji, M., Hosaka, M., Takahashi, H., Shimotohno, K. Hepatitis C virus nonstructural proteins in the probable membranous compartment function in viral genome replication. J. Biol. Chem. 2003;278:50301–50308. doi: 10.1074/jbc.M305684200. [DOI] [PubMed] [Google Scholar]
- Pankraz, A., Thiel, H.-J., Becher, P. Essential and nonessential elements in the 3′ nontranslated region of bovine viral diarrhea virus. J. Virol. 2005;79:9119–9127. doi: 10.1128/JVI.79.14.9119-9127.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parker, L.M., Fierro-Monti, I., Mathews, M.B. Nuclear factor 90 is a substrate and regulator of the eukaryotic initiation factor 2 kinase double-stranded RNA-activated protein kinase. J. Biol. Chem. 2001;276:32522–32530. doi: 10.1074/jbc.M104408200. [DOI] [PubMed] [Google Scholar]
- Pawlotsky, J.M. Therapy of hepatitis C: From empiricism to eradication. Hepatology. 2006;43:S207–S220. doi: 10.1002/hep.21064. [DOI] [PubMed] [Google Scholar]
- Reichman, T.W., Mathews, M.B. RNA binding and intramolecular interactions modulate the regulation of gene expression by nuclear factor 110. RNA. 2003;9:543–554. doi: 10.1261/rna.2181103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reichman, T.W., Muñiz, L.C., Mathews, M.B. The RNA binding protein nuclear factor 90 functions as both a positive and negative regulator of gene expression in mammalian cells. Mol. Cell. Biol. 2002;22:343–356. doi: 10.1128/MCB.22.1.343-356.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saunders, L.R., Barber, G.N. The dsRNA binding protein family: Critical roles, diverse cellular functions. FASEB J. 2003;17:961–983. doi: 10.1096/fj.02-0958rev. [DOI] [PubMed] [Google Scholar]
- Saunders, L.R., Perkins, D.J., Balachandran, S., Michaels, R., Ford, R., Mayeda, A., Barber, G.N. Characterization of two evolutionarily conserved, alternatively spliced nuclear phosphoproteins, NFAR-1 and -2, that function in mRNA processing and interact with the double-stranded RNA-dependent protein kinase, PKR. J. Biol. Chem. 2001;276:32300–32312. doi: 10.1074/jbc.M104207200. [DOI] [PubMed] [Google Scholar]
- Shevchenko, A., Wilm, M., Vorm, O., Mann, M. Mass spectrometric sequencing of proteins from silver-stained polyacrylamide gels. Anal. Chem. 1996;68:850–858. doi: 10.1021/ac950914h. [DOI] [PubMed] [Google Scholar]
- Shi, S.T., Lee, K.J., Aizaki, H., Hwang, S.B., Lai, M.M.C. Hepatitis C virus RNA replication occurs on a detergent-resistant membrane that cofractionates with caveolin-2. J. Virol. 2003;77:4160–4168. doi: 10.1128/JVI.77.7.4160-4168.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sizova, D.V., Kolupaeva, V.G., Pestova, T.V., Shatsky, I.N., Hellen, C.U. Specific interaction of eukaryotic translation initiation factor 3 with the 5′ nontranslated regions of hepatitis C virus and classical swine fever virus RNAs. J. Virol. 1998;72:4775–4782. doi: 10.1128/jvi.72.6.4775-4782.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wells, S.E., Hillner, P.E., Vale, R.D., Sachs, A.B. Circularization of mRNA by eukaryotic translation initiation factors. Mol. Cell. 1998;2:135–140. doi: 10.1016/s1097-2765(00)80122-7. [DOI] [PubMed] [Google Scholar]
- Yanagi, M., St. Claire, M., Emerson, S.U., Purcell, R.H., Bukh, J. In vivo analysis of the 3′ untranslated region of the hepatitis C virus after in vitro mutagenesis of an infectious cDNA clone. Proc. Natl. Acad. Sci. 1999;96:2291–2295. doi: 10.1073/pnas.96.5.2291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yi, M., Lemon, S.M. 3′ Nontranslated RNA signals required for replication of hepatitis C virus RNA. J. Virol. 2003;77:3557–3568. doi: 10.1128/JVI.77.6.3557-3568.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu, H., Grassmann, C.W., Behrens, S.E. Sequence and structural elements at the 3′ terminus of bovine viral diarrhea virus genomic RNA: Functional role during RNA replication. J. Virol. 1999;73:3638–3648. doi: 10.1128/jvi.73.5.3638-3648.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu, H., Isken, O., Grassmann, C.W., Behrens, S.E. A stem–loop motif formed by the immediate 5′ terminus of the bovine viral diarrhea virus genome modulates translation as well as replication of the viral RNA. J. Virol. 2000;74:5825–5835. doi: 10.1128/jvi.74.13.5825-5835.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang, S., Grosse, F. Multiple functions of nuclear DNA helicase II (RNA helicase A) in nucleic acid metabolism. Acta Biochim. Biophys. Sin. (Shanghai) 2004;36:177–183. doi: 10.1093/abbs/36.3.177. [DOI] [PubMed] [Google Scholar]