SUMMARY
Conventional kinesin and class V and VI myosins coordinate the mechanochemical cycles of their motor domains for processive movement of cargo along microtubules or actin filaments. It is widely accepted that this coordination is achieved by allosteric communication or mechanical strain between the motor domains, which controls the nucleotide state and interaction with microtubules or actin. However, questions remain about the interplay between the strain and the nucleotide state. We present an analysis of Saccharomyces cerevisiae Kar3/Vik1, a heterodimeric C-terminal Kinesin-14 containing catalytic Kar3 and the nonmotor protein Vik1. The X-ray crystal structure of Vik1 exhibits a similar fold to the kinesin and myosin catalytic head but lacks an ATP-binding site. Vik1 binds more tightly to microtubules than Kar3, facilitates cooperative microtubule decoration by Kar3/Vik1 heterodimers, and yet allows motility. These results demand communication between Vik1 and Kar3 via a novel mechanism to coordinate their interactions with microtubules.
INTRODUCTION
Most members of the kinesin and myosin superfamilies of linear molecular motors consist of two identical motor head domains held together by a section of coiled-coil. There is clear consensus that this arrangement can allow a single molecular assembly to move processively along either a microtubule or actin filament through cooperative action of the two heads. However, questions remain about the dimeric assemblies that do not appear to exhibit processive movement. Specifically, what purpose does having two heads serve in such assemblies? Is it simply a way of making a more rigid linker between the motor and its cargo in which the heads function independently or is there truly communication between them? These questions are particularly relevant for understanding the molecular and biological function of the minus-end directed kinesins such as Drosophila Ncd and budding yeast Kar3. Both of these motors dimerize via a section of coiled-coil and in the case of Ncd it has been suggested that only one of the two heads binds to the microtubule and is involved in force production during the ATPase cycle (Sosa et al., 1997; Hirose et al., 1998; Wendt et al., 2002; Endres et al., 2006). Kar3, on the other hand, is unusual since, although it forms a homodimer in vitro (Chu et al., 2005), the functional forms of the protein in vivo are a heterodimer with either of two alternative non-motor proteins Cik1 or Vik1 (Page et al., 1994; Manning et al., 1999; Barrett et al., 2000; Manning and Snyder, 2000; Chu et al., 2005; Sproul et al., 2005). While it has been demonstrated that Cik1 and Vik1 differentially regulate the interaction of Kar3 with microtubules, the mechanism by which they accomplish this and how these heterodimers generate force remains uncertain.
Kar3 is one of six kinesins in budding yeast (Meluh and Rose, 1990; Hildebrandt and Hoyt, 2000). Like Drosophila Ncd (Endow et al., 1990; McDonald and Goldstein, 1990; McDonald et al., 1990), Kar3 is classified as a Kinesin-14 because its motor domain is at the carboxy terminus, and it generates minus-end-directed force. Kar3 is the only Kinesin-14 in S. cerevisiae, and it has specific roles during karyogamy (the nuclear fusion event during mating) and vegetative growth that are mediated by the non-motor proteins Cik1 or Vik1 (Page et al., 1994; Manning et al., 1999; Barrett et al., 2000; Manning and Snyder, 2000). In response to mating pheromone, Cik1 targets Kar3 to cytoplasmic or astral microtubules where the Kar3/Cik1 heterodimer generates minus-end directed microtubule shortening during karyogamy (Maddox et al., 2003; Chu et al., 2005; Sproul et al., 2005; Molk et al., 2006). Vik1, on the other hand, is not expressed and has no role in karyogamy (Page and Snyder, 1992; Page et al., 1994; Manning et al., 1999; Manning and Snyder, 2000).
In contrast to karyogamy, the function of Kar3 during vegetative growth is not well understood, in part because both Kar3/Cik1 and Kar3/Vik1 heterodimers have distinct and separate roles during mitosis (Manning et al., 1999). What is currently known is that Vik1, but not Cik1, localizes Kar3 at the mitotic spindle poles, while Cik1, in the absence of Vik1, promotes accumulation of Kar3 along the length of the spindle (Manning et al., 1999; Manning and Snyder, 2000). In the absence of both Vik1 and Cik1, Kar3 appears diffusely throughout the nucleus, suggesting that Vik1 and Cik1 are directly involved in microtubule interactions (Manning et al., 1999).
This study focuses on the structure and function of Vik1 and how it modulates the activities of Kar3. The X-ray crystal structure of the C-terminal globular domain of Vik1 shows that this region is structurally similar to the catalytic motor core of kinesins and myosins but is devoid of a nucleotide-binding site. The structural results also provide evidence that the configuration of the Kar3 and Vik1 motor domains relative to the coiled-coil in their heterodimeric form is similar to that of the Ncd homodimer. In the context of the heterodimer, Vik1 modulates Kar3 behavior by direct interaction with the microtubule and, together, Kar3/Vik1 exhibit the characteristics of a Kinesin-14 motor. In contrast to Kar3/Cik1, Kar3/Vik1 binds the microtubule lattice cooperatively and promotes microtubule stabilization. Based on these structural and functional data, we propose that both Vik1 and Cik1 may have evolved from ancient forms of kinesin where the microtubule-binding and coiled-coil forming capabilities were retained while the nucleotide-binding ability was lost. We also propose that the role of Kar3/Vik1 at the spindle pole bodies is to focus and crosslink microtubules for bipolar spindle assembly and stabilization.
RESULTS
Vik1 Structure
Sequence analysis predicts that Kar3, Cik1, and Vik1 consist of an N-terminal globular domain, a central coiled-coil forming region, and a C-terminal globular domain (Figure 1). For Kar3, the N-terminal globular domain is believed to function in cargo binding, while the coiled-coil region forms the primary site of its interaction with Cik1 and Vik1. The exact function of the N- and C-terminal globular domains in Cik1 and Vik1 are unknown. Through sequence analysis and partial proteolytic digestion, the boundaries for the coiled-coil and C-terminal domain of Cik1 and Vik1 were defined. With this information, the C-terminal globular domain of Vik1 was expressed, purified (Figure S1), and crystallized, allowing a structure determination to a resolution of 1.6 Å (Figure 2A and see Table S1 for data collection and refinement statistics), as well as an evaluation of its functional properties in relation to Kar3 (described later).
Figure 1. Bar Diagrams of the Predicted Structural Domains of Full-length Kar3, Cik1, and Vik1.

The coiled-coil regions (CC) were predicted by PAIRCOIL (Berger et al., 1995). The regions of each protein that were used to make the truncated versions of Kar3, Cik1, and Vik1, as well as the Kar3MD and Vik1MHD constructs are indicated next to each bar diagram.
Figure 2. The Structure of Vik1 and Its Comparison to Kar3.
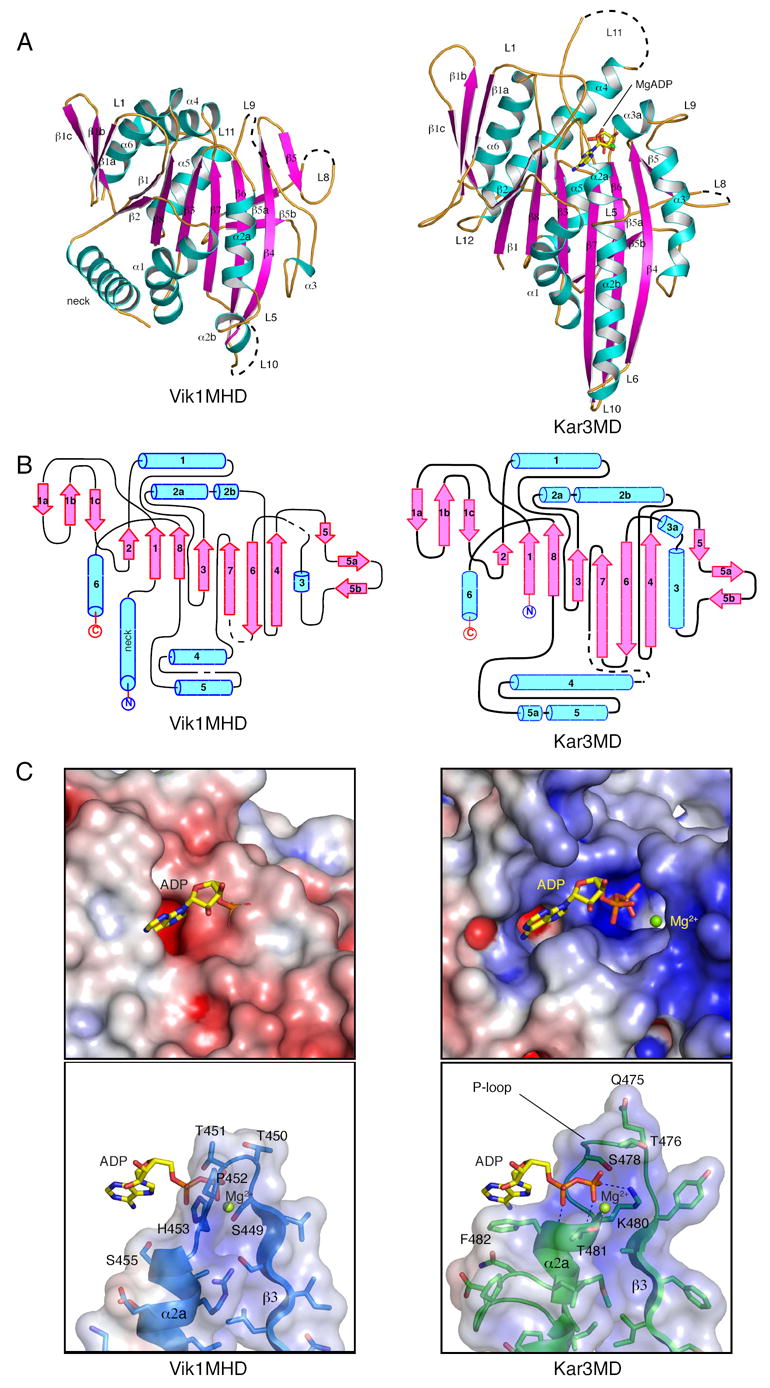
(A) A ribbon representation of the structure of the C-terminal globular domain of Vik1 (left), including a short stretch of the coiled-coil forming region (neck), is shown beside the motor domain of Kar3 (right) (PDB accession: 3KAR) (Gulick et al., 1998). The construct used for crystallization consisted of residues Thr353 to Thr647 from Vik1, however the model has been truncated at Cys640 due to disorder of the C-terminus. Thr353 is preceded by three residues, Gly350, Ala351, and Ser352, respectively, at the N-terminus of the molecule, which are a result of the affinity tag used for protein purification. Poor electron density existed for several loops, and were not built into the final model. These lie between residues Tyr497 and Asp503, Asp537 and Ser545, Ser559 and Pro567, and Lys593 and Ser595. Residues between Ser485 and Ser490 are also located on a loop that is not well-ordered, however, the electron density was of sufficient quality to build this region into the model when an occupancy of 0.5 was applied. (B) Topology diagrams for the Vik1 (left) and Kar3 (right) motor domains were generated with TopDraw based on topology analyses by the TOPS server (Westhead et al., 1999; Bond, 2003). The diagrams are labeled and colored to match the structures in (A). (C) Electrostatic surface representation of the nucleotide-binding pocket of Kar3MD (upper right) and the analogous region of Vik1MHD (upper left) with MgADP superimposed after alignment of the α-carbons of the Vik1MHD (Gly373 to Cys640) onto those of the Kar3MD structure. ADP is shown as yellow sticks and Mg2+ as a green sphere. Mg2+ is obscured by the protein surface in the Vik1MHD figure. The electrostatic surface potential was generated in Pymol using APBS (Baker et al., 2001; DeLano, 2002). A ribbon representation of the P-loop (including relevant side chains) of Kar3 and its interaction with MgADP is shown (lower right). The analogous loop of Vik1 is shown with MgADP superimposed based on the overall alignment of Vik1MHD onto the Kar3MD structure (lower left). All structure figures were generated with Pymol (DeLano, 2002).
The fold of the C-terminal globular domain of Vik1 is remarkably similar to the motor domain of all structurally characterized forms of kinesin, and hence we refer to it as the Vik1 ‘motor homology domain’ (Vik1MHD). The structure of the Kar3 motor domain (Kar3MD) is shown for comparison (Figure 2A, right) (Gulick et al., 1998). The Vik1 structure includes an N-terminal α-helical segment attached to the Vik1MHD, which is also analogous to the ‘neck’ of Ncd (Sablin et al., 1998). Vik1MHD’s structural motif is an α/β fold, where, like kinesin, the central eight-stranded β-sheet is surrounded by six α-helices, three on either side. The strands have been numbered consecutively from the N-terminus of the construct, adopting the system described by Fletterick and co-workers for kinesin heavy chain (KHC) and Ncd (Kull et al., 1996; Sablin et al., 1996). Topology diagrams for the Vik1MHD and Kar3MD structures illustrate the analogous arrangement of the secondary structure elements for these proteins (Figure 2B). The rms deviation for 210 structurally equivalent α-carbons in Vik1MHD and Kar3MD is 2.6 Å (Figure S2), and there is a 71% secondary structure match despite sharing only 11% sequence identity based on a structure-based sequence alignment (Figure S3).
Most of the differences between Vik1 and Kar3 are located in the surface loops, where this type of variability is a characteristic feature of different classes of kinesin (Figure S2). Many of these loops in the kinesins appear flexible and have been proposed to undergo conformational changes during their motile cycle. These loops are often disordered in the crystal structures. Thus, while the overall electron density for the Vik1MHD is well defined, there is some disorder in four of its loops. A comparison of the residue lengths of selected loops for Vik1, Kar3, and Ncd reveals that the largest discrepancies are found in L10, L11, and L12 (Table S2). Importantly, L11 and L12 comprise a significant proportion of the primary microtubule-binding surface in kinesins (Alonso et al., 1998; Woehlke et al., 1997).
The Missing ‘Active Site’
A structural comparison of the MgADP-bound surface cleft of Kar3 with the analogous region of the Vik1MHD reveals that Vik1 does not contain a nucleotide-binding site (Figure 2C and Figure S4). In kinesins, the nucleotide-binding pocket is formed by structural elements that are highly conserved (Sablin et al., 1996). (1) The P-loop (motif GxxxxGKT) is formed by loop L4, which connects strand β3 and helix α2, and wraps around the phosphates of MgADP (Figure 2A and 2C, right). This motif is shared by many other nucleotide-binding proteins (Schulz, 1992). (2) Loop L1 (motif RxRP) interacts with the base. (3) Switch 1 (motif NxxSSR), and (4) Switch 2 (motif DLAGSE) form the remaining nucleotide-binding pocket elements. The structure-based sequence alignment of Kar3 and Vik1 shows that none of these conserved elements exist in Vik1 (Figure S3, boxed sequences). This is not surprising since previous sequence analyses did not find the trademark motifs for nucleotide-binding in Vik1, however, such analyses provided no clues that Vik1 contained a motor domain fold, or that it might bind microtubules.
Vik1 Binds to Microtubules
Vik1MHD binds surprisingly tightly to microtubules relative to Kar3 as demonstrated by equilibrium co-sedimentation (Figure 3A). The Kd,MT for Vik1MHD is 43 nM which indicates tighter binding than that observed for the AMPPNP complex of Kar3MD that has a Kd,MT of 119 nM (Figure 3C). As with all kinesins, AMPPNP serves as an analog of the tightly bound ATP state. Conversely, ADP confers a very weak microtubule-binding state in kinesins. The microtubule binding properties of a truncated version of the Kar3/Vik1 heterodimer in the presence and absence of nucleotides show that there is allosteric communication between the two globular domains since the dissociation constants are not a simple combination of the individual affinities (Figure 3B and 3C). This truncated version of Kar3/Vik1 contains much of the coiled-coil dimerization region and the C-terminal globular domains of Kar3 and Vik1 (Figure 1 and S1). This construct was designed based on sequence analyses and proteolysis experiments of Kar3, Vik1, and Cik1 to identify the minimal length of coiled-coil allowing the C-terminal globular domains of Vik1 and Cik1 to heterodimerize with the motor domain of Kar3. The dimeric state of this complex was confirmed by analytical gel filtration and equilibrium centrifugation (Figure S1 and S5). In the equilibrium co-sedimentation assay, the Kd,MT for the truncated Kar3/Vik1 was 131 nM in the nucleotide-free state (achieved with apyrase), which was similar to its affinity in the presence of AMPPNP (Kd,MT = 138 nM) (Figure 3B and 3C). Surprisingly, the affinity of Kar3/Vik1 for microtubules was tightest in the presence of ADP (Kd,MT = 38 nM), which was similar to the Kd,MT for the Vik1MHD alone (Kd,MT = 43 nM) (Figure 3A and 3C). These results suggest that, in the ADP state, the Kar3/Vik1 complex is tethered to the microtubule by Vik1 as the Kar3 motor would be detached from the microtubule because it contains ADP in its active site. Note too that in the presence of ADP, only 54% of Kar3/Vik1 partitioned with the microtubules, yet in the presence of AMPPNP, this amount approached 100% with 92% Kar3/Vik1 bound to microtubules. These results are intriguing and suggest that the Kar3•ADP /Vik1 configuration on the microtubule prevents additional Kar3/Vik1 motors from binding. However, at this time we do not have structural data of this microtubule-bound configuration to explain the 54% saturation in the presence of ADP. The results also suggest that, in the presence of ATP, Kar3 must be able to modulate the Vik1-microtubule interaction, which raises the question of how Vik1 interacts with microtubules.
Figure 3. Microtubule Binding by Vik1MHD, SeMetVik1MHD, Kar3MD, and Kar3/Vik1.

(A and C) Microtubule•motor cosedimentation was performed to compare the binding of 50 nM Vik1MHD, SeMetVik1MHD, Kar3MD, and (B and C) Kar3/Vik1 at three different nucleotide conditions. The data were plotted as the fraction of motor partitioning to the pellet as a function of microtubule concentration and fit to quadratic equation 2, providing the constants in C. Data are reported as mean +/− SEM. Final concentrations: 0–3 μM tubulin polymer, 40 μM Taxol, and ± 0.1U/ml Apyrase, or 2 mM MgAMPPNP, or 2 mM MgADP. (D and E) Comparison of microtubule-binding surfaces of Vik1MHD and Kar3MD. The shown view is rotated 180° from Figure 2A. Putative microtubule-binding elements of Kar3MD (E) and the analogous regions of Vik1 (D) whose structural properties are significantly different are shown in orange.
The Potential Microtubule-Binding Surface of Vik1
In kinesins, the putative microtubule-binding surface consists of the following structural elements: (1) Loops L7/L8, (2) loop L11 and the N-terminus of helix α4, (3) Loop L12 and the start of helix α5, and (4) helix α6 (see Figure S3 for the sequence of these elements in Kar3) (Sosa et al., 1997; Woehlke et al., 1997; Alonso et al., 1998). While these elements are also found in the Vik1MHD, several of them exhibit major structural differences in comparison to those of kinesin and Ncd. The most conspicuous structural differences are highlighted in Figure 3D and are shown in relation to Kar3MD in Figure 3E. One of the most prominent discrepancies is found in helix α4 of Vik1MHD, which is three α-helical turns shorter than that of Kar3 and is tilted upwards ~60° relative to Kar3. Helix α5 is uninterrupted unlike that of Kar3 and loop L12 contains a short disordered section near its N-terminus and is found roughly 9 Å away from its position in Kar3. Finally, and perhaps most significantly, loop L11 is 13 residues shorter than its counterpart in Kar3 (Table S2). Together, these elements in kinesin (α4, α5, L11, and L12) form a subdomain on the microtubule-binding side of the central β-sheet that undergoes conformational changes in response to the nucleotide state of the motor (Sack et al., 1999; Vale and Milligan, 2000; Kikkawa et al., 2001; Sablin and Fletterick, 2004). Loop L11, in particular, is near the ATP γ-phosphate sensing region, and its interaction with α-tubulin has been suggested to be important during ATP hydrolysis (Song et al., 2001). It has also been implicated in the coordination of conformational changes between the mobile Switch 1/Switch 2 regions (Figure S3) (Song et al., 2001). The surface differences between Kar3 and Vik1 and the absence of an ATP-binding site suggest that Vik1 may bind to the microtubule lattice through a different set of interactions, and possibly in a different orientation, to that of kinesin. However, regardless of the way it binds, the question remains of how Vik1 is released from the microtubule to permit motility.
Kar3/Vik1 is a Kinesin-14 Heterodimeric Motor
The truncated versions of Kar3/Vik1 and Kar3/Cik1 heterodimers displayed minus-end directed movement in ATP-dependent microtubule gliding assays at a speed that is comparable to other mitotic motors (Figure 4A and Supplemental Movies). The steady-state ATPase kinetics show a higher kcat at 3.7 s−1 for Kar3/Vik1 than observed for Kar3/Cik1 at 2.8 s−1 (Figure 4B). However, both the KM,ATP and the K1/2,MT are similar for each motor. In contrast to these kinetic similarities, Kar3/Cik1 and Kar3/Vik1 differ in their ability to depolymerize microtubules. Unlike Kar/Cik1, Kar3/Vik1 did not induce robust microtubule depolymerization in the presence of MgATP (Figure 4C) (Sproul et al., 2005). Furthermore, while Kar3/Cik1 promoted microtubule shortening predominantly from the microtubule plus-end, the same pronounced plus-end specificity was not observed for Kar3/Vik1 (Figure 4C). Rather, the Kar3/Vik1 end-specificity was similar to that observed for Drosophila Ncd-promoted microtubule depolymerization, where ~1/3 of the microtubules exhibited microtubule plus-end shortening with ~2/3 microtubules shortening from both the plus and minus ends (Sproul et al., 2005). Consistent with this behavior, no role for Ncd- or Kar3/Vik1-promoted microtubule depolymerization in vivo has been reported. As the concentration of Kar3/Vik1 or Kar3/Cik1 was increased, the microtubules appeared more stable, however, the microtubule stabilization effect appeared to be more significant for Kar3/Vik1 than Kar3/Cik1 (Figure 4D).
Figure 4. Kar3/Vik1 Is a Kinesin-14 Heterodimeric Motor.

(A) Kar3/Vik1 minus-end-directed motility in the presence of MgATP. Arrowheads denote the bright microtubule minus end, and the asterisks (*), the dim, leading microtubule plus end. Scale bar = 5μm. The table compares the microtubule gliding promoted by Kar3/Vik1, Kar3/Cik1, and squid Kinesin-1. (B) The steady-state ATPase kinetics of Kar3/Vik1 and Kar3/Cik1 as a function of microtubule and MgATP concentrations. Upper panel final concentrations: 0.82 μM Kar3/Vik1 or 1.1 μM Kar3/Cik1, 0–42 μM tubulin polymer, 40 μM Taxol, and 1 mM [α32P] MgATP. Lower panel final concentrations: 0.85 μM Kar3/Vik1 or 1 μM Kar3/Cik1, 20 μM tubulin polymer, 40 μM Taxol, and 0–1 mM [α32P] MgATP. The table shows the steady-state parameters of Kar3/Vik1 and Kar3/Cik1 in comparison to the Kar3MD (Mackey and Gilbert, 2003). (C) ATP-dependent Kar3/Vik1 and Kar3/Cik1 promoted microtubule shortening. Microtubule•motor complexes were preformed in the presence of 1 mM MgAMPPNP and imaged at t = 0, column 1. MgATP at 1.5 mM plus an ATP regeneration system initiated microtubule shortening (Sproul et al., 2005). Column 3 is the merge of t = 0 and the elapsed time (middle column) to show microtubule shortening (red) in comparison to the original length. Polarity-marked microtubules were identified from microtubule•motor populations at both 25 and 50 nM motor incubated with 500 nM microtubules in the presence of MgATP. (D) Increased motor binding to microtubules stabilizes the microtubule lattice against shortening. Upper panel: Kar3/Cik1 and Kar3/Vik1 rates of microtubule shortening plotted as a function of increasing motor concentration. Lower panel: The percentage of microtubules that showed Kar3/Cik1 or Kar3/Vik1-promoted ATP-dependent shortening plotted as a function of motor concentration. Data are reported as mean +/− SEM.
Cooperative Binding of Kar3/Vik1 to Microtubules
Kar3/Vik1 exhibits cooperative binding to the microtubule lattice (Figure 5). This microtubule-binding behavior differs from Kar3/Cik1, which exhibits a preference for binding microtubule plus-ends, and also from Kar3MD alone, which displays stochastic microtubule-lattice binding characteristics (Sproul et al., 2005). In the presence of AMPPNP, microtubule•motor complexes were assembled in solution and then fixed with glutaraldehyde. These complexes were subsequently centrifuged through a glycerol cushion onto coverslips and processed for immunofluorescence using affinity purified antibodies to Kar3MD or Vik1MHD (Figure S1) (Sproul et al., 2005). The results show that one microtubule is completely saturated by Kar3/Vik1 with other nearby microtubules showing no evidence of motor binding (Figure 5A–C). As the concentration of Kar3/Vik1 was increased from 25–100 nM, additional microtubules in the field of view became saturated with Kar3/Vik1 with other nearby microtubules showing no evidence of Kar3/Vik1 immunofluorescence (Figure 5D–F). At 25 nM Vik1MHD, there were some examples of microtubule end binding (Figure 5P–R), however, as the concentration of the Vik1MHD increased, the microtubule lattice showed increased Vik1MHD occupancy, and microtubules became saturated (Figure 5S–X). Note that even at 25 nM Kar3/Vik1, there were few examples of microtubule end or lattice binding. Most microtubules scored exhibited Kar3/Vik1 saturated binding (90–98.5%). This cooperative microtubule binding behavior was also observed for Drosophila Ncd (Wendt et al., 2002), yet conventional Kinesin-1 does not exhibit this binding pattern (Sproul et al., 2005).
Figure 5. Immunolocalization of Kar3/Vik1, Kar3/Cik1, Kar3MD, and Vik1MHD.

Microtubule•motor complexes were preformed in the presence of MgAMPPNP. Final concentrations: 500 nM tubulin polymer, 40 μM Taxol, and 1 mM MgAMPPNP. Rows 1 and 2 represent magnification of a section of the field, whereas the remaining rows show individual microtubules at a higher magnification (scale bars = 5 μm). The microtubule seed (arrowhead) marks the microtubule minus end and (*) denotes the dim microtubule plus end. The first column of each row shows the rhodamine-labeled microtubules, and the second column, the immunofluorescence of affinity-purified Vik1MHD antibodies. The third column is the merge of the two channels to show the co-localization. The table presents the summary of microtubule localization events scored for the three motors using affinity-purified antibodies to the Kar3MD or the Vik1MHD (Figure S1).
The Interaction Between Kar3 and Vik1 is Similar to the Ncd Dimer
The orientation of the neck of Vik1MHD is similar to that observed in one of the motor domains of a motility deficient Ncd homodimer (Figure 6) (Yun et al., 2003). In this structure, the two heads are asymmetrically positioned, one ~75º relative to the other, in relation to their necks. Hence, the motor domain-neck interactions are different for each motor within the dimer. Cryo-electron microscopy studies of Ncd-decorated microtubules have demonstrated that this type of neck rotation (from position A to position B in Figure 6) occurs in the microtubule-bound Ncd motor upon ATP-binding, causing the neck to point toward the minus end of the microtubule (Wendt et al., 2002; Endres et al., 2006). This rotation of the neck appears to be the force-producing conformational change that drives minus-end-directed motility in Ncd (Endres et al., 2006). The pivot point allowing this rotation of the neck occurs at Gly347 in Ncd. This glycine is highly conserved among the kinesin superfamily and in Vik1. The identities and positions of several of the residues that hold the neck in either of its two positions along the motor domain in Ncd are also found in Vik1 and Kar3. This suggests that the same rotation of the neck occurs in Kar3/Vik1 and that, like Ncd, the coiled-coil formed by the Kar3/Vik1 heterodimer extends to their motor domains. What is not yet clear is how rotation of the neck of Vik1 might be incorporated in Kar3’s motile cycle because the structural models for Ncd propose that only a single motor head of the Ncd dimer interacts with the microtubule (Wendt et al., 2002; Endres et al., 2006). However, the Kar3/Vik1 structure and kinetics provide constraints for models that might incorporate this concept.
Figure 6. Comparison of the Neck Orientations of Vik1 and Ncd.

The view shown has been rotated 90° from Figure 2A and highlights the positions of the neck α-helix (red) of Vik1MHD (left) and two different positions (red and blue) adopted by the neck α-helix of the N600K mutant form of Ncd (PDB accession:1N6M) (Yun et al., 2003). For the Ncd structures, only the α-carbons of residues found in the two motor domains have been superimposed. The α-carbons for the Vik1MHD, excluding the neck, were superimposed onto those of the Ncd motor domain. Atoms for the conserved glycine residue that allows neck rotation, Gly373 in Vik1 and Gly347 in Ncd, are shown as cyan and yellow spheres.
DISCUSSION
Mechanism of Motility in the Kar3/Vik1 Heterodimer
There are two fundamentally different models that might explain how Kar3/Vik1 functions as a molecular motor (Figure 7). The first model is based on the presumption that the overall orientation of the globular domains of both Kar3 and Vik1 when bound to microtubules is similar to that seen for processive kinesins (Model A). This requires that the coiled-coil between Kar3 and Vik1 unwinds to accommodate the 8 nm distance required for both Vik1 and Kar3 to bind the same microtubule protofilament. In Step 1 of this model, Vik1 makes the initial microtubule interaction in an orientation on α/β-tubulin similar to kinesin and Ncd, while Kar3 is tethered with ADP in its active site. This brings Kar3 in close proximity to the next α/β-tubulin in the same microtubule protofilament. Binding of the Kar3MD stimulates a conformational change in the motor that induces ADP release and generates strain (Figure 7, black arrows) in the coiled-coil between Kar3 and Vik1 (Step 2). This strain is communicated to the microtubule-binding surface of Vik1MHD, resulting in weakening of the Vik1MHD-microtubule interaction. Subsequently, as in Ncd, the binding of ATP to the active site of Kar3 would result in a large rotation of Kar3’s neck toward the minus end, causing Vik1 to disengage from the microtubule and the coiled-coil of Kar3/Vik1 to be displaced toward the minus end (Step 3). ATP hydrolysis by Kar3 and/or Pi release returns Kar3 to a weakly bound intermediate, resulting in Kar3/Vik1 detachment from the microtubule (Step 4). It is assumed that after ATP hydrolysis Vik1 will be oriented so that it is unable to rebind the microtubule through a backward step. The second model proposes that the coiled-coil between the necks of Kar3 and Vik1 does not unwind, which maintains the two globular domains in close proximity (Model B). This necessitates that the Kar3MD and Vik1MHD interact with adjacent protofilaments, and demands that Vik1 must bind α/β-tubulin in a different orientation than the kinesin motor domains. An altered binding mode is not unreasonable given the substantial differences in the microtubule-binding elements of Vik1 relative to kinesin in general and the similarity in sequence and structure of helices H11 and H12 of α- and β-tubulin (Nogales et al., 1998). In this model, the steps involved in Kar3/Vik1 binding to the microtubule, production of a minus-end-directed rotation of the coiled-coil, and subsequent release from the microtubule are essentially identical to Model A but are more attractive from the standpoint of conformational simplicity for the Kar3/Vik1 heterodimer. One consequence of this model is that the structural change of the N-terminal helix of Vik1MHD will have the opposite sense of rotation in models A and B during the motile cycle.
Figure 7. Models of the Kar3/Vik1 Motility Mechanism.

The schematic drawings show two possible models for the interaction of the Kar3/Vik1 heterodimer with α/β-tubulin protofilaments during the motility cycle. The microtubule is oriented so that the minus end is on the left. In Model A, the coiled-coil (yellow) formed between the necks of Kar3 and Vik1 unwinds to allow both Kar3 and Vik1 to bind the same microtubule protofilament in a similar orientation, analogous to processive kinesins. In Model B, the coiled-coil between Kar3 and Vik1 does not unwind, keeping the globular domains of Kar3 and Vik1 close to each other. This imposes restraints on their interaction with the microtubule, such that the Kar3MD and Vik1MHD would have to interact with adjacent protofilaments and that Vik1 must bind α/β-tubulin in a different orientation than the motor domain of kinesins. In both models, the binding of the Kar3MD to the microtubule stimulates a conformational change in the motor that generates strain in the coiled-coil between Kar3 and Vik1. This strain produces a conformational change in the Vik1MHD that weakens Vik1’s interaction with the microtubule, which allows the large rotation of Kar3’s neck toward the minus end upon entry of ATP into the active site of Kar3 and complete disengagement of Vik1 from the microtubule. The direction of rotation of the coiled-coil and the timing of this event during the motile cycle is based upon cryo-electron microscopy studies of Ncd-decorated microtubules (Wendt et al., 2002; Endres et al., 2006).
While these are only two possible models, it is clear that Vik1 enhances the fidelity of Kar3’s interaction with microtubules and its presence in the heterodimer results in recruitment of many dimers to a specific region (spindle poles) for biopolar spindle assembly and chromosome segregation. It is also apparent that elements for microtubule binding and heterodimerization in Vik1 are ‘wired’ with similar structures to those found in kinesin, but perhaps in a manner that negates the need for its own motor capability. In this respect, communication between Kar3 and Vik1, and microtubules, may be more direct than in kinesin homodimers or heterodimers with two functional nucleotide-binding motors. The structure of Vik1MHD provides an excellent starting point from which to begin dissecting the method of communication between Kar3 and Vik1.
Functional Significance of Kar3/Vik1 Cooperative Binding
The plus-end microtubule localization by Kar3/Cik1 is consistent with this complex functioning to shorten microtubules in order to pull the nuclei together for nuclear fusion during mating (Maddox et al., 2003; Sproul et al., 2005; Molk et al., 2006). On the other hand, based on the studies presented here, Kar3/Vik1 does not possess the hallmarks of an in vivo microtubule depolymerase. In fact, it can be argued that the cooperative binding by Kar3/Vik1 to microtubules would act to stabilize the microtubules thereby inhibiting depolymerization (Figure 4D). Furthermore, the localization of Kar3/Vik1 at the spindle pole bodies is inconsistent with microtubule depolymerization because the microtubule dynamics have been shown to occur predominantly at the microtubule-plus ends in budding yeast (Maddox et al., 1999). We propose that Kar3/Vik1 accumulates at the microtubule minus-ends by ATP-dependent movement, and its function is to cross-link and focus microtubule minus-ends at the spindle poles for bipolar spindle assembly and stabilization as observed for Drosophila Ncd (Kimble and Church, 1983; Hatsumi and Endow, 1992; Sharp et al., 2000). In addition, the genetics suggest that Kar3/Vik1, like fission yeast S. pombe Pkl1 and Drosophila Ncd, would act as an opposing force to the plus-end directed homotetrameric Eg5 motors Cin8 and Kip1 (Saunders and Hoyt, 1992; Pidoux et al., 1996; Manning et al., 1999; Sharp et al., 2000; Troxell et al., 2001). These structural and mechanistic results are consistent with distinct functional roles for Kar3/Vik1 during mitotis.
Origin of Vik1
Paralogs of one or both Vik1 and Cik1 can be found in many of the hemiascomycete yeasts and most likely arose after the massive gene duplication that occurred early the history of these organisms. The similarity in the structural organization of Vik1MHD to Kar3 supports the hypothesis that Vik1 and Cik1 share a common ancestor that was almost certainly a member of the Kinesin-14 family of molecular motors. The existence of both Vik1 and Cik1 in fungi like S. cerevisiae and Candida glabrata suggests a second gene duplication event subsequently occurred that allowed the evolution of two related proteins with distinct functions. It would appear that, Vik1 and Cik1 retained the motor-like fold, allowing them to bind microtubules and form dimeric motor complexes, but in the process lost their nucleotide-binding requirement. However, it is likely that this could only have occurred if communication between the two motor domains was already an inherent property of the original Kinesin-14.
EXPERIMENTAL PROCEDURES
Constructs
The Kar3/Cik1, Kar3/Vik1, Kar3MD, and Vik1 MHD motor constructs used in this study (Figure S1) were amplified from the full-length genes (a gift from Dr. Michael Snyder, Yale University) by PCR (see supplementary material online for primer sequences). The truncated version of Kar3 (Lys268-Lys729) used to make the heterodimer complex with Cik1 or Vik1 was cloned into pET 24d (Novagen, selection kanamycin) using Nco1 and BamH1. This plasmid when expressed yields amino acid residues MetGly-Lys268-Lys729 with a predicted molecular mass Mr of 52,819. The truncated version of Cik1 (Lys252-Asp594) was cloned into pET 15b (ampicillin selection) at the Nde1 and Bam H1 sites. This construct yields residues MGSSH6SSGGLVPRGSHMet-Lys252-Asp594 with predicted Mr = 43,059. Truncated Vik1 (Leu253-Thr648) was cloned into pET 16b (ampicillin selection) at Nde1 and BamH1. When expressed, this construct yields residues MGH9SSGHIEGRHM-Leu253-Thr648 with a predicted Mr = 58,796. The Vik1MHD construct (Thr353-Thr648) was cloned into a modified version of the pET31b vector (ampicillin selection) called pKLD37 at Nhe1 and Blp I sites. This vector incorporates a His6-tag and a rTEV proteolytic cleavage site prior to the N-terminus of the protein. When expressed, this construct yields residues MSYYH6DYDIPTSENLYFQGASThr353-Thr648. After rTEV cleavage, the protein that remains includes GASThr353-Thr648 with a predicted Mr = 34,586. The Kar3MD construct (Met383-Lys729) was cloned as previously described (Gulick et al., 1998). Its predicted Mr = 38,888.
Protein Expression and Purification
The Kar3 and Vik1 or Cik1 plasmids were co-expressed in E. coli BL21-CodonPlus (DE3)-RIL cell line (Stratagene). The Kar3/Vik1 and Kar3/Cik1 heterodimers were purified as described previously (Sproul et al., 2005), followed by gel filtration (Superose 6 10/300 GL, Amersham Biosciences. Native and selenomethionine-labeled Vik1MHD were also expressed in the E. coli BL21-CodonPlus (DE3)-RIL cell line in LB and M-9 minimal media for the cell culture, respectively. Selenomethionine incorporation was performed by growing the cells at 37 °C to an A600 of ~0.9 and then cooling them on ice for 10 min, followed by incubation at 20 °C for 10 min. At this time, each flask was supplemented with 50 mg each of L-Lysine, L-threonine, L-Phenylalanine, and 25 mg each of L-leucine, L-isoleucine, L-valine, and L-selenomethionine. After an additional 30 min the cells were induced with 0.5 mM IPTG and then grown for 16 hr at 20 °C with shaking prior to harvesting by centrifugation. Native and selenomethionine-labeled Vik1MHD were both purified as described in supplemental online material.
Crystallization of Native and Selenomethionine-labeled Vik1MHD
Crystals of both native and selenomethionine-labeled Vik1MHD (SeMetVik1MHD) were grown by hanging drop vapor diffusion at 4 °C by mixing the protein 1:1 with 100 mM Na/MES/Acetate, pH 5.5, 24 % pentaerythritol ethoxylate (Mr 797), 300 mM NaCl, and 5% ethylene glycol. Single crystals grew to maximum dimensions of ~ 0.6 × 0.2 × 0.2 mm in 2 weeks. Prior to data collection the crystals were transferred directly into 100% of the precipitant solution for ~2 min and then frozen in a stream of nitrogen gas. Microtubule binding studies were performed with SeMetVik1MHD in direct comparison to native Vik1MHD. The results in Figure 3, which determined microtubule affinity, show that the functional behavior of SeMetVik1MHD was very similar to native Vik1MHD.
X-ray Data collection and Structure Refinement
X-ray diffraction data for the native and selenomethionine-labeled Vik1MHD crystals were collected at the SBC 19-BM beam line, Advanced Photon Source, Argonne, IL. The data sets were integrated and scaled with the program HKL2000 (Otwinowski and Minor, 1997). X-ray data collection statistics are given in Table S1. The structure of the Vik1MHD was solved by multiwavelength anomalous dispersion. The positions of the 5 selenium atoms in the asymmetric unit were determined and refined with the program SOLVE (Terwilliger and Berendzen, 1999). Solvent flattening with the program RESOLVE (Terwilliger, 2000) yielded a readily interpretable electron density map at 2.0 Å resolution. A model was built with ARP/wARP (Perrakis et al., 2001) and subjected to manual and automated refinement using TURBO (Roussel and Cambillau, 1991) and Refmac5 (Murshudov et al., 1997), respectively. This model was further refined against the native Vik1MHD crystal diffraction data to a resolution of 1.6 Å. Water molecules were added with ARP/wARP and manually verified. Refinement statistics are given in Table S1. The PDB ID code is 2O0A.
Kar3/Vik1 and Kar3/Cik1 Steady State ATPase
The steady state kinetics of Kar3/Vik1 and Kar3/Cik1 were determined by following the hydrolysis of [α32P] ATP to [α32P] ADP•Pi. The steady-state kinetics as a function of microtubule (MT) concentration (Figure 4B) were fit to the quadratic equation:
| (Eq. 1) |
where Rate denotes the amount of product formed per second per active site, kcat is the maximum rate constant of product formed at saturating substrate, E0 is the motor concentration, and MT0 is the tubulin polymer concentration. The quadratic equation is required because the enzyme concentration is not 10-fold less than the K1/2,MT. These conditions represent stoichiometric binding of the motor and microtubules. The steady-state kinetics as a function of MgATP concentration were fit to the Michaelis-Menton equation. Taxol was maintained at 40μM to stabilize the MTs.
Fluorescence Microscopy Assays
The methods used for the microscopy experiments presented in Figures 4 and 5 are described in more detail in Sproul et al. (Sproul et al., 2005). The Taxol concentration required for each experimental design was determined experimentally.
Kar3/Vik1 and Kar3/Cik1 Time Lapse Microtubule Shortening
Motor at 25, 50, or 100 nM was incubated with 500 nM rhodamine microtubules stabilized at 5μM Taxol in the presence of MgAMPPNP in PME (10 mM PIPES, pH 6.9, 5 mM MgCl2, 1 mM EGTA). An 8 μl aliquot of the complex was flowed into an observation chamber. The complex was incubated for 3 min at room temperature to allow the N-termini of the motors containing poly-His-tags to interact with the glass. Unattached microtubule•motor complexes were removed by perfusion of two 8-μl washes of an oxygen scavenging mix (OSM) + MgAMPPNP (Sproul et al., 2005). Microtubule shortening was initiated by MgATP and imaged over 20 min with frames captured every 20 s. Taxol was maintained at 5μM. At this concentration, microtubules were stable and not observed to shorten in the absence of MgATP and Kar3/Cik1 or Kar3/Vik1. However, in the presence of MgATP plus motor, microtubule shortening was observed.
Microtubule•Motor Immunolocalization
Reactions at 10 μl were formed containing the microtubule•motor complex (25 – 400 nM motor, 500 nM tubulin polymer, and 40 μM Taxol) in the presence of 1 mM MgAMPPNP. The reactions were fixed in 10 volumes of 1% glutaraldehyde in PME (10 mM PIPES, pH 6.9, 5 mM MgCl2, 1 mM EGTA) and processed as described in supplemental online material. The primary affinity-purified polyclonal Kar3 or Vik1 antibodies, generated to the native Kar3MD (Sproul et al., 2005) or Vik1MHD (see Figure S1), were used to localize Kar3MD and Kar3Cik1, or Kar3Vik1 and Vik1MHD, respectively.
Microtubule•Motor Equilibrium Binding Assays
Soluble tubulin was adjusted to 1 mM MgGTP, cold depolymerized, clarified, and cycled each morning of the experiment. All concentrations reported are final after mixing. Reactions of 150 μl microtubules (0–3 μM tubulin) were incubated with 50 nM motor for 10 min at room temperature in PME Buffer. MgAMPPNP or MgADP (2 mM final) or 0.1 U/ml apyrase was then added, and the reactions were incubated for 30–60 min to reach equilibrium. The microtubules and associated proteins were sedimented at 100,000 × g for 30 min at 34 °C (Beckman Coulter TLX Ultracentrifuge). Supernatant fractions were analyzed by SDS-PAGE, followed by staining with Sypro Ruby ® (Invitrogen). To quantify the motor or Vik1MHD that cosedimented with microtubules, a standard curve was used with the corresponding protein within a range of concentrations where Sypro Ruby-staining was linear. The protein was quantified using Image J. The data were plotted as the Fraction of Motor/Protein in the Pellet as a function of MT concentration and fit to quadratic equation 2:
| (Eq. 2) |
where MT•E is the fraction of motor or protein sedimenting with the microtubule pellet, E0 is the total motor or Vik1MHD, and Kd is the dissociation constant. Although the data in Figure 5 indicate cooperative binding of Kar3/Vik1 to microtubules, the equilibrium binding studies were not sensitive enough to detect sigmoidal binding behavior. Therefore, the equilibrium binding data were fit to Eq. 2.
Supplementary Material
Supplemental data includes primer sequences, experimental methods, five figures, two tables, and two movies and can be found with this article online at http://www.cell.com/cgi/content/full.
Acknowledgments
We thank Kristen Dennison and David Close for technical assistance. This work was supported by grants from the NIH to I.R. (AR35186) and S.P.G. (GM54141 and Career Development Award K02-AR47841). J.S.A. was supported by a Canadian Institutes of Health Postdoctoral Fellowship (64606). Use of the Structural Biology BM19 beamline Argonne National Laboratory Advanced Photon Source was supported by the U. S. Department of Energy, Office of Energy Research, under Contract No. W-31-109-ENG-38. Analytical ultracentrifugation data were obtained at the UW - Madison Biophysics Instrumentation Facility, which was established with support from the NSF (BIR-9512577) and NIH (S10 RR13790).
Footnotes
The structural coordinates for Vik1MHD have been deposited in the RCSB under accession number 2O0A.
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Alonso MC, van Damme J, Vandekerckhove J, Cross RA. Proteolytic mapping of kinesin/ncd-microtubule interface: nucleotide-dependent conformational changes in the loops L8 and L12. EMBO J. 1998;17:945–951. doi: 10.1093/emboj/17.4.945. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baker NA, Sept D, Joseph S, Holst MJ, McCammon JA. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc Natl Acad Sci USA. 2001;98:10037–10041. doi: 10.1073/pnas.181342398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barrett JG, Manning BD, Snyder M. The Kar3p kinesin-related protein forms a novel heterodimeric structure with its associated protein Cik1p. Mol Biol Cell. 2000;11:2373–2385. doi: 10.1091/mbc.11.7.2373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berger B, Wilson DB, Wolf E, Tonchev T, Milla M, Kim PS. Predicting coiled coils by use of pairwise residue correlations. Proc Natl Acad Sci U S A. 1995;92:8259–8263. doi: 10.1073/pnas.92.18.8259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bond CS. TopDraw: a sketchpad for protein structure topology cartoons. Bioinformatics. 2003;19:311–312. doi: 10.1093/bioinformatics/19.2.311. [DOI] [PubMed] [Google Scholar]
- Chu HM, Yun M, Anderson DE, Sage H, Park HW, Endow SA. Kar3 interaction with Cik1 alters motor structure and function. EMBO J. 2005;24:3214–3223. doi: 10.1038/sj.emboj.7600790. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cochran JC, Sontag CA, Maliga Z, Kapoor TM, Correia JJ, Gilbert SP. Mechanistic analysis of the mitotic kinesin Eg5. J Biol Chem. 2004;279:38861–38870. doi: 10.1074/jbc.M404203200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DeLano WL. The PyMOL Molecular Graphics System. www.pymol.org 2002
- Endow SA, Henikoff S, Soler-Niedziela L. Mediation of meiotic and early mitotic chromosome segregation in Drosophila by a protein related to kinesin. Nature. 1990;345:81–83. doi: 10.1038/345081a0. [DOI] [PubMed] [Google Scholar]
- Endres NF, Yoshioka C, Milligan RA, Vale RD. A lever-arm rotation drives motility of the minus-end-directed kinesin Ncd. Nature. 2006;439:875–878. doi: 10.1038/nature04320. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gulick AM, Song H, Endow SA, Rayment I. X-ray crystal structure of the yeast Kar3 motor domain complexed with Mg. ADP to 23 A resolution. Biochemistry. 1998;37:1769–1776. doi: 10.1021/bi972504o. [DOI] [PubMed] [Google Scholar]
- Hatsumi M, Endow SA. Mutants of the microtubule motor protein, nonclaret disjunctional, affect spindle structure and chromosome movement in meiosis and mitosis. J Cell Sci. 1992;101(Pt 3):547–559. doi: 10.1242/jcs.101.3.547. [DOI] [PubMed] [Google Scholar]
- Hildebrandt ER, Hoyt MA. Mitotic motors in Saccharomyces cerevisiae. Biochim Biophys Acta. 2000;1496:99–116. doi: 10.1016/s0167-4889(00)00012-4. [DOI] [PubMed] [Google Scholar]
- Hirose K, Cross RA, Amos LA. Nucleotide-dependent structural changes in dimeric NCD molecules complexed to microtubules. J Mol Biol. 1998;278:389–400. doi: 10.1006/jmbi.1998.1709. [DOI] [PubMed] [Google Scholar]
- Kikkawa M, Sablin EP, Okada Y, Yajima H, Fletterick RJ, Hirokawa N. Switch-based mechanism of kinesin motors. Nature. 2001;411:439–445. doi: 10.1038/35078000. [DOI] [PubMed] [Google Scholar]
- Kimble M, Church K. Meiosis and early cleavage in Drosophila melanogaster eggs: effects of the claret-non-disjunctional mutation. J Cell Sci. 1983;62:301–318. doi: 10.1242/jcs.62.1.301. [DOI] [PubMed] [Google Scholar]
- Kull FJ, Sablin EP, Lau R, Fletterick RJ, Vale RD. Crystal Structure of the Kinesin Motor Domain Reveals a Structural Similarity to Myosin. Nature. 1996;380:550–555. doi: 10.1038/380550a0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mackey AT, Gilbert SP. The ATPase cross-bridge cycle of the Kar3 motor domain. Implications for single head motility. J Biol Chem. 2003;278:3527–3535. doi: 10.1074/jbc.M206219200. [DOI] [PubMed] [Google Scholar]
- Maddox P, Chin E, Mallavarapu A, Yeh E, Salmon ED, Bloom K. Microtubule dynamics from mating through the first zygotic division in the budding yeast Saccharomyces cerevisiae. J Cell Biol. 1999;144:977–987. doi: 10.1083/jcb.144.5.977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maddox PS, Stemple JK, Satterwhite L, Salmon ED, Bloom K. The minus end-directed motor Kar3 is required for coupling dynamic microtubule plus ends to the cortical shmoo tip in budding yeast. Curr Biol. 2003;13:1423–1428. doi: 10.1016/s0960-9822(03)00547-5. [DOI] [PubMed] [Google Scholar]
- Manning BD, Barrett JG, Wallace JA, Granok H, Snyder M. Differential regulation of the Kar3p kinesin-related protein by two associated proteins, Cik1p and Vik1p. J Cell Biol. 1999;144:1219–1233. doi: 10.1083/jcb.144.6.1219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manning BD, Snyder M. Drivers and passengers wanted! the role of kinesin-associated proteins. Trends Cell Biol. 2000;10:281–289. doi: 10.1016/s0962-8924(00)01774-8. [DOI] [PubMed] [Google Scholar]
- McDonald HB, Goldstein LS. Identification and characterization of a gene encoding a kinesin-like protein in Drosophila. Cell. 1990;61:991–1000. doi: 10.1016/0092-8674(90)90064-l. [DOI] [PubMed] [Google Scholar]
- McDonald HB, Stewart RJ, Goldstein LS. The kinesin-like ncd protein of Drosophila is a minus end-directed microtubule motor. Cell. 1990;63:1159–1165. doi: 10.1016/0092-8674(90)90412-8. [DOI] [PubMed] [Google Scholar]
- Meluh PB, Rose MD. KAR3, a kinesin-related gene required for yeast nuclear fusion. Cell. 1990;60:1029–1041. doi: 10.1016/0092-8674(90)90351-e. [DOI] [PubMed] [Google Scholar]
- Molk JN, Salmon ED, Bloom K. Nuclear congression is driven by cytoplasmic microtubule plus end interactions in S. cerevisiae. J Cell Biol. 2006;172:27–39. doi: 10.1083/jcb.200510032. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murshudov GN, Vagin AA, Dodson EJ. Refinement of Macromolecular Structures by the Maximum-Likelihood Method. Acta Crystallogr D Biol Crystallogr. 1997;53:240–255. doi: 10.1107/S0907444996012255. [DOI] [PubMed] [Google Scholar]
- Nogales E, Wolf SG, Downing KH. Structure of the alpha beta tubulin dimer by electron crystallography. Nature. 1998;391:199–203. doi: 10.1038/34465. [DOI] [PubMed] [Google Scholar]
- Otwinowski Z, Minor W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 1997;276:307–326. doi: 10.1016/S0076-6879(97)76066-X. [DOI] [PubMed] [Google Scholar]
- Page BD, Satterwhite LL, Rose MD, Snyder M. Localization of the Kar3 kinesin heavy chain-related protein requires the Cik1 interacting protein. J Cell Biol. 1994;124:507–519. doi: 10.1083/jcb.124.4.507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Page BD, Snyder M. CIK1: a developmentally regulated spindle pole body-associated protein important for microtubule functions in Saccharomyces cerevisiae. Genes Dev. 1992;6:1414–1429. doi: 10.1101/gad.6.8.1414. [DOI] [PubMed] [Google Scholar]
- Perrakis A, Harkiolaki M, Wilson KS, Lamzin VS. ARP/wARP and molecular replacement. Acta Crystallogr D Biol Crystallogr. 2001;57:1445–1450. doi: 10.1107/s0907444901014007. [DOI] [PubMed] [Google Scholar]
- Pidoux AL, LeDizet M, Cande WZ. Fission yeast pkl1 is a kinesin-related protein involved in mitotic spindle function. Mol Biol Cell. 1996;7:1639–1655. doi: 10.1091/mbc.7.10.1639. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roussel A, Cambillau C. Silicon Graphics Geometry Partners Directory. Mountain View, California: Silicon Graphics; 1991. Turbo Frodo. [Google Scholar]
- Sablin EP, Case RB, Dai SC, Hart CL, Ruby A, Vale RD, Fletterick RJ. Direction determination in the minus-end-directed kinesin motor ncd. Nature. 1998;395:813–816. doi: 10.1038/27463. [DOI] [PubMed] [Google Scholar]
- Sablin EP, Fletterick RJ. Coordination between motor domains in processive kinesins. J Biol Chem. 2004;279:15707–15710. doi: 10.1074/jbc.R300036200. [DOI] [PubMed] [Google Scholar]
- Sablin EP, Kull FJ, Cooke R, Vale RD, Fletterick RJ. Three-Dimensional Structure of the Motor Domain of NCD, a Kinesin-Related Motor with Reversed Polarity of Movement. Nature. 1996;380:555–559. doi: 10.1038/380555a0. [DOI] [PubMed] [Google Scholar]
- Sack S, Kull FJ, Mandelkow E. Motor proteins of the kinesin family. Structures, variations, and nucleotide binding sites. Eur J Biochem. 1999;262:1–11. doi: 10.1046/j.1432-1327.1999.00341.x. [DOI] [PubMed] [Google Scholar]
- Saunders WS, Hoyt MA. Kinesin-related proteins required for structural integrity of the mitotic spindle. Cell. 1992;70:451–458. doi: 10.1016/0092-8674(92)90169-d. [DOI] [PubMed] [Google Scholar]
- Schulz GE. Binding of nucleotides by proteins. Curr Opin Struct Biol. 1992;2:61–67. [Google Scholar]
- Sharp DJ, Brown HM, Kwon M, Rogers GC, Holland G, Scholey JM. Functional coordination of three mitotic motors in Drosophila embryos. Mol Biol Cell. 2000;11:241–253. doi: 10.1091/mbc.11.1.241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Song YH, Marx A, Muller J, Woehlke G, Schliwa M, Krebs A, Hoenger A, Mandelkow E. Structure of a fast kinesin: implications for ATPase mechanism and interactions with microtubules. EMBO J. 2001;20:6213–6225. doi: 10.1093/emboj/20.22.6213. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sosa H, Dias DP, Hoenger A, Whittaker M, Wilson-Kubalek E, Sablin E, Fletterick RJ, Vale RD, Milligan RA. A model for the microtubule-Ncd motor protein complex obtained by cryo-electron microscopy and image analysis. Cell. 1997;90:217–224. doi: 10.1016/s0092-8674(00)80330-x. [DOI] [PubMed] [Google Scholar]
- Sproul LR, Anderson DJ, Mackey AT, Saunders WS, Gilbert SP. Cik1 targets the minus-end kinesin depolymerase kar3 to microtubule plus ends. Curr Biol. 2005;15:1420–1427. doi: 10.1016/j.cub.2005.06.066. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Terwilliger TC. Maximum-likelihood density modification. Acta Crystallogr D Biol Crystallogr. 2000;56:965–972. doi: 10.1107/S0907444900005072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Terwilliger TC, Berendzen J. Automated MAD and MIR structure solution. Acta Crystallogr. 1999;D55:849–861. doi: 10.1107/S0907444999000839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Troxell CL, Sweezy MA, West RR, Reed KD, Carson BD, Pidoux AL, Cande WZ, McIntosh JR. pkl1(+)and klp2(+): Two kinesins of the Kar3 subfamily in fission yeast perform different functions in both mitosis and meiosis. Mol Biol Cell. 2001;12:3476–3488. doi: 10.1091/mbc.12.11.3476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vale RD, Milligan RA. The way things move: looking under the hood of molecular motor proteins. Science. 2000;288:88–95. doi: 10.1126/science.288.5463.88. [DOI] [PubMed] [Google Scholar]
- Wendt TG, Volkmann N, Skiniotis G, Goldie KN, Muller J, Mandelkow E, Hoenger A. Microscopic evidence for a minus-end-directed power stroke in the kinesin motor ncd. EMBO J. 2002;21:5969–5978. doi: 10.1093/emboj/cdf622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Westhead DR, Slidel TW, Flores TP, Thornton JM. Protein structural topology: Automated analysis and diagrammatic representation. Protein Sci. 1999;8:897–904. doi: 10.1110/ps.8.4.897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woehlke G, Ruby AK, Hart CL, Ly B, Hom-Booher N, Vale RD. Microtubule interaction site of the kinesin motor. Cell. 1997;90:207–216. doi: 10.1016/s0092-8674(00)80329-3. [DOI] [PubMed] [Google Scholar]
- Yun M, Bronner CE, Park CG, Cha SS, Park HW, Endow SA. Rotation of the stalk/neck and one head in a new crystal structure of the kinesin motor protein, Ncd. EMBO J. 2003;22:5382–5389. doi: 10.1093/emboj/cdg531. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplemental data includes primer sequences, experimental methods, five figures, two tables, and two movies and can be found with this article online at http://www.cell.com/cgi/content/full.


