Abstract
Human chromosome 11q23 translocations disrupting MLL result in poor prognostic leukemias. It fuses the common MLL N-terminal ∼1400 amino acids in-frame with >60 different partners without shared characteristics. In addition to the well-characterized activity of MLL in maintaining Hox gene expression, our recent studies established an MLL–E2F axis in orchestrating core cell cycle gene expression including Cyclins. Here, we demonstrate a biphasic expression of MLL conferred by defined windows of degradation mediated by specialized cell cycle E3 ligases. Specifically, SCFSkp2 and APCCdc20 mark MLL for degradation at S phase and late M phase, respectively. Abolished peak expression of MLL incurs corresponding defects in G1/S transition and M-phase progression. Conversely, overexpression of MLL blocks S-phase progression. Remarkably, MLL degradation initiates at its N-terminal ∼1400 amino acids, and tested prevalent MLL fusions are resistant to degradation. Thus, impaired degradation of MLL fusions likely constitutes the universal mechanism underlying all MLL leukemias. Our data conclude an essential post-translational regulation of MLL by the cell cycle ubiquitin/proteasome system (UPS) assures the temporal necessity of MLL in coordinating cell cycle progression.
Keywords: MLL, Taspase1, Skp2, Cdc20, cell cycle, leukemia
The eukaryotic cell cycle consists of a tightly orchestrated circular progression of a sequence of distinct phases—namely, G1, S, G2, and M—designated for the execution of inherent genetic programs (Nurse 2000). Key regulatory components of the mammalian cell cycle machinery include E2Fs, Rbs, Cyclins, Cyclin-dependent kinases (CDKs), and CDK inhibitors (CDKIs), which form complex positive and negative epistatic circuits ensuring accurate cell cycle progression (Murray 2004). The molecular blueprint of a normal cell cycle details the central controls held by a series of CDKs that positively influences proliferation through phosphorylating target proteins such as Rbs (Morgan 1997). This releases the repressive activities of Rbs on E2Fs, the major executors of cell cycle gene expression programs (Dyson 1998; Nevins 2001; Trimarchi and Lees 2002; Blais and Dynlacht 2004; Bracken et al. 2004; Giacinti and Giordano 2006). Individual CDKs need to complex with specialized Cyclins to form catalytically active Cyclin/CDKs; therefore, their activities depend on the availability of involved Cyclins. Additional regulations of Cyclin/CDKs come from negative factors, the CDKIs (Sherr and Roberts 1999). Levels of Cyclins and CDKIs undulate during cell division, which in part is due to the defined windows of degradation by the ubiquitin–proteasome system (UPS) (Reed 2003). UPS commences at substrate recognition and the subsequent covalent conjugation of ubiquitin by the responsible E3 ligases, followed by degradation executed by the 26S proteasome (Hershko 2005). The two major E3 complexes involved in proteolyzing core components of the cell cycle machinery are SCF (Skp1–Cul1–F-box protein) and APC (anaphase-promoting complex/cyclosome) complexes, the substrate recognition of which is conducted by the variable components—F-box proteins for SCF, and Cdc20 and Cdh1 for APC (Cardozo and Pagano 2004; Nakayama and Nakayama 2006; Peters 2006). Thus, two types of post-translational modifications—phosphorylation and ubiquitination—constitute the two mainstream controls of the cell cycle.
Polycomb group (PcG) and trithorax group (trxG) proteins are chromatin modifiers required for the epigenetic maintenance of repressive versus active states of cell fate specification genes (Schumacher and Magnuson 1997; Yu et al. 1998; Hanson et al. 1999; Ringrose and Paro 2004; Schuettengruber et al. 2007). The most-characterized targets are the Hox genes in vertebrates and homeotic genes in invertebrates. Beyond perpetuating cellular memory through multiple cell divisions, recent studies on mammals and flies highlighted the active, dynamic participation of PcG proteins in the transcriptional control of cell cycle genes such as the p16Ink4a/ARF locus and Cyclin A (Jacobs et al. 1999a, b; Martinez and Cavalli 2006; Martinez et al. 2006). For example, Bmi-1 can function as an oncogene through its positive influence on cell proliferation by suppressing p16Ink4a (Jacobs et al. 1999b). On the other hand, genetic evidence also supports certain PcG proteins, such as Mel-18, as tumor suppressors in that they negatively regulate proliferation (Guo et al. 2007). Despite ample elucidation on the intimate relationship between PcG and the cell cycle, we are only at the beginning in realizing a similar cell cycle duty carried out by the trxG members.
MLL, the mammalian ortholog of Drosophila trithorax, is the founding member of the trxG proteins (Gu et al. 1992; Tkachuk et al. 1992; Djabali et al. 1993; Domer et al. 1993; Thirman et al. 1993). Genetic studies confirmed the evolutionarily conserved role of trx/MLL in maintaining homeotic/Hox gene expression, therefore specifying segmental identity (Yu et al. 1995; Ringrose and Paro 2004). Recurrent human chromosome band 11q23 translocations disrupting MLL cause leukemias of poor prognosis. MLL leukemic cells carry pathognomonic MLL fusion proteins resulting from an in-frame fusion between the N-terminal ∼1400 amino acids of MLL with a wide spectrum of translocation partners, ranging from nuclear transcription factors to cytoplasmic structural proteins (Rowley 1998; Ayton and Cleary 2001; Canaani et al. 2004; Daser and Rabbitts 2004; Gilliland et al. 2004; Hess 2004; Eguchi et al. 2005). As implicated by the broad diversity of translocation partners, elegant murine models specifically examined their possible dispensability and nonspecificity (Corral et al. 1996; Dobson et al. 2000; Wang et al. 2005). Remarkably, mice engineered to carry the MLL-lacZ but not the MLL-Myc tag allele developed leukemia, confirming the indispensability and, yet, nonspecificity of these partners (Dobson et al. 2000). Although certain shared properties have been recognized among some fusion partners (Slany et al. 1998; Martin et al. 2003; So et al. 2003), there is no universally applicable mechanism identified.
MLL encodes a 3969-amino-acid nuclear protein consisting of multiple conserved domains with distinct biological properties. The most well-characterized biochemical property of MLL is its ability to methylate histone H3 at Lys 4 through the conserved SET domain at the very C terminus (Milne et al. 2002; Nakamura et al. 2002). The complexity of MLL gene regulation was further illustrated when we and others showed that the 500-kDa full-length MLL precursor (MLLFL) undergoes evolutionarily conserved site-specific proteolysis to generate a mature MLLN320/C180 heterodimer, consisting of processed N-terminal 320-kDa and C-terminal 180-kDa fragments (Nakamura et al. 2002; Yokoyama et al. 2002; Hsieh et al. 2003b). Cleavage of MLL at QL(V)D/GXDD sites is mediated by Taspase1 (threonine aspartase 1) (Hsieh et al. 2003a), which is required for the full activity of MLL (Takeda et al. 2006). In other words, unprocessed MLL precursor functions as a hypomorphic allele (Takeda et al. 2006).
The discovery of Taspase1 initiated a novel class of endopeptidases that utilizes its N-terminal threonine of the mature β subunit to proteolyze substrates after an aspartate (Hsieh et al. 2003a). Taspase1 is translated as a proenzyme that undergoes autoproteolysis to generate a mature α/β heterodimer that displays an overall α/β/β/α structure (Khan et al. 2005). In addition to its anticipated roles in Hox gene regulation, our recent studies on Taspase1−/− mice linked MLL proteolysis to the heart of cell cycle gene programming (Takeda et al. 2006). We showed that MLL directly participates in cell cycle progression through an MLL–E2F axis that controls expression of Cyclins E, A, and B. In the absence of Taspase1, MLL remains as a precursor with reduced histone H3 K4 methyltransferase (HMT) activity (Takeda et al. 2006). Here we further examine how MLL controls the cell cycle and whether its activity is as tightly regulated as most of the key cell cycle regulators.
We provide evidence that MLL undergoes a specialized bimodal degradation resulting in its biphasic expression through the cell cycle. This unique expression is conferred by SCFSkp2 and APCCdc20 E3 ligases of the cell cycle UPS. Importantly, deregulation of this organized expression, either through overexpression or knockdown, incurs corresponding cell cycle defects. Therefore, the observed biphasic appearance of MLL is essential in assuring the correct execution of inherent genetic programs underlying every single cell division. Our model predicts that tight control of MLL activity is essential in cell proliferation, and deregulated expression of MLL impedes normal cell proliferation. Thus, it explains published seemingly contradictory positive or negative regulatory activity of MLL/MLL fusion in cell proliferation (Muyrers-Chen et al. 2004; Milne et al. 2005; Xia et al. 2005; Takeda et al. 2006). Furthermore, MLL degradation signals through its N-terminal ∼1400 amino acids, which are universally retained in MLL leukemia fusions. Importantly, tested prevalent MLL fusions—including MLL-AF4, MLL-AF9, MLL-ENL, and MLL-ELL, and leukemogenic MLL-lacZ—all exhibit diminished interaction with Skp2 and Cdc20, resulting in resistance to degradation mediated by the respective cell cycle UPS and a constant presence through the cell cycle phase transition. Here, we uncover a functional commonality among structurally diversified fusion partners and propose a model in which fusion partners interfere with the recognition of respective MLL fusions by the cell cycle E3 ligases, resulting in their nonoscillating expression. We hypothesize that loss of the biphasic expression of MLL fusions represents the long-sought-after universal defect underlying all MLL leukemias. Complementary to our demonstrated MLL–E2F axis utilized by MLL in activating Cyclin expression, the current study recognizes a critical post-translational regulation of MLL by the cell cycle degradation machinery that is lost in MLL fusions.
Results
Biphasic expression of MLL through the cell cycle
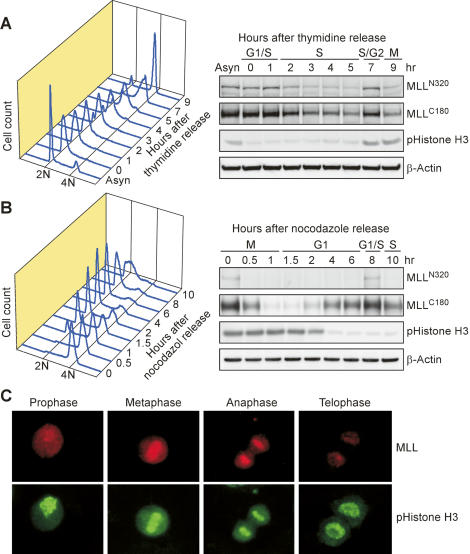
To determine whether the activity of MLL, like other key cell cycle regulators, needs to be temporally regulated, we examined MLL protein expression through progressive phases of the cell cycle. HeLa cells were first synchronized at G1/S and early M phases with double thymidine and nocodazole, respectively, before being subjected to release for the indicated periods of time. Cells at distinct phases of the cell cycle were confirmed by fluorescence-activated cell sorter (FACS) analysis following propidium iodine (PI) staining for DNA contents (Fig. 1A,B, left panels) or by fluorescent microscopy (Fig. 1C). Remarkably, we uncovered a unique biphasic expression pattern of MLL protein when cells progressed through individual phases (Fig. 1A,B, right panels). Both N-terminal (MLLN320) and C-terminal (MLLC180) MLL fragments exhibited a coordinated expression pattern: down-regulation at S and late M phases following respective peaks at G1/S and G2/M transitions (Fig. 1A,B). The degradation of MLL in late M phase was further illustrated by fluorescent microscopy (Fig. 1C). The same biphasic expression pattern was observed in human embryonic kidney 293T cells and human osteosarcoma U2OS cells, irrespective of the synchronization techniques utilized, such as centrifugal elutriation and mimosine block (Supplementary Fig. S1; data not shown). These data unveil a previously unrecognized temporal regulation of MLL activity during the cell cycle progression.
Figure 1.
Biphasic expression of MLL during cell cycle progression. (A) Double thymidine block and release of HeLa cells demonstrated two individual peaks of MLL expression. HeLa cells were synchronized at the G1/S transition by double thymidine block followed by the release into thymidine-free media for the indicated time. (B) Nocodazole block and release of HeLa cells identified the same two peak expressions of MLL as A. (A,B) The levels of MLLN320 and MLLC180 were determined with anti-NT and anti-CT immunoblots, respectively. (A,B, left panels) The successful synchronization and subsequent release from indicated chemical inhibitor was confirmed by FACS analyses of the DNA contents and by immunoblots using the anti-phospho-histone H3 Ser-10 antibody, which marks mitosis. (A,B) The expression of β-actin served as a loading control. (C) MLL protein is decreased at late M phase. Immunofluorescence studies were performed on asynchronous HeLa cells using anti-MLL (red) and anti-phospho-histone H3 S10 (green) antibodies. Individual phase assignments were based on nuclear morphology, with representative photos presented.
MLL is degraded by the UPS during the cell cycle
To investigate the mechanism(s) underlying the biphasic expression of MLL, we examined the MLL mRNA level in cells at different phases of the cell cycle by quantitative RT–PCR (qRT–PCR). The amounts of MLL transcripts did not correlate with its protein levels (Fig. 2A), suggesting a dominating post-transcriptional control. As protein degradation constitutes the major post-translational control of cell cycle regulators, we examined whether cell cycle phase-specific degradation occurs. Immunoprecipitated Flag-MLLN320/C180 was incubated with cellular extracts purified from the indicated phases of the cell cycle in an attempt to recapitulate the in vivo degradation of MLL. Degradation of immunoprecipitated Flag-MLLN320/C180 was then monitored by anti-Flag immunoblots. MLL was degraded by extracts derived from S- and late M-phase cells but not from G1-, G2-, or early M-phase cells (Fig. 2B). These data indicate the utilization of specialized degradation machinery in controlling MLL protein expression. Since a specialized UPS constitutes the major protein degradation machinery of the cell cycle, we tested whether the addition of MG132, a proteasome inhibitor, will increase the protein half-life of MLL in the presence of cyclohexamide, which blocks translation. Indeed, a nearly eightfold extension of its half-life was observed (Fig. 2C). The cell cycle UPS marks proteins for degradation through polymerized ubiquitin conjugation. To determine whether MLL also undergoes UPS-mediated post-translational modifications, HA-Ubiquitin and Flag-MLL were coexpressed in cells followed by anti-Flag immunoprecipitation. Flag-MLL was covalently modified by HA-Ubiquitin and an increased abundance of the ubiquitin-modified high-molecular-weight MLL was observed when MG132 was added (Fig. 2D).
Figure 2.
MLL undergoes bimodal degradation by the UPS. (A) The levels of MLL mRNA do not correlate with the biphasic MLL protein expression through specific phases of the cell cycle. qRT–PCR was performed on HeLa cells that were synchronized by double thymidine treatment and released for the indicated periods of time. MLL expression was normalized against GAPDH. The average MLL expression in asynchronous HeLa cells was arbitrarily assigned as 1.0. Values shown are mean ± 1 SD obtained from three independent experiments. (B) MLL is degraded at specific phases of the cell cycle. On-bead in vitro degradation assays were carried out by incubating purified Flag-MLLN320/C180 with cellular lysates prepared from cells synchronized at the indicated phases of the cell cycle for 30 min at 30°C. The abundance of Flag-MLL was analyzed by anti-Flag immunoblots. (C) The degradation of MLL can be prevented by inhibitors of the 26S proteasome. HeLa cells were treated with 10 μg/mL cycloheximide for the indicated periods of time in the presence of either DMSO as a control or 10 μM MG132 to inactivate 26S proteasome. (Left panel) The expression of MLL was evaluated by anti-MLL immunoblots. The expression of β-actin served as a loading control. (Right panel) The half-life of MLLN320/C180 was determined by quantifying the levels of normalized MLLC180. (D) MLL undergoes polyubiquitination. 293T cells transfected with indicated expression vectors for 48 h were treated with MG132 or DMSO vehicle for an additional 24 h. Cells were lysed in RIPA buffer containing 20 mM N-ethylmaleimide, which inhibits deubiquitination. Cellular extracts were subjected to anti-Flag immunoprecipitation, SDS-PAGE, and immunoblots with the indicated antibodies.
Specialized cell cycle E3 ligases mediate cognate degradation of MLL
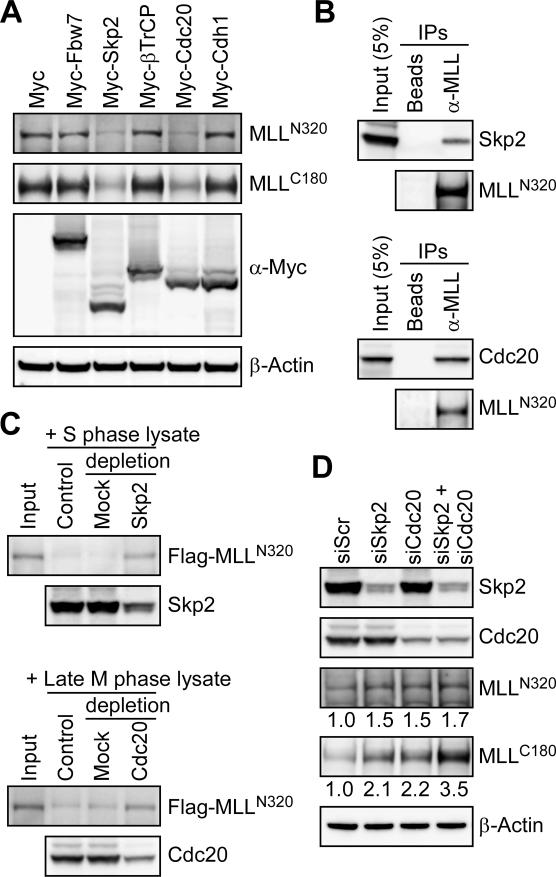
The two major E3 complexes of UPS involved in degradation of core cell cycle regulators are SCF and APC complexes, the substrate recognition of which is conducted by the variable components—F-box proteins for SCF, and Cdc20 and Cdh1 for APC (Nakayama and Nakayama 2006). The bimodal degradation of MLL at S and late M phases suggests the involvement of both SCF and APC for respective degradation. We first examined whether overexpression of any of the major substrate recognition F-box proteins of SCF—including Fbw7, Skp2, and βTrCP—affects the expression of endogenous MLL. Among tested F-box proteins, only Skp2 enhanced the degradation of MLL (Fig. 3A). Similarly, Cdc20 but not Cdh1 degraded MLL in vivo (Fig. 3A). To determine whether Skp2 and Cdc20 directly target MLL for degradation, we probed the direct interactions between these two substrate recognition modules and MLL by coimmunoprecipitation assays in the presence of MG132. Indeed, direct intracellular interactions were identified between endogenous MLL and Skp2 as well as MLL and Cdc20 (Fig. 3B). In addition to the observed in vivo interactions, an in vitro degradation assay was employed to confirm the direct degradation of MLLN320/C180 by SCFSkp2 and APCCdc20. Protein lysates prepared from synchronized S- and late M-phase cells were incubated with equal amounts of immunoprecipitated Flag-MLLN320/C180 for 1 h at 30°C followed by anti-Flag Western blot analysis. Since protein modifications such as phosphorylation are known important signals required for substrate recognition/degradation by the cell cycle UPS, the utilization of immunoprecipitated Flag-MLLN320/C180 as test substrates abrogates the need for in vitro modifications before being subjected to in vitro degradation assays. Both S- and late M-phase lysates efficiently degraded Flag-MLLN320/C180 (Fig. 3C). Importantly, MLL degradation was impaired when SCFSkp2 and APCCdc20 were immunodepleted, confirming the direct degradation of MLLN320/C180 at S and late M phases by SCFSkp2 and APCCdc20, respectively (Fig. 3C). These data demonstratedirect degradation of MLL by SCFSkp2 and APCCdc20, and predict an increase of MLL protein when the expression of Skp2 or Cdc20 is suppressed. To test this hypothesis, small interfering RNA (siRNA)-mediated knockdown of Skp2 and/or Cdc20 was performed to determine whether the MLL protein level is altered. Singular knockdown of Skp2 or Cdc20 led to an increase of the MLL level, and simultaneous knockdown of Skp2 and Cdc20 resulted in a robust ∼3.5-fold increase of MLLC180 and a 1.7-fold increase of MLLN320. The observed lesser increase of MLLN320 is likely due to the known poor gel transfer of MLLN320 onto a membrane. Furthermore, it may also reflect the more unstable nature of the N-terminal MLL (Hsieh et al. 2003b). Taken together, our studies detail the blueprint underlying MLL degradation through the cell cycle, confirming SCFSkp2 and APCCdc20 as two major E3 complexes in orchestrating MLL expression. Demonstrated regulation of MLL by the cell cycle UPS raises the possibility that the temporal necessity of MLL for gene activation is of quintessential importance in executing inherent programs for cell cycle progression.
Figure 3.
The bimodal degradation of MLL is mediated by SCFSkp2 and APCCdc20. (A) Specialized substrate recognition modules (Skp2 and Cdc20) of the cell cycle E3 ligases are required for the degradation of MLL. Specific degradation of the endogenous MLL protein occurred in cells with overexpression of Skp2 or Cdc20. 293T cells were transfected with the indicated Myc-tagged substrate recognition module of cell cycle E3 ligases, and the levels of endogenous MLL were detected with the indicated anti-MLL antibodies. The level of β-actin served as a loading control. (B) MLL interacts with Skp2 and Cdc20 in vivo. Cellular lysates of 293T cells treated with MG132 were subjected to immunoprecipitation (IP) with anti-MLL antibody. Coprecipitated Skp2 and Cdc20 were detected by immunoblots. Protein-A beads served as a negative control. (C) In vitro degradation assays confirm the direct recognition and subsequent destruction of MLL by SCFSkp2 and APCCdc20 in S and late M phases, respectively. Immunoprecipitated Flag-MLL was incubated with the indicated cellular lysates obtained from synchronized HeLa cells at 30°C for 30 min before being subjected to Western blot analyses. Anti-Skp2 or anti-Cdc20 immunoprecipitation was performed to deplete SCFSkp2 or APCCdc20 from the indicated lysates. Mouse IgG was used as mock depletion. (D) Knockdown of Skp2 and/or Cdc20 leads to the accumulation of MLL protein. HeLa cells were transfected with the indicated siRNA specific for Skp2 or Cdc20. Protein levels of endogenous Skp2, Cdc20, and MLL were analyzed by Western blots at 72 h after transfection and quantified using ImageGauge software (FujiFilm). The level of β-actin served as a loading control, and the expression of MLL in control knockdown (siScr, si scramble) was arbitrarily assigned as 1.0.
Concerted expression of MLL is necessary for proper cell cycle progression
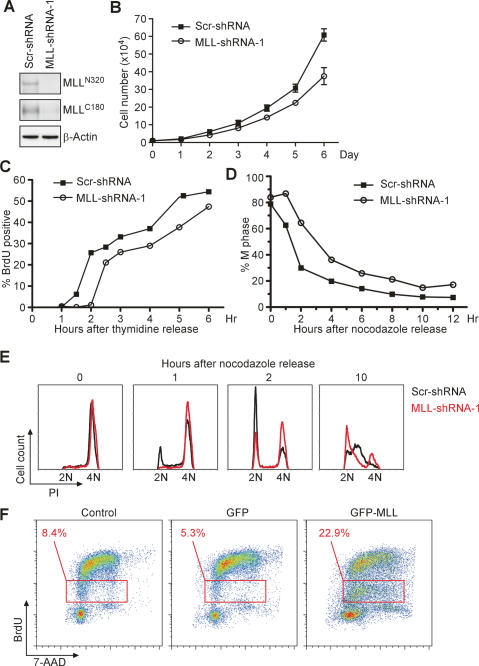
In addition to our prior report, which indicates direct participation of MLL in the cell cycle gene expression program through the MLL–E2F axis (Takeda et al. 2006), our current data highlight a programmed regulation of the MLL level conferred by the cell cycle UPS. The next important questions are why the activity of MLL needs to be temporally controlled and whether deregulation of such an exquisite expression causes any cell cycle aberrations. To tackle these questions, RNA interference (RNAi)-mediated knockdown and overexpression of MLL were performed to interrogate possible cell cycle defects. Retrovirus-mediated stable knockdown of MLL in HeLa cells was performed, and the consequences of failed MLL induction at G1/S and G2/M transitions were examined. Two independent MLL-short hairpin RNA (shRNA) constructs stably reduced the expression of MLL as compared with the scramble shRNA (scr-shRNA) control (Fig. 4A; Supplementary Fig. S2). MLL knockdown cells were impaired in proliferation—a nearly twofold reduction of total cell number was observed after 6 d in culture (Fig. 4B; Supplementary Fig. S2). To specifically pinpoint the defect at G1/S transition, cells were synchronized by double thymidine treatment before being released into standard culture medium for the indicated periods of time. The S-phase entrance rate was evaluated with a 30-min pulse of BrdU, which specifically labels active DNA synthesis. Strikingly, a 1-h delay in S-phase entry was observed in MLL-deficient cells (Fig. 4C), indicating that the G1/S peak of MLL is required for proper G1/S transition. We then examined the defect in M phase following the loss of the second MLL peak. MLL-deficient HeLa cells were synchronized with nocodazole in prometaphase before being released into standard culture medium for the indicated periods of time. Cellular DNA contents were stained with PI and analyzed by FACS. In MLL knockdown cells, the impaired G2/M peak expression resulted in a marked delay in M-phase progression (Fig. 4D,E). Thus, the up-regulation of MLL in G1/S and G2/M transitions is apparently required for S-phase entry and M-phase progression, respectively.
Figure 4.
Deregulation of MLL protein level incurs corresponding cell cycle progression defects. (A) Stable knockdown of MLL in HeLa cells was achieved with shRNA against MLL. HeLa cells were infected with retrovirus-expressing shRNA targeting MLL (MLL-shRNA) or control scramble shRNA (Scr-shRNA). The successful suppression of MLL expression was confirmed by anti-MLL Western blots. (B) Depletion of MLL impairs cell proliferation. The proliferation curves of HeLa cells with or without MLL knockdown are provided. Replicate cultures of 10,000 cells were plated and trypsinized at the indicated time points to obtain cell numbers. Values shown are mean ± 1 SD obtained from three independent experiments. (C) A marked impairment in S-phase entry is observed in MLL knockdown cells. HeLa cells with or without MLL knockdown were synchronized at the G1/S transition by double thymidine block, followed by a release for the indicated time points. Cells were chased with BrdU for 30 min before collections, and S-phase entry was assessed by the positive BrdU staining determined by FACS. (D,E) MLL is required for proper M-phase progression. HeLa cells with MLL or control knockdown were synchronized at the prometaphase by nocodazole block plus shake-off procedure before being released into regular culture medium. Cells were stained with PI for DNA content before being subjected to FACS analyses to monitor phase progression of the cell cycle. Quantification of the M-phase progression from serial graphs presented in E is presented in D. (F) Overexpression of MLL induces an S-phase block. 293T cells were transfected with GFP-MLL or GFP-expressing plasmids for 48 h before being subjected to a 30-min BrdU pulse immediately before analyses. GFP-positive cells were gated for analyses of BrdU incorporation and DNA content. Additional controls using untransfected cells were included. (C–F) Results presented are representative data of three independent experiments.
We next investigated why MLL needs to be degraded. In other words: What are the consequences if MLL continues to express at high levels? To answer this question, GFP-MLL was transiently overexpressed in 293T cells, the cell cycle profile of which was monitored by costaining with 7-AAD for DNA contents and BrdU for active DNA synthesis. An increased appearance (22.9%) of cells with an intermediate incorporation of BrdU was identified in GFP-MLL-expressing cells, suggestive of deterred replication (Fig. 4F). Taken together, these data emphasize the importance of the biphasic expression of MLL in coordinating the cell cycle progression.
The MLL N-terminal ∼1400 amino acids target MLLN320/C180 for degradation
To further characterize how MLL degradation is regulated, the region responsible for the observed degradation was mapped. Flag-tagged constructs representing different regions of MLL were cotransfected with either Skp2 or Cdc20, followed by examinations of their expressions by anti-Flag immunoblots (Fig. 5A). The first ∼1400 amino acids of MLL were found to recapitulate the degradation of MLLN320/C180 (Fig. 5A). In contrast, MLLC180 alone was insensitive to degradation by coexpressed Skp2 or Cdc20 (Fig. 5A). As endogenous MLLC180 derived from proteolytic processing of MLLFL precursor heterodimerizes with MLLN320 (Hsieh et al. 2003b) and is susceptible to degradation, it is possible that MLLC180 is degraded through its interaction with MLLN320; i.e., MLLN320 brings heterodimerized MLLC180 to the 26S proteasome for degradation. To test this hypothesis, MLLN320 (amino acids 1–2664) and MLLC180 (amino acids 2772–3969) were cotransfected with or without specific E3 substrate recognition modules, Skp2 and Cdc20. In accordance with our data on the degradation of endogenous MLLN320/C180 (Fig. 3A), MLLC180 was degraded along with the cotransfected MLLN320 in the presence of either Skp2 or Cdc20 (Fig. 5B). If the MLL C terminus was degraded while complexed with MLLN320, a MLL C-terminal mutant that no longer interacts with MLLN320 would be resistant to such a degradation. We employed a mutant MLL C terminus (MLLC180ΔF/SET) that lacks FYRC and SET domains—a region that mediates its heterodimerization with MLLN320 (Hsieh et al. 2003b). When Flag-MLLC180ΔF/SET (amino acids 2722– 3620) was coexpressed with Flag-MLLN320 (amino acids 1–2664) plus either Myc-Skp2 or Myc-Cdc20, its level remained unchanged (Fig. 5B). This was in stark contrast to the nearly complete disappearance of Flag-MLLC180 (Fig. 5B). Thus far, our data establish a model in which the very N-terminal ∼1400 amino acids of MLL are responsible for the coordinated biphasic control of MLLN320/C180. Since all of the MLL leukemia fusions retain the degradation-prone region of MLL (Rowley 1998; Ayton and Cleary 2001; Canaani et al. 2004; Daser and Rabbitts 2004; Gilliland et al. 2004; Hess 2004), it is possible that the fusion partners confer resistance to degradation to initiate the first universal insult underlying all MLL leukemias.
Figure 5.
The MLL N terminus is responsible for the degradation of MLLN320/C180. (A) The N-terminal 1400 amino acids of MLL are targeted by SCFSkp2 and APCCdc20 for degradation. Various Flag-MLL fragments were coexpressed in 293T cells with either Myc-Skp2 or Myc-Cdc20. The protein levels of various MLL fragments were determined by anti-Flag Western blots. The diagram on the right depicts the domain compositions of transfected MLL fragments. Flag-MLL(1–3969) served as a positive control. (B) MLLN320 brings MLLC180 to degradation. Flag-MLLN320 plus Flag-MLLC180 or Flag-MLLN320 plus Flag-MLLC180ΔF/SET were coexpressed in 293T cells with either Myc-Skp2 or Myc-Cdc20 before being subjected to indicated Western blot analyses.
MLL fusions are refractory to the cell cycle UPS, resulting in stable expression through the cell cycle
The findings that the common MLL N-terminal ∼1400 amino acids are responsible for the degradation of MLLN320/C180 are consistent with our prior observation that MLLN320 is less stable (Hsieh et al. 2003b). These data rekindle a prior proposal that fusion partners may contribute to leukemogenesis in part by stabilizing the N-terminal MLL (Dobson et al. 2000; Hsieh et al. 2003b). We next examined whether leukemogenic MLL fusions are refractory to degradation. Indeed, tested prevalent MLL fusions, including MLL-AF4, MLL-AF9, MLL-ENL and MLL-ELL, were all resistant to SCFSkp2- and APCCdc20-mediated degradation (Fig. 6A). Remarkably, the same resistance to degradation was also observed with the leukemogenic MLL-lacZ but not the nonleukemogenic MLL-Myc tag fusion (Fig. 6A). Subsequent experiments were performed to determine whether endogenous MLL fusions undergo cell cycle-coordinated degradation in various MLL leukemia cell lines. Interestingly, unlike commonly utilized tissue culture cell lines such as HeLa, 293T, and U2OS cells, we were unable to use mimosine, double thymidine, or nocodazole to synchronize available MLL leukemia cells, including MLL-AF4-bearing SEM and RS(4;11) and MLL-AF9-bearing THP-1, NOMO-1, and MOLM13 cell lines. This surprising observation suggests inherent cell cycle defects of these MLL leukemia cells. One possible scenario is that these cells have cycle checkpoint defects that confer resistance to chemical-induced synchronization. Nevertheless, individual populations representing distinct phases of the cell cycle were separated using centrifugal elutriation based on their physical properties. The purity of each population was assessed by FACS analysis of DNA contents (Fig. 6B). Remarkably, we witnessed a loss of the unique biphasic expression of MLL-AF4 in SEM human MLL leukemia cells (Fig. 6B). In contrast to the biphasic expression of wild-type MLL, MLL-AF4 protein did not fluctuate through the cell cycle while its level was not markedly increased (Fig. 6B). As overexpression of MLL fusion is likely to block cell cycle progression and induce apoptosis (Muyrers-Chen et al. 2004; Xia et al. 2005), additional compensatory mechanism(s) must have evolved to maintain the level of MLL fusions within a tolerable range. Since all of the MLL fusions contain the common N-terminal ∼1400 amino acids that signal degradation, it is possible that individual fusion partners directly interfere with the recognition of the MLL N terminus by the cell cycle E3 ligases. Coimmunoprecipitation assays were performed in cells transfected with indicated Flag-MLL fusions to examine this hypothesis. Indeed, diminished interactions between tested MLL fusions and endogenous Skp2 or Cdc20 were observed (Fig. 6C). To further demonstrate the refractoriness of MLL fusions to the cell cycle UPS, in vitro degradation assays were performed on immunoprecipitated MLL-AF4. In contrast to the nearly complete degradation of MLL by S- or late M-phase extracts, no significant degradation of MLL-AF4 was observed (Fig. 6D). In conclusion, we discovered a common defect associated with all of the tested prevalent MLL fusions that were not previously appreciated. The resistance of MLL fusions to preprogrammed degradation likely constitutes the first universal insult committed by all MLL fusions, leading to the ultimate development of frank MLL leukemias.
Figure 6.
MLL fusions are resistant to SCFSkp2- or APCCdc20-mediated degradation. (A) Prevalent MLL fusion proteins, including MLL-AF4, MLL-AF9, MLL-ENL, and MLL-ELL, and leukemogenic MLL-lacZ, are resistant to Skp2- or Cdc20-mediated degradation. Various Flag-tagged MLL fusions were coexpressed in 293T cells with Myc-Skp2 or Myc-Cdc20. The levels of MLL fusions were analyzed by anti-Flag immunoblots. (B) Human MLL leukemia cell line SEM t(4;11) exhibits constant expression of endogenous MLL-AF4 protein through the cell cycle progression. (Bottom panel) To examine the expression of MLL-AF4, we synchronized the cells with centrifugal elutriation and confirmed individual phases of the cell cycle with PI staining of the DNA content followed by FACS analyses. The same amounts of lysates purified from indicated cellular fractions were subjected to immunoprecipitation with anti-AF4 antibody, which recognizes the C terminus of AF4. Immunoprecipitates were resolved by SDS-PAGE and analyzed with an anti-MLL antibody (Bethyl Laboratories) that recognizes amino acids 720–780 of MLL. (C) Fusion partners interfere with the recognition of MLL N terminus by Skp2 and Cdc20. Flag-MLL fusions and Flag-MLL(1–1400) were transfected in 293T cells for 2 d, followed by treatment with MG132 for 4 h. Cellular extracts were subjected to anti-Flag immunoprecipitation assays. Immunoprecipitates were analyzed by Western blots using anti-Flag, anti-Skp2, and anti-Cdc20 antibodies. The abundance of coprecipitated endogenous Skp2 and Cdc20 was determined with anti-Skp2 and anti-Cdc20 Western blots, respectively. Immunoprecipitation with extracts prepared from untransfected cells or Flag-MLL(1–1400) transfected cells served as negative and positive controls, respectively. (D) MLL-AF4 is resistant to degradation by S- or late M-phase lysates. Flag-MLL-AF4 was transiently expressed in 293T cells and purified with anti-Flag antibody. Immunoprecipitated Flag-MLL-AF4 was aliquoted and subjected to in vitro degradation experiments using the indicated cellular lysates. The abundance of Flag-MLL-AF4 was then determined by anti-Flag Western blots.
Discussion
MLL and the cell cycle machinery
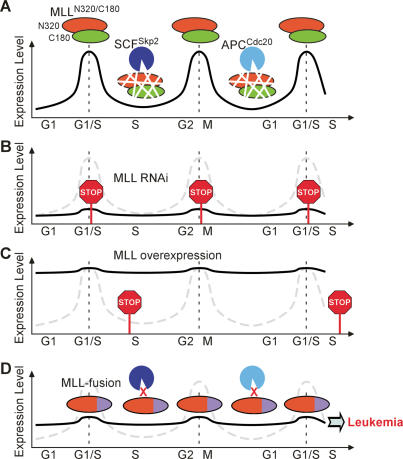
MLL, the mammalian homolog of Drosophila trithorax, is best known for its positive regulation of Hox gene expression. Homozygous disruption of MLL in mice results in early embryonic lethality, and heterozygous deficiency results in homeotic transformation due to impaired maintenance of Hox genes (Yu et al. 1995). The early lethality of MLL−/− mouse embryos precludes detailed investigation of its involvement in other signaling pathways. Our demonstration that MLL is regulated by Taspase1-mediated proteolytic cleavage broadens the avenue in studying MLL in that noncleaved precursor MLL functions as a hypomorphic allele. Although the underlying mechanisms were not further investigated, initial characterizations on MLL−/− mice suggest a role of MLL in proliferation in addition to differentiation. For example, MLL−/− fetal liver or yolk sac hematopoietic cells grow more slowly and form smaller colonies in methyl cellulose assays (Hess et al. 1997). Our subsequent studies on cells deficient for Taspase1 or bearing noncleavable alleles of MLL (MLLNC/NC) recognized a participation of MLL through E2Fs in regulating cell proliferation (Takeda et al. 2006). The active involvement of MLL/trx in the cell cycle is conserved through evolution, in that genetic studies in Drosophila also highlighted a significant role of trx in cell proliferation (Muyrers-Chen et al. 2004). Recent studies, including ours, began to elucidate the downstream targets for MLL in the cell cycle regulation that include Cyclins and CDKIs (Muyrers-Chen et al. 2004; Milne et al. 2005; Xia et al. 2005). We demonstrated a MLL–E2F axis in regulating Cyclin E/A/B expression for progressive cell cycle phase transition (Takeda et al. 2006). Here, we further investigated the participation of MLL in the cell cycle control and discovered a tightly controlled biphasic expression of MLL. This unique expression is conferred by defined windows of degradation mediated by specialized cell cycle E3 ligases: SCFSkp2 and APCCdc20 (Fig. 7A). Importantly, individual peak expressions of MLL precede the induction of Cyclin E/A and Cyclin B to ensure proper G1/S transition and M-phase progression, respectively (Fig. 7A). Deregulation of this unique expression of MLL by shRNA-mediated knockdown causes corresponding defects in G1/S entry and M-phase progression (Fig. 7B). Furthermore, overexpression of MLL incurs specific S-phase defects, indicating the importance of down-regulating its activity in S phase (Fig. 7C). However, whether this resulted from sustained expression of Cyclin E/A and/or invoked not-yet-identified insults remains to be determined. Our data highlight the significance of this biphasic expression of MLL in regulating cell proliferation and uncover a novel mechanism in regulating MLL through protein degradation—another post-translational regulatory scheme in addition to Taspase1-mediated site-specific proteolysis. As MLL directly activates the transcription of Cyclins that exhibit periodic expression during cell proliferation, it is necessary for a cell to incorporate MLL expression into the intricately assembled cell cycle circuitry to ensure correct transition of progressive phases. Current data not only consolidate the role of MLL in activating cell cycle but also discover a built-in program elegantly designed to turn off MLL with an impeccable temporal sequence.
Figure 7.
Models depict a critical biphasic expression of MLL through cell cycle progression, the consequences of its deregulation, and the resistance of MLL fusions to degradation. (A) MLL protein oscillates throughout the cell cycle with two distinct peaks at the G1/S and G2/M transitions. This unique expression was conferred by a bimodal degradation executed by SCFSkp2 and APCCdc20 at S and late M phases, respectively. (B) Cells with constitutively low expression of MLL have defects in S-phase entry and M-phase progression. (C) Constitutively high expression of MLL incurs an S-phase block. (D) In human MLL leukemia cells, MLL fusions remain constant through the cell cycle due to their impaired interactions with Skp2 and Cdc20. This deregulated expression may represent the first initial biological insult contributing to the ultimate development of MLL leukemias.
MLL leukemia
The observed undesirable cell cycle consequences from perturbations of MLL levels led us to postulate that MLL fusions may also interfere with the cell cycle—a mechanism that is not fully appreciated in MLL leukemogenesis. Human chromosome 11q23 aberrations disrupt MLL, leading to infant and chemotherapy-related leukemias. These balanced translocations fuse the N-terminal ∼1400 amino acids of MLL in-frame with a wide spectrum of fusion partners to generate leukemogenic MLL fusions. Mysteriously, among the >60 diverse fusion partners identified so far, there are no commonly shared characteristics identified based on sequence or structure homology (Rowley 1998; Ayton and Cleary 2001; Canaani et al. 2004; Daser and Rabbitts 2004; Gilliland et al. 2004; Hess 2004). Genetic evidence provided by studying mice carrying individual MLL fusions reveals several fundamental aspects of MLL leukemias. First, the fusion partner is indispensable (Corral et al. 1996; Dobson et al. 2000). Second, the fusion partner can be as nonspecific as bacterial galactosidase (lac Z) in that mice bearing MLL-lacZ developed myeloid leukemia after a prolonged latency (Dobson et al. 2000). Third, the fusion partner determines the phenotypes of the resulting leukemia (Corral et al. 1996; Wang et al. 2005; Chen et al. 2006). For example, mice carrying MLL-AF4 or MLL-AF9 develop lymphoid versus myeloid malignancies (Corral et al. 1996; Chen et al. 2006; Metzler et al. 2006), mimicking human counterparts. Detailed analyses of individual MLL fusions using retrovirus-mediated gene transduction of the hematopoietic stem cells provide insightful mechanistic explanations regarding MLL leukemias. These studies established important models, such as transactivation and dimerization (Slany et al. 1998; Martin et al. 2003; So et al. 2003). However, these two models can only explain subsets of MLL leukemias since not all fusion partners contain transactivation or dimerization domains. As mice bearing MLL-lacZ developed leukemia, it had long been postulated that lacZ induces leukemia through either oligomerizing or stabilizing the N terminus of MLL—two non-mutually exclusive mechanisms. Although stabilization may contribute to MLL leukemogenesis, it has not been further examined. Our initial studies on the site-specific proteolysis of MLL indicated that proteolytic cleavage of MLL is not only required for its full activation but also regulates its stability (Hsieh et al. 2003a, b; Takeda et al. 2006). Interestingly, Taspase1-mediated site-specific proteolysis also controls the protein levels of another recently identified Taspase1 substrate, TFIIA (transcription factor II A) (Hoiby et al. 2004; Zhou et al. 2006). As our prior studies implicated that MLL N terminus is unstable, we envisioned that fusion partners must stabilize the common MLL N terminus for downstream gene regulation (Hsieh et al. 2003b). Based on our current observations that levels of MLL need to be tightly monitored during cell proliferation and the degradation of MLLN320/C180 signals through its N-terminal ∼1400 amino acids (the common denominator present in all MLL fusions), we hypothesize that deregulated expression of MLL fusions through cell division may constitute the long-awaited universal insult underlying all MLL leukemias. To interrogate this model, we examined whether overexpression of SCFSkp2 and APCCdc20 affects the levels of leukemogenic MLL fusions, including MLL-AF4, MLL-AF9, MLL-ENL, MLL-ELL, and MLL-lacZ. Remarkably, our in vivo and in vitro degradation assays demonstrated that these MLL fusions, unlike MLLN320/C180, have acquired resistance to specialized cell cycle UPS. Furthermore, when the human t(4;11) MLL leukemia cell line, SEM, was examined, we witnessed a constant expression of MLL-AF4 throughout progressive phases of the cell cycle. Although the consequences of stable expression of MLL fusions through the cell cycle progression remain to be determined, several studies focused on individual MLL fusions such as MLL-AF4 and MLL-AF9 did report associated cell cycle defects (Pession et al. 2003; Caslini et al. 2004; Xia et al. 2005). Most interestingly, we noticed that all of the five tested human MLL leukemia cell lines failed to be synchronized by drugs like mimosine, thymidine, and nocodazole, implicating global defects in cell cycle checkpoints. As shown in several knock-in murine MLL leukemia models—including MLL-AF4 (Chen et al. 2006; Metzler et al. 2006), MLL-AF9 (Corral et al. 1996), MLL-lacZ (Dobson et al. 2000), MLL-ELL(Luo et al. 2002), and MLL-CBP (Wang et al. 2005)—engineered mice only developed leukemia after a long latency or the challenge with carcinogens such as ENU (Luo et al. 2002; Wang et al. 2005), suggesting that MLL fusion alone is necessary but insufficient in MLL leukemogenesis. Therefore, additional mutations are likely to contribute to the full-blown MLL leukemia phenotypes (Felix et al. 1998; Mahgoub et al. 1998; Armstrong et al. 2003; Oguchi et al. 2003). The loss of the biphasic expression of MLL fusions may initiate the first universal mechanistic insult toward the ultimate development of MLL leukemias by disrupting cell cycle checkpoints (Fig. 7D). Future studies focusing on dissecting the mechanisms by which MLL fusions compromise cell cycle checkpoints and evaluating the contribution of such defects in MLL leukemogenesis will certainly provide therapeutic vantage points in treating this deadly illness.
The intertwined relationship between PcG and trxG members in cell fate and cell cycle
Deregulation of essential developmental pathways commonly leads to dire outcomes, including cancer; the PcG and trxG proteins are such examples (van Lohuizen 1999). Initial genetic evidence obtained from flies along with subsequent studies in mammals established the fundamental roles of PcG and trxG proteins in maintaining Hox gene expression, thus ensuring the correct installment of complex body plans in higher organisms. In addition to their essential roles in embryonic development, deregulated PcG and trxG proteins also contribute to various oncogenic processes. For example, Bmi-1, a PcG protein, is well recognized for its ability to accelerate Myc-induced lymphoma in mice (van Lohuizen et al. 1991). Subsequent studies on Bmi-1-associated oncogenesis primarily emphasized the ability of Bmi-1 to suppress the expression of tumor suppressors p16ink4a and ARF, thus promoting caner cell proliferation (Jacobs et al. 1999b). Despite the well-established role in Hox gene regulation, the significance of Bmi-1-induced Hox gene alterations in Bmi-1-mediated oncogenesis remains undetermined. On the contrary, studies on MLL, the founder of trxG proteins, and its associated leukemias recognized the potential mechanistic importance of overexpression of Hox genes including HoxA7, HoxA9, and HoxA11 in leukemogenesis (Armstrong et al. 2002; Ayton and Cleary 2003). However, whether aberrant cell cycle gene expression contributes to MLL leukemogenesis has not been largely questioned. Based on our findings that link MLL/MLL fusions to cell cycle regulation, we propose that a simultaneous deregulation of differentiation and proliferation through Hox and cell cycle genes by Bmi-1 and MLL is of paramount importance in respective carcinogenesis.
Materials and methods
Plasmid constructions, siRNA, and transfections
cDNAs encoding full-length MLL, various MLL fragments, and MLL fusions were fused in-frame after an N-terminal Flag tag (pCMV-3xFlag; Sigma) to generate individual eukaryotic expression constructs for transient transfection assays. Full-length Fbw7, Skp2, βTrCP1, Cdc20, and Cdh1 were cloned from a 293T cDNA library and inserted into an N-terminal Myc tag expression vector (CMV-Myc; Clontech). The pMT123 plasmid encoding HA-Ubiquitin was kindly provided by Dr. Dirk Bohmann (Treier et al. 1994). Lipofectamine and Oligofectamine (Invitrogen) were used to transfect plasmid and siRNA oligos (Dharmacon), respectively, according to the manufactures’ protocols. siRNA oligos against Skp2 (M-003324-03) and Cdc20 (M-003225-03) were purchased from Dharmacon.
Antibodies and Western blots
Anti-CT (C terminus) antibody that recognizes the transactivation domain of MLLC180 has been described previously (Hsieh et al. 2003b). A rabbit polyclonal anti-NT (N terminus) antibody (MO-353) was raised against amino acids 1600–1985 of MLL. Western blots using commercially available antibodies against Skp2 (Santa Cruz Biotechnology), Cdc20 (Santa Cruz Biotechnology), Myc (Santa Cruz Biotechnolgoy), phospho-Histone H3 (Ser-10) (Upstate Biotechnology), MLL1 (recognizing amino acids 720–780 of MLL; Bethyl Laboratories), β-Actin (Sigma), Flag (Sigma), and HA (12CA5) were performed according to the manufacturers’ recommendations. Antibodies were detected using the enhanced chemiluminescence method (Western Lightning, PerkinElmer). Western blot signals were acquired with the LAS-3000 Imaging system (FujiFilm) and were analyzed by ImageGauge software (FujiFilm) as described previously (Kim et al. 2006).
Immunofluorescence
HeLa cells were plated on LabTek II chamber slides (Nunc) overnight, washed with PBS twice, fixed with 4% paraformaldehyde for 15 min, permeabilized with 0.2% Triton X-100 for 10 min, and blocked with 3% BSA in PBS for 1 h. These cells were subsequently incubated with the indicated primary antibodies for 3 h, followed by two PBS washes, and were then incubated with Alexa488-conjugated anti-mouse or Alexa568-conjugated anti-rabbit antibodies (Molecular Probes) for an additional 30 min. The nuclei were counterstained with Hoechst 33342 (Molecular Probes). Fluorescent images were captured by a CCD camera (Diagnostic Instruments) attached to an Olympus IX51 microscope and were analyzed with the SPOT advanced software.
Cell culture and synchronization
HeLa and 293T cells were obtained from American Type Culture Collection and grown in DMEM (Invitrogen) supplemented with 10% fetal bovine serum, nonessential amino acids, L-glutamine, and penicillin/streptomycin. The human leukemia cell line SEM, carrying the t(4;11)(q21;q23) chromosomal translocation, was maintained in RPMI 1640 medium (Invitrogen) containing the same supplements. For double thymidine block and release experiments, HeLa cells were treated with 2 mM thymidine for 12 h and released for 12 h, followed by the second 12-h treatment before being released into thymidine-free medium for the indicated periods of time. As to the nocodazole block and release experiments, HeLa cells were blocked with 200 ng/mL nocodazole for 18 h. Mitotic cells were enriched by shake-off and washed twice with PBS before being released into nocodazole-free medium for the indicated time. Centrifugal elutriation was carried out using a Beckman JE6B elutriation rotor (Beckman). Small aliquots of cells collected at indicated time points or fractions were subjected to FACS analysis of the DNA contents for cell cycle phase assignment before subsequent analyses.
qRT–PCR
Cells were harvested and lysed with Trizol (Invitrogen) for RNA purification using RNeasy (Qiagen). Reverse transcriptions were performed with oligo-dT plus random octamer primers (Ambion) using SuperScript II (Invitrogen). qPCR was performed with Power SYBR Green master mix (Applied Biosystems) in triplicate using the following MLL primer set: MLL-forward, ACACATTCCAGACCAAGAAACGAC, and MLL-reverse, AG GCATCTTCAATACTTTCTGCACAG. Data were acquired using an ABI PRISM 7000 system (Applied Biosystems) and analyzed as described previously (Takeda et al. 2006). GAPDH expression served as a control.
Cell cycle analyses and BrdU incorporation assays
Cells were trypsinized, washed with PBS, treated with 20 μg/mL RNase A, and stained with 25 μg/mL PI for 1 h before being subjected to cell cycle analyses. Flow-cytometric analyses were performed using a FACSCalibur flow cytometer (Becton-Dickinson) to measure DNA contents. Ten-thousand events were collected per each experiment, and the data were analyzed with FlowJo software (Tree Star) or FCS Express (De Novo System). Cells were pulsed with BrdU for 30 min and analyzed by FACSCalibur according to the manufacturer’s manual (BD Pharmingen; FITC/APC BrdU Flow Kit).
shRNA-mediated knockdown of MLL in HeLa cells
Target sequences (GTGCCAAGCACTGTCGAAA [Fig. 4] and (CTACCAACCCTAAACCCTG) [Supplementary Fig. S2]), against human MLL and a control scrambled sequence (GCGC GCTTTGTAGGATTCG) that has no significant homology with the human genome were inserted into the pSUPER. retro.puro vector, according to the manufacturer’s protocol (Oligoengine). Generated retrovirus carrying indicated shRNA was used to infect HeLa cells for 2 d before being subjected to puromycin selection at 2 μg/mL.
Protein extracts and in vitro degradation assays
To obtain G1-phase lysates, HeLa cells were synchronized by mimosine for 16 h. For S-phase lysates, HeLa cells were synchronized by double thymidine block, followed by a 4-h release. To synchronize cells at G2 phase, HeLa cells were released from a double thymidine block for 5 h, followed by a 5-h treatment with 0.5 μM etoposide and extensive washes to remove mitotic cells. As to early M-phase lysates, HeLa cells were synchronized at prometaphase by nocodazole treatment for 18 h. Cells released from the nocodazole block for 1 h were at late M phase. Synchronized cells were resuspended in buffer containing 20 mM Tris-HCl (pH 7.2), 2 mM dithiothreitol (DTT), 0.25 mM EDTA, and Complete Protease Inhibitor Cocktail (Roche) on ice before being transferred to a nitrogen disruption bomb (Parr). The pressure was subsequently brought up to 1000 psi, followed by incubation for 30 min on ice. The pressure was then slowly released to gently disrupt the cells. The homogenized materials were spun down at 10,000g for 10 min. The resulting cellular extracts were aliquoted and stored at −80°C. For immunodepletion experiments, 10 μg of anti-Skp2 or anti-Cdc20 antibodies were first adsorbed to 15 μL of protein-A beads (GE Healthcare) by rocking for 90 min at 4°C. The antibody-bound beads were washed and incubated with 40 μL of HeLa extracts (∼400 μg of protein) for 2 h at 4°C. Bead-bound immunocomplexes were subsequently removed by centrifugation. Immunodepleted extracts were aliquoted and used for the indicated in vitro degradation assays. 293T cells were transfected with the indicated Flag-MLL-expressing constructs for 2 d before being subjected to lysis with RIPA buffer in the presence of Complete Protease Inhibitor Cocktail (Roche). Expressed Flag-MLL was immunoprecipitated using anti-Flag M2 beads (Sigma). Bead-bound Flag-MLL was aliquoted for the indicated experiments. The degradation reaction (10 μL) contained 40 mM Tris-HCl (pH 7.6), 5 mM MgCl2, 1 mM DTT, 10% glycerol, 1 μM ubiquitin aldehyde, 10 mM phosphocreatine, 100 μg/mL creatine phosphokinase, 0.5 mM ATP, 10 μg of the indicated HeLa cellular extracts, 3 μL of Flag-MLL immunoprecipitates, and Complete Protease Inhibitor Cocktail (Roche). Following incubation for 1 h at 30°C, reactions were stopped by the addition of SDS sample buffer before being subjected to SDS-PAGE analyses.
Acknowledgments
We thank Drs. Scott Armstrong and Jorg Faber for providing human MLL leukemia cell lines. MLL-ENL and MLL-ELL expression constructs were kindly provided by Dr. Cleary and Dr. Mitani, respectively. H.L. is supported by NIH CA119008 to J.J.-D.H. E.H.-Y.C. is supported by the Searle Scholars Program, Mallinckrodt Jr. Foundation, NCI Howard Temin Award, and NIH CA125562. This work is supported by Mallinckrodt Jr. Foundation, ASH Scholar Award, NCI Howard Temin Award, and NIH CA119008 to J.J.-D.H.
Footnotes
Supplemental material is available at http://www.genesdev.org.
Article is online at http://www.genesdev.org/cgi/doi/10.1101/gad.1574507
References
- Armstrong S.A., Staunton J.E., Silverman L.B., Pieters R., den Boer M.L., Minden M.D., Sallan S.E., Lander E.S., Golub T.R., Korsmeyer S.J., Staunton J.E., Silverman L.B., Pieters R., den Boer M.L., Minden M.D., Sallan S.E., Lander E.S., Golub T.R., Korsmeyer S.J., Silverman L.B., Pieters R., den Boer M.L., Minden M.D., Sallan S.E., Lander E.S., Golub T.R., Korsmeyer S.J., Pieters R., den Boer M.L., Minden M.D., Sallan S.E., Lander E.S., Golub T.R., Korsmeyer S.J., den Boer M.L., Minden M.D., Sallan S.E., Lander E.S., Golub T.R., Korsmeyer S.J., Minden M.D., Sallan S.E., Lander E.S., Golub T.R., Korsmeyer S.J., Sallan S.E., Lander E.S., Golub T.R., Korsmeyer S.J., Lander E.S., Golub T.R., Korsmeyer S.J., Golub T.R., Korsmeyer S.J., Korsmeyer S.J. MLL translocations specify a distinct gene expression profile that distinguishes a unique leukemia. Nat. Genet. 2002;30:41–47. doi: 10.1038/ng765. [DOI] [PubMed] [Google Scholar]
- Armstrong S.A., Kung A.L., Mabon M.E., Silverman L.B., Stam R.W., Den Boer M.L., Pieters R., Kersey J.H., Sallan S.E., Fletcher J.A., Kung A.L., Mabon M.E., Silverman L.B., Stam R.W., Den Boer M.L., Pieters R., Kersey J.H., Sallan S.E., Fletcher J.A., Mabon M.E., Silverman L.B., Stam R.W., Den Boer M.L., Pieters R., Kersey J.H., Sallan S.E., Fletcher J.A., Silverman L.B., Stam R.W., Den Boer M.L., Pieters R., Kersey J.H., Sallan S.E., Fletcher J.A., Stam R.W., Den Boer M.L., Pieters R., Kersey J.H., Sallan S.E., Fletcher J.A., Den Boer M.L., Pieters R., Kersey J.H., Sallan S.E., Fletcher J.A., Pieters R., Kersey J.H., Sallan S.E., Fletcher J.A., Kersey J.H., Sallan S.E., Fletcher J.A., Sallan S.E., Fletcher J.A., Fletcher J.A., et al. Inhibition of FLT3 in MLL. Validation of a therapeutic target identified by gene expression based classification. Cancer Cell. 2003;3:173–183. doi: 10.1016/s1535-6108(03)00003-5. [DOI] [PubMed] [Google Scholar]
- Ayton P.M., Cleary M.L., Cleary M.L. Molecular mechanisms of leukemogenesis mediated by MLL fusion proteins. Oncogene. 2001;20:5695–5707. doi: 10.1038/sj.onc.1204639. [DOI] [PubMed] [Google Scholar]
- Ayton P.M., Cleary M.L., Cleary M.L. Transformation of myeloid progenitors by MLL oncoproteins is dependent on Hoxa7 and Hoxa9. Genes & Dev. 2003;17:2298–2307. doi: 10.1101/gad.1111603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blais A., Dynlacht B.D., Dynlacht B.D. Hitting their targets: An emerging picture of E2F and cell cycle control. Curr. Opin. Genet. Dev. 2004;14:527–532. doi: 10.1016/j.gde.2004.07.003. [DOI] [PubMed] [Google Scholar]
- Bracken A.P., Ciro M., Cocito A., Helin K., Ciro M., Cocito A., Helin K., Cocito A., Helin K., Helin K. E2F target genes: Unraveling the biology. Trends Biochem. Sci. 2004;29:409–417. doi: 10.1016/j.tibs.2004.06.006. [DOI] [PubMed] [Google Scholar]
- Canaani E., Nakamura T., Rozovskaia T., Smith S.T., Mori T., Croce C.M., Mazo A., Nakamura T., Rozovskaia T., Smith S.T., Mori T., Croce C.M., Mazo A., Rozovskaia T., Smith S.T., Mori T., Croce C.M., Mazo A., Smith S.T., Mori T., Croce C.M., Mazo A., Mori T., Croce C.M., Mazo A., Croce C.M., Mazo A., Mazo A. ALL-1/MLL1, a homologue of DrosophilaTRITHORAX, modifies chromatin and is directly involved in infant acute leukaemia. Br. J. Cancer. 2004;90:756–760. doi: 10.1038/sj.bjc.6601639. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cardozo T., Pagano M., Pagano M. The SCF ubiquitin ligase: Insights into a molecular machine. Nat. Rev. Mol. Cell Biol. 2004;5:739–751. doi: 10.1038/nrm1471. [DOI] [PubMed] [Google Scholar]
- Caslini C., Serna A., Rossi V., Introna M., Biondi A., Serna A., Rossi V., Introna M., Biondi A., Rossi V., Introna M., Biondi A., Introna M., Biondi A., Biondi A. Modulation of cell cycle by graded expression of MLL-AF4 fusion oncoprotein. Leukemia. 2004;18:1064–1071. doi: 10.1038/sj.leu.2403321. [DOI] [PubMed] [Google Scholar]
- Chen W., Li Q., Hudson W.A., Kumar A., Kirchhof N., Kersey J.H., Li Q., Hudson W.A., Kumar A., Kirchhof N., Kersey J.H., Hudson W.A., Kumar A., Kirchhof N., Kersey J.H., Kumar A., Kirchhof N., Kersey J.H., Kirchhof N., Kersey J.H., Kersey J.H. A murine Mll-AF4 knock-in model results in lymphoid and myeloid deregulation and hematologic malignancy. Blood. 2006;108:669–677. doi: 10.1182/blood-2005-08-3498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Corral J., Lavenir I., Impey H., Warren A.J., Forster A., Larson T.A., Bell S., McKenzie A.N., King G., Rabbitts T.H., Lavenir I., Impey H., Warren A.J., Forster A., Larson T.A., Bell S., McKenzie A.N., King G., Rabbitts T.H., Impey H., Warren A.J., Forster A., Larson T.A., Bell S., McKenzie A.N., King G., Rabbitts T.H., Warren A.J., Forster A., Larson T.A., Bell S., McKenzie A.N., King G., Rabbitts T.H., Forster A., Larson T.A., Bell S., McKenzie A.N., King G., Rabbitts T.H., Larson T.A., Bell S., McKenzie A.N., King G., Rabbitts T.H., Bell S., McKenzie A.N., King G., Rabbitts T.H., McKenzie A.N., King G., Rabbitts T.H., King G., Rabbitts T.H., Rabbitts T.H. An Mll-AF9 fusion gene made by homologous recombination causes acute leukemia in chimeric mice: A method to create fusion oncogenes. Cell. 1996;85:853–861. doi: 10.1016/s0092-8674(00)81269-6. [DOI] [PubMed] [Google Scholar]
- Daser A., Rabbitts T.H., Rabbitts T.H. Extending the repertoire of the mixed-lineage leukemia gene MLL in leukemogenesis. Genes & Dev. 2004;18:965–974. doi: 10.1101/gad.1195504. [DOI] [PubMed] [Google Scholar]
- Djabali M., Selleri L., Parry P., Bower M., Young B., Evans G.A., Selleri L., Parry P., Bower M., Young B., Evans G.A., Parry P., Bower M., Young B., Evans G.A., Bower M., Young B., Evans G.A., Young B., Evans G.A., Evans G.A. A trithorax-like gene is interrupted by chromosome 11q23 translocations in acute leukaemias. Nat. Genet. 1993;4:431. doi: 10.1038/ng0893-431. [DOI] [PubMed] [Google Scholar]
- Dobson C.L., Warren A.J., Pannell R., Forster A., Rabbitts T.H., Warren A.J., Pannell R., Forster A., Rabbitts T.H., Pannell R., Forster A., Rabbitts T.H., Forster A., Rabbitts T.H., Rabbitts T.H. Tumorigenesis in mice with a fusion of the leukaemia oncogene Mll and the bacterial lacZ gene. EMBO J. 2000;19:843–851. doi: 10.1093/emboj/19.5.843. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Domer P.H., Fakharzadeh S.S., Chen C.S., Jockel J., Johansen L., Silverman G.A., Kersey J.H., Korsmeyer S.J., Fakharzadeh S.S., Chen C.S., Jockel J., Johansen L., Silverman G.A., Kersey J.H., Korsmeyer S.J., Chen C.S., Jockel J., Johansen L., Silverman G.A., Kersey J.H., Korsmeyer S.J., Jockel J., Johansen L., Silverman G.A., Kersey J.H., Korsmeyer S.J., Johansen L., Silverman G.A., Kersey J.H., Korsmeyer S.J., Silverman G.A., Kersey J.H., Korsmeyer S.J., Kersey J.H., Korsmeyer S.J., Korsmeyer S.J. Acute mixed-lineage leukemia t(4;11)(q21;q23) generates an MLL-AF4 fusion product. Proc. Natl. Acad. Sci. 1993;90:7884–7888. doi: 10.1073/pnas.90.16.7884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dyson N. The regulation of E2F by pRB-family proteins. Genes & Dev. 1998;12:2245–2262. doi: 10.1101/gad.12.15.2245. [DOI] [PubMed] [Google Scholar]
- Eguchi M., Eguchi-Ishimae M., Greaves M., Eguchi-Ishimae M., Greaves M., Greaves M. Molecular pathogenesis of MLL-associated leukemias. Int. J. Hematol. 2005;82:9–20. doi: 10.1532/IJH97.05042. [DOI] [PubMed] [Google Scholar]
- Felix C.A., Megonigal M.D., Chervinsky D.S., Leonard D.G., Tsuchida N., Kakati S., Block A.M., Fisher J., Grossi M., Salhany K.I., Megonigal M.D., Chervinsky D.S., Leonard D.G., Tsuchida N., Kakati S., Block A.M., Fisher J., Grossi M., Salhany K.I., Chervinsky D.S., Leonard D.G., Tsuchida N., Kakati S., Block A.M., Fisher J., Grossi M., Salhany K.I., Leonard D.G., Tsuchida N., Kakati S., Block A.M., Fisher J., Grossi M., Salhany K.I., Tsuchida N., Kakati S., Block A.M., Fisher J., Grossi M., Salhany K.I., Kakati S., Block A.M., Fisher J., Grossi M., Salhany K.I., Block A.M., Fisher J., Grossi M., Salhany K.I., Fisher J., Grossi M., Salhany K.I., Grossi M., Salhany K.I., Salhany K.I., et al. Association of germline p53 mutation with MLL segmental jumping translocation in treatment-related leukemia. Blood. 1998;91:4451–4456. [PubMed] [Google Scholar]
- Giacinti C., Giordano A., Giordano A. RB and cell cycle progression. Oncogene. 2006;25:5220–5227. doi: 10.1038/sj.onc.1209615. [DOI] [PubMed] [Google Scholar]
- Gilliland D.G., Jordan C.T., Felix C.A., Jordan C.T., Felix C.A., Felix C.A. The molecular basis of leukemia. Hematology. 2004;2004:80–97. doi: 10.1182/asheducation-2004.1.80. [DOI] [PubMed] [Google Scholar]
- Gu Y., Nakamura T., Alder H., Prasad R., Canaani O., Cimino G., Croce C.M., Canaani E., Nakamura T., Alder H., Prasad R., Canaani O., Cimino G., Croce C.M., Canaani E., Alder H., Prasad R., Canaani O., Cimino G., Croce C.M., Canaani E., Prasad R., Canaani O., Cimino G., Croce C.M., Canaani E., Canaani O., Cimino G., Croce C.M., Canaani E., Cimino G., Croce C.M., Canaani E., Croce C.M., Canaani E., Canaani E. The t(4;11) chromosome translocation of human acute leukemias fuses the ALL-1 gene, related to Drosophila trithorax, to the AF-4 gene. Cell. 1992;71:701–708. doi: 10.1016/0092-8674(92)90603-a. [DOI] [PubMed] [Google Scholar]
- Guo W.J., Datta S., Band V., Dimri G.P., Datta S., Band V., Dimri G.P., Band V., Dimri G.P., Dimri G.P. Mel-18, a polycomb group protein, regulates cell proliferation and senescence via transcriptional repression of Bmi-1 and c-Myc oncoproteins. Mol. Biol. Cell. 2007;18:536–546. doi: 10.1091/mbc.E06-05-0447. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hanson R.D., Hess J.L., Yu B.D., Ernst P., van Lohuizen M., Berns A., van der Lugt N.M., Shashikant C.S., Ruddle F.H., Seto M., Hess J.L., Yu B.D., Ernst P., van Lohuizen M., Berns A., van der Lugt N.M., Shashikant C.S., Ruddle F.H., Seto M., Yu B.D., Ernst P., van Lohuizen M., Berns A., van der Lugt N.M., Shashikant C.S., Ruddle F.H., Seto M., Ernst P., van Lohuizen M., Berns A., van der Lugt N.M., Shashikant C.S., Ruddle F.H., Seto M., van Lohuizen M., Berns A., van der Lugt N.M., Shashikant C.S., Ruddle F.H., Seto M., Berns A., van der Lugt N.M., Shashikant C.S., Ruddle F.H., Seto M., van der Lugt N.M., Shashikant C.S., Ruddle F.H., Seto M., Shashikant C.S., Ruddle F.H., Seto M., Ruddle F.H., Seto M., Seto M., et al. Mammalian Trithorax and polycomb-group homologues are antagonistic regulators of homeotic development. Proc. Natl. Acad. Sci. 1999;96:14372–14377. doi: 10.1073/pnas.96.25.14372. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hershko A. The ubiquitin system for protein degradation and some of its roles in the control of the cell division cycle. Cell Death Differ. 2005;12:1191–1197. doi: 10.1038/sj.cdd.4401702. [DOI] [PubMed] [Google Scholar]
- Hess J.L. Mechanisms of transformation by MLL. Crit. Rev. Eukaryot. Gene Expr. 2004;14:235–254. doi: 10.1615/critreveukaryotgeneexpr.v14.i4.10. [DOI] [PubMed] [Google Scholar]
- Hess J.L., Yu B.D., Li B., Hanson R., Korsmeyer S.J., Yu B.D., Li B., Hanson R., Korsmeyer S.J., Li B., Hanson R., Korsmeyer S.J., Hanson R., Korsmeyer S.J., Korsmeyer S.J. Defects in yolk sac hematopoiesis in Mll-null embryos. Blood. 1997;90:1799–1806. [PubMed] [Google Scholar]
- Hoiby T., Mitsiou D.J., Zhou H., Erdjument-Bromage H., Tempst P., Stunnenberg H.G., Mitsiou D.J., Zhou H., Erdjument-Bromage H., Tempst P., Stunnenberg H.G., Zhou H., Erdjument-Bromage H., Tempst P., Stunnenberg H.G., Erdjument-Bromage H., Tempst P., Stunnenberg H.G., Tempst P., Stunnenberg H.G., Stunnenberg H.G. Cleavage and proteasome-mediated degradation of the basal transcription factor TFIIA. EMBO J. 2004;23:3083–3091. doi: 10.1038/sj.emboj.7600304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hsieh J.J., Cheng E.H., Korsmeyer S.J., Cheng E.H., Korsmeyer S.J., Korsmeyer S.J. Taspase1: A threonine aspartase required for cleavage of MLL and proper HOX gene expression. Cell. 2003a;115:293–303. doi: 10.1016/s0092-8674(03)00816-x. [DOI] [PubMed] [Google Scholar]
- Hsieh J.J., Ernst P., Erdjument-Bromage H., Tempst P., Korsmeyer S.J., Ernst P., Erdjument-Bromage H., Tempst P., Korsmeyer S.J., Erdjument-Bromage H., Tempst P., Korsmeyer S.J., Tempst P., Korsmeyer S.J., Korsmeyer S.J. Proteolytic cleavage of MLL generates a complex of N- and C-terminal fragments that confers protein stability and subnuclear localization. Mol. Cell. Biol. 2003b;23:186–194. doi: 10.1128/MCB.23.1.186-194.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacobs J.J., Kieboom K., Marino S., DePinho R.A., van Lohuizen M., Kieboom K., Marino S., DePinho R.A., van Lohuizen M., Marino S., DePinho R.A., van Lohuizen M., DePinho R.A., van Lohuizen M., van Lohuizen M. The oncogene and Polycomb-group gene bmi-1 regulates cell proliferation and senescence through the ink4a locus. Nature. 1999a;397:164–168. doi: 10.1038/16476. [DOI] [PubMed] [Google Scholar]
- Jacobs J.J., Scheijen B., Voncken J.W., Kieboom K., Berns A., van Lohuizen M., Scheijen B., Voncken J.W., Kieboom K., Berns A., van Lohuizen M., Voncken J.W., Kieboom K., Berns A., van Lohuizen M., Kieboom K., Berns A., van Lohuizen M., Berns A., van Lohuizen M., van Lohuizen M. Bmi-1 collaborates with c-Myc in tumorigenesis by inhibiting c-Myc-induced apoptosis via INK4a/ARF. Genes & Dev. 1999b;13:2678–2690. doi: 10.1101/gad.13.20.2678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khan J.A., Dunn B.M., Tong L., Dunn B.M., Tong L., Tong L. Crystal structure of human Taspase1, a crucial protease regulating the function of MLL. Structure. 2005;13:1443–1452. doi: 10.1016/j.str.2005.07.006. [DOI] [PubMed] [Google Scholar]
- Kim H., Rafiuddin-Shah M., Tu H.C., Jeffers J.R., Zambetti G.P., Hsieh J.J., Cheng E.H., Rafiuddin-Shah M., Tu H.C., Jeffers J.R., Zambetti G.P., Hsieh J.J., Cheng E.H., Tu H.C., Jeffers J.R., Zambetti G.P., Hsieh J.J., Cheng E.H., Jeffers J.R., Zambetti G.P., Hsieh J.J., Cheng E.H., Zambetti G.P., Hsieh J.J., Cheng E.H., Hsieh J.J., Cheng E.H., Cheng E.H. Hierarchical regulation of mitochondrion-dependent apoptosis by BCL-2 subfamilies. Nat. Cell Biol. 2006;8:1348–1358. doi: 10.1038/ncb1499. [DOI] [PubMed] [Google Scholar]
- Luo R.T., Kebriaei P., Kaberlein J.J., Thirman M.J., Kebriaei P., Kaberlein J.J., Thirman M.J., Kaberlein J.J., Thirman M.J., Thirman M.J. Cooperating mutations are necessary for the development of AML in mll-ELL knock-in mice. Blood. 2002;100:136a. [Google Scholar]
- Mahgoub N., Parker R.I., Hosler M.R., Close P., Winick N.J., Masterson M., Shannon K.M., Felix C.A., Parker R.I., Hosler M.R., Close P., Winick N.J., Masterson M., Shannon K.M., Felix C.A., Hosler M.R., Close P., Winick N.J., Masterson M., Shannon K.M., Felix C.A., Close P., Winick N.J., Masterson M., Shannon K.M., Felix C.A., Winick N.J., Masterson M., Shannon K.M., Felix C.A., Masterson M., Shannon K.M., Felix C.A., Shannon K.M., Felix C.A., Felix C.A. RAS mutations in pediatric leukemias with MLL gene rearrangements. Genes Chromosomes Cancer. 1998;21:270–275. [PubMed] [Google Scholar]
- Martin M.E., Milne T.A., Bloyer S., Galoian K., Shen W., Gibbs D., Brock H.W., Slany R., Hess J.L., Milne T.A., Bloyer S., Galoian K., Shen W., Gibbs D., Brock H.W., Slany R., Hess J.L., Bloyer S., Galoian K., Shen W., Gibbs D., Brock H.W., Slany R., Hess J.L., Galoian K., Shen W., Gibbs D., Brock H.W., Slany R., Hess J.L., Shen W., Gibbs D., Brock H.W., Slany R., Hess J.L., Gibbs D., Brock H.W., Slany R., Hess J.L., Brock H.W., Slany R., Hess J.L., Slany R., Hess J.L., Hess J.L. Dimerization of MLL fusion proteins immortalizes hematopoietic cells. Cancer Cell. 2003;4:197–207. doi: 10.1016/s1535-6108(03)00214-9. [DOI] [PubMed] [Google Scholar]
- Martinez A.M., Cavalli G., Cavalli G. The role of polycomb group proteins in cell cycle regulation during development. Cell Cycle. 2006;5:1189–1197. doi: 10.4161/cc.5.11.2781. [DOI] [PubMed] [Google Scholar]
- Martinez A.M., Colomb S., Dejardin J., Bantignies F., Cavalli G., Colomb S., Dejardin J., Bantignies F., Cavalli G., Dejardin J., Bantignies F., Cavalli G., Bantignies F., Cavalli G., Cavalli G. Polycomb group-dependent Cyclin A repression in Drosophila. Genes & Dev. 2006;20:501–513. doi: 10.1101/gad.357106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Metzler M., Forster A., Pannell R., Arends M.J., Daser A., Lobato M.N., Rabbitts T.H., Forster A., Pannell R., Arends M.J., Daser A., Lobato M.N., Rabbitts T.H., Pannell R., Arends M.J., Daser A., Lobato M.N., Rabbitts T.H., Arends M.J., Daser A., Lobato M.N., Rabbitts T.H., Daser A., Lobato M.N., Rabbitts T.H., Lobato M.N., Rabbitts T.H., Rabbitts T.H. A conditional model of MLL-AF4 B-cell tumourigenesis using invertor technology. Oncogene. 2006;25:3093–3103. doi: 10.1038/sj.onc.1209636. [DOI] [PubMed] [Google Scholar]
- Milne T.A., Briggs S.D., Brock H.W., Martin M.E., Gibbs D., Allis C.D., Hess J.L., Briggs S.D., Brock H.W., Martin M.E., Gibbs D., Allis C.D., Hess J.L., Brock H.W., Martin M.E., Gibbs D., Allis C.D., Hess J.L., Martin M.E., Gibbs D., Allis C.D., Hess J.L., Gibbs D., Allis C.D., Hess J.L., Allis C.D., Hess J.L., Hess J.L. MLL targets SET domain methyltransferase activity to Hox gene promoters. Mol. Cell. 2002;10:1107–1117. doi: 10.1016/s1097-2765(02)00741-4. [DOI] [PubMed] [Google Scholar]
- Milne T.A., Hughes C.M., Lloyd R., Yang Z., Rozenblatt-Rosen O., Dou Y., Schnepp R.W., Krankel C., Livolsi V.A., Gibbs D., Hughes C.M., Lloyd R., Yang Z., Rozenblatt-Rosen O., Dou Y., Schnepp R.W., Krankel C., Livolsi V.A., Gibbs D., Lloyd R., Yang Z., Rozenblatt-Rosen O., Dou Y., Schnepp R.W., Krankel C., Livolsi V.A., Gibbs D., Yang Z., Rozenblatt-Rosen O., Dou Y., Schnepp R.W., Krankel C., Livolsi V.A., Gibbs D., Rozenblatt-Rosen O., Dou Y., Schnepp R.W., Krankel C., Livolsi V.A., Gibbs D., Dou Y., Schnepp R.W., Krankel C., Livolsi V.A., Gibbs D., Schnepp R.W., Krankel C., Livolsi V.A., Gibbs D., Krankel C., Livolsi V.A., Gibbs D., Livolsi V.A., Gibbs D., Gibbs D., et al. Menin and MLL cooperatively regulate expression of cyclin-dependent kinase inhibitors. Proc. Natl. Acad. Sci. 2005;102:749–754. doi: 10.1073/pnas.0408836102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morgan D.O. Cyclin-dependent kinases: Engines, clocks, and microprocessors. Annu. Rev. Cell Dev. Biol. 1997;13:261–291. doi: 10.1146/annurev.cellbio.13.1.261. [DOI] [PubMed] [Google Scholar]
- Murray A.W. Recycling the cell cycle: Cyclins revisited. Cell. 2004;116:221–234. doi: 10.1016/s0092-8674(03)01080-8. [DOI] [PubMed] [Google Scholar]
- Muyrers-Chen I., Rozovskaia T., Lee N., Kersey J.H., Nakamura T., Canaani E., Paro R., Rozovskaia T., Lee N., Kersey J.H., Nakamura T., Canaani E., Paro R., Lee N., Kersey J.H., Nakamura T., Canaani E., Paro R., Kersey J.H., Nakamura T., Canaani E., Paro R., Nakamura T., Canaani E., Paro R., Canaani E., Paro R., Paro R. Expression of leukemic MLL fusion proteins in Drosophilaaffects cell cycle control and chromosome morphology. Oncogene. 2004;23:8639–8648. doi: 10.1038/sj.onc.1207904. [DOI] [PubMed] [Google Scholar]
- Nakamura T., Mori T., Tada S., Krajewski W., Rozovskaia T., Wassell R., Dubois G., Mazo A., Croce C.M., Canaani E., Mori T., Tada S., Krajewski W., Rozovskaia T., Wassell R., Dubois G., Mazo A., Croce C.M., Canaani E., Tada S., Krajewski W., Rozovskaia T., Wassell R., Dubois G., Mazo A., Croce C.M., Canaani E., Krajewski W., Rozovskaia T., Wassell R., Dubois G., Mazo A., Croce C.M., Canaani E., Rozovskaia T., Wassell R., Dubois G., Mazo A., Croce C.M., Canaani E., Wassell R., Dubois G., Mazo A., Croce C.M., Canaani E., Dubois G., Mazo A., Croce C.M., Canaani E., Mazo A., Croce C.M., Canaani E., Croce C.M., Canaani E., Canaani E. ALL-1 is a histone methyltransferase that assembles a supercomplex of proteins involved in transcriptional regulation. Mol. Cell. 2002;10:1119–1128. doi: 10.1016/s1097-2765(02)00740-2. [DOI] [PubMed] [Google Scholar]
- Nakayama K.I., Nakayama K., Nakayama K. Ubiquitin ligases: Cell-cycle control and cancer. Nat. Rev. Cancer. 2006;6:369–381. doi: 10.1038/nrc1881. [DOI] [PubMed] [Google Scholar]
- Nevins J.R. The Rb/E2F pathway and cancer. Hum. Mol. Genet. 2001;10:699–703. doi: 10.1093/hmg/10.7.699. [DOI] [PubMed] [Google Scholar]
- Nurse P. A long twentieth century of the cell cycle and beyond. Cell. 2000;100:71–78. doi: 10.1016/s0092-8674(00)81684-0. [DOI] [PubMed] [Google Scholar]
- Oguchi K., Takagi M., Tsuchida R., Taya Y., Ito E., Isoyama K., Ishii E., Zannini L., Delia D., Mizutani S., Takagi M., Tsuchida R., Taya Y., Ito E., Isoyama K., Ishii E., Zannini L., Delia D., Mizutani S., Tsuchida R., Taya Y., Ito E., Isoyama K., Ishii E., Zannini L., Delia D., Mizutani S., Taya Y., Ito E., Isoyama K., Ishii E., Zannini L., Delia D., Mizutani S., Ito E., Isoyama K., Ishii E., Zannini L., Delia D., Mizutani S., Isoyama K., Ishii E., Zannini L., Delia D., Mizutani S., Ishii E., Zannini L., Delia D., Mizutani S., Zannini L., Delia D., Mizutani S., Delia D., Mizutani S., Mizutani S. Missense mutation and defective function of ATM in a childhood acute leukemia patient with MLL gene rearrangement. Blood. 2003;101:3622–3627. doi: 10.1182/blood-2002-02-0570. [DOI] [PubMed] [Google Scholar]
- Pession A., Martino V., Tonelli R., Beltramini C., Locatelli F., Biserni G., Franzoni M., Freccero F., Montemurro L., Pattacini L., Martino V., Tonelli R., Beltramini C., Locatelli F., Biserni G., Franzoni M., Freccero F., Montemurro L., Pattacini L., Tonelli R., Beltramini C., Locatelli F., Biserni G., Franzoni M., Freccero F., Montemurro L., Pattacini L., Beltramini C., Locatelli F., Biserni G., Franzoni M., Freccero F., Montemurro L., Pattacini L., Locatelli F., Biserni G., Franzoni M., Freccero F., Montemurro L., Pattacini L., Biserni G., Franzoni M., Freccero F., Montemurro L., Pattacini L., Franzoni M., Freccero F., Montemurro L., Pattacini L., Freccero F., Montemurro L., Pattacini L., Montemurro L., Pattacini L., Pattacini L., et al. MLL-AF9 oncogene expression affects cell growth but not terminal differentiation and is downregulated during monocyte–macrophage maturation in AML-M5 THP-1 cells. Oncogene. 2003;22:8671–8676. doi: 10.1038/sj.onc.1207125. [DOI] [PubMed] [Google Scholar]
- Peters J.M. The anaphase promoting complex/cyclosome: A machine designed to destroy. Nat. Rev. Mol. Cell Biol. 2006;7:644–656. doi: 10.1038/nrm1988. [DOI] [PubMed] [Google Scholar]
- Reed S.I. Ratchets and clocks: The cell cycle, ubiquitylation and protein turnover. Nat. Rev. Mol. Cell Biol. 2003;4:855–864. doi: 10.1038/nrm1246. [DOI] [PubMed] [Google Scholar]
- Ringrose L., Paro R., Paro R. Epigenetic regulation of cellular memory by the Polycomb and Trithorax group proteins. Annu. Rev. Genet. 2004;38:413–443. doi: 10.1146/annurev.genet.38.072902.091907. [DOI] [PubMed] [Google Scholar]
- Rowley J.D. The critical role of chromosome translocations in human leukemias. Annu. Rev. Genet. 1998;32:495–519. doi: 10.1146/annurev.genet.32.1.495. [DOI] [PubMed] [Google Scholar]
- Schuettengruber B., Chourrout D., Vervoort M., Leblanc B., Cavalli G., Chourrout D., Vervoort M., Leblanc B., Cavalli G., Vervoort M., Leblanc B., Cavalli G., Leblanc B., Cavalli G., Cavalli G. Genome regulation by polycomb and trithorax proteins. Cell. 2007;128:735–745. doi: 10.1016/j.cell.2007.02.009. [DOI] [PubMed] [Google Scholar]
- Schumacher A., Magnuson T., Magnuson T. Murine Polycomb- and trithorax-group genes regulate homeotic pathways and beyond. Trends Genet. 1997;13:167–170. [PubMed] [Google Scholar]
- Sherr C.J., Roberts J.M., Roberts J.M. CDK inhibitors: Positive and negative regulators of G1-phase progression. Genes & Dev. 1999;13:1501–1512. doi: 10.1101/gad.13.12.1501. [DOI] [PubMed] [Google Scholar]
- Slany R.K., Lavau C., Cleary M.L., Lavau C., Cleary M.L., Cleary M.L. The oncogenic capacity of HRX-ENL requires the transcriptional transactivation activity of ENL and the DNA binding motifs of HRX. Mol. Cell. Biol. 1998;18:122–129. doi: 10.1128/mcb.18.1.122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- So C.W., Lin M., Ayton P.M., Chen E.H., Cleary M.L., Lin M., Ayton P.M., Chen E.H., Cleary M.L., Ayton P.M., Chen E.H., Cleary M.L., Chen E.H., Cleary M.L., Cleary M.L. Dimerization contributes to oncogenic activation of MLL chimeras in acute leukemias. Cancer Cell. 2003;4:99–110. doi: 10.1016/s1535-6108(03)00188-0. [DOI] [PubMed] [Google Scholar]
- Takeda S., Chen D.Y., Westergard T.D., Fisher J.K., Rubens J.A., Sasagawa S., Kan J.T., Korsmeyer S.J., Cheng E.H., Hsieh J.J., Chen D.Y., Westergard T.D., Fisher J.K., Rubens J.A., Sasagawa S., Kan J.T., Korsmeyer S.J., Cheng E.H., Hsieh J.J., Westergard T.D., Fisher J.K., Rubens J.A., Sasagawa S., Kan J.T., Korsmeyer S.J., Cheng E.H., Hsieh J.J., Fisher J.K., Rubens J.A., Sasagawa S., Kan J.T., Korsmeyer S.J., Cheng E.H., Hsieh J.J., Rubens J.A., Sasagawa S., Kan J.T., Korsmeyer S.J., Cheng E.H., Hsieh J.J., Sasagawa S., Kan J.T., Korsmeyer S.J., Cheng E.H., Hsieh J.J., Kan J.T., Korsmeyer S.J., Cheng E.H., Hsieh J.J., Korsmeyer S.J., Cheng E.H., Hsieh J.J., Cheng E.H., Hsieh J.J., Hsieh J.J. Proteolysis of MLL family proteins is essential for taspase1-orchestrated cell cycle progression. Genes & Dev. 2006;20:2397–2409. doi: 10.1101/gad.1449406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thirman M.J., Gill H.J., Burnett R.C., Mbangkollo D., McCabe N.R., Kobayashi H., Ziemin-van der Poel S., Kaneko Y., Morgan R., Sandberg A.A., Gill H.J., Burnett R.C., Mbangkollo D., McCabe N.R., Kobayashi H., Ziemin-van der Poel S., Kaneko Y., Morgan R., Sandberg A.A., Burnett R.C., Mbangkollo D., McCabe N.R., Kobayashi H., Ziemin-van der Poel S., Kaneko Y., Morgan R., Sandberg A.A., Mbangkollo D., McCabe N.R., Kobayashi H., Ziemin-van der Poel S., Kaneko Y., Morgan R., Sandberg A.A., McCabe N.R., Kobayashi H., Ziemin-van der Poel S., Kaneko Y., Morgan R., Sandberg A.A., Kobayashi H., Ziemin-van der Poel S., Kaneko Y., Morgan R., Sandberg A.A., Ziemin-van der Poel S., Kaneko Y., Morgan R., Sandberg A.A., Kaneko Y., Morgan R., Sandberg A.A., Morgan R., Sandberg A.A., Sandberg A.A., et al. Rearrangement of the MLL gene in acute lymphoblastic and acute myeloid leukemias with 11q23 chromosomal translocations. N. Engl. J. Med. 1993;329:909–914. doi: 10.1056/NEJM199309233291302. [DOI] [PubMed] [Google Scholar]
- Tkachuk D.C., Kohler S., Cleary M.L., Kohler S., Cleary M.L., Cleary M.L. Involvement of a homolog of Drosophila trithorax by 11q23 chromosomal translocations in acute leukemias. Cell. 1992;71:691–700. doi: 10.1016/0092-8674(92)90602-9. [DOI] [PubMed] [Google Scholar]
- Treier M., Staszewski L.M., Bohmann D., Staszewski L.M., Bohmann D., Bohmann D. Ubiquitin-dependent c-Jun degradation in vivo is mediated by the δ domain. Cell. 1994;78:787–798. doi: 10.1016/s0092-8674(94)90502-9. [DOI] [PubMed] [Google Scholar]
- Trimarchi J.M., Lees J.A., Lees J.A. Sibling rivalry in the E2F family. Nat. Rev. Mol. Cell Biol. 2002;3:11–20. doi: 10.1038/nrm714. [DOI] [PubMed] [Google Scholar]
- van Lohuizen M. The trithorax-group and polycomb-group chromatin modifiers: Implications for disease. Curr. Opin. Genet. Dev. 1999;9:355–361. doi: 10.1016/s0959-437x(99)80053-7. [DOI] [PubMed] [Google Scholar]
- van Lohuizen M., Verbeek S., Scheijen B., Wientjens E., van der Gulden H., Berns A., Verbeek S., Scheijen B., Wientjens E., van der Gulden H., Berns A., Scheijen B., Wientjens E., van der Gulden H., Berns A., Wientjens E., van der Gulden H., Berns A., van der Gulden H., Berns A., Berns A. Identification of cooperating oncogenes in E μ-myc transgenic mice by provirus tagging. Cell. 1991;65:737–752. doi: 10.1016/0092-8674(91)90382-9. [DOI] [PubMed] [Google Scholar]
- Wang J., Iwasaki H., Krivtsov A., Febbo P.G., Thorner A.R., Ernst P., Anastasiadou E., Kutok J.L., Kogan S.C., Zinkel S.S., Iwasaki H., Krivtsov A., Febbo P.G., Thorner A.R., Ernst P., Anastasiadou E., Kutok J.L., Kogan S.C., Zinkel S.S., Krivtsov A., Febbo P.G., Thorner A.R., Ernst P., Anastasiadou E., Kutok J.L., Kogan S.C., Zinkel S.S., Febbo P.G., Thorner A.R., Ernst P., Anastasiadou E., Kutok J.L., Kogan S.C., Zinkel S.S., Thorner A.R., Ernst P., Anastasiadou E., Kutok J.L., Kogan S.C., Zinkel S.S., Ernst P., Anastasiadou E., Kutok J.L., Kogan S.C., Zinkel S.S., Anastasiadou E., Kutok J.L., Kogan S.C., Zinkel S.S., Kutok J.L., Kogan S.C., Zinkel S.S., Kogan S.C., Zinkel S.S., Zinkel S.S., et al. Conditional MLL-CBP targets GMP and models therapy-related myeloproliferative disease. EMBO J. 2005;24:368–381. doi: 10.1038/sj.emboj.7600521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xia Z.B., Popovic R., Chen J., Theisler C., Stuart T., Santillan D.A., Erfurth F., Diaz M.O., Zeleznik-Le N.J., Popovic R., Chen J., Theisler C., Stuart T., Santillan D.A., Erfurth F., Diaz M.O., Zeleznik-Le N.J., Chen J., Theisler C., Stuart T., Santillan D.A., Erfurth F., Diaz M.O., Zeleznik-Le N.J., Theisler C., Stuart T., Santillan D.A., Erfurth F., Diaz M.O., Zeleznik-Le N.J., Stuart T., Santillan D.A., Erfurth F., Diaz M.O., Zeleznik-Le N.J., Santillan D.A., Erfurth F., Diaz M.O., Zeleznik-Le N.J., Erfurth F., Diaz M.O., Zeleznik-Le N.J., Diaz M.O., Zeleznik-Le N.J., Zeleznik-Le N.J. The MLL fusion gene, MLL-AF4, regulates cyclin-dependent kinase inhibitor CDKN1B (p27kip1) expression. Proc. Natl. Acad. Sci. 2005;102:14028–14033. doi: 10.1073/pnas.0506464102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yokoyama A., Kitabayashi I., Ayton P.M., Cleary M.L., Ohki M., Kitabayashi I., Ayton P.M., Cleary M.L., Ohki M., Ayton P.M., Cleary M.L., Ohki M., Cleary M.L., Ohki M., Ohki M. Leukemia proto-oncoprotein MLL is proteolytically processed into 2 fragments with opposite transcriptional properties. Blood. 2002;100:3710–3718. doi: 10.1182/blood-2002-04-1015. [DOI] [PubMed] [Google Scholar]
- Yu B.D., Hess J.L., Horning S.E., Brown G.A., Korsmeyer S.J., Hess J.L., Horning S.E., Brown G.A., Korsmeyer S.J., Horning S.E., Brown G.A., Korsmeyer S.J., Brown G.A., Korsmeyer S.J., Korsmeyer S.J. Altered Hox expression and segmental identity in Mll-mutant mice. Nature. 1995;378:505–508. doi: 10.1038/378505a0. [DOI] [PubMed] [Google Scholar]
- Yu B.D., Hanson R.D., Hess J.L., Horning S.E., Korsmeyer S.J., Hanson R.D., Hess J.L., Horning S.E., Korsmeyer S.J., Hess J.L., Horning S.E., Korsmeyer S.J., Horning S.E., Korsmeyer S.J., Korsmeyer S.J. MLL, a mammalian trithorax-group gene, functions as a transcriptional maintenance factor in morphogenesis. Proc. Natl. Acad. Sci. 1998;95:10632–10636. doi: 10.1073/pnas.95.18.10632. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou H., Spicuglia S., Hsieh J.J., Mitsiou D.J., Hoiby T., Veenstra G.J., Korsmeyer S.J., Stunnenberg H.G., Spicuglia S., Hsieh J.J., Mitsiou D.J., Hoiby T., Veenstra G.J., Korsmeyer S.J., Stunnenberg H.G., Hsieh J.J., Mitsiou D.J., Hoiby T., Veenstra G.J., Korsmeyer S.J., Stunnenberg H.G., Mitsiou D.J., Hoiby T., Veenstra G.J., Korsmeyer S.J., Stunnenberg H.G., Hoiby T., Veenstra G.J., Korsmeyer S.J., Stunnenberg H.G., Veenstra G.J., Korsmeyer S.J., Stunnenberg H.G., Korsmeyer S.J., Stunnenberg H.G., Stunnenberg H.G. Uncleaved TFIIA is a substrate for taspase 1 and active in transcription. Mol. Cell. Biol. 2006;26:2728–2735. doi: 10.1128/MCB.26.7.2728-2735.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]