Abstract
HIV-1 compartmentalization in the CNS has been demonstrated for gag, pol, and env, genes. However, little is known about tissue compartmentalization of nef genes and their functional characteristics in brain. We have cloned 97 nef genes and characterized 10 Nef proteins from autopsy brain and lymphoid tissues from 2 patients with AIDS and HIV-1-associated dementia. Distinct compartmentalization of brain versus lymphoid nef genes was demonstrated within each patient. CD4 and MHC-I downregulation were conserved in all tissue-derived Nefs. However, MHC-I downregulation by brain-derived Nefs was weaker than downregulation by lymphoid-derived Nefs. The motifs KEEE- or EKEE- at the PACS-1 binding site represented brain-specific signature patterns in these 2 patients and contributed to the reduced MHC-I downregulation activity of brain-derived Nefs from these patients. Pak2 association was highly variable in Nefs from both patients. Three of 10 tissue-derived Nefs coimmunoprecipitated activated Pak2, with strong association demonstrated for only 2 Nefs. The ability of Nef to associate with activated Pak2 did not correlate with brain or lymphoid tissue origin. Nef genes from viruses isolated from brain by coculture with PBMC were not closely related to sequences amplified directly from brain tissue, suggesting that viral selection or adaptation occurred during coculture. This study of tissue-derived HIV-1 Nefs demonstrates that CD4 and MHC-I downregulation are highly conserved Nef functions, while Pak2 association is variable in late stage AIDS patients.
Keywords: Human Immunodeficiency Virus Type 1 (HIV-1), Nef, brain, lymph node, CD4, MHC-I, Pak2
Introduction
The Nef protein of human and simian immunodeficiency viruses (HIV-1, HIV-2 and SIV) is an important determinant of progression to AIDS. Nef is required for maintenance of high viral load and disease induction in SIV-infected rhesus monkeys (Du et al., 1995; Kestler et al., 1991). Furthermore, expression of the HIV-1 nef gene in transgenic mice produces AIDS-like disease (Hanna et al., 1998; Skowronski et al., 1993). Nef-defective HIV-1 has been associated with long-term nonprogression in infected individuals (Dyer et al., 1999; Kirchhoff et al., 1995; Learmont et al., 1992). These findings suggest that Nef is important for viral replication and pathogenicity in vivo.
HIV-1 Nef is a 27 kDa membrane-associated cytoplasmic protein expressed during early stages of viral replication. Well-studied functions of Nef include downregulation of CD4 and MHC-I, enhancement of viral infectivity, and modulation of cellular signaling pathways (reviewed in (Arold and Baur, 2001; Bour and Strebel, 2000; Fackler and Baur, 2002; Geyer et al., 2001; Wei et al., 2003)). Nef downregulates CD4 by linking cell surface CD4 to the adaptor protein complex of clathrin-coated pits (Bresnahan et al., 1998; Greenberg et al., 1998a) and transferring CD4 to COP-I for transport to lysosomes (Piguet et al., 1999a). Downregulation of cell surface CD4 enhances HIV-1 replication and infectivity, virion production and release, and prevents superinfection of infected cells (reviewed in (Harris, 1999; Piguet et al., 1999b) and (Lama, 2003). MHC-I downregulation is achieved by targeting of MHC-I to the endosome-to-Golgi sorting pathway governed by PACS-1 (phospho-furin acidic cluster sorting protein –1) (Doms and Trono, 2000). Nef motifs critical for MHC-I downregulation are the acidic EEEE62–65 motif, which binds PACS-1 (Piguet et al., 2000), and the proline-rich SH3-binding domain (PxxP domain) (Blagoveshchenskaya et al., 2002; Greenberg et al., 1998b; Mangasarian et al., 1999). MHC-I downregulation may be important for protection of infected cells against lysis mediated by HIV-I-specific cytotoxic T lymphocytes in vivo (Collins et al., 1998; Swigut et al., 2004; Tomiyama et al., 2002; Yang et al., 2002). The mechanism by which Nef enhances viral infectivity is still controversial.
Nef expression enhances HIV-1 replication when resting peripheral blood mononuclear cells (PBMC) are infected and subsequently stimulated (Miller et al., 1995; Spina et al., 1994), most likely by activation of resting T cells through modulation of cellular signaling pathways (Alexander et al., 1997; Baur et al., 1994; Du et al., 1995; Schrager and Marsh, 1999; Simmons et al., 2001; Wang et al., 2000; Wu and Marsh, 2001). In vivo, the SIVmac239 nef gene enhances productive infection of resting memory CD4+ T cells in the paracortical regions of lymph nodes, thereby increasing viral replication and pathogenicity (Sugimoto et al., 2003). The interaction between Nef and p21-activated kinase 2 (Pak2), a cellular serine/threonine kinase, has been proposed to play an important role in T cell activation and AIDS progression in vivo (Arora et al., 2000; Lu et al., 1996; Nunn and Marsh, 1996; Renkema et al., 1999; Renkema et al., 2001; Sawai et al., 1996; Wei et al., 2005; Wolf et al., 2001). Pak2 is involved in the regulation of several cellular processes including cytoskeleton rearrangement, cell morphology, motility, apoptosis and gene transcription and is activated in response to a variety of cellular stresses (reviewed in (Bagrodia and Cerione, 1999; Daniels and Bokoch, 1999)). Nef-Pak2 association has also been demonstrated in primary macrophages (Brown et al., 2004). The PxxP domain of Nef, which is required for interaction with the SH3 domains of Src tyrosine kinases and other signaling molecules, is required for Pak2 activation (Khan et al., 1998; Manninen et al., 1998; Wiskerchen and Cheng-Mayer, 1996). A cluster of residues on the surface of the Nef core domain near F191 and H89 is also critical for association with activated Pak2 (Agopian et al., 2006; Foster et al., 2001; O’Neill et al., 2006), and may form part of a unique binding surface specifically involved in Nef-Pak2 association.
In addition to enhancing T cell activation, Nef also activates signaling pathways in monocyte-derived macrophages. Macrophages play an important role in HIV-1 transmission and AIDS pathogenesis, and are the primary cellular reservoir for HIV-1 infection in the CNS (Dunfee et al., 2006; Gonzalez-Scarano and Martin-Garcia, 2005; Verani et al., 2005). HIV-1-infected macrophages release inflammatory cytokines including MIP-1α/CCL3, MIP-1β/CCL4, and other factors (e.g., IL-16 and soluble forms of CD23 and ICAM-1) that contribute to leukocyte recruitment and activation in a Nef-dependent manner (Olivetta et al., 2003; Swingler et al., 2003; Swingler et al., 1999; Zhao et al., 2004). The PxxP domain and motifs involved in CD4 downregulation in Nef are important for efficient viral replication in macrophages (Brown et al., 2004). Nef-expressing macrophages transplanted into the brain of rats induce monocyte/macrophage recruitment, TNF-α expression, astrogliosis, and cognitive defects (Mordelet et al., 2004). Furthermore, SCID/NOD mice injected with Nef-expressing viral vectors exhibited evidence of neuronal injury and neurological disease (van Marle et al., 2004). HIV-1- infected and/or activated macrophages release soluble neurotoxic factors that induce neuronal injury and neurocognitive impairment (Gonzalez-Scarano and Martin-Garcia, 2005; Kaul et al., 2005). Nef may regulate production of some of these neurotoxic factors by stimulating macrophage activation (Mahlknecht et al., 2000; Olivetta et al., 2005; Vilhardt et al., 2002)
HIV-1 enters the CNS in the early stages of infection by trafficking across the blood- brain barrier within infected monocytes and possibly lymphocytes (reviewed in (Williams and Hickey, 2002) and (Dunfee et al., 2006)). However, productive infection of macrophages and microglia is typically detected only after progression to AIDS. Genetic evolution of HIV-1 within the brain is distinct from that in lymphoid tissues and other organs (Chang et al., 1998; Donaldson et al., 1994; Gartner et al., 1997; Gorry et al., 2001; Hughes et al., 1997; Korber et al., 1994; Shapshak et al., 1999; van’t Wout et al., 1998; Wong et al., 1997). Previous studies demonstrated CNS compartmentalization of the HIV-1 gag, pol, and env genes. Only one previous study analyzed the genetic compartmentalization of full-length nef genes in the CNS (van Marle et al., 2004). This study also found that 3 brain-derived nef genes differentially induced pro-inflammatory gene expression in astrocytes, indicating that Nef sequence variability could affect pathogenesis. However, little is known about functional differences of Nefs from brain versus Nefs from other tissues. Nefs in brain may be functionally divergent from those in blood or lymphoid tissues as a consequence of adaptation to different target cell populations and reduced immune surveillance. In particular, low CD4 expression on macrophages/microglia and reduced CTL surveillance in the brain could result in reduced selection pressure to maintain strong CD4 or MHC-I downregulation activity, respectively.
To investigate genetic and functional characteristics of Nefs in brain versus lymphoid tissues, full-length nef genes were amplified from the genomic DNA of autopsy brain and lymphoid tissues from 2 patients with late stage AIDS and HIV-1-associated dementia (HAD). Distinct compartmentalization of brain versus lymphoid nef sequences was demonstrated within each patient. CD4 and MHC-I downregulation activities were highly conserved among primary Nefs. However, MHC-I downregulation by brain-derived Nefs was weaker than downregulation by lymphoid-derived Nefs. The motifs EKEE65 or KEEE65 in the PACS-1 binding site were identified as brain-specific signature patterns in both patients, whereas most lymphoid Nefs had the motif EEEEE65. Mutagenesis demonstrated that Nefs with EKEE65 or KEEE65 had a reduced capacity to downregulate MHC-I Nefs with EEEE65. Higher frequencies of EKEE65 or KEEE65 motifs and reduced capacity to downregulate MHC-I in brain-derived Nefs from these 2 AIDS patients may reflect reduced immune selection pressures in the CNS compared to non-CNS tissues. Of 10 brain- and lymphoid tissue-derived Nefs, only 3 associated with activated Pak2. The ability of Nef to associate with Pak2 did not correlate brain or lymphoid tissue origin. These findings indicate that CD4 and MHC-I downregulation are conserved in primary Nef alleles derived from brain and lymphoid tissues from late stage AIDS patients, while Pak2 activation is highly variable.
Results
Amplification of nef alleles from brain and lymphoid tissues and brain- and lymph node-derived viral isolates
To compare genetic and functional characteristics of Nef sequences from brain versus lymphoid tissues, we amplified nef genes from the genomic DNA of autopsy brain and lymph node tissues of 2 patients (MACS2 and MACS3) with late stage AIDS and HIV-1-associated dementia (Gorry et al., 2001). Nef genes were also amplified from the genomic DNA of spleen of patient MACS2. Phylogenetic analysis showed that sequences from both bulk and limiting dilution PCRs were well interleaved and did not show evidence of amplification bias (data not shown). To reduce potential effects of primer bias, 2 different primer sets were used in limiting dilution PCR reactions. Although some sequences were more commonly amplified by one or the other primer set, both primer sets amplified at least one nef sequence from every major branch of the phylogenetic trees (data not shown). A total of 51 nef sequences were amplified directly from MACS2 tissue (18 from brain, 22 from lymph node, and 11 from spleen) and 46 from MACS3 tissue (12 from brain, and 34 from lymph node). All nef sequences amplified directly from MACS2 and MACS3 tissues encoded full-length open reading frames.
In addition to amplifying nef genes directly from the tissue samples as discussed above, we also amplified nef genes from viral isolates obtained from coculture of PBMC with the same tissues. These isolates, MACS2-br, MACS2-LN, MACS3-br and MACS3-LN, were described previously (Gorry et al., 2001). For each viral isolate, we sequenced the bulk PCR product and 3 to 7 nef genes amplified from genomic DNA isolated from PBMC on day 7 post-infection with virus stock. Within each viral isolate, Nef clones were highly similar (98.3% – 100% Nef predicted amino acid identity). All nef genes from viral isolate MACS2-br, including representative Nef clone 4G, contain an early frameshift that introduced a stop codon after only 61 amino acids. Clone MACS2-br-4G, could not be detected by Western blot and was nonfunctional in assays for CD4 and MHC-I downregulation and Pak2 activation (data not shown). Representative Nef clones matching the consensus sequence of each of the other viral isolates were chosen for further functional analysis. Sequences and functional analyses of representative viral isolate clones MACS2-LN-5C, MACS3-br-6I, and MACS3-LN-7D have been published (Agopian et al., 2006).
Nef sequences from viral isolates derived from brain-PBMC co-cultures are not representative of those in brain tissue
To determine whether coculture of brain tissue with PBMC resulted in viral isolates representative of virus in the brain in vivo, we used phylogenetic analysis to compare the nucleotide sequences of nef genes from PBMC/brain coculture-derived viral isolates with sequences of nefs amplified directly from the brain tissues. We found that nef sequences amplified from viral isolates derived by brain-PBMC coculture were not representative of nef sequences in brain tissue. Even after repair of the frameshift, the nucleotide sequence of nef clone 4G from viral isolate MACS2-br (shown in pink in Figure 1A) is highly divergent from all other MACS2 sequences, including sequences amplified directly from the brain of MACS2. The sequence of nef clone 6I from viral isolate MACS3-Br is closely related to nef sequences from lymph node tissue and is highly divergent from nef sequences amplified directly from MACS3 brain. In contrast, the sequences of nef clones 5C and 7D from viral isolates MACS2-LN and MACS3-LN were closely related to nef sequences amplified directly from the DNA of MACS2 or MACS3 lymph node tissue. Together, these data indicate that nef sequences from primary viruses isolated by brain-PBMC coculture are not necessarily representative of nef sequences in the brain and suggest that viral selection or adaptation to PBMC may have occurred during brain coculture.
Figure 1. Phylogenetic analysis of full-length HIV-1 nef nucleotide sequences from autopsy tissue samples and viral isolates from AIDS patients MACS2 (A) and MACS3 (B).

Nef sequences amplified directly from frontal lobe, spleen, and lymph node tissues are color-coded pink, green and black, respectively. Nef sequences from brain and lymph node viral isolates obtained by PBMC co-culture are color-coded orange and blue, respectively. *MACS2-br isolate clones all contained a frameshift resulting in truncation after 60 amino acids and undetectable protein expression. To determine the phylogenetic relationship of viral isolate MACS2-br to other MACS2 sequences, the entire region of the MACS2-br 4G sequence corresponding to the intact nef ORF was used in the alignment. Numbers associated with each branch are bootstrap values, which represent the number of trees, out of 1000 replicates performed, in which the same branching order was found. Only values above 700 for the major branches are shown. Branch lengths are proportional to amount of sequence divergence. Scale bars indicate 1% sequence divergence. An ellipse indicates tissue-derived clones used in functional analyses. Isolate-derived clones analyzed previously (Agopian et al., 2006) are indicated by a rectangle.
Nef sequences in brain versus lymphoid tissue are compartmentalized
Phylogenetic analysis revealed distinct compartmentalization of tissue-derived brain versus lymphoid nef sequences within each patient (Figure 1). Nef sequences amplified directly from 2 different sections of MACS2 frontal lobe tissue showed very little sequence diversity (Figure 1A). The low amount of MACS2 brain nef sequence diversity is consistent with the low HIV-1 copy numbers in these MACS2 brain tissue samples. Nef sequences from MACS2 lymph node and spleen were more diverse and were well interleaved, indicating a lack of compartmentalization between these 2 lymphoid tissues. Thus, in patient MACS2, nef sequences are compartmentalized between brain and lymphoid tissues. In patient MACS3, 8 nef sequences amplified directly from brain were highly divergent from sequences amplified from lymph node tissue (Figure 1B). These sequences comprise a branch of the MACS3 phylogenetic tree that is represented solely by nef sequences from brain tissue. Therefore we designated these 8 sequences as ‘true’ or ‘unique’ brain Nefs. Four nef sequences amplified from MACS3 brain tissue were identical or nearly identical to sequences amplified from lymph node. Thus, patient MACS3 demonstrates both compartmentalized brain nef sequences and nef sequences shared by lymph node and brain. Together, phylogenetic data from both late-stage AIDS patients MACS2 and MACS3 demonstrate compartmentalization of nef sequences in the brain versus lymphoid tissues.
Conservation of motifs in representative tissue-derived nef sequences
To test for functional differences between Nef sequences from brain versus lymphoid tissue, 2 to 3 representative nef sequences from each tissue were selected (circled in Figure 1). These Nefs were chosen so that all major branches of each phylogenetic tree were represented. Figure 2 shows an amino acid sequence alignment of these representative Nefs. Well-conserved motifs in these Nefs include the myristoylation signal (Geyer et al., 2001), the PxxP SH3-binding region (Saksela et al., 1995), the putative CD4 binding site (Grzesiek et al., 1996), and motifs in the flexible loop proposed to be important for association with the endocytic machinery including β-COP (Piguet et al., 1999a), adaptor proteins AP-1/2/3 (Bresnahan et al., 1998; Craig et al., 1998; Greenberg et al., 1998a) and vacuolar ATPase (Lu et al., 1998). More variable motifs in our primary Nefs are the PACS-1 binding motif (Piguet et al., 2000), residues 85, 89, 187, 188, and 191 important for Pak2 association (Agopian et al., 2006; O’Neill et al., 2006), and the FPD121 motif implicated in thioesterase binding (Cohen et al., 2000). Analysis of Nef expression in 293T cells by Western blotting suggests that except for 16N, these Nef proteins are expressed at levels similar to those of control Nef proteins SF2 and NL4-3 (see Figures 3B, 4B, and 5A and B). Nef 16N contains the variation T80I, which disrupts a consensus phosphate acceptor target for protein kinase C (Shugars et al., 1993). Mutation of this highly conserved threonine has been shown to reduce protein stability (Aiken et al., 1996). Although Western blotting with three different polyclonal sera suggests that 16N is expressed at lower levels compared to the other Nef alleles, we cannot rule out the possibility that 16N is well-expressed but poorly recognized by these sera. All five MACS3 brain Nef sequences closely related to 16N carry this T80I variation.
Figure 2. Amino acid alignment of primary HIV-1 Nef clones with the Clade B consensus.
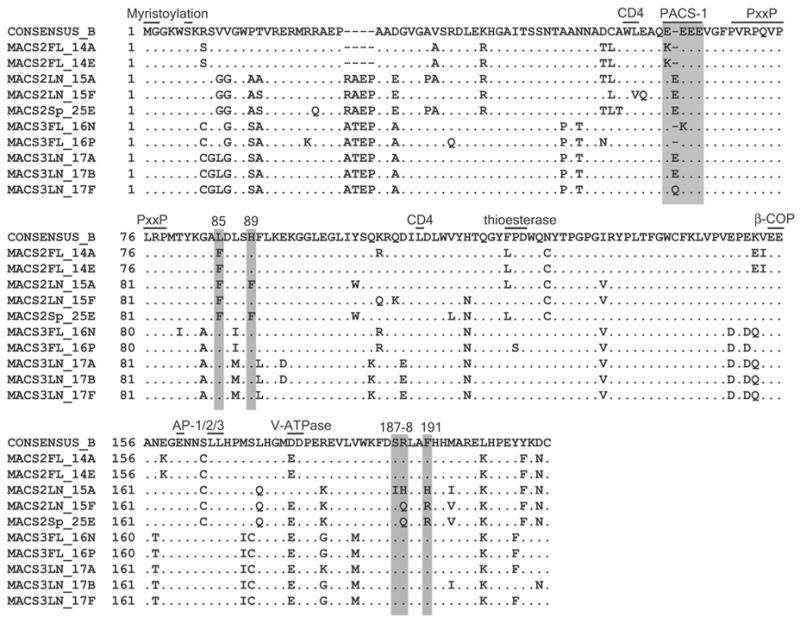
The PACS-1 binding site, important for MHC downregulation (Piguet et al., 2000), and residues 85, 89, 187, 188 and 191, important for Pak2 association (Agopian et al., 2006; O’Neill et al., 2006) are shaded. Bars mark the locations of other proposed Nef domains, including the myristoylation signal (Geyer et al., 2001), the PxxP SH3-binding region (Saksela et al., 1995), the putative CD4 binding site (Grzesiek et al., 1996), and the putative binding sites for thioesterase (Cohen et al., 2000), and the endocytic machinery components β-COP (Piguet et al., 1999a), adaptor proteins AP-1/2/3 (Bresnahan et al., 1998; Craig et al., 1998; Greenberg et al., 1998a) and vacuolar ATPase (Lu et al., 1998). The Clade B consensus is from the LANL sequence database (August 2004).
Figure 3. Downregulation of cell surface CD4 is conserved in primary Nef alleles from brain and lymphoid tissues.

(A) Quantitation of flow cytometric analysis of cell surface CD4 in HIJ cells co-transfected with pCDNA3-EGFP and either pCR3.1 (Vector) or pCR3.1-Nef plasmids as indicated. Relative surface CD4 was calculated from the geometric mean PE fluorescence in GFP-positive cells as described in Materials and Methods and is shown relative to GFP-positive vector-transfected cells. Averages are from 3 independent experiments. Error bars represent standard deviations. (B) Nef expression in HIJ cells assayed for CD4 downregulation is confirmed by Western blot. Lysates were split and identical membranes were blotted in parallel with either rabbit anti-Nef #188 (raised to Bru) or rabbit anti-Nef #2949. (C) Primary FACS data for flow cytometric analysis of cell surface CD4 in transfected HIJ cells. Vertical lines indicate the threshold for GFP positive cells. Data shown are representative of three independent experiments.
Figure 4. Downregulation of cell surface MHC-I by primary Nef proteins from brain and lymphoid tissues.
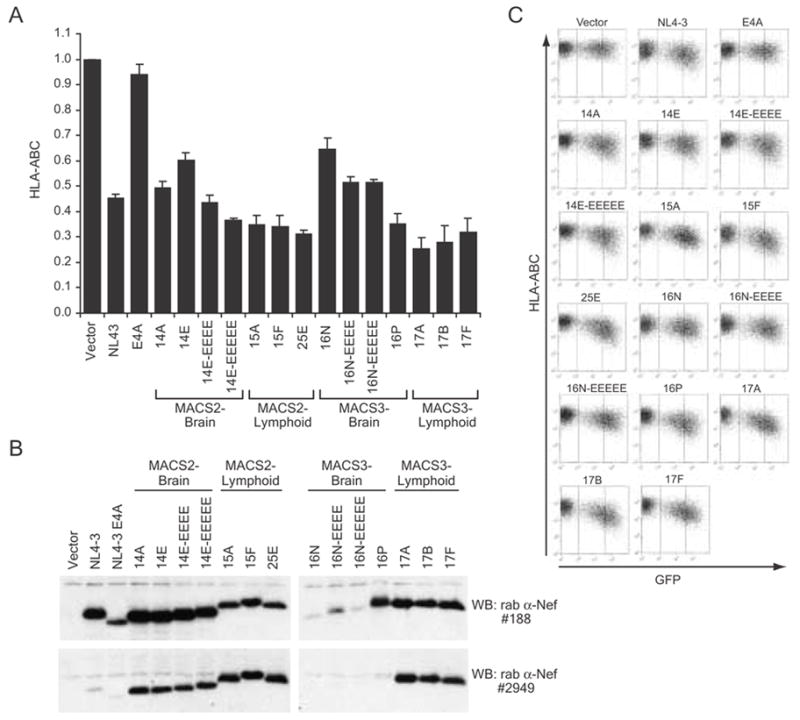
(A) Quantitation of flow cytometric analysis of cell surface HLA-ABC in Jurkat T-Antigen cells co-transfected with pCDNA3-EGFP and either pCR3.1 (Vector) or pCR3.1-Nef plasmids as indicated. Relative surface HLA-ABC was calculated from the geometric mean PE fluorescence in GFP-high-expressing cells as described in Materials and methods and is shown relative to GFP-high vector-transfected cells. Averages are from 2 to 5 independent experiments. Error bars represent standard deviations. (B) Nef expression in Jurkat T-Antigen cells used for MHC-I downregulation assays confirmed by Western blot. Lysates were split and identical membranes were blotted in parallel with either rabbit anti-Nef #188 (raised to Bru) or rabbit anti-Nef #2949. (C) Primary FACS data for flow cytometric analysis of cell surface MHC-I in transfected Jurkat T-Antigen cells. Vertical lines indicate the thresholds for GFP-negative, GFP-low and GFP-high expressing cells. Data shown are representative of two to five independent experiments.
Figure 5. Association of primary HIV-1 Nef proteins with activated Pak2 is highly variable.

(A) In vitro kinase assay (IVKA). 293T cells transiently expressing GFP and the indicated Nef proteins were immunoprecipitated with sheep-anti-Nef (raised to SF2 Nef) and assayed by IVKA (top panel) as described in Materials and methods. SF2 Nef and MACS2 5C Nef, previously demonstrated to strongly activate Pak2 (Arora et al., 2000)and (Agopian et al., 2006), are included as positive controls. An arrowhead indicates the 62 kDa band corresponding to Pak2. Whole cell lysates used for IVKA immunoprecipitations were immunoblotted with rabbit-anti-Nef antibody #2949 or sheep anti-Nef (bottom panels). (B) 293T cells transiently expressing GFP and the indicated Nef proteins were immunoprecipitated with sheep-anti-Nef and Western blotted with rabbit anti-Nef #2949 to confirm efficient Nef immunoprecipitation (top panel). Input whole cell lysates were immunoblotted with anti-Nef antibody #2949 (middle panel) or anti-GFP (bottom panel).
CD4 downregulation is highly conserved
To examine the ability of brain and lymphoid Nef proteins to downregulate CD4, we co-transfected Nef and GFP plasmids into HIJ cells (MAGI cells expressing high levels of CD4) and analyzed cells for surface expression of CD4 by flow cytometry (Figure 3). The LL164AA mutation, previously shown to abolish CD4 downregulation, was made in the background of NL4-3 Nef and was included as a negative control (Bresnahan et al., 1998; Craig et al., 1998; Greenberg et al., 1998a). We also included the EEEE65/AAAA (E4A) mutant of NL4-3 Nef which is defective for MHC-I downregulation but efficiently downregulates CD4 (Piguet et al., 2000). CD4 downregulation activity was highly conserved among primary Nefs, with all 10 clones showing at least some CD4 downregulation activity. Furthermore, 9 out of 10 primary clones downregulated CD4 at least as strongly as NL4-3 Nef. Six of these clones downregulated CD4 more strongly than NL4-3 Nef (p< 0.05 by Student’s two-tailed t test). Only one primary clone, MACS3 brain-derived Nef clone 16P, had weaker CD4 downregulation activity than NL4-3 Nef (p<0.05). Nef 16P also had less than half the activity of the other primary Nef clones (p< 0.001). MACS3 brain-derived Nef clone 16N strongly downregulated CD4, despite an apparent lower expression level suggested by Western blot (Figure 3B). Moderately lower levels of Nef 16N may be sufficient for strong CD4 downregulation activity, since this function is induced even at lower concentrations of Nef (Figure 3C and (Liu et al., 2001)). Alternatively, Nef 16N may be expressed at levels comparable to those of the other Nef alleles but poorly recognized by the polyclonal sera. Overall, there was no significant difference in CD4 downregulation activity between brain and lymphoid Nef clones. Together, these data demonstrate that CD4 downregulation is a highly conserved phenotype of primary Nef proteins from brain and lymphoid tissues of late stage AIDS patients.
All tissue-derived Nefs downregulate MHC-I but brain-derived Nefs have reduced MHC-I downregulation
To examine the ability of brain and lymphoid Nefs to downregulate MHC-I, we co-transfected Nef and GFP plasmids into Jurkat-T-Antigen cells and analyzed cells for surface expression of HLA-ABC by flow cytometry. MHC-I downregulation by Nef is generally less efficient than CD4 downregulation, with higher Nef expression levels required to induce MHC-I downregulation. Therefore, we only analyzed high-expressing cells, marked by high GFP levels. As a negative control, we included the EEEE65/AAAA (E4A) mutant of NL4-3, which has been shown to abolish MHC-I downregulation (Piguet et al., 2000). All primary Nef clones downregulated MHC-I (Figure 4). However, as a group, brain-derived Nef clones (14A, 14E, 16N, 16P) induced less MHC-I downregulation than lymphoid-derived Nef clones (15A, 15F, 25E, 17A, 17B, 17F) (Wilcoxon rank sum test, two-sided p-value = 0.01). Furthermore, all lymphoid-derived Nef clones downregulated MHC-I more strongly than did NL4-3 Nef (p values < 0.05 by Student’s t test for each Nef versus NL4-3). MACS2 brain Nef alleles were expressed at levels comparable to MACS2 lymphoid Nef alleles, suggesting that their reduced capacity to downregulate MHC-I is due to a specific functional defect (Figure 4B). However, MACS3 brain Nef expression levels appeared to be lower than MACS3 lymphoid Nef expression levels (Figure 4B and data not shown). Therefore, it is unclear whether the reduced ability of MACS3 brain Nef clones to downregulate MHC-I is due to a specific functional defect or simply to lower expression levels. Nevertheless, these data demonstrate that while MHC-I downregulation is conserved in primary brain- and lymphoid-derived Nef clones, brain-derived Nefs are reduced in their ability to downregulate MHC-I relative to lymphoid Nefs in these 2 late stage AIDS patients.
Motifs KEEE- or EKEE- at the PACS-1 binding site represent brain-specific signature patterns and contribute to reduced MHC-I downregulation
Subtle differences in the PACS-1 binding site, which is important for MHC-I downregulation (Blagoveshchenskaya et al., 2002; Piguet et al., 2000), were observed in brain-versus lymphoid-derived Nef clones (Figure 2). Comparison of all sequences across the data set from both patients using Viral Epidemiology Signature Pattern Analysis (VESPA) identified only one brain-specific pattern in Nef clones from both patients. A high frequency of Nef clones unique to brain had only 3 glutamic acids (EKEE-65 or KEEE-65) in the PACS-1 binding site (Table 1). In contrast, the majority of Nef sequences from lymphoid tissue had 5 glutamic acids (EEEEE65). Representative of this difference, brain-derived Nef clones 14A, 14E and 16N have either the KEEE- or EKEE- motifs in their PACS-1 binding site, while 5 of 6 representative lymphoid Nef clones have 5 glutamic acids. The KEEE- or EKEE- sequences found in these brain-derived Nef clones are rare among Clade B Nef sequences, with only 0.88% or 0.44% of Nef sequences in the LANL database containing these sequences, respectively (Table 2). Therefore, we hypothesized that the KEEE- or EKEE- motifs represent a rare brain-specific signature pattern in these 2 patients that results in reduced ability to downregulate MHC-I.
Table 1.
Frequency of Motifs at PACS-1 Binding Site in Tissue-Derived Nef Sequences a
| Patient | Brain | Lymphoid |
|---|---|---|
| MACS2 | 18/18 KEEE- (14A, 14E) | 1/24 EEEEK
23/24 EEEEE (15A, 15F, 25E) |
| MACS3 | 6/8 EKEE- (16N)
2/8 EEEE- (16P) |
19/34 EQEEE (17F)
15/34 EEEEE (17A, 17B) |
| Total | 18/26 KEEE-
6/26 EKEE- 2/26 EEEE- |
38/58 EEEEE
19/58 EQEEE 1/58 EEEEK |
Nef sequences unique to brain were compared to sequences from lymph node and spleen. VESPA analysis indicated signature sequences at positions 62–66 which distinguished brain-derived Nef sequences from lymphoid-derived Nef sequences. Sequence frequencies in each tissue are shown. Representative Nef clones used in functional assays are in parentheses.
Table 2.
PACS-1 Binding Site Motifs in Clade B Nef Sequencesa
| Nef 62–65 | HIV Clade B(n=455) |
|---|---|
| EEEE- | 62.7% |
| EDEE- | 12.1% |
| KEEE- | 0.88% |
| EKEE- | 0.44% |
| EEEEE | 9.27% |
| Otherb | 14.61% |
Clade B Nef sequences deposited in the LANL database
Includes approximately 23 low-frequency variants
To determine if the EKEE or KEEE motifs influence the ability of the primary Nef clones to downregulate MHC-I, mutations were made in the MACS2FL-14E and MACS3FL-16N backgrounds to restore the more common EEEE motif to these Nef proteins. Additionally, a fifth glutamic acid was inserted into the PACS-1 binding sites of these Nef clones to determine if an increased number of glutamic acids in the PACS-1 binding site enhances MHC-I downregulation activity. Nef mutant 14E-EEEE downregulated MHC-I more strongly than Nef 14E (p< 0.01 by Student’s t Test) and Nef 14E-EEEEE was even more potent relative to parental MACS2FL-14E (p< 0.001) (Figure 4A). In fact, there was no significant difference in downregulation of MHC-I by 14E-EEEEE versus MACS2 lymphoid-derived Nef clones 15A and 15F, which contain the EEEEE motif in their PACS-1 binding sites. Nef 16N-EEEE also downregulated MHC-I more strongly than did the parental clone 16N. However, Western blot analyses suggest that this Nef is expressed at higher levels than clone 16N. Therefore, it is not possible to conclude whether the mutation at the PACS-1 binding site specifically increased MHC-I downregulation activity. However, Nef 16N-EEEEE appeared to be expressed at levels similar to those of 16N, but downregulated MHC-I more strongly than 16N (p< 0.01). Since the expression of 16N appears to be low, it is difficult to compare its MHC-I downregulation activity directly to that of the MACS3 lymphoid-derived Nef clones. Nevertheless, the 14E and 16N mutagenesis data support the hypothesis that 5 glutamic acids in the PACS-1 binding site result in the strongest downregulation of MHC-1, while the KEEE or EKEE motifs reduce MHC-I downregulation relative to the more common EEEE motif. Therefore, the reduced MHC-I downregulation activity of the brain-derived Nef clones in these 2 patients is likely a consequence of their KEEE or EKEE motifs and can be enhanced by mutation of these motifs to the more potent EEEE or EEEEE motifs.
Pak2 association is variable, and does not correlate with brain or lymphoid origin of Nef sequences
To determine whether association with activated Pak2 is conserved in primary brain and lymphoid-derived Nef proteins, representative Nef clones were tested for their ability to immunoprecipitate activated Pak2 by in vitro kinase assay (IVKA) (Figure 5A). SF2 Nef and MACS2-LN isolate Nef clone 5C were used as positive controls (Agopian et al., 2006; Foster et al., 2001). 5C Nef associates with a very high amount of activated Pak2, while SF2 Nef is commonly used as a positive control for this assay (Arora et al., 2000; Foster et al., 2001; Luo and Garcia, 1996; Sawai et al., 1994). Of 10 tissue-derived Nef clones tested, 7 clones did not coimmunoprecipitate detectable Pak2 activity (Figure 5A). Two tissue-derived Nef clones (MACS2LN-15A and MACS3FL-16P) coimmunoprecipitated more activated Pak2 than did the positive control SF2 Nef. Nef clone 16N coimmunoprecipitated a small amount of activated Pak2. Although Western blots with three different polyclonal antibodies suggest that Nef clone 16N has a lower expression level relative to the other Nef clones (see Figs. 3B, 4B, and 5A), lower expression alone cannot account entirely for the small amount of activated Pak2 immunoprecipitated by Nef 16N. To confirm that all Nef proteins were efficiently immunoprecipitated, anti-Nef Western blots were performed on anti-Nef immunoprecipitates (Figure 5B). All Nef clones except 16N appeared to be efficiently immunoprecipitated. Since 16N appears to have a lower expression level than that of the other Nef alleles and is poorly recognized by the rabbit polyclonal antibody, it is likely that 16N was efficiently immunoprecipitated but its detection by Western blot was limited by a lower concentration of 16N in the cell lysate. Out of 10 tissue-derived Nef clones, we consider 2 to strongly associate with activated Pak2 and 1 to weakly associate with activated Pak2. Of MACS2 tissue-derived Nef clones, only 1 lymphoid-derived and no brain-derived Nef associated with activated Pak2. For MACS3, both brain-derived Nef clones but none of the lymphoid-derived clones associated with activated Pak2. These results demonstrate that the ability to coimmunoprecipitate activated Pak2 is not well conserved in primary Nef alleles from late stage AIDS patients. Furthermore, the ability of Nef sequences to associate with activated Pak2 did not correlate with either brain or lymphoid tissue origin in these 2 patients.
The ability of tissue-derived Nef clones to associate with activated Pak2 appeared to segregate with distinct branches of the phylogenetic trees (Figure 6). MACS2 Nef alleles 15A and 5C immunoprecipitated a very high amount of activated Pak2 and share motif F89I187H188H191 at residues critical for Pak2 association (Figure 2 and (Agopian et al., 2006)), while clones 14A, 14E, 15F and 25E differ at some or all of these residues. Unlike most Clade B Nefs, all Nef clones from MACS2 have a phenylalanine at position 85, which is also critical for Pak2 association (Agopian et al., 2006). It is possible that in the context of F85 only Nef clones with the motif F89I187H188H191 associate with activated Pak2. If so, we predict that all MACS2 Nef alleles with motif F89I187H188H191 (in black in Figure 6A) would associate with activated Pak2, while other Nefs that differ at these residues (in blue in Figure 6B) would be defective for this function. In patient MACS3, Nef clones 17A, 17B, and 17F have no variations in residues previously identified as specifically critical for Pak2 activation (Figure 2). However, these alleles, like many Nef alleles from patient MACS3, have the variation F90L. Residue 90 of Nef is a phenylalanine in the clade B consensus sequence. F90 interacts with the RT loop of SH3 domains (Lee et al., 1996; Lim et al., 1997) and mutation of this residue or its corresponding residue in SIV, F122, reduced or abolished association of Nef with Hck and NAK/Pak2 (Khan et al., 1998; Manninen et al., 1998). Therefore, the F90L variation in most MACS3 Nefs (shown in blue in Figure 6B) would be expected to abolish Pak2 association. Barring other deleterious mutations, only the MACS3 alleles with F90 (shown in black in Figure 6B) have the potential for Pak2 association. Thus, about one-third of all Nefs from patients MACS2 and MACS3 are predicted to associate with activated Pak2 based on sequence analysis.
Figure 6. Phylogenetic analysis of HIV-1 Nef amino acid sequences from AIDS patients MACS2 (A) and MACS3 (B).

(A) MACS2 Nef proteins containing the F89I187H188H191 motif are colored black. Nef clones containing other motifs at these residues are colored blue. (B) MACS3 Nef proteins containing the Clade B consensus phenylalanine at residue 90 are colored black and Nef clones with a leucine at position 90 are colored blue. Bootstrap values above 700 (out of 1000) are shown. Scale bars represent 1% amino acid sequence divergence. Ellipses or rectangles indicate clones analyzed for CD4 and MHC-I downregulation and Pak2 association in this study or previously (Agopian et al., 2006).
Discussion
This study provides evidence for distinct compartmentalization of nef genes from brain and lymphoid tissues of 2 patients with AIDS and HIV-1-associated dementia. CD4 downregulation activity was highly conserved among primary Nefs from brain and lymphoid tissues; 9 of 10 Nef clones downregulated CD4 strongly, in most cases more strongly than NL4-3 Nef, and one downregulated CD4 weakly. MHC-I downregulation was also conserved in these primary Nef clones. However, MHC-I downregulation by brain-derived Nefs was weaker than downregulation by lymphoid-derived Nefs. The motifs KEEE- or EKEE- at the PACS-1 binding site represented brain-specific signature patterns in these 2 patients that contributed to the reduced MHC-I downregulation ability of brain-derived Nefs that contained these motifs. In contrast to CD4 and MHC-I downregulation, Pak2 association was not well conserved. Three of 10 tissue-derived Nefs coimmunoprecipitated detectable activated Pak2, and only 2 of these were strongly associated with Pak2. Thus, CD4 and MHC-I downregulation are highly conserved Nef functions, while Pak2 association is variable in Nef clones from these 2 late stage AIDS patients.
Nef genes from viruses isolated from brain by coculture with PBMC were more closely related to sequences amplified from lymphoid tissue than from brain tissue, suggesting that viral selection or adaptation occurred during coculture. Thus, future studies of brain-specific genes should avoid isolating virus by co-culture with PBMC. Four brain-derived MACS3 sequences were identical to sequences from lymph node. These nef sequences could represent HIV-1 neuroinvasion during late stage AIDS after the blood-brain barrier has been disrupted, contamination by blood vessels that contain blood-derived or lymphoid Nef sequences, or archived ancestral sequences. Two main hypotheses have been proposed regarding the source of HIV-1 divergence within the CNS (McCrossan et al., 2006): 1) divergence is a result of early entry followed by isolation and genetic divergence; and 2) HIV-1 in the brain is a subpopulation of macrophage-tropic virus present in macrophages throughout the body, representing a macrophage-specific rather than brain-specific viral subpopulation. Thus, these 4 nef sequences amplified from brain that are similar to lymphoid-derived Nefs could represent archival HIV sequences in the brain present prior to genetic divergence or sequences from replicating macrophage-tropic subset of virus that is also present in lymphoid tissues.
CD4 downregulation activity was highly conserved in brain and lymphoid Nef alleles, consistent with previous reports that found 75–100% conservation of CD4 downregulation activity in Nefs from rapid progressors and late stage AIDS patients (Casartelli et al., 2003; Michael et al., 1995; Ratner et al., 1996; Tobiume et al., 2002). Brain-derived MACS3 Nef 16P was the only Nef with reduced CD4 downregulation. In Nef 16P, the WL58 CD4 binding site and the motifs in the flexible loop critical for association with the endocytic machinery are all conserved (Geyer et al., 2001). However, 16P contains the variation P122S. Mutation of proline122 dramatically affects both downregulation of CD4 and association with thioesterase (Cohen et al., 2000; Liu et al., 2000). Therefore, the P122S change may account for the reduced CD4 downregulation activity of Nef 16P.
By increasing the efficiency of viral particle release and subsequent engagement of CD4 receptors on target cells, downregulation of cell surface CD4 is one mechanism through which Nef enhances viral infectivity (Lama, 2003; Lama et al., 1999; Ross et al., 1999). CD4 downregulation is the most dramatic Nef function, and possibly the most important. Therefore, it is not surprising that this function is so highly conserved in our primary Nef alleles. It is important to note the strong conservation of CD4 downregulation in the brain, where target cells such as macrophages and microglia have very low CD4 levels. Our data, together with previous reports showing that even low levels of CD4, similar to those found in primary macrophages, significantly reduce the infectivity of HIV particles (Cortes et al., 2002) and that viruses expressing Nef alleles defective in CD4 downregulation exhibit delayed replication kinetics and reduced infectivity in primary macrophages (Brown et al., 2004) suggest that Nef-mediated CD4 downregulation is a critically important function in both high-CD4-expressing lymphocytes and low-CD4-expressing macrophages. These data underscore the importance of CD4 downregulation in the HIV life cycle.
All tissue-derived Nef alleles downregulated MHC-I. This finding is consistent with previous reports showing 90–100% conservation of this activity in progressing and late stage AIDS patients (Casartelli et al., 2003; Tobiume et al., 2002). The preservation of MHC-I downregulation in Nefs from these late stage AIDS patients suggests that MHC –I downregulation is an important function in the life cycle of HIV-1. However, in the 2 AIDS patients studied here, MHC-I downregulation by brain-derived Nefs was weaker than downregulation by lymphoid-derived Nefs. Furthermore, we demonstrated that motifs KEEE- or EKEE- at the PACS-1 binding site represented brain-specific signature patterns in these patients. Mutational analysis confirmed that KEEE-65 or EKEE-65 contributed to the reduced MHC-I downregulation ability of brain-derived Nefs from patients MACS2 and MACS3. To our knowledge, this is the first report of a functional difference between brain- and lymphoid-derived Nefs.
The motifs KEEE-65 or EKEE-65 are found in only 1.32% of 455 Clade B Nefs in the LANL database and only 1.2% of 1643 Clade B Nefs in the database of Foster and colleagues (O’Neill et al., 2006). Since these rare motifs were found at a high frequency in the brain but not lymphoid tissues of the 2 AIDS patients studied here, we hypothesized that these motifs might represent brain-specific signatures in some individuals with AIDS. In one previous report, 5 Nef clones derived from in vitro infection of PBMC with HIV-1 isolated from the cerebrospinal fluid of an AIDS dementia patient were found to have the motif KEEE (McPhee et al., 1998). In contrast, all 12 sequences amplified directly from uncultured brain tissue by Saksena et al. and the 2 sequences amplified directly from uncultured brain tissue by Li et al. contain the common EEEE motif (Li et al., 1992; Saksena et al., 1997). Furthermore, 10 of 11 sequences amplified directly from brain by Van Marle et. al. contain EEEE or EDEE motifs (van Marle et al., 2004). Therefore, EKEE and KEEE are associated with brain-derived Nefs in only a subset of AIDS patients. A search of the database of 1643 Nef alleles (O’Neill et al., 2006) revealed that 6 of 11 Nef sequences with motif KEEE were isolated from PBMC or plasma RNA (Asamitsu et al., 1999; Fang et al., 2001; Fang et al., 1999; Hahn et al., 2003; Kirchhoff et al., 1999), and 3 of 9 sequences with motif EKEE were isolated from PBMC (Asamitsu et al., 1999), while the source of the other 6 was not specified (Cho, and Ahn, direct submission, 2005 and Ashton, L et al unpublished, 2002). Therefore, the EKEE and KEEE motifs are not excluded from blood or lymphoid tissues nor are they a common brain signature pattern. Since immune surveillance is reduced in the CNS compared to non-CNS tissues, the brain may be a more permissive tissue site for maintenance of Nef alleles with weaker MHC-I downregulation.
In contrast to CD4 and MHC-I downregulation, Pak2 association was not well conserved in primary Nef alleles from 2 late stage AIDS patients. Only 3 of 10 tissue-derived Nefs immunoprecipitated Pak2. We found a higher frequency of Nef alleles positive for Pak2 activation (5 of 7) in Nef alleles from primary viral isolates UK1-Br, MACS1-Br, MACS1-Sp, MACS2-LN (5C), MACS3-br (6I), MACS3-LN (7D) and slow progressor Δ32-Blood [(Agopian et al., 2006; Gorry et al., 2001) and unpublished data]. Other studies also found higher rates of association with activated Pak2 among primary Nef alleles. In one study, 3 of 5 subtype B Nefs were positive for Pak2 activation and a second study found that 1 of 2 subtype B and 5 of 5 subtype E HIV-1 Nefs activated Pak2 (Foster et al., 2001; Huang et al., 1995; O’Neill et al., 2006). Furthermore, 5 of 5 Group O and N and 3 of 6 SIVcpz Nefs were strongly positive for Pak2 activation (Kirchhoff et al., 2004). The methods used by Kirchhoff et al appear to be more sensitive for detecting Pak2 association, since NL4-3 Nef coimmunoprecipitates a stronger band of Pak2 activity in their assay compared to ours (Arora et al., 2000; Luo and Garcia, 1996). Our IVKA detects Nef association with endogenous Pak2, whereas the IVKA used by Kirchhoff et. al. detects association with exogenous Pak2 in cells cotransfected with activated Cdc42V12 (Pulkkinen et al., 2004).
The significance of the low frequency of Pak2 association in primary Nef alleles in our study remains unclear. Compilation of results of previous studies of Pak2 association suggests a frequency of 80% (16/20) positive in HIV-1 Nef alleles and 100% (7/7) positive in SIV Nef alleles (Foster et al., 2001; Kirchhoff et al., 2004; Luo and Garcia, 1996; O’Neill et al., 2006). An important distinction between our study and previous studies is the number of independent nef alleles amplified by limiting dilution from each patient and our selection, based on phylogenetics, of representative alleles to be tested in functional assays. Other studies that analyzed Nefs from several patients typically selected only one or a few alleles representing each patient. Further studies examining large numbers of Nef clones from several patients are needed to determine the true proportion of Nef alleles that efficiently associate with and/or activate Pak2 in vivo.
If the true frequency of Nef alleles positive for Pak2 association in vivo is indeed low, then what is the significance of Nef-Pak2 association during the course of HIV infection? Evidence of compensation and co-variation between Nef residues critical for association with Pak2 suggests that Nef-mediated activation of Pak2 confers a selective advantage during the course of HIV infection (Agopian et al., 2006; O’Neill et al., 2006). Nef-mediated activation of Pak2 might be critical for infection of particular cell types during a specific stage(s) of infection. In the present study, tissue samples from patients MACS2 and MACS3 were collected during autopsy and therefore represent end-stage AIDS. If Pak2 activation is indeed critical for enhancement of T cell activation, as previously proposed (Fackler et al., 1999; Lu et al., 1996; Vilhardt et al., 2002; Wei et al., 2005; Wolf et al., 2001), then this function might be particularly important during transmission and/or early stages of infection when resting T cells are important targets for infection in genital mucosa and gut-associated lymphoid tissues (Pope and Haase, 2003). Nef residues important for Pak2 activation are also targets for CTLs (Los Alamos National Laboratories Database). Thus, the high level of variation at these sites in MACS2 could also reflect CTL escape. Further studies are needed to determine if the frequency of Nef alleles positive for Pak2 activation differs between specific types of infected cells or during the course of infection. The primary Nef alleles described here will facilitate future studies to investigate the importance of Nef-Pak2 association and other Nef functions for HIV-1 replication and pathogenesis.
Materials and Methods
Amplification of HIV-1 Nef alleles from autopsy tissue
Genomic DNA was extracted from autopsy samples of uncultured brain, lymph node and spleen tissue from patients MACS2 and MACS3 using the Gentra Puregene DNA Purification kit (Gentra). Patients MACS2 and MACS3 were participants in the Chicago Component of the Multicenter AIDS Cohort Study who had late stage AIDS and HIV-1-associated dementia (Gorry et al., 2001). Quantification of HIV-1 gag copy number by real-time PCR was performed using the TaqMan Universal PCR Master Mix (Applied Biosystems, Foster City, CA) and primers CCR5-933R (5′-TGAAGATAAGCCTCACAGCCCT- 3′), CCR5-778F (5′-AGGTCTTCATTACACCTGCAGCTC-3′), CCR5 probe (5′-CAGTCAGTATCAATTCTGGAAGAATTTCCAGACAT-3′), 793-gag F (5′-GGTGCGAGAGCGTCAGTATTAAG-3′), 911-gag R (5′-AGCTCCCTGCTTGCCCATA-3′), and 835-gag probe (5′ TGGGAAAAAATTCGGTTAAGGCCAGGG-3′). Cycling conditions were 95C° (15 min), followed by 45 cycles of 95°C (15 sec), 60°C (1 min), 25°C (2 min). Copies of HIV-1 gag per million cells were 3,460 and 2,713 in 2 different sections of MACS2 frontal lobe, 21,683 in MACS2 lymph node, 6,129 in MACS2 spleen, 1,645 in MACS3 frontal lobe and 252,104 in MACS3 lymph node. The HIV-1 nef gene was amplified by nested PCR performed on 1 μl of undiluted genomic DNA (bulk PCR) or at limiting dilution. Dilutions were determined to be limiting when an average of 3 of 10 PCR reactions were positive. For brain genomic DNA, limiting dilutions ranged from 1:2–1:8. For lymph node and spleen DNA, dilutions ranged from 1:50–1:200. Maximally conserved primer binding sites were selected. To reduce effects of selective amplification, 2 different sets of nested PCR primers were used. Positions of the primer sequences in the HIV-1 HXB2 genome are given in parentheses. For the first set, outer primers were Nef1 (5′-GCCCGAAGGAATAGAAGAAG-3′) (8411–8430) and Nef2 (5′-GCACTCAAGGCAAGCTTTATTGAGGC-3′) (9631–9605). The Nef2 primer was previously published (Huang et al., 1995). Inner primers included the restriction sites BamHI and ClaI (underlined) and were Nef3b.5 (5′-TTAGTGAACGGATCCTTAGCACTTATCTGGG-3′) (8466–8496) and Nef4.5 (5′-GCGGAAAGTCCCTTGTAGCAACATCGATGTCA-3′) (9446–9415). For the second set, outer primers were Nef5′b (5′-GCAGTAGCTGAGGGGACAGATAGGGTTATAG) (8685–8705) and Nef2. Inner primers were HI5′ (5′-CGTCTAGAACATACCTAGAAGAATAAGACAGG (8748–8768) and NFkB3′ (5′-AAAGTCCCCAGCGGAAAGTCCC) (9456–9435). All reactions were performed with Pfu polymerase (Stratagene). The HI5′ primer was previously published (Huang et al., 1995). The cycling conditions for the 50 μl first-round reaction were 94° (2 min), followed by 30 cycles of 94° (15 sec), 57°C (15 sec), 68°C (3 min), and then the final elongation step at 72°C (7 min). 1 μl of the first-round PCR product was used as template for the second-round reaction. The cycling conditions for the second-round primer set were the same except that hybridization was performed at 55°C. PCR products were purified using the Qiaquick PCR purification kit and were either sequenced directly (represented by numbers, e.g. LN-15) or cloned into the pCR3.1 expression vector and sequenced (represented by numbers and letters, e.g. LN-15A). From each tissue, 11 to 34 independent sequences were obtained.
The nucleotide and amino acid sequences from MACS2 frontal lobe (XX), MACS2 lymph node (XX), MACS2 spleen (DQ358015-DQ358047), MACS3 frontal lobe (XX), and MACS3 lymph node (XX) have been submitted to the GenBank sequence database.
Amplification of HIV-1 Nef alleles from viral isolates
The primary HIV-1 viruses MACS2-LN, MACS3-br and MACS3-LN were isolated from patients MACS2 and MACS3 and characterized as described (Gorry et al., 2001). Briefly, autopsy lymph node and brain tissue samples were homogenized and cocultured with CD8-depleted PBMC. Supernatants testing positive for reverse transcriptase activity were filtered and stored at −80°C. From cell pellets of cultured PBMC infected with viral supernatants, DNA was extracted using a Qiagen DNeasy kit. Nef alleles were amplified and cloned from 1 μl undiluted genomic DNA as described above.
The nucleotide and amino acid Nef sequences of viral isolate-derived clones MACS2LN-5C (DQ357219), MACS2LN-5F (DQ358012), MACS2LN-5G (DQ358013), MACS2LN-5H DQ358014), MACS3br-6I (DQ357220), and MACS3LN-7D (DQ357221) have been submitted to GenBank.
Phylogenetic analysis
Sequences from each independent limiting dilution PCR reaction and each clone from bulk PCR were included in the phylogenetic analyses. Alignments were made using Clustal W. Bootstrapped phylogenetic trees were created by the neighbor-joining method using Clustal X.
Plasmids and mutagenesis
The NL4-3 nef allele was amplified from the pNL4-3 proviral plasmid by PCR with primers Nef3b.5 and Nef 4.5 and inserted into pCR3.1 by TA cloning. The SF2 nef allele was cloned into the EcoRI site of vector PCDNA3.1 (Foster et al., 2001). The pCR3.1 vector lacking an insert was used for mock transfections. Mutations were created using the QuikChange site-directed mutagenesis kit (Stratagene) and all mutants were confirmed by sequencing.
Cell culture
293T cells and HIJ cells (HeLa cells retrovirally transduced to express high levels of CD4), provided by David Kabat (Kabat et al., 1994), were maintained in Dulbecco’s modified Eagle medium (DMEM) supplemented with 10% fetal bovine serum and 100 μg/ml penicillin/streptomycin. Jurkat-T-Antigen (J-TAg) cells (Northrop et al., 1993), provided by Heinrich Gottlinger, were maintained in RPMI 1640 supplemented with 10% fetal bovine serum and 100 μg/ml penicillin/streptomycin.
CD4 and MHC-I downregulation assays
For CD4 downregulation assays, HIJ cells were co-transfected with nef and pCDNA3-EGFP expression plasmids using Lipofectamine 2000 (Invitrogen). Cells were harvested after 28 hours and stained for 30 minutes at 4°C with saturating amounts of PE-conjugated mouse anti-CD4 antibody (BD Pharmingen). Stained cells were analyzed by flow cytometry and dead cells were excluded by forward and side scatter (Becton Dickinson FACScan). The geometric mean of CD4-PE fluorescence intensity was calculated for transfected (GFPpos) and untransfected (GFPneg) cell populations. To calculate the relative mean fluorescence intensity (relMFI) of each transfected sample, the following formula was used: . This formula corrects for slight staining variation between stained samples and expresses the CD4 surface levels relative to the mock-transfected negative control.
MHC-I downregulation assays were performed by co-transfecting Jurkat T-Antigen cells with nef and pCDNA3-EGFP plasmids using Lipofectamine 2000. Cells were activated 4 hours later with 1 μg/ml PHA-L and 50 ng/ml PMA. Cells were harvested after 28 hours and stained with a saturating amount of PE-conjugated mouse anti-HLA-ABC antibody (BD Pharmingen). Stained cells were analyzed by flow cytometry as described above. The relative MFI of HLA-ABC-PE fluorescence was calculated for high-GFP-expressing (GFPhigh) using the formula described above.
Western blots
Aliquots of cells used for CD4 and MHC-I downregulation assays were lysed in Cold Lysis Buffer (50 mM Tris HCl pH 7.5, 150 mM NaCl, 10% glycerol, 1% Triton X-100, Complete protease inhibitors (Roche)). Lysates were centrifuged at 14,000 rpm for 20 min at 4°C to remove cell debris. Equal amounts of total protein were run on 12% SDS-PAGE gels and analyzed by Western blotting with rabbit anti-Nef #2949 (AIDS Research and Reference Reagent Program), rabbit anti-Nef Bru #188 (Zazopoulos and Haseltine, 1992), or sheep anti-Nef SF2 (Foster et al., 2001). Aliquots of cell lysates used for the IVKAs were tested for Nef expression by Western blot with sheep polyclonal anti-Nef SF2 serum (1:2000 dilution) as described (Foster et al., 2001) or rabbit anti-Nef #2949 (1:2000 dilution).
In vitro kinase assays (IVKAs)
IVKAs were performed as previously described (Foster et al., 2001). Briefly, 293T cells transfected (Lipofectamine 2000, Invitrogen) with Nef and pN1GFP plasmids were lysed in IVKA Lysis Buffer (50 mM Tris pH 8.0, 100 mM NaCl, 10% glycerol, 0.5% IGEPAL CA-630 (Sigma), 1 mM EDTA, 25 mM NaF, 2 mM Na3VO4, 20 mM beta-glycerophosphate, 1 mM PMSF, 25 mM benzamidine and Complete protease inhibitors (Roche). Cleared whole cell lysates (600 μg) were immunoprecipitated with sheep-anti-Nef serum. Kinase reactions were performed on immunoprecipitates in the presence of [γ32P] ATP and separated by 10% SDS-PAGE gels. Dried gels were exposed to a phosphoimager screen (Packard).
Acknowledgments
We thank R. Desrosiers, H. Gottlinger, M. Farzan, J. Sodroski, J. Wang, and P. Gorry for helpful discussions and technical advice. We also thank D. Kabat for providing HIJ cells, H. Gottlinger for Jurkat T-Antigen cells, and the NIH AIDS Reagent Program for Nef antibody 2949 (donated by R. Swanstrom). This work was supported by NIH NS35734 and NS37277. K.A. was supported in part by a fellowship from the Cancer Research Institute. B.W. and J.V.G. were supported by NIH AI33331 to J.V.G. Core facilities were supported by Center for AIDS Research and DFCI/Harvard Center for Cancer Research grants.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Agopian K, Wei BL, Garcia JV, Gabuzda D. A Hydrophobic Binding Surface on the Human Immunodeficiency Virus Type 1 Nef Core Is Critical for Association with p21-Activated Kinase 2. J Virol. 2006;80(6):3050–61. doi: 10.1128/JVI.80.6.3050-3061.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aiken C, Krause L, Chen YL, Trono D. Mutational analysis of HIV-1 Nef: identification of two mutants that are temperature-sensitive for CD4 downregulation. Virology. 1996;217(1):293–300. doi: 10.1006/viro.1996.0116. [DOI] [PubMed] [Google Scholar]
- Alexander L, Du Z, Rosenzweig M, Jung JU, Desrosiers RC. A role for natural simian immunodeficiency virus and human immunodeficiency virus type 1 nef alleles in lymphocyte activation. J Virol. 1997;71(8):6094–9. doi: 10.1128/jvi.71.8.6094-6099.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arold ST, Baur AS. Dynamic Nef and Nef dynamics: how structure could explain the complex activities of this small HIV protein. Trends Biochem Sci. 2001;26(6):356–63. doi: 10.1016/s0968-0004(01)01846-1. [DOI] [PubMed] [Google Scholar]
- Arora VK, Molina RP, Foster JL, Blakemore JL, Chernoff J, Fredericksen BL, Garcia JV. Lentivirus Nef specifically activates Pak2. J Virol. 2000;74(23):11081–7. doi: 10.1128/jvi.74.23.11081-11087.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Asamitsu K, Morishima T, Tsuchie H, Kurimura T, Okamoto T. Conservation of the central proline-rich (PxxP) motifs of human immunodeficiency virus type 1 Nef protein during the disease progression in two hemophiliac patients. FEBS Lett. 1999;459(3):399–404. doi: 10.1016/s0014-5793(99)01288-0. [DOI] [PubMed] [Google Scholar]
- Bagrodia S, Cerione RA. Pak to the future. Trends Cell Biol. 1999;9(9):350–5. doi: 10.1016/s0962-8924(99)01618-9. [DOI] [PubMed] [Google Scholar]
- Baur AS, Sawai ET, Dazin P, Fantl WJ, Cheng-Mayer C, Peterlin BM. HIV-1 Nef leads to inhibition or activation of T cells depending on its intracellular localization. Immunity. 1994;1(5):373–84. doi: 10.1016/1074-7613(94)90068-x. [DOI] [PubMed] [Google Scholar]
- Blagoveshchenskaya AD, Thomas L, Feliciangeli SF, Hung CH, Thomas G. HIV-1 Nef downregulates MHC-I by a PACS-1- and PI3K-regulated ARF6 endocytic pathway. Cell. 2002;111(6):853–66. doi: 10.1016/s0092-8674(02)01162-5. [DOI] [PubMed] [Google Scholar]
- Bour S, Strebel K. HIV accessory proteins: multifunctional components of a complex system. Adv Pharmacol. 2000;48:75–120. doi: 10.1016/s1054-3589(00)48004-x. [DOI] [PubMed] [Google Scholar]
- Bresnahan PA, Yonemoto W, Ferrell S, Williams-Herman D, Geleziunas R, Greene WC. A dileucine motif in HIV-1 Nef acts as an internalization signal for CD4 downregulation and binds the AP-1 clathrin adaptor. Curr Biol. 1998;8(22):1235–8. doi: 10.1016/s0960-9822(07)00517-9. [DOI] [PubMed] [Google Scholar]
- Brown A, Moghaddam S, Kawano T, Cheng-Mayer C. Multiple human immunodeficiency virus type 1 Nef functions contribute to efficient replication in primary human macrophages. J Gen Virol. 2004;85(Pt 6):1463–9. doi: 10.1099/vir.0.79946-0. [DOI] [PubMed] [Google Scholar]
- Casartelli N, Di Matteo G, Potesta M, Rossi P, Doria M. CD4 and major histocompatibility complex class I downregulation by the human immunodeficiency virus type 1 nef protein in pediatric AIDS progression. J Virol. 2003;77(21):11536–45. doi: 10.1128/JVI.77.21.11536-11545.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang J, Jozwiak R, Wang B, Ng T, Ge YC, Bolton W, Dwyer DE, Randle C, Osborn R, Cunningham AL, Saksena NK. Unique HIV type 1 V3 region sequences derived from six different regions of brain: region-specific evolution within host-determined quasispecies. AIDS Res Hum Retroviruses. 1998;14(1):25–30. doi: 10.1089/aid.1998.14.25. [DOI] [PubMed] [Google Scholar]
- Cohen GB, Rangan VS, Chen BK, Smith S, Baltimore D. The human thioesterase II protein binds to a site on HIV-1 Nef critical for CD4 down-regulation. J Biol Chem. 2000;275(30):23097–105. doi: 10.1074/jbc.M000536200. [DOI] [PubMed] [Google Scholar]
- Collins KL, Chen BK, Kalams SA, Walker BD, Baltimore D. HIV-1 Nef protein protects infected primary cells against killing by cytotoxic T lymphocytes. Nature. 1998;391(6665):397–401. doi: 10.1038/34929. [DOI] [PubMed] [Google Scholar]
- Cortes MJ, Wong-Staal F, Lama J. Cell surface CD4 interferes with the infectivity of HIV-1 particles released from T cells. J Biol Chem. 2002;277(3):1770–9. doi: 10.1074/jbc.M109807200. [DOI] [PubMed] [Google Scholar]
- Craig HM, Pandori MW, Guatelli JC. Interaction of HIV-1 Nef with the cellular dileucine-based sorting pathway is required for CD4 down-regulation and optimal viral infectivity. Proc Natl Acad Sci U S A. 1998;95(19):11229–34. doi: 10.1073/pnas.95.19.11229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Daniels RH, Bokoch GM. p21-activated protein kinase: a crucial component of morphological signaling? Trends Biochem Sci. 1999;24(9):350–5. doi: 10.1016/s0968-0004(99)01442-5. [DOI] [PubMed] [Google Scholar]
- Doms RW, Trono D. The plasma membrane as a combat zone in the HIV battlefield. Genes Dev. 2000;14(21):2677–88. doi: 10.1101/gad.833300. [DOI] [PubMed] [Google Scholar]
- Donaldson YK, Bell JE, Holmes EC, Hughes ES, Brown HK, Simmonds P. In vivo distribution and cytopathology of variants of human immunodeficiency virus type 1 showing restricted sequence variability in the V3 loop. J Virol. 1994;68(9):5991–6005. doi: 10.1128/jvi.68.9.5991-6005.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Du Z, Lang SM, Sasseville VG, Lackner AA, Ilyinskii PO, Daniel MD, Jung JU, Desrosiers RC. Identification of a nef allele that causes lymphocyte activation and acute disease in macaque monkeys. Cell. 1995;82(4):665–74. doi: 10.1016/0092-8674(95)90038-1. [DOI] [PubMed] [Google Scholar]
- Dunfee R, Thomas E, Gorry PR, Wang J, Ancuta P, Gabuzda D. Mechanisms of HIV-1 Neurotropism. Current HIV Research. 2006 doi: 10.2174/157016206777709500. in press. [DOI] [PubMed] [Google Scholar]
- Dyer WB, Ogg GS, Demoitie MA, Jin X, Geczy AF, Rowland-Jones SL, McMichael AJ, Nixon DF, Sullivan JS. Strong human immunodeficiency virus (HIV)-specific cytotoxic T-lymphocyte activity in Sydney Blood Bank Cohort patients infected with nef-defective HIV type 1. J Virol. 1999;73(1):436–43. doi: 10.1128/jvi.73.1.436-443.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fackler OT, Baur AS. Live and let die: Nef functions beyond HIV replication. Immunity. 2002;16(4):493–7. doi: 10.1016/s1074-7613(02)00307-2. [DOI] [PubMed] [Google Scholar]
- Fackler OT, Luo W, Geyer M, Alberts AS, Peterlin BM. Activation of Vav by Nef induces cytoskeletal rearrangements and downstream effector functions. Mol Cell. 1999;3(6):729–39. doi: 10.1016/s1097-2765(01)80005-8. [DOI] [PubMed] [Google Scholar]
- Fang G, Burger H, Chappey C, Rowland-Jones S, Visosky A, Chen CH, Moran T, Townsend L, Murray M, Weiser B. Analysis of transition from long-term nonprogressive to progressive infection identifies sequences that may attenuate HIV type 1. AIDS Res Hum Retroviruses. 2001;17(15):1395–404. doi: 10.1089/088922201753197060. [DOI] [PubMed] [Google Scholar]
- Fang G, Weiser B, Visosky A, Moran T, Burger H. PCR-mediated recombination: a general method applied to construct chimeric infectious molecular clones of plasma-derived HIV-1 RNA. Nat Med. 1999;5(2):239–42. doi: 10.1038/5607. [DOI] [PubMed] [Google Scholar]
- Foster JL, Molina RP, Luo T, Arora VK, Huang Y, Ho DD, Garcia JV. Genetic and functional diversity of human immunodeficiency virus type 1 subtype B Nef primary isolates. J Virol. 2001;75(4):1672–80. doi: 10.1128/JVI.75.4.1672-1680.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gartner S, McDonald RA, Hunter EA, Bouwman F, Liu Y, Popovic M. Gp120 sequence variation in brain and in T-lymphocyte human immunodeficiency virus type 1 primary isolates. J Hum Virol. 1997;1(1):3–18. [PubMed] [Google Scholar]
- Geyer M, Fackler OT, Peterlin BM. Structure--function relationships in HIV-1 Nef. EMBO Rep. 2001;2(7):580–5. doi: 10.1093/embo-reports/kve141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gonzalez-Scarano F, Martin-Garcia J. The neuropathogenesis of AIDS. Nat Rev Immunol. 2005;5(1):69–81. doi: 10.1038/nri1527. [DOI] [PubMed] [Google Scholar]
- Gorry PR, Bristol G, Zack JA, Ritola K, Swanstrom R, Birch CJ, Bell JE, Bannert N, Crawford K, Wang H, Schols D, De Clercq E, Kunstman K, Wolinsky SM, Gabuzda D. Macrophage tropism of human immunodeficiency virus type 1 isolates from brain and lymphoid tissues predicts neurotropism independent of coreceptor specificity. J Virol. 2001;75(21):10073–89. doi: 10.1128/JVI.75.21.10073-10089.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Greenberg M, DeTulleo L, Rapoport I, Skowronski J, Kirchhausen T. A dileucine motif in HIV-1 Nef is essential for sorting into clathrin-coated pits and for downregulation of CD4. Curr Biol. 1998a;8(22):1239–42. doi: 10.1016/s0960-9822(07)00518-0. [DOI] [PubMed] [Google Scholar]
- Greenberg ME, Iafrate AJ, Skowronski J. The SH3 domain-binding surface and an acidic motif in HIV-1 Nef regulate trafficking of class I MHC complexes. Embo J. 1998b;17(10):2777–89. doi: 10.1093/emboj/17.10.2777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grzesiek S, Stahl SJ, Wingfield PT, Bax A. The CD4 determinant for downregulation by HIV-1 Nef directly binds to Nef. Mapping of the Nef binding surface by NMR. Biochemistry. 1996;35(32):10256–61. doi: 10.1021/bi9611164. [DOI] [PubMed] [Google Scholar]
- Hahn T, Ramakrishnan R, Ahmad N. Evaluation of genetic diversity of human immunodeficiency virus type 1 NEF gene associated with vertical transmission. J Biomed Sci. 2003;10(4):436–50. doi: 10.1007/BF02256435. [DOI] [PubMed] [Google Scholar]
- Hanna Z, Kay DG, Rebai N, Guimond A, Jothy S, Jolicoeur P. Nef harbors a major determinant of pathogenicity for an AIDS-like disease induced by HIV-1 in transgenic mice. Cell. 1998;95(2):163–75. doi: 10.1016/s0092-8674(00)81748-1. [DOI] [PubMed] [Google Scholar]
- Harris M. HIV: a new role for Nef in the spread of HIV. Curr Biol. 1999;9(12):R459–61. doi: 10.1016/s0960-9822(99)80282-6. [DOI] [PubMed] [Google Scholar]
- Huang Y, Zhang L, Ho DD. Characterization of nef sequences in long-term survivors of human immunodeficiency virus type 1 infection. J Virol. 1995;69(1):93–100. doi: 10.1128/jvi.69.1.93-100.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hughes ES, Bell JE, Simmonds P. Investigation of the dynamics of the spread of human immunodeficiency virus to brain and other tissues by evolutionary analysis of sequences from the p17gag and env genes. J Virol. 1997;71(2):1272–80. doi: 10.1128/jvi.71.2.1272-1280.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kabat D, Kozak SL, Wehrly K, Chesebro B. Differences in CD4 dependence for infectivity of laboratory-adapted and primary patient isolates of human immunodeficiency virus type 1. J Virol. 1994;68(4):2570–7. doi: 10.1128/jvi.68.4.2570-2577.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaul M, Zheng J, Okamoto S, Gendelman HE, Lipton SA. HIV-1 infection and AIDS: consequences for the central nervous system. Cell Death Differ 12 Suppl. 2005;1:878–92. doi: 10.1038/sj.cdd.4401623. [DOI] [PubMed] [Google Scholar]
- Kestler HW, 3rd, Ringler DJ, Mori K, Panicali DL, Sehgal PK, Daniel MD, Desrosiers RC. Importance of the nef gene for maintenance of high virus loads and for development of AIDS. Cell. 1991;65(4):651–62. doi: 10.1016/0092-8674(91)90097-i. [DOI] [PubMed] [Google Scholar]
- Khan IH, Sawai ET, Antonio E, Weber CJ, Mandell CP, Montbriand P, Luciw PA. Role of the SH3-ligand domain of simian immunodeficiency virus Nef in interaction with Nef-associated kinase and simian AIDS in rhesus macaques. J Virol. 1998;72(7):5820–30. doi: 10.1128/jvi.72.7.5820-5830.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kirchhoff F, Easterbrook PJ, Douglas N, Troop M, Greenough TC, Weber J, Carl S, Sullivan JL, Daniels RS. Sequence variations in human immunodeficiency virus type 1 Nef are associated with different stages of disease. J Virol. 1999;73(7):5497–508. doi: 10.1128/jvi.73.7.5497-5508.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kirchhoff F, Greenough TC, Brettler DB, Sullivan JL, Desrosiers RC. Brief report: absence of intact nef sequences in a long-term survivor with nonprogressive HIV-1 infection. N Engl J Med. 1995;332(4):228–32. doi: 10.1056/NEJM199501263320405. [DOI] [PubMed] [Google Scholar]
- Kirchhoff F, Schindler M, Bailer N, Renkema GH, Saksela K, Knoop V, Muller-Trutwin MC, Santiago ML, Bibollet-Ruche F, Dittmar MT, Heeney JL, Hahn BH, Munch J. Nef proteins from simian immunodeficiency virus-infected chimpanzees interact with p21-activated kinase 2 and modulate cell surface expression of various human receptors. J Virol. 2004;78(13):6864–74. doi: 10.1128/JVI.78.13.6864-6874.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Korber BT, Kunstman KJ, Patterson BK, Furtado M, McEvilly MM, Levy R, Wolinsky SM. Genetic differences between blood- and brain-derived viral sequences from human immunodeficiency virus type 1-infected patients: evidence of conserved elements in the V3 region of the envelope protein of brain-derived sequences. J Virol. 1994;68(11):7467–81. doi: 10.1128/jvi.68.11.7467-7481.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lama J. The physiological relevance of CD4 receptor down-modulation during HIV infection. Curr HIV Res. 2003;1(2):167–84. doi: 10.2174/1570162033485276. [DOI] [PubMed] [Google Scholar]
- Lama J, Mangasarian A, Trono D. Cell-surface expression of CD4 reduces HIV-1 infectivity by blocking Env incorporation in a Nef- and Vpu-inhibitable manner. Curr Biol. 1999;9(12):622–31. doi: 10.1016/s0960-9822(99)80284-x. [DOI] [PubMed] [Google Scholar]
- Learmont J, Tindall B, Evans L, Cunningham A, Cunningham P, Wells J, Penny R, Kaldor J, Cooper DA. Long-term symptomless HIV-1 infection in recipients of blood products from a single donor. Lancet. 1992;340(8824):863–7. doi: 10.1016/0140-6736(92)93281-q. [DOI] [PubMed] [Google Scholar]
- Lee CH, Saksela K, Mirza UA, Chait BT, Kuriyan J. Crystal structure of the conserved core of HIV-1 Nef complexed with a Src family SH3 domain. Cell. 1996;85(6):931–42. doi: 10.1016/s0092-8674(00)81276-3. [DOI] [PubMed] [Google Scholar]
- Li Y, Hui H, Burgess CJ, Price RW, Sharp PM, Hahn BH, Shaw GM. Complete nucleotide sequence, genome organization, and biological properties of human immunodeficiency virus type 1 in vivo: evidence for limited defectiveness and complementation. J Virol. 1992;66(11):6587–600. doi: 10.1128/jvi.66.11.6587-6600.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lim EM, Lagranderie M, Le Grand R, Rauzier J, Gheorghiu M, Gicquel B, Winter N. Recombinant Mycobacterium bovis BCG producing the N-terminal half of SIVmac251 Env antigen induces neutralizing antibodies and cytotoxic T lymphocyte responses in mice and guinea pigs. AIDS Res Hum Retroviruses. 1997;13(18):1573–81. doi: 10.1089/aid.1997.13.1573. [DOI] [PubMed] [Google Scholar]
- Liu LX, Heveker N, Fackler OT, Arold S, Le Gall S, Janvier K, Peterlin BM, Dumas C, Schwartz O, Benichou S, Benarous R. Mutation of a conserved residue (D123) required for oligomerization of human immunodeficiency virus type 1 Nef protein abolishes interaction with human thioesterase and results in impairment of Nef biological functions. J Virol. 2000;74(11):5310–9. doi: 10.1128/jvi.74.11.5310-5319.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu X, Schrager JA, Lange GD, Marsh JW. HIV Nef-mediated cellular phenotypes are differentially expressed as a function of intracellular Nef concentrations. J Biol Chem. 2001;276(35):32763–70. doi: 10.1074/jbc.M101025200. [DOI] [PubMed] [Google Scholar]
- Lu X, Wu X, Plemenitas A, Yu H, Sawai ET, Abo A, Peterlin BM. CDC42 and Rac1 are implicated in the activation of the Nef-associated kinase and replication of HIV-1. Curr Biol. 1996;6(12):1677–84. doi: 10.1016/s0960-9822(02)70792-6. [DOI] [PubMed] [Google Scholar]
- Lu X, Yu H, Liu SH, Brodsky FM, Peterlin BM. Interactions between HIV1 Nef and vacuolar ATPase facilitate the internalization of CD4. Immunity. 1998;8(5):647–56. doi: 10.1016/s1074-7613(00)80569-5. [DOI] [PubMed] [Google Scholar]
- Luo T, Garcia JV. The association of Nef with a cellular serine/threonine kinase and its enhancement of infectivity are viral isolate dependent. J Virol. 1996;70(9):6493–6. doi: 10.1128/jvi.70.9.6493-6496.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mahlknecht U, Deng C, Lu MC, Greenough TC, Sullivan JL, O’Brien WA, Herbein G. Resistance to apoptosis in HIV-infected CD4+ T lymphocytes is mediated by macrophages: role for Nef and immune activation in viral persistence. J Immunol. 2000;165(11):6437–46. doi: 10.4049/jimmunol.165.11.6437. [DOI] [PubMed] [Google Scholar]
- Mangasarian A, Piguet V, Wang JK, Chen YL, Trono D. Nef-induced CD4 and major histocompatibility complex class I (MHC-I) down-regulation are governed by distinct determinants: N-terminal alpha helix and proline repeat of Nef selectively regulate MHC-I trafficking. J Virol. 1999;73(3):1964–73. doi: 10.1128/jvi.73.3.1964-1973.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manninen A, Hiipakka M, Vihinen M, Lu W, Mayer BJ, Saksela K. SH3-Domain binding function of HIV-1 Nef is required for association with a PAK-related kinase. Virology. 1998;250(2):273–82. doi: 10.1006/viro.1998.9381. [DOI] [PubMed] [Google Scholar]
- McCrossan M, Marsden M, Carnie FW, Minnis S, Hansoti B, Anthony IC, Brettle RP, Bell JE, Simmonds P. An immune control model for viral replication in the CNS during presymptomatic HIV infection. Brain. 2006;129(Pt 2):503–16. doi: 10.1093/brain/awh695. [DOI] [PubMed] [Google Scholar]
- McPhee DA, Greenway AL, Holloway G, Smith K, Deacon N, Pemberton L, Brew BJ. Anomalies in Nef expression within the central nervous system of HIV-1 positive individuals/AIDS patients with or without AIDS dementia complex. J Neurovirol. 1998;4(3):291–300. doi: 10.3109/13550289809114530. [DOI] [PubMed] [Google Scholar]
- Michael NL, Chang G, d’Arcy LA, Tseng CJ, Birx DL, Sheppard HW. Functional characterization of human immunodeficiency virus type 1 nef genes in patients with divergent rates of disease progression. J Virol. 1995;69(11):6758–69. doi: 10.1128/jvi.69.11.6758-6769.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller MD, Warmerdam MT, Page KA, Feinberg MB, Greene WC. Expression of the human immunodeficiency virus type 1 (HIV-1) nef gene during HIV-1 production increases progeny particle infectivity independently of gp160 or viral entry. J Virol. 1995;69(1):579–84. doi: 10.1128/jvi.69.1.579-584.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mordelet E, Kissa K, Cressant A, Gray F, Ozden S, Vidal C, Charneau P, Granon S. Histopathological and cognitive defects induced by Nef in the brain. Faseb J. 2004;18(15):1851–61. doi: 10.1096/fj.04-2308com. [DOI] [PubMed] [Google Scholar]
- Northrop JP, Ullman KS, Crabtree GR. Characterization of the nuclear and cytoplasmic components of the lymphoid-specific nuclear factor of activated T cells (NF-AT) complex. J Biol Chem. 1993;268(4):2917–23. [PubMed] [Google Scholar]
- Nunn MF, Marsh JW. Human immunodeficiency virus type 1 Nef associates with a member of the p21-activated kinase family. J Virol. 1996;70(9):6157–61. doi: 10.1128/jvi.70.9.6157-6161.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olivetta E, Percario Z, Fiorucci G, Mattia G, Schiavoni I, Dennis C, Jager J, Harris M, Romeo G, Affabris E, Federico M. HIV-1 Nef induces the release of inflammatory factors from human monocyte/macrophages: involvement of Nef endocytotic signals and NF-kappa B activation. J Immunol. 2003;170(4):1716–27. doi: 10.4049/jimmunol.170.4.1716. [DOI] [PubMed] [Google Scholar]
- Olivetta E, Pietraforte D, Schiavoni I, Minetti M, Federico M, Sanchez M. HIV-1 Nef regulates the release of superoxide anions from human macrophages. Biochem J. 2005;390(Pt 2):591–602. doi: 10.1042/BJ20042139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O’Neill E, Kuo LS, Krisko JF, Tomchick DR, Garcia JV, Foster JL. Dynamic evolution of the human immunodeficiency virus type 1 pathogenic factor, Nef. J Virol. 2006;80(3):1311–20. doi: 10.1128/JVI.80.3.1311-1320.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piguet V, Gu F, Foti M, Demaurex N, Gruenberg J, Carpentier JL, Trono D. Nef-induced CD4 degradation: a diacidic-based motif in Nef functions as a lysosomal targeting signal through the binding of beta-COP in endosomes. Cell. 1999a;97(1):63–73. doi: 10.1016/s0092-8674(00)80715-1. [DOI] [PubMed] [Google Scholar]
- Piguet V, Schwartz O, Le Gall S, Trono D. The downregulation of CD4 and MHC-I by primate lentiviruses: a paradigm for the modulation of cell surface receptors. Immunol Rev. 1999b;168:51–63. doi: 10.1111/j.1600-065x.1999.tb01282.x. [DOI] [PubMed] [Google Scholar]
- Piguet V, Wan L, Borel C, Mangasarian A, Demaurex N, Thomas G, Trono D. HIV-1 Nef protein binds to the cellular protein PACS-1 to downregulate class I major histocompatibility complexes. Nat Cell Biol. 2000;2(3):163–7. doi: 10.1038/35004038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pope M, Haase AT. Transmission, acute HIV-1 infection and the quest for strategies to prevent infection. Nat Med. 2003;9(7):847–52. doi: 10.1038/nm0703-847. [DOI] [PubMed] [Google Scholar]
- Pulkkinen K, Renkema GH, Kirchhoff F, Saksela K. Nef associates with p21-activated kinase 2 in a p21-GTPase-dependent dynamic activation complex within lipid rafts. J Virol. 2004;78(23):12773–80. doi: 10.1128/JVI.78.23.12773-12780.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ratner L, Joseph T, Bandres J, Ghosh S, Vander Heyden N, Templeton A, Hahn B, Powderly W, Arens M. Sequence heterogeneity of Nef transcripts in HIV-1-infected subjects at different stages of disease. Virology. 1996;223(1):245–50. doi: 10.1006/viro.1996.0474. [DOI] [PubMed] [Google Scholar]
- Renkema GH, Manninen A, Mann DA, Harris M, Saksela K. Identification of the Nef-associated kinase as p21-activated kinase 2. Curr Biol. 1999;9(23):1407–10. doi: 10.1016/s0960-9822(00)80086-x. [DOI] [PubMed] [Google Scholar]
- Renkema GH, Manninen A, Saksela K. Human immunodeficiency virus type 1 Nef selectively associates with a catalytically active subpopulation of p21-activated kinase 2 (PAK2) independently of PAK2 binding to Nck or beta-PIX. J Virol. 2001;75(5):2154–60. doi: 10.1128/JVI.75.5.2154-2160.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ross TM, Oran AE, Cullen BR. Inhibition of HIV-1 progeny virion release by cell-surface CD4 is relieved by expression of the viral Nef protein. Curr Biol. 1999;9(12):613–21. doi: 10.1016/s0960-9822(99)80283-8. [DOI] [PubMed] [Google Scholar]
- Saksela K, Cheng G, Baltimore D. Proline-rich (PxxP) motifs in HIV-1 Nef bind to SH3 domains of a subset of Src kinases and are required for the enhanced growth of Nef+ viruses but not for down-regulation of CD4. Embo J. 1995;14(3):484–91. doi: 10.1002/j.1460-2075.1995.tb07024.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saksena NK, Wang B, Ge YC, Chang J, Dwyer DE, Xiang SH, Packham DR, Randle C, Cunningham AL. Region-specific changes, gene duplications, and random deletions in the nef gene from HIV type 1-infected brain tissues and blood of a demented patient. AIDS Res Hum Retroviruses. 1997;13(1):111–6. doi: 10.1089/aid.1997.13.111. [DOI] [PubMed] [Google Scholar]
- Sawai ET, Baur A, Struble H, Peterlin BM, Levy JA, Cheng-Mayer C. Human immunodeficiency virus type 1 Nef associates with a cellular serine kinase in T lymphocytes. Proc Natl Acad Sci U S A. 1994;91(4):1539–43. doi: 10.1073/pnas.91.4.1539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sawai ET, Khan IH, Montbriand PM, Peterlin BM, Cheng-Mayer C, Luciw PA. Activation of PAK by HIV and SIV Nef: importance for AIDS in rhesus macaques. Curr Biol. 1996;6(11):1519–27. doi: 10.1016/s0960-9822(96)00757-9. [DOI] [PubMed] [Google Scholar]
- Schrager JA, Marsh JW. HIV-1 Nef increases T cell activation in a stimulus-dependent manner. Proc Natl Acad Sci U S A. 1999;96(14):8167–72. doi: 10.1073/pnas.96.14.8167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shapshak P, Segal DM, Crandall KA, Fujimura RK, Zhang BT, Xin KQ, Okuda K, Petito CK, Eisdorfer C, Goodkin K. Independent evolution of HIV type 1 in different brain regions. AIDS Res Hum Retroviruses. 1999;15(9):811–20. doi: 10.1089/088922299310719. [DOI] [PubMed] [Google Scholar]
- Shugars DC, Smith MS, Glueck DH, Nantermet PV, Seillier-Moiseiwitsch F, Swanstrom R. Analysis of human immunodeficiency virus type 1 nef gene sequences present in vivo. J Virol. 1993;67(8):4639–50. doi: 10.1128/jvi.67.8.4639-4650.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simmons A, Aluvihare V, McMichael A. Nef triggers a transcriptional program in T cells imitating single-signal T cell activation and inducing HIV virulence mediators. Immunity. 2001;14(6):763–77. doi: 10.1016/s1074-7613(01)00158-3. [DOI] [PubMed] [Google Scholar]
- Skowronski J, Parks D, Mariani R. Altered T cell activation and development in transgenic mice expressing the HIV-1 nef gene. Embo J. 1993;12(2):703–13. doi: 10.1002/j.1460-2075.1993.tb05704.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spina CA, Kwoh TJ, Chowers MY, Guatelli JC, Richman DD. The importance of nef in the induction of human immunodeficiency virus type 1 replication from primary quiescent CD4 lymphocytes. J Exp Med. 1994;179(1):115–23. doi: 10.1084/jem.179.1.115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sugimoto C, Tadakuma K, Otani I, Moritoyo T, Akari H, Ono F, Yoshikawa Y, Sata T, Izumo S, Mori K. nef gene is required for robust productive infection by simian immunodeficiency virus of T-cell-rich paracortex in lymph nodes. J Virol. 2003;77(7):4169–80. doi: 10.1128/JVI.77.7.4169-4180.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swigut T, Alexander L, Morgan J, Lifson J, Mansfield KG, Lang S, Johnson RP, Skowronski J, Desrosiers R. Impact of Nef-mediated downregulation of major histocompatibility complex class I on immune response to simian immunodeficiency virus. J Virol. 2004;78(23):13335–44. doi: 10.1128/JVI.78.23.13335-13344.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swingler S, Brichacek B, Jacque JM, Ulich C, Zhou J, Stevenson M. HIV-1 Nef intersects the macrophage CD40L signalling pathway to promote resting-cell infection. Nature. 2003;424(6945):213–9. doi: 10.1038/nature01749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swingler S, Mann A, Jacque J, Brichacek B, Sasseville VG, Williams K, Lackner AA, Janoff EN, Wang R, Fisher D, Stevenson M. HIV-1 Nef mediates lymphocyte chemotaxis and activation by infected macrophages. Nat Med. 1999;5(9):997–103. doi: 10.1038/12433. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tobiume M, Takahoko M, Yamada T, Tatsumi M, Iwamoto A, Matsuda M. Inefficient enhancement of viral infectivity and CD4 downregulation by human immunodeficiency virus type 1 Nef from Japanese long-term nonprogressors. J Virol. 2002;76(12):5959–65. doi: 10.1128/JVI.76.12.5959-5965.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tomiyama H, Akari H, Adachi A, Takiguchi M. Different effects of Nef-mediated HLA class I down-regulation on human immunodeficiency virus type 1-specific CD8(+) T-cell cytolytic activity and cytokine production. J Virol. 2002;76(15):7535–43. doi: 10.1128/JVI.76.15.7535-7543.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Marle G, Henry S, Todoruk T, Sullivan A, Silva C, Rourke SB, Holden J, McArthur JC, Gill MJ, Power C. Human immunodeficiency virus type 1 Nef protein mediates neural cell death: a neurotoxic role for IP-10. Virology. 2004;329(2):302–18. doi: 10.1016/j.virol.2004.08.024. [DOI] [PubMed] [Google Scholar]
- van’t Wout AB, Ran LJ, Kuiken CL, Kootstra NA, Pals ST, Schuitemaker H. Analysis of the temporal relationship between human immunodeficiency virus type 1 quasispecies in sequential blood samples and various organs obtained at autopsy. J Virol. 1998;72(1):488–96. doi: 10.1128/jvi.72.1.488-496.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Verani A, Gras G, Pancino G. Macrophages and HIV-1: dangerous liaisons. Mol Immunol. 2005;42(2):195–212. doi: 10.1016/j.molimm.2004.06.020. [DOI] [PubMed] [Google Scholar]
- Vilhardt F, Plastre O, Sawada M, Suzuki K, Wiznerowicz M, Kiyokawa E, Trono D, Krause KH. The HIV-1 Nef protein and phagocyte NADPH oxidase activation. J Biol Chem. 2002;277(44):42136–43. doi: 10.1074/jbc.M200862200. [DOI] [PubMed] [Google Scholar]
- Wang JK, Kiyokawa E, Verdin E, Trono D. The Nef protein of HIV-1 associates with rafts and primes T cells for activation. Proc Natl Acad Sci U S A. 2000;97(1):394–9. doi: 10.1073/pnas.97.1.394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wei BL, Arora VK, Foster JL, Sodora DL, Garcia JV. In vivo analysis of Nef function. Curr HIV Res. 2003;1(1):41–50. doi: 10.2174/1570162033352057. [DOI] [PubMed] [Google Scholar]
- Wei BL, Arora VK, Raney A, Kuo LS, Xiao GH, O’Neill E, Testa JR, Foster JL, Garcia JV. Activation of p21-activated kinase 2 by human immunodeficiency virus type 1 Nef induces merlin phosphorylation. J Virol. 2005;79(23):14976–80. doi: 10.1128/JVI.79.23.14976-14980.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams KC, Hickey WF. Central nervous system damage, monocytes and macrophages, and neurological disorders in AIDS. Annu Rev Neurosci. 2002;25:537–62. doi: 10.1146/annurev.neuro.25.112701.142822. [DOI] [PubMed] [Google Scholar]
- Wiskerchen M, Cheng-Mayer C. HIV-1 Nef association with cellular serine kinase correlates with enhanced virion infectivity and efficient proviral DNA synthesis. Virology. 1996;224(1):292–301. doi: 10.1006/viro.1996.0531. [DOI] [PubMed] [Google Scholar]
- Wolf D, Witte V, Laffert B, Blume K, Stromer E, Trapp S, d’Aloja P, Schurmann A, Baur AS. HIV-1 Nef associated PAK and PI3-kinases stimulate Akt-independent Bad-phosphorylation to induce anti-apoptotic signals. Nat Med. 2001;7(11):1217–24. doi: 10.1038/nm1101-1217. [DOI] [PubMed] [Google Scholar]
- Wong JK, Ignacio CC, Torriani F, Havlir D, Fitch NJ, Richman DD. In vivo compartmentalization of human immunodeficiency virus: evidence from the examination of pol sequences from autopsy tissues. J Virol. 1997;71(3):2059–71. doi: 10.1128/jvi.71.3.2059-2071.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu Y, Marsh JW. Selective transcription and modulation of resting T cell activity by preintegrated HIV DNA. Science. 2001;293(5534):1503–6. doi: 10.1126/science.1061548. [DOI] [PubMed] [Google Scholar]
- Yang OO, Nguyen PT, Kalams SA, Dorfman T, Gottlinger HG, Stewart S, Chen IS, Threlkeld S, Walker BD. Nef-mediated resistance of human immunodeficiency virus type 1 to antiviral cytotoxic T lymphocytes. J Virol. 2002;76(4):1626–31. doi: 10.1128/JVI.76.4.1626-1631.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zazopoulos E, Haseltine WA. Mutational analysis of the human immunodeficiency virus type 1 Eli Nef function. Proc Natl Acad Sci U S A. 1992;89(14):6634–8. doi: 10.1073/pnas.89.14.6634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao ML, Si Q, Lee SC. IL-16 expression in lymphocytes and microglia in HIV-1 encephalitis. Neuropathol Appl Neurobiol. 2004;30(3):233–42. doi: 10.1046/j.0305-1846.2003.00527.x. [DOI] [PubMed] [Google Scholar]


