Abstract
Oomycete plant pathogens deliver effector proteins inside host cells to modulate plant defense circuitry and to enable parasitic colonization. These effectors are defined by a conserved motif, termed RXLR (for Arg, any amino acid, Leu, Arg), that is located downstream of the signal peptide and that has been implicated in host translocation. Because the phenotypes of RXLR effectors extend to plant cells, their genes are expected to be the direct target of the evolutionary forces that drive the antagonistic interplay between pathogen and host. We used the draft genome sequences of three oomycete plant pathogens, Phytophthora sojae, Phytophthora ramorum, and Hyaloperonospora parasitica, to generate genome-wide catalogs of RXLR effector genes and determine the extent to which these genes are under positive selection. These analyses revealed that the RXLR sequence is overrepresented and positionally constrained in the secretome of Phytophthora relative to other eukaryotes. The three examined plant pathogenic oomycetes carry complex and diverse sets of RXLR effector genes that have undergone relatively rapid birth and death evolution. We obtained robust evidence of positive selection in more than two-thirds of the examined paralog families of RXLR effectors. Positive selection has acted for the most part on the C-terminal region, consistent with the view that RXLR effectors are modular, with the N terminus involved in secretion and host translocation and the C-terminal domain dedicated to modulating host defenses inside plant cells.
INTRODUCTION
A diverse number of plant pathogens deliver effector proteins inside host cells to modulate plant defense circuitry and enable parasitic colonization (Birch et al., 2006; Chisholm et al., 2006; Grant et al., 2006; Huang et al., 2006a, 2006b; Jones and Dangl, 2006; Kamoun, 2006; O'Connell and Panstruga, 2006). Because these so-called cytoplasmic effectors function inside plant cells and produce phenotypes that extend to plant cells and tissues, their genes are expected to be the direct target of the evolutionary forces that drive the antagonistic interplay between pathogen and host (Dawkins and Krebs, 1979; Dawkins, 1999). For instance, as predicted by the arms-race model, several phytopathogen effector genes and their plant targets are known to be under positive selection (Allen et al., 2004; Dodds et al., 2004, 2006; Rohmer et al., 2004; Schurch et al., 2004; Liu et al., 2005; Rehmany et al., 2005; Ma et al., 2006).
Oomycetes form a phylogenetically distinct group of eukaryotic microorganisms that includes several of the most devastating pathogens of plants (Kamoun, 2003). Some oomycetes, such as the soybean (Glycine max) root and stem rot agent Phytophthora sojae and the potato (Solanum tuberosum) and tomato (Solanum lycopersicum) late blight agent Phytophthora infestans, have caused longstanding problems for agriculture, whereas others, such as the sudden oak (Quercus spp) death pathogen Phytophthora ramorum, have surfaced in recent epidemics. Other significant oomycetes include downy mildews, such as Hyaloperonospora parasitica, a natural pathogen of Arabidopsis thaliana that figures prominently in research on disease resistance in this model plant. These oomycetes establish intimate associations with plants and typically require living host cells to complete their infection cycle, a process known as biotrophy (O'Connell and Panstruga, 2006). Here, we describe adaptive evolution (positive selection) in the cytoplasmic effectors of three recently sequenced oomycete plant pathogens (Tyler et al., 2006) (Genome Sequencing Center at Washington University).
Little is known about the translocation of filamentous pathogen effectors into host cells, although specialized infection structures like haustoria are thought to be involved (Birch et al., 2006; Ellis et al., 2006; Kamoun, 2006; O'Connell and Panstruga, 2006). This contrasts with the well-studied specialized secretory machineries, such as the type III secretion system, that bacterial pathogens use to deliver effectors inside plant cells (Cornelis, 2006; Galan and Wolf-Watz, 2006). Nonetheless, significant insight has resulted from the recent identification of cytoplasmic oomycete effectors with avirulence activity (Birch et al., 2006; Ellis et al., 2006; Kamoun, 2006). These effectors from the oomycetes H. parasitica, P. infestans, and P. sojae carry a conserved motif, termed RXLR (for Arg, any amino acid, Leu, Arg), that is located downstream of the signal peptide and that has been implicated in host translocation (Rehmany et al., 2005; Bhattacharjee et al., 2006; Birch et al., 2006; Kamoun, 2006).
Several independent lines of evidence suggest that the RXLR motif defines a domain that functions in the delivery of effector proteins into host cells. First, the RXLR motif is similar in sequence and position to the plasmodial host translocation (HT)/Pexel motif that functions in the delivery of parasite proteins into the red blood cells of mammalian hosts (Hiller et al., 2004; Marti et al., 2004). Indeed, an ∼30–amino acid region encompassing the RXLR motif of the P. infestans RXLR proteins AVR3a and PH001D5 mediates the export of the green fluorescent protein from the Plasmodium falciparum parasite to the host red blood cell, suggesting that the RXLR and HT/Pexel domains are functionally interchangeable (Bhattacharjee et al., 2006). Second, the RXLR motif is not required for the effector activities of P. infestans AVR3a when this protein is directly expressed inside plant cells, consistent with a role in targeting rather than effector activity (Bos et al., 2006). Third, higher levels of polymorphism were observed in the C-terminal regions of H. parasitica ATR1 and ATR13, which have coevolved with host resistance proteins (Allen et al., 2004; Rehmany et al., 2005). Altogether, these findings led to the view that oomycete RXLR effectors are modular proteins with two major functional domains (Kamoun, 2006). While the N-terminal domain encompassing the signal peptide and RXLR leader functions in secretion and targeting, the remaining C-terminal region carries the effector activity and operates inside plant cells. Such a modular structure is reminiscent of that of bacterial type III secretion system effectors and suggests that the two domains might be under different selection pressures (Stavrinides et al., 2006).
The most reliable indicator of positive selection at the molecular level is a higher nonsynonymous nucleotide substitution rate (dN) than synonymous nucleotide substitution rate (dS) between two protein-coding DNA sequences (ratio ω = dN:dS > 1) (Yang et al., 2000). Based on this criterion, statistical methods, such as the approximate (counting) method and the maximum likelihood (ML) method, have been developed and implemented into the PAML 3.15 software package (Yang, 1997; Nielsen and Yang, 1998; Yang et al., 2000). Typically, comparisons are performed with data from intraspecific populations or sibling species. However, in the absence of such data, an acceptable alternative approach is to test for selection among families of closely related paralogs (Thomas et al., 2005; Thomas, 2006).
In this study, we used the draft genome sequences of three oomycete plant pathogens, P. sojae, P. ramorum, and H. parasitica (Tyler et al., 2006) (Genome Sequencing Center at Washington University), to generate genome-wide catalogs of RXLR effector genes and to determine the extent to which these genes are under positive selection. We obtained robust evidence of positive selection in more than two-thirds of the examined paralog families of RXLR effectors. Positive selection has for the most part targeted the C-terminal region of the RXLR effectors, consistent with the view that this domain is dedicated to executing the effector activity inside plant cells and has been involved in coevolutionary arms races with host factors.
RESULTS
Defining Features of Oomycete RXLR Effectors
To determine the defining features of oomycete RXLR effectors and develop criteria for identifying candidate genes from genome sequences, we put together an unbiased list of 43 oomycete RXLR proteins. The control set consisted of all 13 RXLR proteins with known avirulence or effector activity (Kamoun, 2006) (our unpublished data) and 30 additional sequences with significant homology with these validated effectors (BLASTP or TBLASTN cutoff E value < 10−4) (Table 1, Figure 1). The 43 proteins are relatively small, averaging 158 amino acids in length (range, 83 to 409 amino acids). They contain typical signal peptides (Signal P v2.0 NN score > 0.65, hidden Markov model [HMM] score > 0.93) that range in length from 15 to 28 amino acids. The RXLR sequence starts at positions 31 to 57 from the N terminus (average, 45). Thirty of the 43 sequences contain the EER sequence, following the RXLR, starting at positions 51 to 76 (average, 62). None of the 43 genes contains introns, as is typically the case in small oomycete genes (Kamoun, 2003).
Table 1.
Validated RXLR Effectors Used as a Control Set in This Study
| Description | Accession Numbera | Species | Evidence | Lengthb | Signal Peptide Lengthb | SignalP v2.0 HMM Score | SignalP v2.0 NN Score | RXLR Positionc | EER Positionc |
|---|---|---|---|---|---|---|---|---|---|
| Avirulence protein ATR1 | AAX51205 | H. parasitica | Avr effector | 324 | 15 | 0.995 | 0.944 | 48 | 60 |
| Unknown protein similar to ATR1 | Contig204.2_F52 | H. parasitica | Homolog | 290 | 15 | 0.979 | 0.901 | 48 | 59 |
| Unknown protein similar to ATR1 | Contig214.3_F41 | H. parasitica | Homolog | 409 | 16 | 0.987 | 0.688 | 52 | 63 |
| Avirulence protein ATR13 | AAW63766 | H. parasitica | Avr effector | 187 | 18 | 0.939 | 0.751 | 38 | |
| Unknown protein similar to ATR13 | Contig166.8_F9 | H. parasitica | Homolog | 196 | 24 | 0.983 | 0.741 | 38 | |
| Avirulence protein Avr3a | CAI72345 | P. infestans | Avr effector | 147 | 21 | 0.996 | 0.716 | 44 | 57 |
| Pex147-2 similar to Avr3a | CAI72329 | P. infestans | Homolog | 148 | 21 | 0.992 | 0.742 | 44 | 57 |
| Pex147-3 similar to Avr3a | CAI72350 | P. infestans | Avr effector | 147 | 21 | 0.991 | 0.725 | 44 | 57 |
| Unknown protein similar to Avr3a | CONTIG_3976_R2 | P. infestans | Homolog | 135 | 21 | 0.963 | 0.712 | 54 | 64 |
| Unknown protein similar to Avr3a | CONTIG_7218_R1 | P. infestans | Homolog | 153 | 23 | 1 | 0.93 | 47 | 60 |
| Unknown protein similar to Avr3a | CONTIG_9053_R1 | P. infestans | Homolog | 118 | 21 | 0.999 | 0.777 | 50 | 63 |
| Avirulence protein PiAvr4 | CV920942 | P. infestans | Avr effector | 287 | 24 | 0.999 | 0.836 | 42 | 53 |
| IPIO1 | AAY43376 | P. infestans | Alters defense | 152 | 21 | 1 | 0.968 | 51 | 70 |
| Unknown protein similar to IPIO1 | CONTIG_5858_R7 | P. infestans | Homolog | 167 | 21 | 1 | 0.953 | 57 | 76 |
| Unknown protein similar to IPIO1 | CONTIG_6141_R58 | P. infestans | Homolog | 140 | 23 | 0.999 | 0.932 | 56 | 70 |
| Unknown protein similar to IPIO1 | CONTIG_11746_R2 | P. infestans | Homolog | 153 | 20 | 1 | 0.956 | 56 | 75 |
| Unknown protein similar to IPIO1 | CONTIG_9802_R3 | P. infestans | Homolog | 83 | 21 | 1 | 0.924 | 57 | 73 |
| Unknown protein similar to IPIO1 | CONTIG_9926_R1 | P. infestans | Homolog | 137 | 19 | 0.992 | 0.835 | 53 | 69 |
| Pex-RD1 PH001D5 | CV917954 | P. infestans | Alters defense | 213 | 19 | 0.989 | 0.654 | 50 | 75 |
| Pex-RD2 | CV960315 | P. infestans | Alters defense | 121 | 20 | 0.998 | 0.913 | 38 | |
| Unknown protein similar to Pex-RD2 | CONTIG_10479_F49 | P. infestans | Homolog | 132 | 22 | 1 | 0.904 | 35 | 51 |
| Unknown protein similar to Pex-RD2 | CONTIG_10693_R5 | P. infestans | Homolog | 132 | 20 | 0.992 | 0.843 | 38 | 56 |
| Unknown protein similar to Pex-RD2 | CONTIG_1385_R18 | P. infestans | Homolog | 139 | 20 | 1 | 0.895 | 42 | 57 |
| Unknown protein similar to Pex-RD2 | CONTIG_2820_R8 | P. infestans | Homolog | 132 | 20 | 0.994 | 0.87 | 38 | 56 |
| Unknown protein similar to Pex-RD2 | CONTIG_3370_F60 | P. infestans | Homolog | 128 | 22 | 1 | 0.826 | 42 | 58 |
| Unknown protein similar to Pex-RD2 | CONTIG_3385_R23 | P. infestans | Homolog | 137 | 20 | 1 | 0.935 | 42 | 58 |
| Unknown protein similar to Pex-RD2 | CONTIG_3388_F56 | P. infestans | Homolog | 126 | 20 | 1 | 0.905 | 42 | |
| Unknown protein similar to Pex-RD2 | CONTIG_4905_F130 | P. infestans | Homolog | 125 | 20 | 0.999 | 0.871 | 31 | |
| Unknown protein similar to Pex-RD2 | CONTIG_6452_R6 | P. infestans | Homolog | 130 | 20 | 0.99 | 0.831 | 38 | 54 |
| Unknown protein similar to Pex-RD2 | CONTIG_7461_F10 | P. infestans | Homolog | 241 | 28 | 0.992 | 0.885 | 44 | 60 |
| Unknown protein similar to Pex-RD2 | CONTIG_8507_R1 | P. infestans | Homolog | 117 | 20 | 1 | 0.902 | 40 | 56 |
| Pex-RD39 | CV970797 | P. infestans | Alters defense | 100 | 22 | 1 | 0.864 | 43 | |
| Pex-RD45 | CONTIG_3612_R6 | P. infestans | Alters defense | 106 | 22 | 0.999 | 0.812 | 44 | |
| Pex-RD46 | CONTIG_3963_R4 | P. infestans | Alters defense | 105 | 21 | 1 | 0.854 | 39 | |
| Elicitor Avr1b | AAM20937 | P. sojae | Avr effector | 138 | 21 | 0.997 | 0.836 | 41 | |
| Elicitor Avh1b | AAM20941 | P. sojae | Alters defense | 138 | 21 | 0.995 | 0.833 | 41 | |
| Unknown protein similar to Avr1b | Ps_scaffold_108_R437 | P. sojae | Homolog | 150 | 21 | 0.998 | 0.928 | 55 | 69 |
| Unknown protein similar to Avr1b | Ps_scaffold_1152_F10 | P. sojae | Homolog | 110 | 20 | 1 | 0.919 | 51 | |
| Unknown protein similar to Avr1b | Ps_scaffold_31_R3 | P. sojae | Homolog | 109 | 20 | 1 | 0.919 | 51 | 63 |
| Unknown protein similar to Avr1b | Ps_scaffold_33_F1590 | P. sojae | Homolog | 150 | 24 | 1 | 0.841 | 51 | 70 |
| Unknown protein similar to Avr1b | Ps_scaffold_39_F1383 | P. sojae | Homolog | 128 | 22 | 1 | 0.926 | 40 | |
| Unknown protein similar to Avr1b | Ps_scaffold_39_F1389 | P. sojae | Homolog | 135 | 19 | 0.999 | 0.925 | 43 | |
| Unknown protein similar to PiAvr4 | AAO24646 | P. sojae | Homolog | 191 | 20 | 1 | 0.895 | 39 | 53 |
| Means | 158.3 | 20.7 | 0.99 | 0.86 | 45.0 | 62.1 |
GenBank accession number is provided for previously described sequences. Otherwise, accession numbers correspond to sequences listed in Supplemental Table 2 online.
Length in amino acids.
Position counting from the N terminus.
Figure 1.
Features of Validated RXLR Effectors.
(A) Position distribution of signal peptide cleavage site (SP), RXLR motif start position (RxLR), and EER start position (EER) among a control set of 43 RXLR effectors.
(B) Consensus sequence pattern of the RXLR/RXLR-EER motif was calculated using WebLogo based on an alignment of the 43 control sequences. The bigger the letter, the more conserved the amino acid site. This consensus served as a basis for the HMM described in the text.
The RXLR Sequence Is Overrepresented and Positionally Constrained in the Secretome of Phytophthora Relative to Other Eukaryotes
To further investigate the features of oomycete RXLR effectors, we looked for biases in the distribution of the RXLR motif in the proteomes of the two sequenced Phytophthora species (Tyler et al., 2006) compared with 46 other eukaryotes. For this purpose, we developed an exhaustive proteome database of 571,249 proteins corresponding to 48 eukaryotic species from 10 major taxonomic groups and consisting almost exclusively of complete proteomes (see Supplemental Table 1 online). Of these 571,249 proteins, 85,257 contained the sequence RXLR in any position, indicating that on average 1 eukaryotic protein out of 6.7 has the motif RXLR. The frequency of proteins with the RXLR sequence ranged from 1.72% (Plasmodium chabaudi) to 37.3% (Cyanidioschyzon merolae) in the 48 examined species (average, 14.9%). We then examined the occurrence of the RXLR sequence in putative extracellular proteins (referred to here as PEX and defined by the presence of a signal peptide using SignalP v2.0 [Nielsen et al., 1997]) (see Methods) compared with the remainder of the proteome (non-PEX). Of the 52,749 PEX proteins, 6410 or 12.2% contained RXLR. The frequency of PEX proteins with an RXLR sequence ranged from 0.95% (P. chabaudi) to 26.3% (C. merolae) in the 48 examined species compared with 1.73% (P. chabaudi) to 37.9% (C. merolae) for non-PEX proteins. The two Phytophthora species exhibited some of the highest frequencies of RXLR-containing proteins for both PEX proteins (19.3 and 19.7% for P. sojae and P. ramorum, respectively) and non-PEX proteins (24.2 and 20.5%, respectively). However, other species, notably C. merolae (red algae), human, rice (Oryza sativa), Leishmania major, and Trypanosoma brucei, also showed elevated frequencies of RXLR-containing proteins at levels similar to or higher than the two Phytophthora species.
Previous reports (Bhattacharjee et al., 2006) and the analyses described above of the 43 control sequences suggest that the RXLR motif is positionally conserved in the N-terminal region following the signal peptide. Consequently, we performed additional calculations of the frequency of proteins carrying RXLR at positions 10 to 110 from the N terminus and also at positions 30 to 60, which corresponds to the range identified in the validated set (31 to 57). The latter parameter resulted in a clear distributional bias in the Phytophthora species (Figure 2). The frequency of PEX proteins with an RXLR sequence at positions 30 to 60 ranged from 0% (four taxa) to 3.5% (Schizosaccharomyces pombe) in the 46 eukaryotes examined; by contrast, their frequency in P. sojae (6.8%) and P. ramorum (6.4%) was significantly higher than in the other species examined (randomization test, P < 0.005). Similar patterns were less pronounced or undetected for the PEX proteins with RXLR starting at positions 10 to 110 and for the non-PEX proteins (Figure 2).
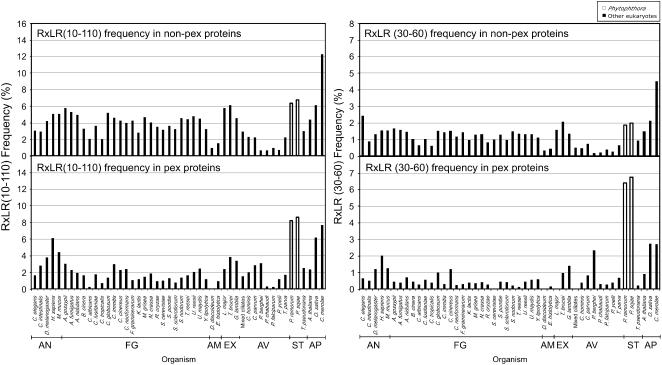
Figure 2.
The RXLR Sequence Is Overrepresented in the Secretome of Phytophthora Relative to Other Eukaryotic Proteomes.
The RXLR motif distribution in proteomes of eukaryotes is shown. RXLR distribution is shown in nonextracellular proteins (non-PEX proteins) in the top panels and in putative extracellular proteins (PEX proteins) in the bottom panels. RXLR motifs were counted only if they fell within the first 10 to 110 amino acids (left panels) or the first 30 to 60 amino acids (right panels) starting from the N termini of the proteins. The frequency for each eukaryote was calculated as a percentage of either all non-PEX proteins or all PEX proteins. RXLR frequency for each eukaryote was calculated as a percentage of either all non-PEX proteins or all PEX proteins. A full list of species is shown in Supplemental Table 1 online. AN, animals; FG, fungi; AM, amoebozoa; EX, excavata; AV, alveolates; ST, stramenopiles; AP, archaeplastida.
To further explore positional biases of the RXLR sequence in Phytophthora proteins, we compared the distribution of RXLR position in PEX and non-PEX proteins (Figure 3). The analysis clearly illustrated a nonrandom distribution of the RXLR position in Phytophthora PEX proteins relative to non-PEX proteins and PEX proteins from other species (χ2, P < 0.001) (Figure 3).
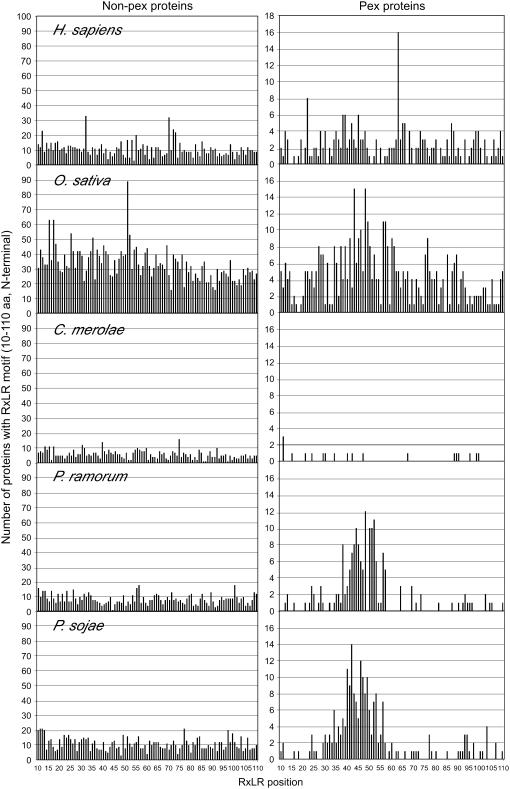
Figure 3.
The RXLR Sequence Is Positionally Constrained in the Secretome of Phytophthora.
Positional distribution of the RXLR motif within the first 10 to 110 amino acids (aa) in non-PEX proteins (left panels) and PEX proteins (right panels) of selected eukaryotes.
Identification of the RXLR Effector Secretome of P. sojae, P. ramorum, and H. parasitica
Based on the above features, we developed an algorithm for ab initio identification of RXLR effector genes from assembled genome sequences (Figure 4). Briefly, the genome sequences were scanned for all possible ATG start codons, translated, and then evaluated for the presence of signal peptides and the RXLR sequence starting at positions 30 to 60 from the N terminus (see Methods for full details). The three genome assemblies yielded variable numbers of candidate RXLR effectors, ranging from 149 for H. parasitica to 531 for P. ramorum and 672 for P. sojae (Table 2; see Supplemental Table 2 online for a full description of the 1352 candidates). Of these, 42 H. parasitica, 189 P. sojae, and 214 P. ramorum predicted proteins had the EER sequence following the RXLR. We also analyzed the sequences using a HMM from an alignment of the RXLR-EER region generated with the 43 control sequences. In total, 19 H. parasitica, 158 P. sojae, and 181 P. ramorum proteins were positive with the HMM (E value cutoff < 10) (Table 2; see Supplemental Table 2 online).
Figure 4.
An Algorithm for ab Initio Mining of RXLR Effectors from Oomycete Genomes.
Table 2.
Number of RXLR Candidate Effectors Identified in Each Genome
| Species | Reference | Genome Size (Mb) | Total RXLRa | RXLR + EERb | HMMc |
|---|---|---|---|---|---|
| P. ramorum | Tyler et al. (2006) | 65 | 531 | 214 | 181 |
| P. sojae | Tyler et al. (2006) | 95 | 672 | 189 | 158 |
| H. parasitica | Washington University | 75 | 149 | 42 | 19 |
| Total | 1352 | 445 | 358 |
Total number of candidate RXLR effectors predicted by the presence of a signal peptide and the RXLR sequence at amino acids 30 to 60.
Number of candidates with RXLR and EER sequences.
We performed comparative analyses between the three RXLR secretomes using bidirectional best BLASTP hit searches (E value cutoff < 10−10). Only limited overlap was observed between the three sets of sequences, with only four trios of genes showing 1:1:1 orthology relationships (see Supplemental Figure 1 online). The H. parasitica candidates were most divergent, with only 14 of the 149 genes (9%) showing similarity to the Phytophthora RXLR effectors. Overlap between the two Phytophthora species was more significant but was limited nonetheless to 192 of 531 (36%) of the P. ramorum candidates and 151 of 672 (22%) of the P. sojae genes, with the similarity often involving the signal peptide and RXLR domains rather than the C-terminal effector region.
Not all of the identified candidate genes appeared to be structurally intact, and several are likely to correspond to pseudogenes with insertion/deletion, missense, and/or nonsense mutations. In the absence of a validated set of intact genes as a reference, we had no robust approach to systematically catalog pseudogenes. Nonetheless, among the 1352 candidate effectors, we recognized 125 open reading frames (ORFs) that do not extend over 30 amino acids from the RXLR or RXLR-EER motif (see Supplemental Table 2 online). This suggests that relaxed selection has been a factor in the evolution of this family.
In summary, the small overlap between the three RXLR secretomes and the presence of pseudogenes suggest that the majority of RXLR effector genes are undergoing birth and death evolution, resulting in divergent sets of effectors in the three species. In addition, accelerated evolution may have also contributed to the extreme divergence of the RXLR effector genes.
Higher Rates of Closely Related Paralogs in P. ramorum
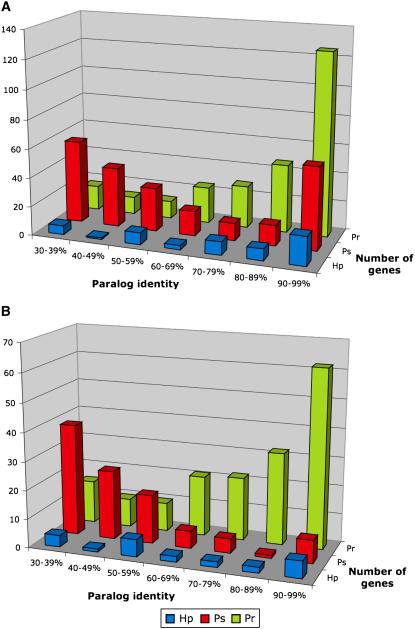
We used BLASTP searches (E value cutoff < 10−10) to identify the extent to which the RXLR effector genes have paralogs and to classify them into paralogous gene groups (PGGs). A total of 269 of 531 (51%) P. ramorum candidates, 235 of 672 (35%) P. sojae candidates, and 57 of 149 (38%) H. parasitica candidates showed similarity to at least one other effector from the same species, suggesting that P. ramorum exhibits a larger proportion of paralog families among the RXLR effectors. Higher rates of closely related paralogs were also observed in P. ramorum when we examined the distributions of the similarity levels of the closest paralogs (Figure 5). In P. ramorum, 202 of 269 (75%) RXLR effectors with a paralog had 70 to 99% identity to their closest paralogs, whereas only 83 of 235 (35%) were this similar in P. sojae (Figure 5A). This pattern was even more obvious when we analyzed the RXLR-EER subset of effectors (Figure 5B). A total of 116 of 172 (67%) P. ramorum RXLR-EER effectors with a paralog had 70 to 99% identity to their closest paralogs, versus only 14 of 100 (14%) in P. sojae. Overall, no obvious pattern was detected for H. parasitica RXLR and RXLR-EER effectors. In summary, these observations suggest significantly higher rates of closely related paralogs among the RXLR effector genes of P. ramorum, most likely as a result of a larger number of recent gene duplications.
Figure 5.
Higher Rates of Closely Related Paralogs among the RXLR Effectors of P. ramorum.
(A) Distribution of percentage identity of the closest paralog among the RXLR effectors of P. ramorum (Pr), P. sojae (Ps), and H. parasitica (Hp).
(B) Closest paralog identity among the RXLR-EER effectors.
Positive Selection in RXLR Effector Genes
We identified a subset of 99 PGGs comprising 212 sequences (range, 2 to 9 per PGG) that could be used in standard tests of selection (Table 3). This approach is an acceptable alternative to intraspecific or sibling species analyses and seeks evidence for positive selection among relatively recently duplicated paralogs (Thomas et al., 2005; Thomas, 2006). To minimize the impact of the pitfalls of positive selection analyses, such as gap-induced misalignments and relaxed selection in pseudogenes, we applied three criteria for selecting the PGGs: (1) similarity throughout the majority of the paralog coding sequences; (2) no or few gaps across the aligned sequences; and (3) at least 50% amino acid identity between the paralogs. The 99 PGGs were subjected to several tests of positive selection using the approximate (counting) method and the ML method implemented in the PAML 3.15 software package (Yang, 1997; Nielsen and Yang, 1998; Yang et al., 2000). First, we calculated dN and dS values across the entire ORF sequences. We found that the dN value was significantly greater than dS (ω = dN:dS > 1.2) in at least one pairwise comparison in 43 of the 99 PGGs (Tables 3 and 4; see Supplemental Table 3 online for full details). We detected ω > 1.0 in 119 and ω > 1.2 in 97 of the 175 pairwise sequence comparisons with the 43 PGG sequences. The average ω was 1.4, with the highest observed ω = 8.8 (Ps PGG20). We still detected 33 and 22 of the 99 PGGs using the more stringent ω cutoffs of 1.5 and 2.0, respectively (see Supplemental Table 4 online). Two-sample t tests indicated that the differences between the dN and dS values are significant, with P < 0.05 (ω > 1.0), P < 0.02 (ω > 1.2), and P < 0.005 (ω > 1.5) (see Supplemental Table 5 online). Although calculating dN:dS ratios over entire ORF sequences is considered an insensitive method, our results suggest that this method is sufficient to provide evidence that positive selection has acted on 43 of the examined 99 PGGs.
Table 3.
Summary of Positive Selection Analyses of PGGs of P. ramorum, P. sojae, and H. parasitica
| PGGsa
|
dN:dS > 1.2b
|
|||||
|---|---|---|---|---|---|---|
| Species | Total | Positive Selection | Full Sequences | N-Terminal | C-Terminal | MLc |
| P. ramorum | 59 | 41 | 26 | 22 | 26 | 15 |
| P. sojae | 28 | 18 | 11 | 9 | 16 | 1 |
| H. parasitica | 12 | 11 | 6 | 6 | 7 | 2 |
| Total | 99 | 70 | 43 | 37 | 49 | 18 |
Total and positively selected number of PGGs examined.
Number of PGGs with at least one pairwise ω = dN:dS > 1.2 over the full length, N-terminal, or C-terminal region of the sequences.
Number of PGGs under positive selection based on the ML method. Note that only 31 of the 99 PGGs could be tested using this method.
Table 4.
PGGs under Positive Selection
| Number of Sequencesa | Positive Using Both Methodsb | Highest dN:dS among Pairwise Comparisonsc
|
||||
|---|---|---|---|---|---|---|
| PGG | Full Sequences | N-Terminal | C-Terminal | MLd | ||
| Pr PGG44 | 9 | 0.31 | 0.35 | 0.28 | Yes | |
| Ps PGG6 | 8 | 3.26, ∞ | 1.92, ∞ | 2.85, ∞ | No | |
| Hp PGG13 | 7 | + | 1.02 | 2.17, ∞ | 1.08 | Yes |
| Pr PGG11 | 7 | + | 1.57, ∞ | 1.22 | 1.49, ∞ | Yes |
| Pr PGG59 | 7 | + | 2.53 | 1.31 | 3.07 | Yes |
| Pr PGG5 | 5 | 0.97 | 0.90 | 0.96 | Yes | |
| Pr PGG21 | 5 | + | 2.38 | 2.87, ∞ | 3.49 | Yes |
| Pr PGG22 | 5 | 0.89 | 1.33 | 1.05 | No | |
| Pr PGG31 | 5 | + | 1.74, ∞ | 1.55, ∞ | 1.24 | Yes |
| Hp PGG1 | 4 | + | 1.72 | 1.85 | 1.75 | Yes |
| Pr PGG7 | 4 | + | 1.02 | 0.76, ∞ | 0.99 | Yes |
| Pr PGG13 | 4 | + | 1.09 | 1.38 | 1.22 | Yes |
| Pr PGG14 | 4 | 0.68 | 1.06, ∞ | 0.52 | No | |
| Pr PGG17 | 4 | + | 1.50 | 0.82, ∞ | 1.57 | Yes |
| Pr PGG24 | 4 | + | 1.85 | 1.24 | 5.47 | Yes |
| Pr PGG27 | 4 | 1.72 | 0.49 | 1.59 | No | |
| Pr PGG39 | 4 | + | 1.46 | 1.24 | 1.73 | Yes |
| Ps PGG16 | 4 | 1.53, ∞ | 0.76 | 3.68, ∞ | No | |
| Ps PGG19 | 4 | 3.93 | 1.86, ∞ | 2.27, ∞ | No | |
| Pr PGG1 | 3 | 0.98 | 0.84 | 0.75 | Yes | |
| Pr PGG16 | 3 | + | 0.91 | 0.42 | 2.14 | Yes |
| Pr PGG20 | 3 | 1.87 | 1.57 | 2.04 | No | |
| Pr PGG23 | 3 | + | 2.08 | 0.61 | 4.00 | Yes |
| Pr PGG47 | 3 | 1.69 | ∞ | 1.58 | No | |
| Pr PGG54 | 3 | 0.87 | 0.42 | 0.86 | Yes | |
| Ps PGG3 | 3 | 0.74 | 1.01, ∞ | 0.18, ∞ | No | |
| Ps PGG4 | 3 | 0.37, ∞ | 0.23 | 0.44, ∞ | No | |
| Ps PGG10 | 3 | 0.80 | 0.61 | ∞ | No | |
| Ps PGG18 | 3 | 2.68 | 1.62 | ∞ | No | |
| Ps PGG21 | 3 | + | 0.98 | 0.95, ∞ | 1.19 | Yes |
| Ps PGG23 | 3 | ∞ | ∞ | ∞ | No | |
| Hp PGG2 | 2 | NA | ∞ | 0 | ∞ | NA |
| Hp PGG3 | 2 | NA | 3.96 | 0 | 4.08 | NA |
| Hp PGG4 | 2 | NA | 1.76 | 0.56 | 1.42 | NA |
| Hp PGG6 | 2 | NA | 1.35 | 0.47 | 1.39 | NA |
| Hp PGG7 | 2 | NA | 0.94 | ∞ | 0.87 | NA |
| Hp PGG8 | 2 | NA | 0.53 | ∞ | 0.29 | NA |
| Hp PGG9 | 2 | NA | 2.55 | 2.73 | 2.58 | NA |
| Hp PGG10 | 2 | NA | 0.44 | 0.05 | 2.49 | NA |
| Hp PGG12 | 2 | NA | 1.01 | ∞ | 0.96 | NA |
| Pr PGG3 | 2 | NA | ∞ | ∞ | ∞ | NA |
| Pr PGG4 | 2 | NA | 0.62 | 0.27 | 1.43 | NA |
| Pr PGG6 | 2 | NA | 1.23 | 1.52 | 0 | NA |
| Pr PGG8 | 2 | NA | 0.67 | 1.79 | 0.54 | NA |
| Pr PGG10 | 2 | NA | 1.57 | ∞ | 0.37 | NA |
| Pr PGG15 | 2 | NA | 1.46 | 1.14 | 1.22 | NA |
| Pr PGG18 | 2 | NA | 0.88 | 1.87 | 0.714 | NA |
| Pr PGG19 | 2 | NA | 1.24 | 0.93 | 1.11 | NA |
| Pr PGG25 | 2 | NA | 0.91 | 0.45 | 1.25 | NA |
| Pr PGG34 | 2 | NA | 1.33 | 0.71 | ∞ | NA |
| Pr PGG35 | 2 | NA | 0.70 | 0 | ∞ | NA |
| Pr PGG37 | 2 | NA | 1.64 | 0.62 | 2.03 | NA |
| Pr PGG38 | 2 | NA | 0.75 | ∞ | 0.56 | NA |
| Pr PGG41 | 2 | NA | ∞ | ∞ | ∞ | NA |
| Pr PGG42 | 2 | NA | ∞ | ∞ | 0 | NA |
| Pr PGG46 | 2 | NA | ∞ | 0 | ∞ | NA |
| Pr PGG48 | 2 | NA | ∞ | 0 | ∞ | NA |
| Pr PGG49 | 2 | NA | ∞ | ∞ | 0 | NA |
| Pr PGG51 | 2 | NA | ∞ | ∞ | 0 | NA |
| Pr PGG57 | 2 | NA | ∞ | 0 | ∞ | NA |
| Pr PGG58 | 2 | NA | ∞ | 0 | ∞ | NA |
| Ps PGG1 | 2 | NA | 2.04 | 1.13 | ∞ | NA |
| Ps PGG11 | 2 | NA | 1.45 | 1.18 | 1.61 | NA |
| Ps PGG12 | 2 | NA | 0.49 | 0 | ∞ | NA |
| Ps PGG13 | 2 | NA | 1.18 | ∞ | 0.80 | NA |
| Ps PGG14 | 2 | NA | 7.00 | ∞ | 5.43 | NA |
| Ps PGG17 | 2 | NA | ∞ | 0 | ∞ | NA |
| Ps PGG20 | 2 | NA | 8.77 | ∞ | 5.88 | NA |
| Ps PGG22 | 2 | NA | 0.17 | 0 | ∞ | NA |
| Ps PGG28 | 2 | NA | 0.89 | 0.67 | 1.33 | NA |
Number of sequences in the listed PGGs.
+, positive using both the counting and ML methods; NA, ML method not applicable (number of sequences < 3).
Highest calculated dN:dS ratio among all pairwise comparisons within one PGG; dN/dS > 1.2 is highlighted in boldface; ∞ refers to pairwise comparisons in which dN > 0 and dS = 0. In such cases and whenever applicable, the second higher ratio is also shown.
Yes, positive selection detected using the ML method; No, no evidence for positive selection.
Elevated dN:dS Ratios in the C-Terminal Domain of the RXLR Effectors
Positive selection typically acts on particular domains or amino acid residues within a given protein. To test for deviation in the substitution pattern of the different domains of the RXLR effectors and to identify additional PGGs under positive selection, we partitioned the sequences into N-terminal regions (from the start codon to RXLR or RXLR-EER sequences) and C-terminal regions (from RXLR or RXLR-EER to the end). We detected positive selection in an additional 23 PGGs that showed at least one pairwise comparison with ω > 1.2 for either the N- or C-terminal region (Tables 3 and 4; see Supplemental Table 3 online). More PGGs showed ω > 1.2 for the C-terminal domains (49) than for the N-terminal domains (37). In the analyses of the N-terminal domains, we found ω > 1.0 in 107 and ω > 1.2 in 91 of the 251 pairwise comparisons with the 66 PGG sequences. The average ω was 0.8, with the highest observed ω = 2.9 (Pr PGG21). By contrast, in the C-terminal domains, we found ω > 1.0 in 147 and ω > 1.2 in 130 of the 251 pairwise sequence comparisons. The average ω was 1.2, with the highest observed ω = 5.9 (Ps PGG20). When more stringent ω cutoffs of 1.5 and 2.0 were used, we also observed higher numbers of positive PGGs for the C-terminal regions (40 and 34, respectively) versus the N-terminal regions (30 and 26, respectively) (see Supplemental Table 4 online). Two-sample t tests of the ω values calculated for the C- and N-terminal domains of the 66 PGGs were highly significant (P < 0.0005). These differences in the distribution of ω between the N- and C-terminal domains are also apparent in the dN:dS plots shown in Figure 6. In summary, these results suggest that positive selection has more greatly affected the C-terminal domain than the N-terminal domain of the examined RXLR effectors.
Figure 6.
Elevated dN:dS Ratios in the C-Terminal Domain of the RXLR Effectors.
(A) Comparison of dN:dS (ω) values calculated for the full-length sequences versus the N-terminal region.
(B) dN:dS plots for the full-length sequences versus the C-terminal region.
(C) dN:dS plots for the N-terminal region versus the C-terminal region.
The lines demark a dN:dS value of 1, above which positive selection is inferred.
Positively Selected Amino Acid Sites Localize to the C-Terminal Domain of the RXLR Effectors
We applied the ML method (Nielsen and Yang, 1998; Yang et al., 2000) to identify additional PGGs under positive selection, to validate the results obtained with the approximate method using an independent approach, and to identify some of the amino acid residues that are under positive selection. The ML method can only be implemented to sets of three or more sequences and thus could only be applied to 31 of the 99 PGGs. Of these, we detected statistically significant evidence of positive selection in 18 of the 31 PGGs using the M8/M7 models of the ML method (Tables 3 and 5; see Methods). Fourteen of these were previously identified with the approximate method and thus were validated using the independent ML approach (Tables 3 and 4). The remaining four PGGs (Pr PGG1, Pr PGG5, Pr PGG44, and Pr PGG54) were not detected using the approximate method, bringing the total of positively selected PGGs to 70 of the 99 PGGs examined.
Table 5.
Likelihood Ratio Statistics and Distribution of Positively Selected Sites Based on the ML Method
| N-Terminal Domain
|
C-Terminal Domain
|
||||||
|---|---|---|---|---|---|---|---|
| PGGa | 2ΔLb | P | Total Number of Sites | Positively Selected Sites | Total Number of Sites | Positively Selected Sites | Positively Selected Sitesc |
| Hp PGG1 | 65.17 | <0.0001 | 71 | 0 | 58 | 30 | 73 F 75 I 76 A 77 T 80 K 84 F 86 V 87 D 88 M 89 W 92 L 95 M 98 D 100 W 101 E 102 F 103 S 105 N 106 R 107 Q 108 L 109 T 110 L 112 G 113 F 114 V 119 V 120 K 124 V 129 R |
| Hp PGG13 | 12.53 | 0.002 | 51 | 0 | 320 | 3 | 284 T 355 M 358 E |
| Pr PGG1 | 17.213 | 0.0002 | 71 | 0 | 67 | 2 | 97 R 130 A |
| Pr PGG11 | 57.92 | <0.0001 | 57 | 0 | 176 | 18 | 109 W 119 D 120 W 122 E 123 E 128 K 129 D 133 D 137 A 138 G 142 A 152 Q 170 S 176 V 177 Y 198 K 200 W 218 T |
| Pr PGG13 | 9.18 | 0.01 | 69 | 0 | 181 | 2 | 239 Y 250 Y |
| Pr PGG16 | 12.16 | 0.002 | 75 | 0 | 64 | 2 | 78 P 92 N |
| Pr PGG17 | 10.84 | 0.004 | 63 | 0 | 199 | 2 | 77 F 212 A |
| Pr PGG21 | 12.88 | 0.002 | 69 | 0 | 67 | 3 | 80 K 107 L 133 I |
| Pr PGG23 | 19.05 | <0.0001 | 68 | 0 | 67 | 11 | 71 I 76 R 82 S 89 V 91 D 95 L 98 K 101 A 103 I 117 Y 130 N |
| Pr PGG24 | 13.66 | 0.001 | 69 | 0 | 145 | 2 | 148 T 151 E |
| Pr PGG31 | 12.62 | 0.002 | 64 | 0 | 72 | 2 | 117 A 134 W |
| Pr PGG39 | 9.35 | 0.009 | 76 | 0 | 95 | 4 | 80 E 108 Q 133 K 145 F |
| Pr PGG44 | 17.92 | 0.0001 | 51 | 0 | 184 | 2 | 69 L 215 S |
| Pr PGG5 | 30.72 | <0.0001 | 58 | 0 | 66 | 7 | 62 M 63 L 66 L 67 N 70 I 114 I 122 K |
| Pr PGG54 | 9.97 | 0.007 | 51 | 0 | 336 | 1 | 326 K |
| Pr PGG59 | 54.27 | <0.0001 | 73 | 0 | 77 | 22 | 82 G 90 L 91 S 93 S 95 S 96 A 97 A 101 R 102 L 105 F 106 S 108 I 109 M 112 K 116 M 118 E 123 V 124 Q 128 R 129 A 130 I 143 A |
| Pr PGG7 | 11.77 | 0.003 | 54 | 0 | 146 | 4 | 91 E 137 R 142 K 146 S |
| Ps PGG21 | 47.23 | <0.0001 | 34 | 2 | 71 | 21 | 45 G 65 T 71 L 72 P 73 P 75 S 77 R 78 S 79 A 81 G 82 N 84 S 85 A 86 R 88 V 94 G 97 T 98 C 102 T 104 R 105 R |
PGG accession number.
Likelihood ratio test: 2(lnL alternative hypothesis − lnL null hypothesis).
P > 95% values are shown in lightface, and P > 99% values are shown in boldface.
The ML method enabled the identification of positively selected amino acid residues (Table 5). Remarkably, the overwhelming majority of positively selected sites localized to the C-terminal domain of the RXLR effectors. A total of 138 positively selected sites (range, 1 to 30 per PGG) were identified in the C-terminal effector domain of the 18 PGGs, while only 2 sites were identified in the N-terminal targeting domain. In fact, no positively selected sites were detected in the N terminus in 17 of the 18 PGGs. In Ps PGG21, 2 of 34 sites (5.9%) were detected in the N-terminal domain versus 21 of 71 sites (29.6%) in the C-terminal domain. These findings confirm our earlier observation that positive selection has for the most part targeted the C-terminal region of the RXLR effectors. The data also indicate that the two main domains of the RXLR proteins of plant pathogenic oomycetes have been exposed to distinct types of selective pressures.
The Positively Selected Ps PGG20 Consists of Two Functionally Distinct Genes
Positively selected Ps PGG20 includes P. sojae Avr1b-1, an RXLR effector that mediates avirulence to the soybean Rps1-b disease resistance gene (Shan et al., 2004). This prompted us to test whether the paralog Avh1b has a distinct activity. Because Avr1b-1 and Avh1b show sequence similarity to P. infestans Avr3a (Armstrong et al., 2005), we coexpressed these effectors with Solanum demissum R3a in Nicotiana benthamiana using Agrobacterium tumefaciens and potato virus X transient expression assays (Figures 7A and 7B; see Methods for details). Interestingly, Avh1b, but not Avr1b-1, induced hypersensitivity-like cell death only in the presence of R3a. The cell death induced by Avh1b was weaker compared with that induced by P. infestans AVR3aKI (∼25% of inoculated sites versus ∼80%, respectively) but nonetheless was consistently observed in repeated experiments as well as on a transgenic N. benthamiana line expressing R3a (Figure 7C). Similar assays with Avr1bAAM20939 (GenBank accession number AAM20939), a third member of the Avr1b family that was reported from a different isolate of P. sojae (Shan et al., 2004), failed to reveal activation of R3a (Figure 7). Overall, these results suggest that Avr1b-1 and Avh1b, which display a dN:dS ratio of 10.95, are recognized by distinct R genes, suggesting that positive selection may have driven the functional divergence of these paralogs.
Figure 7.
The Positively Selected Ps PGG20 Consists of Two Functional Genes, Avr1b-1 and Avh1b, with Distinct Effector Activities.
(A) Percentages of wound-inoculation sites showing R3a-dependent hypersensitive cell death at 3 to 7 d after inoculation of N. benthamiana leaves transiently expressing R3a with Agrobacterium strains expressing AVR3aKI (green circles), Avh1b (gray squares), and Avr1b-1, Avr1bAAM20939, AVR3aEM, and Avh1bT101K mutants (purple asterisks; no response). Percentages are based on the averages of two replicated experiments with 18 and 36 wound-inoculated sites per construct. Error bars indicate se. dpi, days after inoculation.
(B) Symptoms of R3a hypersensitive cell death at wound-inoculated sites of N. benthamiana leaves transiently expressing R3a. Photographs were taken at 7 d after inoculation.
(C) Symptoms of R3a hypersensitive cell death at wound-inoculated sites of N. benthamiana leaves from a transgenic line expressing R3a.
Avr1b-1 and Avr1bAAM20939 share nine identical amino acids in the C-terminal region that are polymorphic in Avh1b. All nine sites were predicted to be under positive selection using the codeml ML method implemented in the PAML 3.15 package (Yang et al., 2005) (see Supplemental Figure 2 online). To test whether the positively selected sites affect the cell death elicitor activity of Avh1b, we mutated Thr-101 into a Lys residue, the residue observed in Avr1b-1. Mutant Avh1bT101K did not activate R3a in wound-inoculation assays on leaves transiently expressing R3a or on transgenic R3a N. benthamiana (Figures 7B and 7C). Therefore, positively selected Thr-101 of Avh1b is critical for the cell death elicitor activity of this effector.
Clustering of Positively Selected RXLR Effector Genes
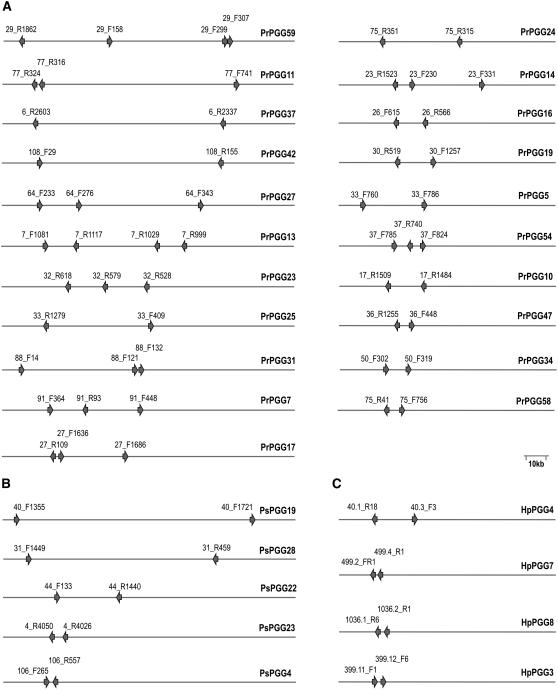
To investigate the genetic organization of the positively selected RXLR effectors, we examined the identified genes for clustering in the three genomes. A total of 30 PGGs corresponding to 72 genes were found clustered in genomic regions of <100 kb (Figure 8). In the P. ramorum genome, 21 PGGs corresponding to 54 genes were clustered, versus 5 PGGs and 10 genes in P. sojae and 4 PGGs and 8 genes in H. parasitica.
Figure 8.
RXLR Effector Gene Clusters.
Gene clusters in P. ramorum (A), P. sojae (B), and H. parasitica (C). The gene names start with the scaffold/supercontig number. The PGG groups are shown at right. Only clusters corresponding to positively selected genes in regions of <100 kb are shown.
Annotation of RXLR Effector Secretome
The C-terminal effector domains of the 1352 predicted RXLR proteins were annotated by sequence similarity searches. A total of 218 sequences (16%) showed significant similarities to known protein sequences (E value < 10−5) based on BLASTP searches to the GenBank nonredundant database (see Supplemental Table 2 online). However, 134 of these hits were to oomycete proteins, suggesting that the great majority of identified RXLR proteins (1268 of 1352, or 94%) have no obvious similarity to known proteins from other organisms. Previously described RXLR proteins, such as H. parasitica ATR1 and ATR13 as well as P. sojae Avr1b and Avh1b, were detected along with proteins with similarity to P. infestans RXLR proteins, such as IPIO1 and IPIO2.
Additionally, we examined the similarity of the predicted RXLR proteins to the HMM profiles in the Pfam protein motif database (E value < 10−3). Pfam searches enabled more sensitive and accurate predictions about the presence of known functional domains in the putative RXLR effectors and revealed 59 sequences with statistically significant hits (see Supplemental Table 6 online). Most of these show similarity to known enzymes (reductases, kinases, and peptidases), transporters (ABC transporter, sugar transporter, amino acid transporter, and ion transporter), or nucleotide binding pockets (cyclic nucleotide binding).
H. parasitica Proteins with Overlapping RXLR and LXLFLAK Motifs
Sixteen H. parasitica RXLR proteins, including the seven sequences that constitute the positively selected Hp PGG13, showed similarity to the Crinkler (CRN) protein family, a distinct class of cytoplasmic effectors first described in P. infestans (Torto et al., 2003; Kamoun, 2006). Multiple alignments of these H. parasitica proteins with 16 members of the CRN family identified in P. infestans (Win et al., 2006) revealed that the RXLR sequence overlaps a different motif, LXLFLAK, that is highly conserved in the CRN proteins, resulting in the sequence RXLRLFLAK (Figure 9A). To identify additional oomycete CRN-like sequences, we searched for signal peptide and LXLFLAK in the first 60 amino acids using an algorithm similar to the one shown in Figure 4. This resulted in a total of 105 sequences. None of the predicted Phytophthora CRNs have an RXLR associated with the LXLFLAK motif; however, there are putative H. parasitica CRNs that have LXLFLAK but also lack RXLR.
Figure 9.
H. parasitica Proteins with Similarity to CRN Effectors and Overlapping RXLR and LXLFLAK Motifs.
(A) Multiple alignment of the H. parasitica CRN-like RXLR proteins and P. infestans CRN1 to CRN16. The signal peptide region is indicated by a gray line. The LXLFLAK typical of CRN proteins is indicated by a black line. Note that the H. parasitica sequences contain the overlapping motif RXLRLFLAK.
(B) Consensus sequence pattern illustrating the LXLFLAK motif was calculated using WebLogo based on an alignment of the 105 oomycete CRN-like sequences. The bigger the letter, the more conserved the amino acid site.
(C) Neighbor-joining tree illustrating the relationship between the H. parasitica CRN-like proteins and the P. infestans CRNs. Bootstrap values corresponding to the neighbor-joining, ML, and parsimony methods are indicated above the branch separating the two major clusters.
To determine the evolutionary relationship between RXLR-containing CRNs from H. parasitica and known P. infestans CRNs, we built a phylogenetic tree using the neighbor-joining method based on the alignment in Figure 9A. The phylogenetic analyses indicate that the H. parasitica CRNs form a distinct class of CRN-like genes (Figure 9B). Shorter average branch lengths in the H. parasitica clade compared with the P. infestans clade indicate that the emergence of the RXLR-containing CRN family might be a more recent evolutionary event.
DISCUSSION
In this study, we developed genome-wide catalogs of RXLR effector genes from three oomycete species and investigated their molecular evolution. These analyses resulted in three major findings. First, the RXLR sequence is overrepresented and positionally constrained in the secretome of Phytophthora relative to other eukaryotes. Second, plant pathogenic oomycetes carry complex and diverse sets of RXLR effector genes that have undergone relatively rapid birth and death evolution. Third, positive selection has acted on paralogous RXLR gene families targeting for the most part the C-terminal region. These findings are consistent with the view that RXLR effectors are modular proteins, with the N terminus involved in secretion and host translocation and the C-terminal domain dedicated to modulating host defenses inside plant cells. Also, we propose that the positively selected genes identified here are more likely functionally important effectors and that the selection criterion can be used to augment other selection criteria for prioritizing candidate effector genes for functional studies.
Although anecdotal evidence that the RXLR motif is overrepresented in Phytophthora relative to other eukaryotes has been reported (Rehmany et al., 2005; Bhattacharjee et al., 2006), a systematic survey of the distribution of this sequence in eukaryotes as described in Figure 2 has not been performed. We found that among the examined eukaryotes, Phytophthora exhibits the highest frequencies of association between a signal peptide and the RXLR sequence. The related and functionally equivalent HT/Pexel motif RXLX(E/Q) is also overrepresented and positionally constrained in Plasmodium (Hiller et al., 2004; Marti et al., 2004; Bhattacharjee et al., 2006). However, it differs from the RXLR motif in the fourth and fifth positions, and searches for the RXLR sequence did not reveal a large number of Plasmodium effectors.
As with other computational predictions, the identification of RXLR effectors from sequence data involves a certain error rate and undoubtedly includes false-positives. We complemented the searches for RXLR at positions 30 to 60 with more stringent methods, such as the occurrence of an EER sequence and similarity to an HMM based on an alignment of the validated controls. However, these two methods most likely eliminate genuine effectors. For example, 13 of the 43 control sequences do not have EER and 3 of the 43 were not identified with the HMM profile (E value cutoff < 10). This suggests that the actual number of RXLR effectors is probably somewhere between the numbers obtained with the HMM/RXLR-EER predictions and those obtained with RXLR only.
Why does H. parasitica have a smaller number of predicted RXLR effectors than P. ramorum and P. sojae? H. parasitica is a highly specialized obligate biotroph that is part of a large monophyletic lineage of downy mildews that infect almost exclusively plants of the Brassicaceae family (Goker et al., 2007). The genus Hyaloperonospora may include >100 valid species, suggesting that specialization on Brassicaceae hosts is an ancestral trait in this lineage (Constantinescu and Fatehi, 2002; Goker et al., 2007). By contrast, Phytophthora species tend to be more ubiquitous in terms of host range (Erwin and Ribeiro, 1996). For instance, P. ramorum is known to infect a diverse number of woody plants (Rizzo et al., 2002, 2005), and P. sojae is closely related to P. sinensis and P. vignae, which infect plants unrelated to soybean (Cooke et al., 2000; Kroon et al., 2004). One hypothetical explanation is that H. parasitica carries a more specialized set of effectors as a consequence of losing unutilized effector genes (the use-it-or-lose-it rule of evolution [Carroll, 2006]). The broader arsenal of effectors in Phytophthora may reflect recent historical associations with phylogenetically diverse host plants and an ability to colonize new hosts. In such a case, it might prove easier to assign functional activities to H. parasitica effectors, compared with those of Phytophthora, if indeed they are specialized to function on Arabidopsis and related Brassicaceae plants. Future functional analyses of the RXLR effectors will help test this possibility. Nonetheless, it is already interesting that the number of RXLR effectors does not directly correlate with genome size in plant pathogenic oomycetes and that the effector genes have likely been shaped by the pathogenic lifestyle and degree of host specialization.
We noted a striking similarity between 13 of the H. parasitica candidate RXLR effectors and members of the large CRN family of Phytophthora (Torto et al., 2003; Kamoun, 2006). CRN1 and CRN2 were identified following an in planta functional expression screen of candidate secreted proteins of P. infestans based on a vector derived from Potato virus X (Torto et al., 2003). Expression of both genes in Nicotiana species and in the host plant tomato results in a leaf-crinkling and cell-death phenotype accompanied by an induction of defense-related genes. Torto et al. (2003) proposed that CRN1 and CRN2 function as effectors that perturb host cellular processes, based on analogy to bacterial effectors, which typically cause macroscopic phenotypes such as cell death, chlorosis, and tissue browning when expressed in host cells (Kjemtrup et al., 2000). In planta expression of a collection of deletion mutants of CRN2 indicates that this protein activates defense responses inside the plant cytoplasm, suggesting that the CRNs form a second class of oomycete cytoplasmic effectors (T. Torto-Alalibo and S. Kamoun, unpublished data). Unlike the H. parasitica proteins, none of the Phytophthora CRNs carry an RXLR sequence; instead, they have a distinct conserved N-terminal motif characterized by the consensus LXLFLAK (Figure 9). The finding that the RXLR sequence overlaps the LXLFLAK motif, resulting in RXLRLFLAK, in the 13 H. parasitica proteins suggests that these motifs might fulfill related host-targeting functions.
What factors drove the burst in effector gene duplication in P. ramorum? Host shifts coupled with the absence of sexual reproduction during the recent evolutionary history of P. ramorum may have favored clonal lineages with expanded effector gene families. However, in the absence of additional oomycete genome sequences, it is premature to speculate further on the biological significance of this finding.
In most proteins, neutral and purifying selection are the dominant evolutionary forces, with a high proportion of amino acid sites conserved as a result of structural and functional constraints (Li, 1997). Under these circumstances, the approximate method is usually not sensitive enough to detect diversifying selection, because it averages ω ratios over all codon sites in the protein (Yang and Bielawski, 2000). Nevertheless, we readily detected positive selection in >40% of the examined PGGs by measuring dN:dS ratios across the entire ORF sequences. This is likely due to the relatively small size and simple structure of the RXLR effectors, which average 100 to 200 amino acids in length, combined with the extreme selective pressures that probably have acted on these genes.
Remarkably, the overwhelming majority of the positively selected sites (138 of 140) localized to the C-terminal effector domain (Table 5). Similarly, higher dN:dS ratios were observed in the C-terminal region relative to the N terminus using the approximate method (Table 3, Figure 6). These findings suggest that the two main domains of the RXLR proteins have been exposed to different types of selective pressures and are consistent with the distinct functional roles of these two domains (Bos et al., 2003; Rehmany et al., 2005; Kamoun, 2006). Positive selection appears to have acted primarily on the C-terminal effector domain that functions inside plant cells and mediates the in planta activities of the effectors, while having less impact on the secretion and host-targeting sequences. Similar observations of uneven patterns of diversification have been made for several polymorphic oomycete effector genes, such as ATR1 and ATR13 of H. parasitica (Allen et al., 2004; Rehmany et al., 2005) and scr74 of P. infestans (Liu et al., 2005).
Although functional divergence of duplicated genes is an important evolutionary force for the emergence of new gene functions, accelerated evolution in paralog families can be driven by either positive selection or a relaxation of functional constraints due to gene redundancy (Zhang, 2003). Most likely, a combination of positive selection and a relaxation of selective constraints drove the evolution and functional divergence of the RXLR effector genes. The detection of putative pseudogenes among the RXLR effectors argues that relaxed selection has been a factor in the evolution of this family. However, the higher number of selected sites observed in the C-terminal domain over the N terminus argues in favor of positive selection for at least a subset of the RXLR effectors and is inconsistent with relaxed selection. Also, 25 of the 29 H. parasitica RXLR effectors deemed to be under positive selection are expressed in planta (A. Rougon and J. Jones, personal communication), indicating that they are unlikely to be under relaxed selection. Finally, the finding that the two paralogs of positively selected Ps PGG20 display distinct effector activities is consistent with positive selection driving functional divergence following gene duplication. In addition, mutation of the positively selected Thr-101 of Avh1b abolished the cell death elicitor activity of this effector, suggesting that this selected residue is critical for function.
Various genetic events may also affect the evolution of the RXLR genes besides the fixation of positively selected sites or relaxed selection. Several RXLR effector genes are clustered (Figure 8), suggesting that recombination and related types of genetic processes could contribute to the evolution of these genes. In plants, disease resistance genes frequently occur in clusters that span up to several megabases (Michelmore and Meyers, 1998; Meyers et al., 2003; Kuang et al., 2004). Extensive sequence exchange between clustered paralogs is known to give rise to genes with novel specificities and is thought to drive the evolution of resistance genes in concert with diversifying selection.
Pathogen effectors produce phenotypes that extend to plant cells and tissues and therefore should be key players in the arms races that drive the antagonistic coevolution between microbes and their hosts (Dawkins and Krebs, 1979; Dawkins, 1999). The high rates of gene loss, duplication, and diversification in the RXLR effectors of plant pathogenic oomycetes are consistent with their roles as both modulators and targets of host innate immunity. Our findings add to a growing body of literature on patterns of positive selection in plant and pathogen genes and the resulting accelerated amino acid substitution rates in sites that determine recognition by the host or the pathogen (Leckie et al., 1999; Gotesson et al., 2002; Allen et al., 2004; Bishop et al., 2004; Rose et al., 2004; Dodds et al., 2006; Ma et al., 2006). For instance, diversifying selection has acted on the flax rust AvrL567, a cytoplasmic effector that binds to the flax (Linum usitatissimum) L5, L6, and L7 resistance proteins to activate hypersensitivity and defense (Dodds et al., 2004). Positively selected sites in AvrL567 alter binding to plant resistance protein receptors, providing evidence that natural selection has acted on modifying the binding affinity between pathogen and plant proteins (Dodds et al., 2006). As with Avh1b, the positively selected sites identified in this study may alter the biochemical activity or the substrate binding affinity of the RXLR effectors. The next step is to identify the host targets in order to test this hypothesis and gain further insight into the evolutionary forces that shaped the effector secretome of oomycetes.
METHODS
Sequence Analysis
Similarity searches and the majority of the other bioinformatics analyses were performed locally on Mac OSX workstations using a combination of standard bioinformatics programs and customized Perl scripts. The SignalP v2.0 program (Nielsen et al., 1997) was ran on an Intel Linux workstation. Note that SignalP v2.0 predictions have been validated for oomycete secreted proteins using proteomics (Torto et al., 2003) and a yeast secretion assay (Lee et al., 2006). The main search program was BLAST (Altschul et al., 1997) and was typically ran with the filter off, unless indicated otherwise. The hmmpfam program (HMMer software; http://hmmer.wustl.edu) (Eddy, 1998) was used to search the Pfam HMM profile database of protein domains (Bateman et al., 2004). Multiple alignments were conducted using the programs ClustalW and ClustalX (Thompson et al., 1997), adjusted manually as necessary, and visualized with BOXSHADE (http://bioweb.pasteur.fr/seqanal/interfaces/boxshade.html) or Belvu alignment editor (Sonnhammer and Hollich, 2005). Sequence alignments were submitted to the WebLogo server (http://weblogo.berkeley.edu) to generate a sequence logo that graphically displays the consensus, as depicted in Figure 1B. Phylogenetic analyses of the CRNs was performed using the PHYLIP 3.66 package (available at evolution.genetics.washington.edu/phylip.html). We used the neighbor-joining, ML, and parsimony methods as implemented in PHYLIP. Signal peptide predictions were performed following the methods of Torto et al. (2003), except that a SignalP v2.0 HMM (Nielsen et al., 1997) score cutoff of >0.9 was used. Standard data analyses, tabulations of motif frequencies and positions, and graphical representations of the data were produced using the spreadsheet program Microsoft Excel.
To construct a hidden HMM profile of the RXLR domain, we manually aligned the RXLR or RXLR-EER domains of the 43 validated oomycete RXLR effectors (Table 1), including 5 to 10 amino acids before and after the RXLR/RXLR-EER sequence. The resulting alignment (Figure 1B) was used as an input to the hmmbuild program (HMMer software) (Eddy, 1998) to generate the RdEER HMM profile (see Supplemental Text online). We then used the RdEER HMM profile to search the collection of RXLR candidates using the hmmsearch program (Eddy, 1998) with default parameters.
Data Sets
A control data set of oomycete RXLR effectors corresponding to 43 oomycete genes (see Supplemental Table 1 online) was compiled from 13 validated effectors with avirulence or effector activities (Allen et al., 2004; Shan et al., 2004; Armstrong et al., 2005; Rehmany et al., 2005; Guo et al., 2006; our unpublished data) and 30 homologs of the validated effectors. The homologs were identified by BLASTP and TBLASTN searches (Altschul et al., 1997) against GenBank protein and nucleotide databases (Benson et al., 2007) using an E value cutoff of 10−4.
The genome sequence assemblies of Phytophthora ramorum and Phytophthora sojae (Tyler et al., 2006) were generated by the Department of Energy Joint Genome Institute and obtained from the Joint Genome Institute website (http://genome.jgi-psf.org) or GenBank (accession numbers AAQX01000000 and AAQY01000000). The draft genome sequence assemblies of Hyaloperonospora parasitica were produced by the Genome Sequencing Center at Washington University School of Medicine and were obtained from its web server at http://genome.wustl.edu/pub/organism/Fungi/Hyaloperonospora_parasitica/assembly/draft/Hyaloperonospora_parasitica-2.0.
To develop a curated database of eukaryotic proteomes, we collected the predicted proteomes of 48 eukaryotic species, including P. sojae and P. ramorum, from 10 major taxonomic groups from various sources and developed an exhaustive proteome database of 571,249 proteins, named darwin_571249.faa (for a full description, see Supplemental Table 1 online). The data included the complete proteomes of 47 eukaryotic species. This is an update of the database reported by Win et al. (2006).
RXLR Effector Identification Pipeline
A pipeline for the identification of candidate RXLR effectors from genome assemblies (Figure 4) was developed as follows. First, we translated all possible ORFs (defined here as either from ATG to a stop codon or from ATG to the end of a sequence) that encode 70 or more amino acids from both strands of the genomic DNA sequences. We then used SignalP v2.0 (Nielsen et al., 1997) to identify putative extracellular proteins following the criteria of Torto et al. (2003), except that a SignalP HMM score cutoff of >0.9 was used. Candidate RXLR effectors were selected from these secretome proteins using the following criteria: (1) the RXLR position must be between 30 and 60 amino acids; (2) the RXLR position must be downstream of the signal peptide cleavage site; and (3) SignalP v2.0 NN predicted cleavage site of <30 amino acids. There were redundant protein sequences in this set of RXLR candidates resulting from the translation of overlapping ORFs and potential sequence duplication due to assembly artifacts. These were identified by BLASTP searches and removed. For this purpose, sequences that showed 100% identity with an E value < 10−5 were deemed to be redundant, and only the sequence with the highest SignalP HMM score was retained. The obtained RXLR effector sets were also double-checked against the predicted proteome obtained from the current genome annotations of P. ramorum and P. sojae (Tyler et al., 2006). The identified 1352 candidate RXLR effector sequences and associated features are reported in detail in Supplemental Table 2 online.
Positive Selection Analyses
PGGs were identified from BLASTP searches of the RXLR effectors using three criteria: (1) similarity throughout the majority of the paralog coding sequences; (2) no or few gaps across the aligned sequences; and (3) at least 50% amino acid identity between the paralog sequences. These criteria helped to minimize the impact of the pitfalls of positive selection analyses, such as gap-induced misalignments and relaxed selection in pseudogenes, and resulted in robust alignments.
The resulting 99 PGGs were then used in positive selection analyses. We first aligned the protein sequences in each PGG using ClustalW (Thompson et al., 1997), extracted the coding DNA sequences, and aligned the codons corresponding to the amino acid sequence alignments using CodonAlign 2.0 (distributed by B.G. Hall, Bellingham Research Institute, Bellingham, WA; http://homepage.mac.com/barryghall/CodonAlign.html). We calculated the rates of nonsynonymous nucleotide substitutions per nonsynonymous site (dN) and the rates of synonymous nucleotide substitutions per synonymous site (dS) across all possible pairwise comparisons within each of the 99 PGGs using the approximate methods of Yang and Nielsen (2000) and Nei and Gojobori (1986) implemented in the YN00 program in the PAML 3.15 software package (Yang, 1997). The calculations were made using the full-length sequences, the N-terminal regions (from the start codon to the RXLR or RXLR-EER sequence), or the C-terminal regions (from the RXLR or RXLR-EER sequence to the end). The resulting dN, dS, and ω pairwise calculations are shown in Supplemental Table 3 online for all of the PGGs with at least one pairwise calculation showing ω > 1.2.
We also applied the ML method to 31 of the 99 PGGs with at least three sequences using the computer program codeml from the PAML 3.15 package (Yang, 1997). We used the codon substitution models M7 and M8. Model M8 allows for heterogeneous selection pressures across codon sites, while the null model M7 only allows ratio classes with ω < 1. Model M8 was implemented with at least two different starting ω values. Statistical significance was tested by comparing the null model M7 with the alternative M8 model using a likelihood ratio test. Twice the difference in log likelihood ratio between M7 and M8 was compared with a χ2 distribution with two degrees of freedom. The likelihood ratio test assesses whether the M8 alternative model fits the data better than the null M7 model and is known to be conservative in simulation tests (Anisimova et al., 2001; Thomas, 2006). Positively selected sites were identified using Bayes empirical Bayes analysis (Yang et al., 2005) implemented in codeml. Table 5 lists the likelihood ratio test and Bayes empirical Bayes statistics for the 18 PGGs that tested positive using the ML method.
The entire data sets, including the alignments and the outputs of the YN00 and codeml programs, were compiled in a FileMaker database and are available upon request.
Statistical Analyses
Statistical analyses to test the hypothesis that the frequency of RXLR proteins in both Phytophthora secretomes is significantly higher than that seen in the secretomes of other eukaryotes were performed using randomization techniques (Manly, 2007), because the frequency of RXLR-containing PEX proteins was not normally distributed among the 48 species examined. This involved randomly sampling 2 of the 48 secretomes with replacement and then computing the mean of the two RXLR frequencies. This random sampling was then repeated 999 times to provide a distribution of random values for comparison. The percentage of RXLR frequency means greater than or equal to the Phytophthora RXLR frequency mean was then calculated to determine the P value. Randomization tests were performed using the random number generation analysis tool of Microsoft Excel.
χ2 tests were used to examine whether the distribution of RXLR motifs in PEX proteins of Phytophthora species differed significantly from that seen in non-PEX proteins of the same species or in PEX proteins of other species. Each protein with an N-terminal RXLR motif (between positions 10 and 110) was placed in a decapeptide bin, depending on the location of the motif (e.g., if the motif was located at position 36, the protein was placed into bin 4). Then the number of proteins in each bin was determined for the PEX proteins or non-PEX proteins of each Phytophthora species and for the PEX proteins of all species combined. A χ2 contingency table test was then performed (Zar, 1999).
To determine whether the differences in observed dN and dS values from full-length pairwise comparisons that showed ω > 1 were significant, we performed two-sample t tests using the dN and dS values from these comparisons. The t tests were performed with dN and dS values from pair-wise comparisons with ω > 1, ω > 1.2, and ω > 1.5 (see Supplemental Table 5 online). To determine whether the ω values from pairwise comparisons of C- and N-terminal regions were significantly different, we performed a two-sample t test on their ω values after removing the values with dN = 0 or dS = 0.
Microbial Strains and Growth Conditions
Agrobacterium tumefaciens strains GV3101 and AGL0 (Hellens et al., 2000) were used in molecular cloning experiments and were routinely cultured at 28°C in Luria-Bertani medium using appropriate antibiotics (Sambrook and Russell, 2001). All bacterial DNA transformations were conducted by electroporation using standard protocols (Sambrook and Russell, 2001).
Plasmid Construction
We amplified both Avr1b-1 and Avh1b from genomic DNA from P. sojae isolate P6497 using oligonucleotide Avh1b/Avrb1-NSP-F-Cla (5′-GGAATCGATGACTGAGTACTCCGACGAAAC-3′) in combination with Avr1b-R-Not (5′-GGCGGCCGCTCAGCTCTGATACAGGTGAAAGGTG-3′) for Avr1b-1 and in combination with Avh1b-R-Not (5′-GGAGCGGCCGCTCAGTTCTGATACAGGTGAAAG-3′) for Avh1b. In addition, we amplified Avr1bAAM20939 from genomic DNA from a P. sojae field isolate kindly provided by Anne Dorrance (Ohio State University Ohio Agricultural Research and Development Center) using Avr1b-NSP-F-Cla and Avr1b-R-Not. The Avh1bT101K mutant was generated by overlap extension PCR using the strategy described by Kamoun et al. (1999). The first PCR was performed using oligonucleotides Avh1b/Avr1b-NSP-F-Cla and Avh1b-T101K-N (5′-CAGGGTGTACCCGTTCTTATCCCACTTCTC-3′), and a second PCR was performed using Avh1b-T101K-C (5′-AACGGGTACACCCTGCAGAAGATCAAGGAC-3′) and Av1h-R-Not. Reaction products were mixed in a 1:1 ratio and subjected to a third PCR with oligonucleotides Avh1b/Avr1b-NSP-F-Cla and Avh1b-R-Not. All amplicons were ligated into ClaI- and NotI-digested pGR106 (Lu et al., 2003).
Agrobacterium Transient Expression Assays
Recombinant Agrobacterium strains were grown as described elsewhere (Van der Hoorn et al., 2000), except that culturing steps were performed in Luria-Bertani medium supplemented with 50 μg of kanamycin. Agroinfiltration (Agrobacterium infiltration) and agroinfection (delivery of Potato virus X via Agrobacterium) experiments were performed on 4- to 6-week-old Nicotiana benthamiana plants (Van der Hoorn et al., 2000; Torto et al., 2003). Plants were grown and maintained throughout the experiments in a greenhouse with an ambient temperature of 22 to 25°C and high light intensity.
To assay for the activation of R3a, leaves of N. benthamiana were infiltrated with Agrobacterium strain AGL0 carrying the pBINplus-R3a construct (Huang et al., 2005) at a final OD600 of 0.3 in MMA induction buffer (1 liter of MMA, 5 g of Murashige and Skoog salts, 1.95 g of MES, 20 g of sucrose, and 200 μM acetosyringone, pH 5.6). One day after infiltration, leaves were challenged by wound-inoculation of Agrobacterium strains expressing Avr1b or Avh1b wild-type or mutant sequences. As a control, each leaf was wound-inoculated with Agrobacterium strains carrying pGR106-AVR3aKI_23-147 or pGR106-AVR3aEM_23-147 (Bos et al., 2006). For the expression of Avh1b in transgenic N. benthamiana expressing R3a, Agrobacterium strains were wound-inoculated on leaves of a transgenic lines produced by Edwin van der Vossen. Cell death symptoms around the inoculation site, indicative of R3a activation, were scored at 4 to 7 d after inoculation.
Supplemental Data
The following materials are available in the online version of this article.
Supplemental Figure 1. Bidirectional Best BLASTP Hit Searches of the RXLR Effectors (E Value Cutoff < 10−10).
Supplemental Figure 2. Positive Selection in Avr1b-1-Like Genes of P. sojae.
Supplemental Table 1. Protein Data Sets Used for RXLR Motif Analysis.
Supplemental Table 2. Description of the 1352 Candidate RXLR Effectors Identified in the Three Oomycetes.
Supplemental Table 3. dN, dS, and dN:dS Calculations for All Pairwise Comparisons of the Examined PGGs.
Supplemental Table 4. Extended Summary of Positive Selection Analyses of Paralogous Gene Groups of P. ramorum, P. sojae, and H. parasitica.
Supplemental Table 5. Summary of t Tests for dN and dS Values.
Supplemental Table 6. Top Pfam Hits of the Candidate RXLR Effectors.
Supplemental Text. Hidden Markov Model File Based on the Consensus Shown in Figure 1B.
Supplementary Material
Acknowledgments
We thank Rick Lehtinen and Michael Collins for advice on the statistical analyses; Soledad Benitez, Jorunn Bos, Carla Garzon, and Sang-Keun Oh for comments on the manuscript; Anne Dorrance and Edwin van der Vossen for providing biomaterial; Kerilynn Jagger for technical assistance; and Juan Valdez and Mr. Roboto for their stimulating influence. This research was supported by National Science Foundation Plant Genome Grant DBI-0211659 and by state and federal funds appropriated to the Ohio State University Ohio Agricultural Research and Development Center.
The author responsible for distribution of materials integral to the findings presented in this article in accordance with the policy described in the Instructions for Authors (www.plantcell.org) is: Sophien Kamoun (kamoun.1@osu.edu).
Online version contains Web-only data.
References
- Allen, R.L., Bittner-Eddy, P.D., Grenville-Briggs, L.J., Meitz, J.C., Rehmany, A.P., Rose, L.E., and Beynon, J.L. (2004). Host-parasite coevolutionary conflict between Arabidopsis and downy mildew. Science 306 1957–1960. [DOI] [PubMed] [Google Scholar]
- Altschul, S.F., Madden, T.L., Schaffer, A.A., Zhang, J., Zhang, Z., Miller, W., and Lipman, D.J. (1997). Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 17 3389–3402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anisimova, M., Bielawski, J.P., and Yang, Z. (2001). Accuracy and power of the likelihood ratio test in detecting adaptive molecular evolution. Mol. Biol. Evol. 18 1585–1592. [DOI] [PubMed] [Google Scholar]
- Armstrong, M.R., et al. (2005). An ancestral oomycete locus contains late blight avirulence gene Avr3a, encoding a protein that is recognized in the host cytoplasm. Proc. Natl. Acad. Sci. USA 102 7766–7771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bateman, A., et al. (2004). The Pfam protein families database. Nucleic Acids Res. 32 D138–D141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Benson, D.A., Karsch-Mizrachi, I., Lipman, D.J., Ostell, J., and Wheeler, D.L. (2007). GenBank. Nucleic Acids Res. 35 D21–D25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bhattacharjee, S., Hiller, N.L., Liolios, K., Win, J., Kanneganti, T.D., Young, C., Kamoun, S., and Haldar, K. (2006). The malarial host-targeting signal is conserved in the Irish potato famine pathogen. PLoS Pathog. 2 e50. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Birch, P.R., Rehmany, A.P., Pritchard, L., Kamoun, S., and Beynon, J.L. (2006). Trafficking arms: Oomycete effectors enter host plant cells. Trends Microbiol. 14 8–11. [DOI] [PubMed] [Google Scholar]
- Bishop, J.G., Ripoll, D.R., Bashir, S., Damasceno, C.M., Seeds, J.D., and Rose, J.K. (2004). Selection on Glycine beta-1,3-endoglucanase genes differentially inhibited by a Phytophthora glucanase inhibitor protein. Genetics 169 1009–1019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bos, J.I., Kanneganti, T.D., Young, C., Cakir, C., Huitema, E., Win, J., Armstrong, M.R., Birch, P.R., and Kamoun, S. (2006). The C-terminal half of Phytophthora infestans RXLR effector AVR3a is sufficient to trigger R3a-mediated hypersensitivity and suppress INF1-induced cell death in Nicotiana benthamiana. Plant J. 48 165–176. [DOI] [PubMed] [Google Scholar]
- Bos, J.I.B., Armstrong, M., Whisson, S.C., Torto, T., Ochwo, M., Birch, P.R.J., and Kamoun, S. (2003). Intraspecific comparative genomics to identify avirulence genes from Phytophthora. New Phytol. 159 63–72. [DOI] [PubMed] [Google Scholar]
- Carroll, S.B. (2006). The Making of the Fittest: DNA and the Ultimate Forensic Record of Evolution. (New York: W.W. Norton).
- Chisholm, S.T., Coaker, G., Day, B., and Staskawicz, B.J. (2006). Host-microbe interactions: Shaping the evolution of the plant immune response. Cell 124 803–814. [DOI] [PubMed] [Google Scholar]
- Constantinescu, O., and Fatehi, J. (2002). Peronospora-like fungi (Chromista, Peronosporales) parasitic on Brassicaceae and related hosts. Nova Hedwigia 74 291–338. [Google Scholar]
- Cooke, D.E., Drenth, A., Duncan, J.M., Wagels, G., and Brasier, C.M. (2000). A molecular phylogeny of Phytophthora and related oomycetes. Fungal Genet. Biol. 30 17–32. [DOI] [PubMed] [Google Scholar]
- Cornelis, G.R. (2006). The type III secretion injectisome. Nat. Rev. Microbiol. 4 811–825. [DOI] [PubMed] [Google Scholar]
- Dawkins, R. (1999). The Extended Phenotype: The Long Reach of the Gene. (Oxford, UK: Oxford University Press).
- Dawkins, R., and Krebs, J.R. (1979). Arms races between and within species. Proc. R. Soc. Lond. B Biol. Sci. 205 489–511. [DOI] [PubMed] [Google Scholar]
- Dodds, P.N., Lawrence, G.J., Catanzariti, A.M., Ayliffe, M.A., and Ellis, J.G. (2004). The Melampsora lini AvrL567 avirulence genes are expressed in haustoria and their products are recognized inside plant cells. Plant Cell 16 755–768. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dodds, P.N., Lawrence, G.J., Catanzariti, A.M., Teh, T., Wang, C.I., Ayliffe, M.A., Kobe, B., and Ellis, J.G. (2006). Direct protein interaction underlies gene-for-gene specificity and coevolution of the flax resistance genes and flax rust avirulence genes. Proc. Natl. Acad. Sci. USA 103 8888–8893. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eddy, S.R. (1998). Profile hidden Markov models. Bioinformatics 14 755–763. [DOI] [PubMed] [Google Scholar]
- Ellis, J., Catanzariti, A.M., and Dodds, P. (2006). The problem of how fungal and oomycete avirulence proteins enter plant cells. Trends Plant Sci. 11 61–63. [DOI] [PubMed] [Google Scholar]
- Erwin, D.C., and Ribeiro, O.K. (1996). Phytophthora Diseases Worldwide. (St. Paul, MN: APS Press).
- Galan, J.E., and Wolf-Watz, H. (2006). Protein delivery into eukaryotic cells by type III secretion machines. Nature 444 567–573. [DOI] [PubMed] [Google Scholar]
- Goker, M., Voglmayr, H., Riethmuller, A., and Oberwinkler, F. (2007). How do obligate parasites evolve? A multi-gene phylogenetic analysis of downy mildews. Fungal Genet. Biol. 44 105–122. [DOI] [PubMed] [Google Scholar]
- Gotesson, A., Marshall, J.S., Jones, D.A., and Hardham, A.R. (2002). Characterization and evolutionary analysis of a large polygalacturonase gene family in the oomycete plant pathogen Phytophthora cinnamomi. Mol. Plant Microbe Interact. 15 907–921. [DOI] [PubMed] [Google Scholar]
- Grant, S.R., Fisher, E.J., Chang, J.H., Mole, B.M., and Dangl, J.L. (2006). Subterfuge and manipulation: Type III effector proteins of phytopathogenic bacteria. Annu. Rev. Microbiol. 60 425–449. [DOI] [PubMed] [Google Scholar]
- Guo, J., Jiang, R.H., Kamphuis, L.G., and Govers, F. (2006). A cDNA-AFLP based strategy to identify transcripts associated with avirulence in Phytophthora infestans. Fungal Genet. Biol. 43 111–123. [DOI] [PubMed] [Google Scholar]
- Hellens, R., Mullineaux, P., and Klee, H. (2000). Technical focus. A guide to Agrobacterium tumefaciens binary Ti vectors. Trends Plant Sci. 5 446–451. [DOI] [PubMed] [Google Scholar]
- Hiller, N.L., Bhattacharjee, S., van Ooij, C., Liolios, K., Harrison, T., Lopez-Estrano, C., and Haldar, K. (2004). A host-targeting signal in virulence proteins reveals a secretome in malarial infection. Science 306 1934–1937. [DOI] [PubMed] [Google Scholar]
- Huang, G., Allen, R., Davis, E.L., Baum, T.J., and Hussey, R.S. (2006. a). Engineering broad root-knot resistance in transgenic plants by RNAi silencing of a conserved and essential root-knot nematode parasitism gene. Proc. Natl. Acad. Sci. USA 103 14302–14306. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang, G., Dong, R., Allen, R., Davis, E.L., Baum, T.J., and Hussey, R.S. (2006. b). A root-knot nematode secretory peptide functions as a ligand for a plant transcription factor. Mol. Plant Microbe Interact. 19 463–470. [DOI] [PubMed] [Google Scholar]
- Huang, S., van der Vossen, E.A.G., Kuang, H., Vleeshouwers, V.G.A.A., Zhang, N., Borm, T.J.A., van Eck, H.J., Baker, B., Jacobsen, E., and Visser, R.G.F. (2005). Comparative genomics enabled the isolation of the R3a late blight resistance gene in potato. Plant J. 42 251–261. [DOI] [PubMed] [Google Scholar]
- Jones, J.D., and Dangl, J.L. (2006). The plant immune system. Nature 444 323–329. [DOI] [PubMed] [Google Scholar]
- Kamoun, S. (2003). Molecular genetics of pathogenic oomycetes. Eukaryot. Cell 2 191–199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kamoun, S. (2006). A catalogue of the effector secretome of plant pathogenic oomycetes. Annu. Rev. Phytopathol. 44 41–60. [DOI] [PubMed] [Google Scholar]
- Kamoun, S., Honee, G., Weide, R., Lauge, R., Kooman-Gersmann, M., de Groot, K., Govers, F., and de Wit, P.J.G.M. (1999). The fungal gene Avr9 and the oomycete gene inf1 confer avirulence to Potato virus X on tobacco. Mol. Plant Microbe Interact. 12 459–462. [Google Scholar]
- Kjemtrup, S., Nimchuk, Z., and Dangl, J.L. (2000). Effector proteins of phytopathogenic bacteria: Bifunctional signals in virulence and host recognition. Curr. Opin. Microbiol. 3 73–78. [DOI] [PubMed] [Google Scholar]
- Kroon, L.P., Bakker, F.T., van den Bosch, G.B., Bonants, P.J., and Flier, W.G. (2004). Phylogenetic analysis of Phytophthora species based on mitochondrial and nuclear DNA sequences. Fungal Genet. Biol. 41 766–782. [DOI] [PubMed] [Google Scholar]
- Kuang, H., Woo, S.S., Meyers, B.C., Nevo, E., and Michelmore, R.W. (2004). Multiple genetic processes result in heterogeneous rates of evolution within the major cluster disease resistance genes in lettuce. Plant Cell 16 2870–2894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leckie, F., Mattei, B., Capodicasa, C., Hemmings, A., Nuss, L., Aracri, B., De Lorenzo, G., and Cervone, F. (1999). The specificity of polygalacturonase-inhibiting protein (PGIP): A single amino acid substitution in the solvent-exposed beta-strand/beta-turn region of the leucine-rich repeats (LRRs) confers a new recognition capability. EMBO J. 18 2352–2363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee, S.J., Kelley, B.S., Damasceno, C.M., St. John, B., Kim, B.S., Kim, B.D., and Rose, J.K. (2006). A functional screen to characterize the secretomes of eukaryotic pathogens and their hosts in planta. Mol. Plant Microbe Interact. 19 1368–1377. [DOI] [PubMed] [Google Scholar]
- Li, W.-H. (1997). Molecular Evolution. (Sunderland, MA: Sinauer Associates).
- Liu, Z., Bos, J.I.B., Armstrong, M., Whisson, S.C., da Cunha, L., Torto-Alalibo, T., Win, J., Avrova, A.O., Wright, F., Birch, P.R., and Kamoun, S. (2005). Patterns of diversifying selection in the phytotoxin-like scr74 gene family of Phytophthora infestans. Mol. Biol. Evol. 22 659–672. [DOI] [PubMed] [Google Scholar]
- Lu, R., Malcuit, I., Moffett, P., Ruiz, M.T., Peart, J., Wu, A.J., Rathjen, J.P., Bendahmane, A., Day, L., and Baulcombe, D.C. (2003). High throughput virus-induced gene silencing implicates heat shock protein 90 in plant disease resistance. EMBO J. 22 5690–5699. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ma, W., Dong, F.F., Stavrinides, J., and Guttman, D.S. (2006). Type III effector diversification via both pathoadaptation and horizontal transfer in response to a coevolutionary arms race. PLoS Genet. 2 e209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manly, B.F.J. (2007). Randomization, Bootstrap and Monte Carlo Methods in Biology, 3rd ed. (Boca Raton, FL: Chapman & Hall/CRC).
- Marti, M., Good, R.T., Rug, M., Knuepfer, E., and Cowman, A.F. (2004). Targeting malaria virulence and remodeling proteins to the host erythrocyte. Science 306 1930–1933. [DOI] [PubMed] [Google Scholar]
- Meyers, B.C., Kozik, A., Griego, A., Kuang, H., and Michelmore, R.W. (2003). Genome-wide analysis of NBS-LRR-encoding genes in Arabidopsis. Plant Cell 15 809–834. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Michelmore, R.W., and Meyers, B.C. (1998). Clusters of resistance genes in plants evolve by divergent selection and a birth-and-death process. Genome Res. 8 1113–1130. [DOI] [PubMed] [Google Scholar]
- Nei, M., and Gojobori, T. (1986). Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol. Biol. Evol. 3 418–426. [DOI] [PubMed] [Google Scholar]
- Nielsen, H., Engelbrecht, J., Brunak, S., and von Heijne, G. (1997). Identification of prokaryotic and eukaryotic signal peptides and prediction of their cleavage sites. Protein Eng. 10 1–6. [DOI] [PubMed] [Google Scholar]
- Nielsen, R., and Yang, Z. (1998). Likelihood models for detecting positively selected amino acid sites and applications to the HIV-1 envelope gene. Genetics 148 929–936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Connell, R.J., and Panstruga, R. (2006). Tete a tete inside a plant cell: Establishing compatibility between plants and biotrophic fungi and oomycetes. New Phytol. 171 699–718. [DOI] [PubMed] [Google Scholar]
- Rehmany, A.P., Gordon, A., Rose, L.E., Allen, R.L., Armstrong, M.R., Whisson, S.C., Kamoun, S., Tyler, B.M., Birch, P.R., and Beynon, J.L. (2005). Differential recognition of highly divergent downy mildew avirulence gene alleles by RPP1 resistance genes from two Arabidopsis lines. Plant Cell 17 1839–1850. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rizzo, D.M., Garbelotto, M., Davidson, J.M., Slaughter, G.W., and Koike, S.T. (2002). Phytophthora ramorum as the cause of extensive mortality of Quercus spp. and Lithocarpus densiflorus in California. Plant Dis. 86 205–214. [DOI] [PubMed] [Google Scholar]
- Rizzo, D.M., Garbelotto, M., and Hansen, E.M. (2005). Phytophthora ramorum: Integrative research and management of an emerging pathogen in California and Oregon forests. Annu. Rev. Phytopathol. 43 309–335. [DOI] [PubMed] [Google Scholar]
- Rohmer, L., Guttman, D.S., and Dangl, J.L. (2004). Diverse evolutionary mechanisms shape the type III effector virulence factor repertoire in the plant pathogen Pseudomonas syringae. Genetics 167 1341–1360. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rose, L.E., Bittner-Eddy, P.D., Langley, C.H., Holub, E.B., Michelmore, R.W., and Beynon, J.L. (2004). The maintenance of extreme amino acid diversity at the disease resistance gene, RPP13, in Arabidopsis thaliana. Genetics 166 1517–1527. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sambrook, J., and Russell, D.W. (2001). Molecular Cloning. (Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press).
- Schurch, S., Linde, C.C., Knogge, W., Jackson, L.F., and McDonald, B.A. (2004). Molecular population genetic analysis differentiates two virulence mechanisms of the fungal avirulence gene NIP1. Mol. Plant Microbe Interact. 17 1114–1125. [DOI] [PubMed] [Google Scholar]
- Shan, W., Cao, M., Leung, D., and Tyler, B.M. (2004). The Avr1b locus of Phytophthora sojae encodes an elicitor and a regulator required for avirulence on soybean plants carrying resistance gene Rps1b. Mol. Plant Microbe Interact. 17 394–403. [DOI] [PubMed] [Google Scholar]
- Sonnhammer, E.L., and Hollich, V. (2005). Scoredist: A simple and robust protein sequence distance estimator. BMC Bioinformatics 6 108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stavrinides, J., Ma, W., and Guttman, D.S. (2006). Terminal reassortment drives the quantum evolution of type III effectors in bacterial pathogens. PLoS Pathog. 2 e104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas, J.H. (2006). Adaptive evolution in two large families of ubiquitin-ligase adapters in nematodes and plants. Genome Res. 16 1017–1030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas, J.H., Kelley, J.L., Robertson, H.M., Ly, K., and Swanson, W.J. (2005). Adaptive evolution in the SRZ chemoreceptor families of Caenorhabditis elegans and Caenorhabditis briggsae. Proc. Natl. Acad. Sci. USA 102 4476–4481. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson, J.D., Gibson, T.J., Plewniak, F., Jeanmougin, F., and Higgins, D.G. (1997). The CLUSTAL_X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 25 4876–4882. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Torto, T., Li, S., Styer, A., Huitema, E., Testa, A., Gow, N.A.R., van West, P., and Kamoun, S. (2003). EST mining and functional expression assays identify extracellular effector proteins from Phytophthora. Genome Res. 13 1675–1685. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tyler, B.M., et al. (2006). Phytophthora genome sequences uncover evolutionary origins and mechanisms of pathogenesis. Science 313 1261–1266. [DOI] [PubMed] [Google Scholar]
- Van der Hoorn, R.A.L., Laurent, F., Roth, R., and De Wit, P.J.G.M. (2000). Agroinfiltration is a versatile tool that facilitates comparative analyses of Avr9/Cf-9-induced and Avr4/Cf-4-induced necrosis. Mol. Plant Microbe Interact. 16 669–680. [DOI] [PubMed] [Google Scholar]
- Win, J., Kanneganti, T.D., Torto-Alalibo, T., and Kamoun, S. (2006). Computational and comparative analyses of 150 full-length cDNA sequences from the oomycete plant pathogen Phytophthora infestans. Fungal Genet. Biol. 43 20–33. [DOI] [PubMed] [Google Scholar]
- Yang, Z. (1997). PAML: A program package for phylogenetic analysis by maximum likelihood. Comput. Appl. Biosci. 13 555–556. [DOI] [PubMed] [Google Scholar]
- Yang, Z., and Bielawski, J.P. (2000). Statistical methods for detecting molecular adaptation. Trends Ecol. Evol. 15 496–503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang, Z., and Nielsen, R. (2000). Estimating synonymous and nonsynonymous substitution rates under realistic evolutionary models. Mol. Biol. Evol. 17 32–43. [DOI] [PubMed] [Google Scholar]
- Yang, Z., Nielsen, R., Goldman, N., and Pedersen, A.M. (2000). Codon-substitution models for heterogeneous selection pressure at amino acid sites. Genetics 155 431–449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang, Z., Wong, W.S., and Nielsen, R. (2005). Bayes empirical Bayes inference of amino acid sites under positive selection. Mol. Biol. Evol. 22 1107–1118. [DOI] [PubMed] [Google Scholar]
- Zar, J.H. (1999). Biostatistical Analysis, 4th ed. (Englewood Cliffs, NJ: Prentice-Hall).
- Zhang, J. (2003). Evolution by gene duplication: An update. Trends Ecol. Evol. 18 292–298. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.