Abstract
We describe the first functional insertion sequence (IS) element in Lactobacillus plantarum. ISLpl1, an IS30-related element, was found on the pLp3 plasmid in strain FB335. By selection of spontaneous mutants able to grow in the presence of uracil, it was demonstrated that the IS had transposed into the uracil phosphoribosyltransferase-encoding gene upp on the FB335 chromosome. The plasmid-carried IS element was also sequenced, and a second potential IS element was found: ISLpl2, an IS150-related element adjacent to ISLpl1. When Southern hybridization was used, the copy number and genome (plasmid versus chromosome) distribution data revealed different numbers and patterns of ISLpl1-related sequences in different L. plantarum strains as well as in Pediococcus strains. The ISLpl1 pattern changed over many generations of the strain L. plantarum NCIMB 1406. This finding strongly supports our hypothesis that ISLpl1 is a mobile element in L. plantarum. Database analysis revealed five quasi-identical ISLpl1 elements in Lactobacillus, Pediococcus, and Oenococcus strains. Three of these elements may be cryptic IS, since point mutations or 1-nucleotide deletions were found in their transposase-encoding genes. In some cases, ISLpl1 was linked to genes involved in cold shock adaptation, bacteriocin production, sugar utilization, or antibiotic resistance. ISLpl1 is transferred among lactic acid bacteria (LAB) and may play a role in LAB genome plasticity and adaptation to their environment.
Lactic acid bacteria (LAB) are gram-positive bacteria involved in fermentation of a wide variety of plants, meats, fish, and dairy products. The genus Lactobacillus comprises one of the largest groups of organisms used in fermentation processes. In industrial fermentations, starter cultures containing mixtures of LAB are used. Promiscuity between different members of microbial populations has facilitated intergenomic rearrangements, and mobile elements have contributed to genetic polymorphism and DNA dynamics (2, 7, 14). Strains of Streptococcus thermophilus have been selected for exopolysaccharide synthesis to improve fermented food texture. S. thermophilus exopolysaccharide locus variability was linked to a 14-kb region with seven insertion sequences (IS) acquired by horizontal transfer from Lactococcus lactis, a LAB used in starter cultures with S. thermophilus (3, 14). Genetic exchange between Lactobacillus plantarum and Pediococcus pentosaceus was also suggested by homologous sucrase genes (98% nucleic acid identity) found in raffinose-sucrose gene clusters flanked by IS elements (18). The comparison of increasing numbers of sequenced bacterial genomes can highlight the extent of gene exchange between microorganisms and help identify potential mobile elements.
Few mobile IS elements have been characterized in Lactobacillus. These include ISL2 (30) in Lactobacillus helveticus, ISL3 (13) in Lactobacillus delbrueckii, IS1223 (28) in Lactobacillus johnsonii, and IS1163 (22) and IS1520 (23) in Lactobacillus sakei. The functional L. plantarum IS element characterized in this work was quasi-identical to LAB IS elements found in databases. This prompted us to study its distribution among other LAB and to discuss gene exchange within LAB.
MATERIALS AND METHODS
Bacterial strain cultivation and physiological tests.
The sources of the lactobacilli are listed in Table 1. Strains were propagated in MRS broth (Difco Laboratories) at 30°C in a 4% CO2-enriched atmosphere with a water-jacketed CH/P incubator (Forma Scientific). Glycine (2.5%) was added when cells were lysed for DNA extractions. Independent spontaneous uracil-resistant variants of L. plantarum FB335 were selected as follows. Each of 40 MRS broth-grown colony isolates was suspended in 1 ml of sterile physiological water (water containing NaCl at 9 g/liter) to obtain around 108 cells/ml, and 30 μl of the suspension was spread on agar plates of defined DLA medium (4) supplemented with uracil (50 μg/ml). After 4 days of incubation in the dark at 30°C in 4% CO2-enriched air, only one clone of each plate was taken for further analysis. All 40 of the independent isolates conserved the ability to grow on DLA medium supplemented with uracil after growth in rich MRS medium.
TABLE 1.
ISLp11 distribution in LAB
| Strainsa
|
Southern hybridization profile(s) detected with an ISLp11 probe (detected bands in kb)
|
ISLp11 genetic location (source) | ||
|---|---|---|---|---|
| Name | Source and comment | HaeIII digest(s) | HindIII digest(s) | |
| L. plantarum FB335 | ΔcarAB mutant of ATCC 8014 (see reference 20) | 2 | 3.4 | Plasmid pLp3 (this work) |
| L. plantarum HN38 | upp::ISLp11; spontaneous mutant of FB335 (see this work) | 2.5; 2 | 11.5; 3.4 | Plasmid pLp3 and chromosome (this work) |
| L. plantarum ATCC 14917T | ATCC; pickled cabbage | 2.6 | 5 | Plasmid pLF1 (EMBL accession no. AF508808) |
| L. plantarum NCIMB 1406 | NCIMB; dental carries | 3.8; 3.3; 2.8; 2.6; 2.5; 1.9; 1.8; 1.6; 1.4; 1.3; 0.8 | 18.5; 12.5; 11; 9.0; 8.4; 7.7; 6.9; 5.9; 5.5; 5.3; 4.9; 4.1; 3.4; 3.0; 2.8; 2.1; 1.7b; 1.2b | Plasmid and chromosome (this work) |
| L. plantarum NCIMB 8826 | NCIMB; human saliva | None | None | None |
| L. plantarum CNRZ 1891 | Munster-type cheese (see reference 12a) | 1.6; 1.4; 0.9; 0.7 | 10; 7.5; 6.2b; 4.7; 3.1; 2.0b | Not tested |
| Pediococcus acidilactici H | See reference 12a | Not tested | 15; 6.2; 4.8 | Not tested |
ATCC, American Type Culture Collection, Manassas, Va.; CNRZ, Centre National de Recherches Zootechniques, Jouy-en-Josas, France; NCIMB, National Collection of Industrial and Marine Bacteria, Aberdeen, Scotland. The Lactobacillus strains belong to the species L. plantarum (6).
Bands detected at lower stringency only (see Materials and Methods).
The genetic stability of the IS element patterns was evaluated over time. MRS broth (12.5 ml) from a single-colony isolate of strain NCIMB 1406 on an MRS agar plate was inoculated at 30°C with no agitation in 4% CO2-enriched air. After the stationary-growth phase was reached, 5 μl of the cell suspension (containing around 106 CFU) was inoculated in 12.5 ml of fresh MRS broth and left until the stationary-growth phase was reached once again. The number of generations was deduced from the viable bacterial counts on MRS plates in the inoculum and after growth. When 11 serial passages in MRS broth had been performed, MRS agar colony isolates were selected and stored at −80°C. Total DNAs from four clones were extracted and probed with ISLpl1-specific DNA to compare their Southern hybridization profiles.
DNA purification.
Total DNA (plasmid and chromosome) was extracted as described previously (6). Two protocols were used to purify plasmid DNA. For all the LAB strains except L. plantarum FB335, after cell lysis (26) the chromosomal DNA was separated from the superhelical plasmid DNA by equilibrium centrifugation in a CsCl-ethidium bromide gradient (21). To purify FB335 plasmid DNA, a QIA filter plasmid Midi kit protocol (QIAGEN) was modified to facilitate cell lysis. Cells from a 0.5-liter MRS broth culture in late exponential phase were washed with sterile water, stored overnight at −20°C, and then treated for 1 h at 37°C in 10 ml of lysis buffer containing 20 mg of lysozyme/ml.
Southern hybridization.
DNA was cut with the restriction enzyme HaeIII or HindIII (which did not cut within the insertion element) and then transferred onto Hybond positively charged nylon membranes (Amersham Pharmacia Biotech) after agarose gel electrophoresis. A specific ISLpl1 fragment was PCR amplified from the copy interrupting the uracil phosphoribosyltransferase-encoding gene upp in strain HN38 (Fig. 1) and labeled (using a nonradioactive DNA labeling and detection kit [Roche Diagnostics]) with digoxigenin-11-dUTP. The composition of the hybridization solution was as follows: 0.75 M NaCl, 0.08 M sodium citrate (pH 7.0), 0.1% (wt/vol) N-lauroylsarcosine sodium salt (Sigma), 0.02% (wt/vol) sodium dodecyl sulfate, and 1% (wt/vol) blocking agent (Roche Diagnostics) dissolved at 60°C. High- and low-hybridization stringencies were performed overnight at 68 and 42°C, respectively. Nucleic acid hybrids were detected using the alkaline phosphatase chemiluminescent substrate CDP-star (Roche Diagnostics). Stripping and reprobing the blots were performed as recommended by the manufacturer.
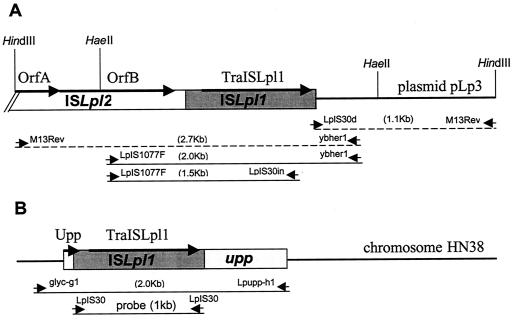
FIG. 1.
Genetic organization of two ISLpl1 loci. ISLpl1 is represented by a gray rectangle, and adjacent white rectangles represent ISLpl2 (A) or the upp gene (B). The ORF orientations are designated with thick arrows. Proposed initiation codons were TTG for ISLpl1 transposase and GTG for ISLpl2 OrfB. Only the C-terminal part of OrfA was determined. Primer names and orientations are indicated with small arrows for each PCR product; continuous lines and broken lines represent PCR products amplified directly from native DNA templates or by inverse PCR, respectively. Dashed lines represent inverse PCR-amplified fragments. (A) In the pLp3 plasmid in strains FB335 and HN38, ISLpl1 is contiguous to ISLpl2 (GenBank accession no. AF459445). Relevant restriction enzyme sites are indicated. (B) In the chromosome of HN38, ISLpl1 is also inserted in the upp gene. The location of the 1-kb specific ISLpl1 probe used in the Southern hybridization is shown.
Standard PCR amplification.
The PCR was catalyzed with Taq DNA polymerase (Q-Biogene) (1 U) using 100 ng of template DNA in a 50-μl final reaction volume consisting of 10 mM Tris HCl (pH 9.0), 50 mM KCl, 1.5 mM MgCl2, 0.1% Triton X-100, 0.2 mg of bovine serum albumin/ml, 50 pmol of each specific primer, and a 312 μM concentration of each deoxynucleoside triphosphate. The PCR cycle consisted of 1 min of denaturation at 95°C followed by 35 cycles of 35 s at 94°C, 35 s of annealing at 55°C, and 5 min of elongation at 72°C in a DNA engine PTC-200 Peltier thermal cycler (MJ Research). Finally, the PCR was completed by a postelongation treatment for 20 min at 72°C. The sequences of the primer are listed in Table 2, and the sizes of the PCR products obtained with different primer sets are shown in Fig. 1. The primer Lp-IS30 (hybridizing to both ISLp11 ends) was used with a modified PCR cycle (52°C for annealing and 2 min for elongation).
TABLE 2.
Primers used in PCR experiments
| Primer | Sequence (5′→3′)a | Nucleotide position | EMBL database accession no. |
|---|---|---|---|
| glyc-g1 | CGGTCAACAAAGAAGCGATTCCG (F) | 2144349-2144372 | AL935263 |
| Lpupp-h1 | TATCCATCGCCATGATGGCTGAACC (R) | 698-722 | AJ012720 |
| Lp-IS30b | GGTAGATTGYAAAWTTAATCCGA | 1219-1241 | AF459445 |
| 2239-2261 | AF459445 | ||
| LpIS1077F | GCCTTTAATAAGCAAACTCGC (F) | 553-573 | AF459445 |
| LpIS30in | GATTAAAGCGTTCATTACTACC (R) | 2062-2083 | AF459445 |
| LpIS1077F | GCCTTTAATAAGCAAACTCGC (F) | 553-573 | AF459445 |
| ybher1 | GAAGTAGCCAAAGAGCACGG (R) | 2625-2644 | AF459445 |
| Lp-IS30d | GGTTATGCTGACAAATTTGTCC (R) | 2211-2232 | AF459445 |
| M13Rev | CAGGAAACAGCTATGACCATGATTAC (F) | 456-481 | X02514 |
F, forward primer; R, reverse primer.
This primer binds to both ends of ISLp11. Y = C or T; W = A or T.
Inverse PCR amplifications.
To determine the ISLpl1 locus in strain FB 335, PCR was performed on ligation mixtures. DNA was digested by the restriction enzyme HindIII (which did not cut within the insertion element) and ligated into HindIII-linearized pUC19 plasmid (29). The 1.1-kb right-junction fragment was obtained (using an annealing temperature of 57°C and primer set M13Rev-Lp-IS30d) and subsequently sequenced with the same primers. Primer ybher1 was deduced (employing the 1.1-kb sequence) for use in a second inverse PCR to amplify the complete IS element. When the same HindIII ligation mixture was used with primer set M13Rev-ybher1, the inverse PCR amplification generated a 2.7-kb fragment. The 1.1- and 2.7-kb inverse PCR-amplified fragments were partially overlapping (Fig. 1A).
DNA sequence.
Prior to sequencing, PCR fragments were purified using S-400 HR MicroSpin columns (Amersham Pharmacia Biotech). DNA sequencing was performed with an Applied Biosystems 373 DNA sequencer. Ribosome binding sites were identified by analyzing their complementarity to the 3′ end of L. plantarum 16S rRNA (GenBank accession no. M58827). Sequences were analyzed using BLAST programs (1) by comparisons with sequences listed in DNA and protein databases.
Nucleotide sequence accession numbers.
The DNA sequences of ISLpl1-ISLpl2 have been deposited in the GenBank database under accession no. AF459445. ISLpl1 and ISLpl2 were named by IS database curators (http://www-is.biotoul.fr).
RESULTS
Discovery of the insertion element ISLpl1.
In our study of pyrimidine metabolism in L. plantarum, we searched for deregulated mutants, which may have lost pyrimidine repression by exogenous pyrimidines. A positive screening with spontaneous selection of uracil-resistant clones was set up to isolate deregulated mutants (19) (Fig. 2), and the genetic lesions were characterized in 37 independent mutants (H. Nicoloff and F. Bringel, unpublished data). To find the mutations within the upp gene encoding uracil phosphoribosyltransferase (EC 2.4.2.9), the upp locus was amplified in the different mutants with the primer set glyc-g1-Lpupp-h1 (Fig. 1B). A fragment that was 1 kb larger than the fragments found in the other mutants was found in mutant HN38 and subsequently sequenced. A 1,043-bp sequence (called ISLpl1) was inserted 32 nucleotides (nt) after the upp initiation codon (data not shown); this deactivated upp in mutant HN38 (Fig. 1B).
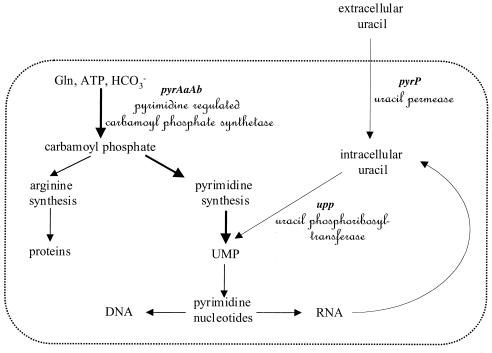
FIG. 2.
Simplified diagram of the metabolic pathways involved in the selection of the spontaneous mutant HN38. The intracellular UMP pool depends on pyrimidine synthesis and utilization of preformed pyrimidine supplied from RNA degradation or pumped from the culture medium. The upp gene catalyzed the key reaction in preformed uracil utilization (19). The UMP pool controls expression of the pyr genes involved in the de novo pyrimidine biosynthesis (shown with thick arrows). PyrR, an RNA binding protein, senses the concentration of UMP in the cell and regulates pyr gene expression through an attenuator mechanism. This regulation mechanism was demonstrated for Bacillus subtilis (24) and has been proposed for L. plantarum (11). Carbamoyl phosphate (CP) is a common intermediate for both arginine and pyrimidine biosynthesis. In L. plantarum, two CP synthetases (CPS) are present: a pyrimidine-regulated CPS encoded by the pyrAaAb genes (12) and an arginine-repressed CPS encoded by the carAB genes (20). In strain FB335, the carAB genes were deleted (20); growth relied on pyrAaAb expression to provide CP for both pathways. When uracil is provided, the pyr genes are not transcribed and FB335 cannot grow since it lacks CP for arginine biosynthesis. However, spontaneous uracil-resistant derivatives of FB335 were selected and one of these, mutant HN38, harbored the upp::ISLpl1 allele. HN38 grew in the presence of uracil, since the inactivation of upp may have generated a low UMP pool, which favored pyrAaAb expression.
Sequence analysis of ISLpl1.
The insert was bordered by 24-bp-long inverted repeats (IR). The IR at the left (IRL) was 5′-GGTAGATTGtAAAaTTAATCCGAA-3′. As indicated with lowercase characters, the IR at the right (IRR) had two mismatches compared to IRL (t and a replaced by c and t, respectively). A unique open reading frame (ORF) spanned nearly the entire IS. Its stop codon (TAA) was located within IRR. A ribosome binding site (AGGAGcTGATC with a calculated free energy ΔG of −15.8 kcal) preceded a potential TTG initiation codon; this would generate a 309-amino-acid (aa) protein. The protein identity between the 309-aa protein and the transposase of Escherichia coli IS30 (GenBank accession no. U70214) (383 aa) was 30%. The characteristics of IS30 elements have been reviewed previously (17), and the conserved DD(33)E consensus motif defined for functional IS30 transposases was found in the 309-aa-long protein (data not shown). Thus, the 1,043-bp insert belonged to the IS30 IS family. This insertion element from L. plantarum was named ISLpl1, and its encoded putative transposase was named TraISLpl1.
Characteristics of ISLpl1-related elements.
We searched in the nucleic acid databases for sequences similar to that of ISLpl1. Five insertion elements with high percentages (98 to 83%) of nucleic acid identity were found (Table 3). These ISLpl1-related elements had similar sizes of between 1,041 and 1,051 bp. These IS elements were bordered by well-conserved 24- to 28-bp IR. After comparing these IR, a 25-bp consensus sequence with 22 conserved nucleotides (shown in uppercase characters) and 3 nt found in at least half of the IR (shown in lowercase characters; the two conserved mismatches between the IRL and IRR are underlined) was proposed as follows: 5′-tGGTAGATTGTAAAATTAATCCgAa-3′ (Table 3). We searched for target duplication at the IS junctions (Table 3); three cases of 4-bp direct repeats and two cases without target sequence duplication were found. In the cases of the two iso-IS30 elements, the A nucleotide repetition did not allow us to discriminate between the absence of duplication and the occurrence of a 1-nt duplication. To be self-transposable, IS code for a functional transposase. This was not the case in three out of the five ISLpl1-related elements due to point mutation or 1-nt deletion (9) (Table 3). IS30 family transposases are between 293 and 383 aa long (17), and TraISLpl1 (the transposase of ISLpl1) was predicted to be 309 aa long. On the other hand, L. plantarum ISLP1, P. pentosaceus iso-IS30, and L. plantarum IS125 code for putative truncated transposases of 233, 276, and 170 aa, respectively (Table 3); this suggests that these elements are cryptic.
TABLE 3.
Characteristics of ISLp11-related elements
| IS characteristics
|
Nucleic acid characteristicsa
|
Transposase size (aa) and/or comment | Source, reference, and/or database accession no. | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Name | Genetic linkage | Bacterial host | % Identity with ISLp11 | Size (bp) | Adjacent sequence | IR sequence (5′→3′) | IR size | Proposed target duplication (bp) | |||
| ISLp11 | ISLp12 on plasmid pLp3 | Lactobacillus plantarum HN38 | 100 | 1,043 | gtgt | GGTAGATTGTAAAATTAATCCGAA (IRL) | 24 | None | 309; functional | This work; EMBL AF459445 | |
| attc | GGTAGATTGCAAATTTAATCCGAA (IRR) | ||||||||||
| Chromosomal upp::ISLp11 | attc | 4 | This work | ||||||||
| gaat | |||||||||||
| iso-IS30 | tet(M) gene on plasmid pMD5057 | Lactobacillus plantarum 5057 | 99 | 1,051 | ttta | ATAT GGTAGATTGTAAAATTAATCCGAA (IRL) | 28 | ? | 309 | 8; GenBank AF440277 | |
| atct | ATAT GGTAGATTGCAAATTTAATCCAAA (IRR) | ||||||||||
| ISLP1 | Cold shock protein on plasmid pLF1 | Lactobacillus plantarum ATCC 14917 | 99 | 1,049 | ttga | AAT GGTAGATTGTAAAATTAATCCGAA (IRL) | 27 | None | 233; truncation due to TGA stop codon (positions 3139-3141) | Unpublished data; EMBL AF508808 | |
| aatg | AAT GGTAGATTGCAAATTTAATCCGAA (IRR) | ||||||||||
| Unnamed | Unknown | Oenococcus oeni MCW | 98 | 1,043 | tttt | GGTAGATTGTAAAATTAATCCGAA (IRL) | 24 | 4 | 309 | MCBI microbial genome annotation project | |
| aaaa | GGTAGATTGCAAATTTAATCCGAA (IRR) | ||||||||||
| iso-IS30 | Sucrose and raffinose operons | Pediococcus pentosaceus PPE1.0 | 98 | 1,041 | taca | TGGTAGATTGTAAAATTAATCCGA (IRL) | 24 | ? | 276; truncation due to 1-nt deletion in position 12233 | 18; EMBL Z32771 | |
| taat | TGGTAGATTGCAAATTTAATCCGA (IRR) | ||||||||||
| IS125 | Plantaricin 1.25β-encoding cluster | Lactobacillus plantarum TMW1.25 | 83 | 1,041 | attc | GGTAGATTGTAAAATTAATCCGAA (IRL) | 24 | 4 | 170; truncation due to 1-nt deletion in position 1841 | 9; EMBL AJ011094 | |
| gaat | GGTAGATTGCAAATTTAATCCGAA (IRR) | ||||||||||
| t GGTAGATTGTAAAATTAATCCGAa (IRL consensus) | |||||||||||
| t GGTAGATTGCAAATTTAATCCgAa (IRR consensus) | |||||||||||
The length of the IR was determined by analysis of the nucleotide identity between the ends of a given IS (nucleotides in upper case); nucleotides at the junction with the IS are indicated in lowercase italic characters; IR conserved nucleotides are indicated in bold; nucleotides which systematically differed between IRL and IRR are indicated by underlining in the consensus sequences; and the nucleotides found in at least half of the IS are indicated in lowercase bold characters.
Copy numbers and distribution of ISLpl1 within other L. plantarum and Pediococcus strains.
The presence of ISLpl1-related elements in other LAB was assessed using Southern hybridization with a specific ISLpl1 probe. To estimate the number of copies, DNA digests were tested with restriction enzymes HindIII and HaeIII, which do not cut within ISLpl1. The relatedness between ISLpl1 and other insertion elements detected in five L. plantarum strains and one Pediococcus strain was estimated using Southern hybridization under two sets of stringency conditions (see Materials and Methods). The results of low-stringency hybridization are shown on Fig. 3; from 0 to 18 bands were detected. The detection of a single band in strain FB335 with the two restriction enzymes (Fig. 3, lanes 6 and 10) strongly suggested that this strain had only one ISLpl1 locus. In its derivative, strain HN38, two copies of the ISLpl1 were found, since an additional band (11.5 kb; Fig. 3, lane 9) corresponding to the upp::ISLpl1 insertion was detected with HindIII. ISLpl1-related insertion elements were found in P. acidilactici (three HindIII bands detected; Fig. 3, lane 12). The absence of ISLpl1-related elements in NCIMB 8826 was confirmed by PCR amplification (no amplification with the ISLpl1-specific primer Lp-IS30 occurred, unlike the results of upp control amplification with primers glyc-g1-Lpupp-h1; data not shown). Each tested strain had a specific pattern, since their band sizes and band numbers were different (Table 1).
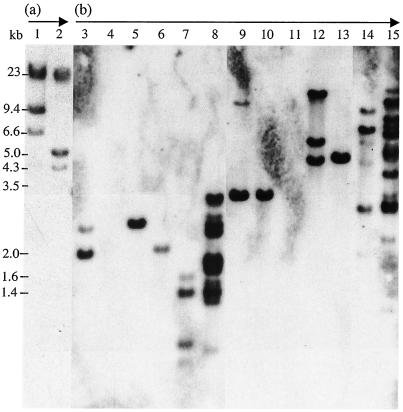
FIG. 3.
ISLpl1-related element distribution in L. plantarum and Pediococcus strains. Southern hybridization blots with probe ISLpl1 (arrow b) or with probe Lambda DNA (arrow a). Lane 1, Lambda DNA digested with HindIII; lane 2, Lambda digested with HindIII and EcoRI. LAB genomes were digested with restriction enzymes HaeIII (lanes 3 to 8) and HindIII (lanes 9 to 15). Lanes 3 and 9, L. plantarum strain HN38; lanes 4 and 11, L. plantarum NCIMB 8826; lanes 5 and 13, L. plantarum ATCC 14917T; lanes 6 and 10, L. plantarum FB335; lanes 7 and 14, L. plantarum CNRZ 1891; lanes 8 and 15, L. plantarum NCFB 1406; lane 12, Pediococcus strain H.
ISLpl1 is plasmidborne in FB335 and other L. plantarum strains.
Few L. plantarum strains are plasmid free, and two ISLpl1-related elements were found on L. plantarum plasmids (Table 3). ISLpl1 might be common in plasmids, so we identified the locations of ISLpl1 in two strains, one harboring a single copy and the other harboring 18 copies (strains FB335 and NCIMB 1406, respectively; Table 1). As a native plasmid DNA marker, we used strain NCIMB 8826, which had three sequenced plasmids (36, 2.3, and 1.9 kb) (16) and no detected ISLpl1-related elements (Table 1). Different plasmid DNA profiles were found in these strains, suggesting different plasmid contents (Fig. 4A, lanes 1, 3, and 5).
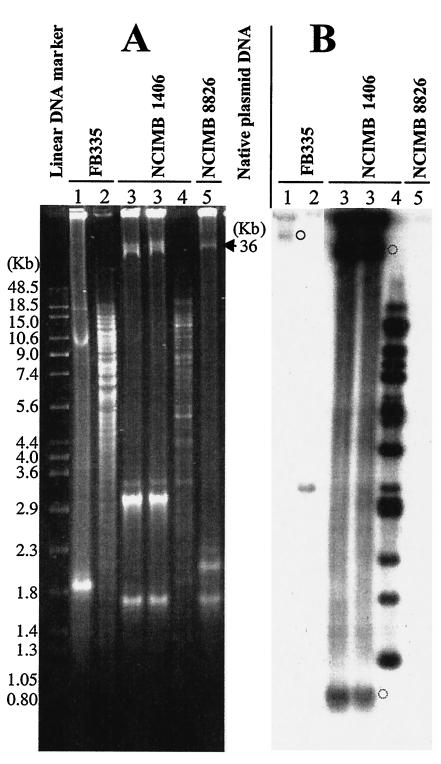
FIG. 4.
Plasmid and chromosomal localization of ISLpl1. (A) Ethidium bromide-stained DNA after agarose gel electrophoresis; (B) Southern blot DNA bands hybridized with the digoxigenin-labeled specific ISLpl1 probe. Lanes 1, 3, and 5, native plasmid DNA extracted from L. plantarum strains FB335, NCIMB 1406, and NCIMB 8826, respectively; lanes 2 and 4, HindIII-digested CsCl gradient linear DNA preparations of strain FB335 and NCIMB 1406, respectively. Linear DNA molecular mass markers (Raoul marker; Appligene) are indicated in kilobases. Native plasmid band sizes were deduced from the three plasmids harbored by strain NCIMB 8826 (36, 2.3, and 1.9 kb). Native plasmid bands detected by the IS probe are shown by circles representing strains FB335 (circle in panel B, lane 1) and NCIMB 1406 (circles in panel B, lane 3).
The presence of ISLpl1-related elements on the extracted plasmids was assessed by Southern hybridization with an ISLpl1 probe. As expected, no hybridization was detected in NCIMB 8826 (Fig. 4B, lane 5). In NCIMB 1406, two plasmid DNA bands hybridized (Fig. 4B, lanes 3); these may correspond to two supercoiling forms of the same plasmids. Only the larger band was visible in the ethidium bromide-stained agarose gel (Fig. 4A, lanes 3). With purified plasmid DNA from strain FB335, the IS probe hybridized to a plasmid called pLp3 (Fig. 4B, lane 1), whose position in the gel suggested that it may be close in size to a 36-kb plasmid in NCIMB 8826. Unlike the sequenced 2.1-kb cryptic plasmid pLp1 (5) and the 10-kb plasmid pLp2, the larger plasmid pLp3 was not efficiently extracted with the QIAGEN plasmid purification protocol; the detected band was barely visible (FB335; Fig. 4A, lane 1). (Native linear chromosomal DNA migrated around 48 kb, which is lower than the distance at which the native pLp3 plasmid migrated [data not shown]). The presence of a weak band in the lane representing CsCl-purified DNA enriched with the chromosome DNA (Fig. 4B, lane 2) suggests that a nicked circular form of pLp3 was contaminating the chromosomal-enriched preparation (Fig. 4B, lane 2). The presence of ISLpl1 on a plasmid explained why the intensity of the chromosomal upp::ISLpl1 band (one copy) was lower than that of ISLpl1 located on plasmid pLp3 (multiple copies) in strain HN38 (DNA preparation with no plasmid or chromosomal separation; Fig. 3, lanes 3 and 9). We concluded that ISLpl1-related elements were present on plasmids in strains FB335 and NCIMB 1406. IS were also present on the chromosome of strain NCIMB 1406 (Fig. 4B, lane 4).
Sequence analysis of ISLpl2, an IS150-related element contiguous to ISLpl1 on plasmid pLp3
. The ISLpl1 locus in plasmid pLp3 was analyzed using inverse PCR to determine the sequences adjacent to the IS (see Materials and Methods) (Fig. 1A). The resulting 3,027-bp sequence was deposited in the GenBank database (accession no. AF459445). The IS present on plasmid pLp3 (positions 1219 to 2261) was identical to the ISLpl1 chromosomal copy. No target duplication was detected at the ISLpl1 junctions (Table 3). The 963-bp sequence downstream of ISLpl1 had no similarities with sequences found in the GenBank, EMBL, DDBJ, and PDB nucleotide databases. Upstream of ISLpl1, two consecutive reading frames, OrfA (C-terminal partial sequence; 88 aa) and OrfB (295 aa [when translation initiated at a GTG codon]), were present. An OrfB stop codon (TAA) was only 66 nt from ISLpl1. OrfA and OrfB shared 22 and 37% identity with E. coli IS150 OrfA and OrfB, respectively (GenBank accession no. X07037). The OrfA translation stop codon (in lowercase characters) overlapped with the putative OrfB initiation codon (Gtg) in the following sequence: 5′-AA AAA TCA GCT CAC GAG AAA CCG tga-3′ (nt positions 243 to 268; GenBank accession no. AF459445). An OrfAB fusion protein might be generated by programmed translational frameshifting, as described for IS150 elements (27). Such a frameshift site may include a stretch of 5 As preceding a potential stem-loop-forming sequence (with a ΔG of −5.8 kcal) (underlined). We called this insertion element ISLpl2 (i.e., the IS150-related insertion element contiguous to ISLpl1 on plasmid pLp3).
Genetic stability over time of the ISLpl1-related patterns in NCIMB 1406.
We chose to study the genetic stability of the IS element in the genetic context of L. plantarum NCIMB 1406 for two reasons. First, of all the tested LAB, the NCIMB 1406 strain harbors the most ISLpl1-related elements (Table 1). Second, NCIMB 1406 grows slowly in liquid medium. Thus, if IS elements are mobile, they might improve NCIMB 1406 growth over time. NCIMB 1406 was grown for 110 generations in MRS broth (described in Materials and Methods). After four rounds of serial growth, the culture had adapted to the imposed laboratory conditions; overnight incubation (rather than the 2-day-long incubation observed with the parent strain) was sufficient for the colonies to reach the stationary-growth phase. After 12 serial growth cycles, four clones isolated on MRS plates were compared to the parent strain. Restricted total DNA was separated by electrophoresis in a 0.8% agarose gel (Fig. 5A) and hybridized with a specific ISLpl1 probe. In clone 1, two additional bands around 4.5 kb were detected. In clone 2, a band of 8 kb was lost and new bands of 17 and 7 kb were detected. In fact, in all tested clones the IS patterns differed from those of the parent strain, whose 18 bands were clearly present (Fig. 5B). Since different ISLpl1-related patterns were found after 110 generations of NCIMB 1406 were grown in MRS broth, these ISLpl1 patterns were not stable over time.
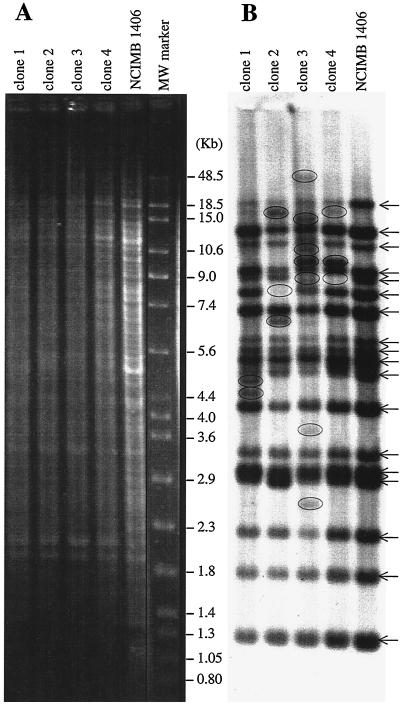
FIG. 5.
Stability over time of the ISLpl1-related element patterns in L. plantarum strain NCIMB 1406. Insertion sites were evaluated with ISLpl1 DNA probed against HaeIII digests of NCIMB 1406-derived strains, using Southern hybridization under low-stringency conditions. The IS profile was analyzed in four clones isolated after 110 generations of NCIMB 1406 grown in liquid MRS medium. Arrows indicate the 18 bands detected in strain NCIMB 1406 at the beginning of the experiment. Clones 1 to 4 were isolated from a liquid MRS culture obtained by serial cultivation of strain NCIMB 1406 as described in Materials and Methods. Circles show detected bands, which differ from those of the parent strain. (A) DNA after agarose gel electrophoresis. (B) Southern blot. MW marker, molecular mass marker (Raoul marker; Appligene) bands (indicated in kilobases).
DISCUSSION
In the spontaneous mutant HN38 derived from L. plantarum ATCC 8014 (Table 1), the ISLpl1 1,043-bp element inactivated the chromosomal upp gene. Among the 37 mutants characterized, HN38 was the only one mutated by the insertion of a mobile element (Nicoloff and Bringel, unpublished). HN38 harbored two identical ISLpl1 copies, one in the chromosome (upp::ISLpl1) and one on plasmid pLp3, as found in the parent strain. Short 4-bp direct repeats (CTTA) of the target DNA flanking the chromosomal insertion were observed. Although most IS30-related insertion elements generate 2- to 3-bp direct repeats (17), IS30-related insertion elements (IS658) found in the Bacillus halodurans genome (25), in L. plantarum (IS125) (9), and in Oenococcus oeni (Table 3) also have 4-bp-long target site sequence duplications. However, no direct-repeat target duplications were observed with other ISLpl1-related elements such as iso-IS30 (8) and ISLp11 on plasmid pLp3 (Table 3).
Evidence of ISLpl1 activity in L. plantarum FB335 includes (i) the isolation of the upp::ISLpl1-mutated allele; (ii) the presence of the direct repeats of the target DNA flanking the IS; (iii) the presence of a single ORF (called TraISLpl1) with the characteristics of a functional IS30 transposase [i.e., the correct size range and the presence of the essential DD(E) amino acids found in these integrases] (17); and (iv) the finding that ISLpl1 was bordered by 24-bp-long IR with only two mismatches between the IRL and IRR. The functionality of ISLpl1 in other L. plantarum strains is suggested by the presence of up to 18 copies of iso-ISLpl1 elements in different strains as well as by the variations in the ISLpl1 patterns seen after 110 generations of strain NCIMB 1406 grown in MRS medium. Even if only 1 of the 18 copies of iso-ISLpl1 detected in NCIMB 1406 was functional, such changes in the IS bands would be expected. Another possibility is that no transposition occurred and that ISLpl1-related elements might have been passively used as templates in bacterial genome rearrangements. The instability of the genetic IS patterns observed in NCIMB 1406 may be the result of both mechanisms. Thus, ISLpl1 is a mobile IS in L. plantarum FB335 and probably in other L. plantarum strains as well. Since no mutation insertion tools are available for L. plantarum, ISLpl1 may be a good candidate for the development of such tools for use with LAB and especially with L. plantarum NCIMB 8826, whose genome has been sequenced (16) and lacks this element.
Among the six ISLpl1-related elements studied, three may be cryptic elements due to putative truncated transposases (9) (Table 3) resulting from point mutation or 1-nt deletions within their transposase-encoding genes. If cryptic ISLpl1-related elements are common in LAB (and since ISLpl1-related elements were found in different LAB genera) (Table 1 and Table 3), ISLpl1 fingerprinting might be a useful tool for LAB intraspecies identification, as described for other IS elements in a report of a study of Lactobacillus (10). IS may participate in genome plasticity as duplicated sequences. Evidence of DNA rearrangements involving ISLpl1 was found in P. pentosaceus (GenBank accession no. Z32771); two partially overlapping sequences were 100% identical to that of the ISLpl1 IRL (Fig. 6). A possible scenario is the occurrence of a deletion event between 6-bp directly repeated sequences present in two ISLpl1 elements with opposite orientations, as shown in Fig. 6. ISLpl1 seems to be implicated in LAB DNA plasticity.
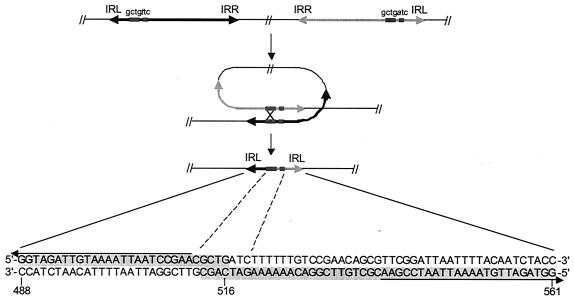
FIG. 6.
Proposed role of ISLp1l in LAB genome plasticity. In a P. pentosaceus isolate, ISLpl1 homologous sequences were present 9 kb upstream of an iso-IS30 (GenBank accession no. Z32771) region. Short sequences shaded in gray (shown at the bottom of the figure) are identical to those of an ISLpl1 left arm. The arrows delimit the IRL defined for ISLpl1. The proposed genetic rearrangement at the origin of this structure is a recombination between two short directly repeated sequences (indicated by small boxes at the top and in the middle of the figure) of two divergently oriented IS. The sequence of the repeats is GCTGt/aTC (shown at the top of the figure), with only one mismatch (lowercase characters) between the two IS elements.
In some cases, ISLpl1 was linked to genetic determinants, such as a bacteriocin-encoding cluster (9), a sugar utilization cluster (18), a tetracycline resistance gene (8), and a cold shock protein gene (GenBank accession no. AF508808) (unpublished data), which may be beneficial for bacterial survival in industrial fermentations and in natural niches. ISLpl1 may be transferred between different LAB genera, as suggested by the following findings: (i) high (98%) degrees of iso-ISLpl1 nucleic acid sequence identity in different LAB genera (these quasi-identical sequences contrast with the lower sequence identities seen between 16S rRNA genes [85 and 91% identity between L. plantarum and O. oeni and between L. plantarum and P. pentosaceus, respectively]) and (ii) ISLpl1-related element genetic linkage to the sucrose-raffinose utilization locus in L. plantarum strain ATCC 14917T and P. pentosaceus (18). Plasmid location may facilitate gene exchange between LAB, and ISLpl1-related elements were associated to different plasmids in L. plantarum isolates (8) (Table 3 and Fig. 4B). So far, no natural transformations have been described in LAB, and only parts of the genes required for natural competence were found in the genome of L. plantarum (16). On the other hand, horizontal gene transfer of different IS in LAB has been documented (see the introduction). Genomic investigations of 15 LAB species (15) will provide an excellent opportunity to investigate genetic exchange between these microorganisms and to further investigate the role of ISLpl1-related elements in LAB genome plasticity.
Acknowledgments
We thank P. Hammann for DNA sequencing, V. Mathé for technical assistance, and B. Johnson, J.-C. Hubert, and M.-C. Lett for comments about the paper.
REFERENCES
- 1.Altschul, S. F., T. L. Madden, A. A. Schaffer, J. Zhang, Z. Zhang, W. Miller, and D. J. Lipman. 1997. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25:3389-3402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Blot, M. 1994. Transposable elements and adaptation of host bacteria. Genetica 93:5-12. [DOI] [PubMed] [Google Scholar]
- 3.Bourgoin, F., A. Pluvinet, B. Gintz, B. Decaris, and G. Guedon. 1999. Are horizontal transfers involved in the evolution of the Streptococcus thermophilus exopolysaccharide synthesis loci? Gene 233:151-161. [DOI] [PubMed] [Google Scholar]
- 4.Bringel, F., L. Frey, S. Boivin, and J. C. Hubert. 1997. Arginine biosynthesis and regulation in Lactobacillus plantarum: the carA gene and the argCJBDF cluster are divergently transcribed. J. Bacteriol. 179:2697-2706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Bringel, F., L. Frey, and J. C. Hubert. 1989. Characterization, cloning, curing, and distribution in lactic acid bacteria of pLP1, a plasmid from Lactobacillus plantarum CCM 1904 and its use in shuttle vector construction. Plasmid 22:193-202. [DOI] [PubMed] [Google Scholar]
- 6.Bringel, F., P. Quénée, and P. Tailliez. 2001. Polyphasic investigation of the diversity within Lactobacillus plantarum related strains revealed two L. plantarum subgroups. Syst. Appl. Microbiol. 24:561-571. [DOI] [PubMed] [Google Scholar]
- 7.Burrus, V., G. Pavlovic, B. Decaris, and G. Guedon. 2002. Conjugative transposons: the tip of the iceberg. Mol. Microbiol. 46:601-610. [DOI] [PubMed] [Google Scholar]
- 8.Danielsen, M. 2002. Characterization of the tetracycline resistance plasmid pMD5057 from Lactobacillus plantarum 5057 reveals a composite structure. Plasmid 48:98-103. [DOI] [PubMed] [Google Scholar]
- 9.Ehrmann, M. A., A. Remiger, V. G. Eijsink, and R. F. Vogel. 2000. A gene cluster encoding plantaricin 1.25β and other bacteriocin-like peptides in Lactobacillus plantarum TMW1.25. Biochim. Biophys. Acta 1490:355-361. [DOI] [PubMed] [Google Scholar]
- 10.Ehrmann, M. A., and R. F. Vogel. 2001. Characterisation of IS153, an IS3-family insertion sequence isolated from Lactobacillus sanfranciscensis and its use for strain differentiation. Syst. Appl. Microbiol. 24:443-450. [DOI] [PubMed] [Google Scholar]
- 11.Elagöz, A. 1997. Structure et régulation de l'expression des gènes de la voie de biosynthèse des pyrimidines de Lactobacillus plantarum CCM 1904. Ph.D. thesis. Université Louis-Pasteur, Strasbourg, France.
- 12.Elagöz, A., A. Abdi, J. C. Hubert, and B. Kammerer. 1996. Structure and organisation of the pyrimidine biosynthesis pathway genes in Lactobacillus plantarum: a PCR strategy for sequencing without cloning. Gene 182:37-43. [DOI] [PubMed] [Google Scholar]
- 12a.Ennahar, S., D. Aoude-Werner, O. Sorokine, A. Van Dorsselaer, F. Bringel, J.-C. Hubert, and C. Hasselmann. 1996. Production of pediocin AcH by Lactobacillus plantarum WHE 92 isolated from cheese. Appl. Environ. Microbiol. 62:4381-4387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Germond, J. E., L. Lapierre, M. Delley, and B. Mollet. 1995. A new mobile genetic element in Lactobacillus delbrueckii subsp. bulgaricus. Mol. Gen. Genet. 248:407-416. [DOI] [PubMed] [Google Scholar]
- 14.Guedon, G., F. Bourgoin, M. Pebay, Y. Roussel, C. Colmin, J. M. Simonet, and B. Decaris. 1995. Characterization and distribution of two insertion sequences, IS1191 and iso-IS981, in Streptococcus thermophilus: does intergeneric transfer of insertion sequences occur in lactic acid bacteria co-cultures? Mol. Microbiol. 16:69-78. [DOI] [PubMed] [Google Scholar]
- 15.Klaenhammer, T., E. Altermann, F. Arigoni, A. Bolotin, F. Breidt, J. Broadbent, R. Cano, S. Chaillou, J. Deutscher, M. Gasson, M. van de Guchte, J. Guzzo, A. Hartke, T. Hawkins, P. Hols, R. Hutkins, M. Kleerebezem, J. Kok, O. Kuipers, M. Lubbers, E. Maguin, L. McKay, D. Mills, A. Nauta, R. Overbeek, H. Pel, D. Pridmore, M. Saier, D. van Sinderen, A. Sorokin, J. Steele, D. O'Sullivan, W. de Vos, B. Weimer, M. Zagorec, and R. Siezen. 2002. Discovering lactic acid bacteria by genomics. Antonie Leeuwenhoek 82:29-58. [DOI] [PubMed] [Google Scholar]
- 16.Kleerebezem, M., J. Boekhorst, R. Van Kranenburg, D. Molenaar, O. P. Kuipers, R. Leer, R. Tarchini, S. A. Peters, H. M. Sandbrink, M. W. Fiers, W. Stiekema, R. M. Lankhorst, P. A. Bron, S. M. Hoffer, M. N. Groot, R. Kerkhoven, M. De Vries, B. Ursing, W. M. De Vos, and R. J. Siezen. 2003. Complete genome sequence of Lactobacillus plantarum WCFS1. Proc. Natl. Acad. Sci. USA 100:1990-1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Mahillon, J., and M. Chandler. 1998. Insertion sequences. Microbiol. Mol. Biol. Rev. 62:725-774. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Naumoff, D. G., and V. A. Livshits. 2001. Molecular structure of the locus for sucrose utilization by Lactobacillus plantarum: comparison with Pediococcus pentosaceus. Mol. Biol. (Engl. Trans. Mol. Biol. [Moscow]) 35:19-27. [PubMed] [Google Scholar]
- 19.Nicoloff, H. 2001. Ph.D. thesis. Université Louis-Pasteur, Strasbourg, France.
- 20.Nicoloff, H., J. C. Hubert, and F. Bringel. 2000. In Lactobacillus plantarum, carbamoyl phosphate is synthesized by two carbamoyl-phosphate synthetases (CPS): carbon dioxide differentiates the arginine-repressed from the pyrimidine-regulated CPS. J. Bacteriol. 182:3416-3422. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Sambrook, J., E. F. Fritsch, and T. Maniatis (ed.). 1989. Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y.
- 22.Skaugen, M., and I. F. Nes. 1994. Transposition in Lactobacillus sake and its abolition of lactocin S production by insertion of IS1163, a new member of the IS3 family. Appl. Environ. Microbiol. 60:2818-2825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Skaugen, M., and I. F. Nes. 2000. Transposition in Lactobacillus sakei: inactivation of a second lactocin S operon by the insertion of IS1520, a new member of the IS3 family of insertion sequences. Microbiology 146:1163-1169. [DOI] [PubMed] [Google Scholar]
- 24.Switzer, R. L., R. J. Turner, and Y. Lu. 1999. Regulation of the Bacillus subtilis pyrimidine biosynthetic operon by transcriptional attenuation: control of gene expression by an mRNA-binding protein. Prog. Nucleic Acid Res. Mol. Biol. 62:329-367. [DOI] [PubMed] [Google Scholar]
- 25.Takami, H., C. G. Han, Y. Takaki, and E. Ohtsubo. 2001. Identification and distribution of new insertion sequences in the genome of alkaliphilic Bacillus halodurans C-125. J. Bacteriol. 183:4345-4356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Vanderslice, P., W. C. Copeland, and J. D. Robertus. 1986. Cloning and nucleotide sequence of wild type and a mutant histidine decarboxylase from Lactobacillus 30a. J. Biol. Chem. 261:15186-15191. [PubMed] [Google Scholar]
- 27.Vögele, K., E. Schwartz, C. Welz, E. Schiltz, and B. Rak. 1991. High-level ribosomal frameshifting directs the synthesis of IS150 gene products. Nucleic Acids Res. 19:4377-4385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Walker, D. C., and T. R. Klaenhammer. 1994. Isolation of a novel IS3 group insertion element and construction of an integration vector for Lactobacillus spp. J. Bacteriol. 176:5330-5340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Yanisch-Perron, C., J. Vieira, and J. Messing. 1985. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene 33:103-119. [DOI] [PubMed] [Google Scholar]
- 30.Zwahlen, M. C., and B. Mollet. 1994. ISL2, a new mobile genetic element in Lactobacillus helveticus. Mol. Gen. Genet. 245:334-338. [DOI] [PubMed] [Google Scholar]