Abstract
The metagenomes of complex microbial communities are rich sources of novel biocatalysts. We exploited the metagenome of a mixed microbial population for isolation of more than 15 different genes encoding novel biocatalysts by using a combined cultivation and direct cloning strategy. A 16S rRNA sequence analysis revealed the presence of hitherto uncultured microbes closely related to the genera Pseudomonas, Agrobacterium, Xanthomonas, Microbulbifer, and Janthinobacterium. Total genomic DNA from this bacterial community was used to construct cosmid DNA libraries, which were functionally searched for novel enzymes of biotechnological value. Our searches in combination with cosmid sequencing resulted in identification of four clones encoding 12 putative agarase genes, most of which were organized in clusters consisting of two or three genes. Interestingly, nine of these agarase genes probably originated from gene duplications. Furthermore, we identified by DNA sequencing several other biocatalyst-encoding genes, including genes encoding a putative stereoselective amidase (amiA), two cellulases (gnuB and uvs080), an α-amylase (amyA), a 1,4-α-glucan branching enzyme (amyB), and two pectate lyases (pelA and uvs119). Also, a conserved cluster of two lipase genes was identified, which was linked to genes encoding a type I secretion system. The novel gene aguB was overexpressed in Escherichia coli, and the enzyme activities were determined. Finally, we describe more than 162 kb of DNA sequence that provides a strong platform for further characterization of this microbial consortium.
Modern biotechnology has a steadily increasing demand for novel biocatalysts, which has prompted the development of novel experimental approaches to find and identify novel biocatalyst-encoding genes. Only recently were studies initiated to investigate the metagenome (28), which represents the genomes of uncultured microbes, as a rich source for isolation of many novel genes. One gram of soil may contain up to 4,000 different species (30); however, current estimates indicate that less than 1% of these organisms are readily culturable with known cultivation techniques (2). This problem can be solved by direct isolation and cloning of metagenomic DNA, thereby circumventing cultivation, which may result in the loss of major portions of the microbial communities because of the different growth requirements of the different microbes (4).
Several different laboratories have successfully isolated novel genes encoding different enzymes and secondary metabolites from microbial communities and their metagenomes without cultivation of the microbes (3, 9, 10, 24, 28). The microbial niches studied were highly diverse and ranged from moderate environments, such as river soil (9), to rather extreme environments, like the deep sea (3). However, a major difficulty associated with the methods employed is related to contamination of purified DNA with polyphenolic compounds which are copurified with the DNA. These compounds are difficult to remove, and it is well known that polyphenols also interfere with enzymatic modifications of isolated DNA (31). As a result, construction of environmentally derived DNA libraries with large inserts is hindered due to the poor quality of the isolated DNA. These known difficulties associated with the construction of libraries directly derived from environmental DNA samples forced us to isolate DNA from the metagenome of a microbial community after precultivation in the laboratory. Although it is expected that laboratory enrichment cultures have only limited biodiversity, this technique has proven to be highly efficient for rapid isolation of large DNA fragments and for cloning of operons and genes with great biotechnological value (7). Additionally, laboratory enrichments allow preselection of microbes that already carry the desired traits, resulting in a high frequency of gene detection and isolation (7, 8, 21).
In this study, we exploited the metagenome of an enrichment culture on agar plates for isolation of genes encoding a variety of different biocatalysts, including β-agarases, amylolytic enzymes, cellulases, and lipases. A detailed DNA analysis of selected cosmid clones resulted in identification of the sequences of four conserved clusters encoding β-agarases, a cluster of two lipase genes, and many other enzymes with high biotechnological potential. Furthermore, we report here that the majority of the microbes found in our mixed consortium have not been characterized previously and therefore represent highly intriguing possible sources for isolation of novel genes for biotechnology.
MATERIALS AND METHODS
Bacterial strains, plasmids, growth conditions, and environmental samples.
Bacterial strains and plasmids used in the present work are listed in Table 1. Escherichia coli was grown at 37°C on Luria-Bertani (LB) medium (29) supplemented with appropriate antibiotics. To obtain agarolytic microbial consortia, cultures were inoculated with a few grains of soil on agar plates containing defined medium and incubated at room temperature until agarolytic activity was observed. The soil samples used for inoculation of the mixed cultures were collected from an unplanted field near the Department of Microbiology of the University of Göttingen. The defined medium employed for growth of the agarolytic consortia contained (per liter) 1.0 g of NaNO3 · 3H2O, 1.0 g of K3HPO4 · 3H2O, 0.25 g of MgSO4 · 7H2O, 0.015 g of CaCl2 · 7H2O, 0.14 g of FeSO4 · 7H2O, 0.02 g of EDTA, and 10 ml of a trace element solution. The trace element solution contained (per liter) 15 mg of Co(NO3)2 · 6H2O, 60 mg of H3BO4, 200 mg of MnCl2 · 4H2O, 60 mg of ZnSO4 · 7H2O, 50 mg of NiCl2 · H2O, 2.5 mg of CuSO4 · 5H2O, 2.5 mg of Na2MoO4, and 4.0 mg of CaCl2 · 6H2O. The medium contained 1.5% agar and 0.2% glucose as combined carbon sources.
TABLE 1.
Microbiological materials
| Material | Relevant trait(s) | Source or reference |
|---|---|---|
| E. coli strains | ||
| VCS257 | DP50 derivative | Stratagene |
| DH5α | Cloning strain, recA1 endA1 gyrA96 thi1 relA1 hsdR17 supE44 lac | Gibco-BRL |
| BL21(DE3) | Expression host for pET vectors | Novagene |
| Plasmids and cosmids | ||
| pBSK+ | pBluescriptSK+ multicopy cloning vector, Ampr | Stratagene |
| pTZ19R | High-copy-number cloning vector, Cmr | Amersham, Pharmacia |
| pWE15 | Cosmid cloning vector for construction of DNA libraries, Ampr | Stratagene |
| pIIIE5 | pWE15 containing a 20.7-kb insert with possible agarase genes from microbial consortium | This study |
| pXE1 | pWE15 containing a 25-kb insert with possible lipase genes from microbial consortium | This study |
| pVIIC10 | pWE15 containing a 34.7-kb insert with possible agarase genes from microbial consortium | This study |
| pXB11 | pWE15 containing a ca. 20-kb insert with possible agarase genes from microbial consortium | This study |
| pXIVA3 | pWE15 containing a 38.1-kb insert with possible agarase genes from microbial consortium | This study |
| pIIIF2 | pWE15 containing a 28-kb insert with potential pectate lyase genes from microbial consortium | This study |
| pAGUB | pET19B (Novagene) carrying the aguB gene in the NdeI site | This study |
Molecular techniques.
The method used for DNA isolation from the agarolytic consortium was based on a method described previously (7), with minor modifications. After bacteria were pelleted, they were resuspended in Tris-EDTA (TE) buffer containing 20% (wt/vol) sucrose and lysed in DNA extraction buffer (100 mM Tris-HCl, 100 mM EDTA, 100 mM Na2HPO4, 1.5 M NaCl, 1% [wt/vol] sodium dodecyl sulfate [SDS]) for several hours. RNA was degraded with RNase A (10 mg/ml). The resulting DNA extracts were incubated with protease and sarcosyl (5%, wt/vol) in TE buffer overnight. Total genomic DNA was then repeatedly extracted with chloroform-phenol (1:1, vol/vol), washed once with chloroform, and dialyzed against 2 liters of TE buffer at 4°C overnight. Finally, an aliquot of the DNA was analyzed on a 0.8% agarose gel to ensure that the DNA was not degraded.
Cosmid libraries were prepared in pWE15 (Stratagene, La Jolla, Calif.) by using standard protocols (7). DNA fragments (20 to 40 kb) obtained after partial Sau3A digestion were ligated into BamHI restriction sites of the cosmid vector. Phage packaging mixtures were obtained from Stratagene, and infection of E. coli VCS257 was performed by using the manufacturer's protocol. The vectors used for subcloning are listed in Table 1; when required, gaps in the DNA sequences were filled by PCR. Automated DNA sequencing was performed by using an ABI 377 sequencer and dye terminator chemistry and following the manufacturer's instructions.
Screening for genes conferring agarolytic and lipolytic activities.
Novel agarases were identified by first growing the cosmid clones in 3-ml microplates in LB medium overnight. Bacteria were pelleted, resuspended in 1 ml of lysis buffer (50 mM Tris-HCl, 5 mM MgCl2; pH 7.5), and lysed by addition of lysozyme (1 mg/ml). Crude cell extracts were centrifuged, and 120-μl portions of the extracts were added to 0.2% solutions of low-melting-point agarose (Biozyme, Bad Oldesloh, Germany) and incubated at 37°C overnight. Clones that produced agarolytic activity were identified in microplate wells containing liquefied agarose, as detected by inversion of the plates after cooling to room temperature. False-positive clones were excluded by reisolation and transformation of DNA from positive clones and detection of agarolytic activity prior to shotgun cloning. Clones conferring lipolytic activity were identified by the formation of clear halos after growth on agar plates containing tributyrin as the indicator substrate (22).
Cloning and expression of AguB as a histidine-tagged fusion protein.
The aguB gene was amplified from pVIIC10 DNA by using 35 cycles and primers AGUB1 (5′-GGCATATGAAAAAGCATATTTCA-3′) and AGUB2 (5′-GGATCCAAGTGATCCAACCCTGGTCAC-3′) designed to introduce a 3′ NdeI restriction site and a 5′ BamHI site. To increase the cloning efficiency, the PCR fragments were first ligated into pBSK+ and then excised with NdeI and ligated into pET19B. The plasmid carrying the aguB sequence was designated pAGUB. The nucleotide sequences of the PCR fragments cloned in pET19B were determined by DNA sequencing. Freshly transformed E. coli BL21(DE3) cells carrying pAGUB were used to overproduce the AguB protein. Cultures were grown to an optical density at 600 nm of 0.5 at 37°C, and production of the recombinant protein was induced by addition of 1 mM isopropyl-β-d-thiogalactopyranoside (IPTG). After 3 h, cells were harvested and disrupted in a French pressure cell, and AguB was purified from the soluble fraction. Extracts were loaded onto 1-ml Ni-nitrilotriacetic agarose (Ni-NTA) columns (Qiagen, Hilden, Germany), and chromatography was performed as recommended by Qiagen. The resulting protein fractions were analyzed by SDS-10% polyacrylamide gel electrophoresis and Western immunoblotting by using six-His-specific antibodies. The enzyme activity of β-agarase was quantified by using 3,5-dinitrosalicylic acid as the substrate to detect reducing sugars, and galactose was used as the standard (27).
Phylogenetic analysis.
The ribosomal gene sequences of the microbial consortium were amplified by PCR by using standard procedures. For this purpose universal bacterial 16S rRNA gene-specific reverse primer 16SRV (5′-GGTTACCTTGTTACGACTT-3′) and forward primer 16SFW (5′-CGTGCCAGAAGCATCGGTG-3′) were employed. The resulting PCR products were cloned by using a TOPO TA cloning kit (Invitrogen, Karlsruhe, Germany), and plasmid DNAs containing inserts were sequenced by using standard protocols with an ABI 377 DNA sequencer. Initially, 40 cloned 16S rRNA gene sequences were compared with reference sequences in the National Center for Biotechnology Information Nucleotide Sequence Database by using the FASTA program. For construction of a phylogenetic tree, all the ambiguous positions were excluded from similarity calculations. The dendrogram shown in Fig. 1B was generated by aligning the cloned sequences with 16S rRNA gene sequences representative of the main bacterial divisions. Sequences were aligned with 16S rRNA sequences of other bacteria obtained from the Ribosomal Database Project (RDP-II) (25). Matrices of evolutionary distance were computed by using the program DNADIST implemented in the software package Phylip (http://evolution.genetics.washington.edu/phylip.html) (version 3.5). The different 16S rRNA gene sequences were used to construct the phylogenetic tree.
FIG.1.
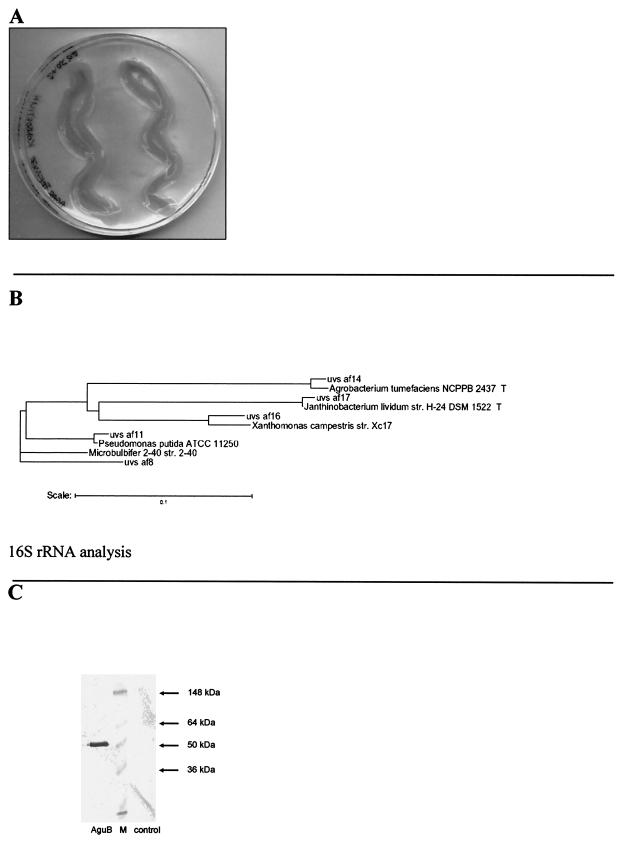
(A) Agarolytic activities of the microbial community studied on 1.5% agar plates. The plates were incubated at room temperature for 4 to 5 days. (B) Dendrogram of the 16S rRNA genes identified in the agarolytic bacterial community, showing the relationship to the closest known relatives. Evolutionary distances and the phylogenetic relationships were calculated from the aligned sequences and were corrected for multiple base changes at single positions by the method of Jukes and Cantor (18). The phylogenetic tree was calculated by using the software package Phylip. Only high-quality DNA sequences from the 16S rRNA clones were included in the calculations, and the hypervariable regions in the 16S rRNA molecule were excluded. Scale bar = 10 nucleotide substitutions per 100 nucleotides. (C) Immunological detection of the agarase AguB: Western blot of Ni-NTA-purified AguB obtained by using a six-His-specific polyclonal antibody. Lane AguB, protein extract of E. coli AguB after Ni-NTA purification; lane control, E. coli BL21(DE3) with empty pET19B after Ni-NTA purification; lane M, marker (prestained SeeBlue Plus2; Invitrogen, San Diego, Calif.). Approximately 3.0 μg of total protein per lane was loaded onto the corresponding SDS-polyacrylamide gel.
Nucleotide sequence accession numbers.
The nucleotide sequences obtained for larger contigs and complete cosmids have been deposited in the GenBank database under the accession numbers shown in Table 2. The 16S rRNA gene sequences have been deposited in the GenBank database under the following accession numbers: clone uvs_af8, AY206866; clone uvs_af11, AY206867; clone uvs_af14, AY206868; clone uvs_af16, AY206869; and clone uvs_af17, AY206870.
TABLE 2.
ORFs and genes identified for the sequenced contigs of cosmid clones pVIIC10, pXIVA3, pIIIE5, pXB11, pXE1, and pIIIF2a
| Cosmid | ORF or gene | Possible function | |
|---|---|---|---|
| pXE1 | aprE | AprE | |
| aprF | AprF | ||
| lipA | Hypothetical protein | ||
| lipB | Lipase | ||
| uvs005 | Hypothetical protein | ||
| uvs006 | Possible DNA helicase | ||
| uvs007 | Hypothetical protein | ||
| uvs008 | Possible sensor or response regulator | ||
| uvs009 | Possible sensor or response regulator | ||
| uvs010 | Possible Na+/H+ antiporter | ||
| uvs011 | Putative aminotransferase | ||
| uvs012 | Hypothetical protein | ||
| uvs013 | Hypothetical protein | ||
| aprD | AprD | ||
| uvs015 | Cytochrome c biogenesis protein CcmG | ||
| uvs016 | Cytochrome c biogenesis protein CycK | ||
| uvs017 | β-Lactamase | ||
| amiA | Enantiomer-selective amidase | ||
| pVIIC10 | uvs031 | Transporter protein | |
| uvs032 | RNA polymerase, σH | ||
| uvs033 | Peptidoglycan binding protein | ||
| aguA | β-Agarase B | ||
| aguB | β-Agarase I | ||
| uvs036 | Hypothetical protein | ||
| uvs037 | Hypothetical protein | ||
| uvs038 | Membrane protein | ||
| uvs039 | Hypothetical protein | ||
| uvs040 | Transcription regulator | ||
| uvs044 | Hypothetical protein | ||
| uvs045 | CBS domain protein | ||
| amyA | α-Amylase | ||
| uvs047 | Hypothetical protein | ||
| uvs048 | Glutamyl transpeptidase | ||
| uvs049 | Hypothetical protein | ||
| uvs050 | Transcriptional regulator | ||
| uvs051 | Hypothetical protein | ||
| uvs052 | Hypothetical protein | ||
| uvs053 | Hypothetical protein | ||
| gnuB | Endoglucanase | ||
| exbD | Biopolymer transport protein | ||
| exbB | Biopolymer transport protein | ||
| uvs054 | Hypothetical protein | ||
| uvs055 | Hypothetical protein | ||
| pXIVA3 | uvs061 | Hypothetical protein | |
| uvs062 | GTP binding protein | ||
| uvs063 | GTPase | ||
| uvs064 | Hypothetical protein | ||
| uvs065 | Hypothetical protein | ||
| uvs066 | Hypothetical protein | ||
| uvs067 | Hypothetical protein | ||
| uvs068 | Hypothetical protein | ||
| uvs069 | Hypothetical protein | ||
| actP | Atu037 | ||
| aguC | β-Agarase | ||
| uvs072 | Hypothetical protein | ||
| aguD | β-Agarase I | ||
| aguE | β-Agarase | ||
| uvs075 | Hypothetical protein | ||
| uvs076 | RNA polymerase, σH | ||
| uvs077 | Membrane protein | ||
| uvs078 | Hypothetical protein | ||
| uvs079 | Hypothetical protein | ||
| uvs080 | β-Glucanase | ||
| aguF | MS115 β-agarase | ||
| uvs082 | Regulatory protein | ||
| pXB11 | uvs083 | Hypothetical protein | |
| aguJ | β-Agarase | ||
| aguK | β-Agarase I | ||
| uvs086 | Hypothetical protein | ||
| aguL | β-Agarase | ||
| pIIIE5 | aguG | β-Agarase | |
| aguH | β-Agarase I | ||
| uvs093 | Hypothetical protein | ||
| aguI | β-Agarase | ||
| uvs095 | Hypothetical protein | ||
| uvs96 | Hypothetical protein | ||
| uvs097 | Hypothetical protein | ||
| uvs099 | Hypothetical protein | ||
| uvs100 | Hypothetical protein | ||
| uvs101 | Electron transfer flavoprotein, alpha subunit | ||
| uvs102 | Hypothetical protein | ||
| uvs103 | Hypothetical protein | ||
| uvs104 | Type III secretory protein PscC | ||
| amyB | 1,4-α-Glucan branching enzyme | ||
| uvs098 | Hypothetical protein | ||
| uvs106 | Glucose-1-phosphate adenylyltransferase | ||
| pIIIF2 | uvs110 | Hypothetical protein | |
| uvs111 | Hypothetical protein (possible lipid A biosynthesis protein) | ||
| uvs112 | Hypothetical protein | ||
| Uvs113 | Hypothetical protein | ||
| uvs114 | Hypothetical protein | ||
| glyQ | Glycyl-tRNA synthetase, alpha chain | ||
| glyS | Glycyl-tRNA synthetase, beta chain | ||
| uvs117 | HisB-like phosphatase | ||
| uvs118 | Probable acyltransferase | ||
| uvs119 | Hypothetical protein (possible pectate lyase II) | ||
| pelA | Pectate lyase B | ||
| uvs121 | Hypothetical protein | ||
| uvs122 | Hypothetical protein | ||
| uvs123 | Metallo beta-lactamase | ||
| uvs124 | Monophenol monooxygenase | ||
| uvs125 | Transmembrane protein | ||
| uvs126 | Hypothetical protein | ||
| uvs127 | Hypothetical protein | ||
| uvs128 | 2-Acyl-glycerophospho-ethanolamine acyltransferase |
DNA sequences were deposited in the GenBank database under accession numbers AY212801, AY212800, AY236226, AY236224, AY236225, AY240957, AY240958, and AY236223. Identified ORFs were designated by a gene name if the e-value was less than 10−80. Additional data, including the results of similarity searches and the lengths of the predicted ORFs, are available on the web at http://www.gwdg.de/∼biofilm/genemining.htm.
RESULTS
Agarolytic and mixed cultures.
The mixed cultures were inoculated with samples derived from soil collected from an unplanted field. The samples were directly applied onto agar plates containing the defined medium, and the plates were incubated until degradation of the agar was clearly visible (Fig. 1A). After two rounds of transfer onto fresh plates, colonies were removed from the agar plates for DNA isolation. Interestingly, agarolytic activity was not observed when the cultures were grown in LB liquid medium or on LB agar plates. Attempts to isolate individual microbes with agarolytic activity from the consortium failed. Although several different isolates were obtained, none of them liquefied the agar after growth on complex or defined medium (data not shown).
The phylogenetic diversity of the selected microbes was assessed by isolation of DNA, amplification, cloning, and DNA sequence analysis of 16S rRNA genes. Altogether, sequences of 24 different clones carrying the 16S rRNA gene sequences were analyzed; this resulted in isolation of five different clones, all of which grouped in the beta and gamma subclasses of the class Proteobacteria. The phylogenetic tree shown in Fig. 1B represents the diversity of the five different clones and their closest bacterial relatives. The microbes present in this bacterial community were closely related to Pseudomonas putida, Agrobacterium tumefaciens, Xanthomonas campestris, Janthinobacterium lividum, and Microbulbifer degradans.
Construction of cosmid DNA libraries from the microbial consortium.
Genes associated with the hydrolytic activities of the mixed culture were identified by isolation of DNA from the bacteria by direct lysis, size fractionation to obtain fragments whose sizes ranged from 20 to 40 kb, and ligation into the cosmid vector pWE15 (see Materials and Methods). After the DNA was packaged and after transfection into the E. coli host strain, a library consisting of several thousand clones was generated; 1,532 of these clones were screened for agarolytic and lipolytic activities. The majority of the clones analyzed contained inserts with an average size of approximately 25 to 40 kb, as determined by examining 24 randomly picked clones. In addition, restriction analysis of randomly chosen cosmid clones revealed a high level of diversity of the DNA fragments cloned into pWE15.
Screening revealed novel agarolytic enzymes.
The environmentally derived cosmid library was grown in microplates, the bacteria were pelleted and lysed, and agarose-encoding clones were identified as described in Materials and Methods. Altogether, we isolated four different cosmid clones which appeared to contain agarolytic activities. A DNA restriction analysis ensured that each clone was unique (data not shown). These clones, designated pIIIE5, pVIIC10, pXB11, and pXIVA3, were further characterized by isolation of DNA, restriction with different enzymes, and partial or complete sequencing. Restriction analysis and shotgun sequencing confirmed that the insert sizes in pWE15 were approximately 25 to 38 kb and allowed detailed mapping of the agarolytic cosmids. Figure 2 and Table 2 summarize the genes identified and the similarities observed after comparison of the deduced amino acid sequences with the sequences available in the National Center for Biotechnology Information databases.
FIG. 2.
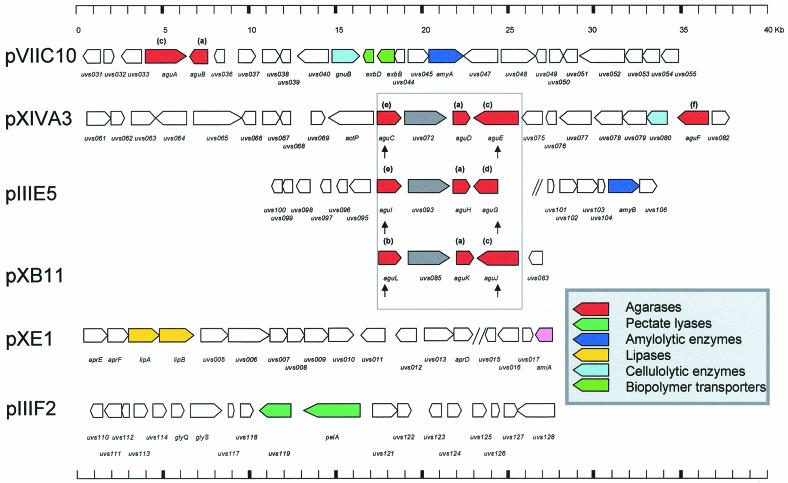
Physical maps of the central parts of five of the different cosmids isolated in this work. The arrows indicate the locations and the directions of transcription of the ORFs on the different cosmids (pVIIC10, pXIVA3, pIIIE5, pXB11, pXE1, and pIIIF2). ORFs encoding potential agarases are indicated by red arrows. Identical agarase genes are indicated by the same letters (a to f) in parentheses; the agarases designated e and d differ in their N-terminal domains but are identical at their C termini. Possible genes with potential value in biotechnology are shown in color. Possible functions of the indicated ORFs are shown in Table 2, together with the GenBank accession numbers. A conserved agarolytic gene cluster is enclosed in a box. ORFs uvs072, uvs093, and uvs085 in this conserved cluster code for the same 785-amino-acid protein and are indicated by gray arrows; contig ends are indicated by two slashes, and the vertical arrows indicate the positions of the two 31-bp inverted repeats. Additional data, including the results of similarity searches and the lengths of the predicted ORFs, are available on the web at http://www.gwdg.de/∼biofilm/genemining.htm.
The complete nucleotide sequence of pVIIC10 was determined, and the insert size was 34.7 kb. Two genes coding for agarases were identified and designated aguA and aguB. These two genes were linked, but they appeared to be transcribed in opposite directions. AguA exhibited 50% identity to the corresponding protein from Vibrio sp., and AguB was 77% identical to the corresponding protein of Pseudoalteromonas atlantica. In addition to these β-agarase genes, several genes with high biotechnological potential were detected on the same cosmid; these genes included a gene that was very similar to a gene encoding an α-amylase (amyA) in Dictyoglomus thermophilum, a gene encoding an endo-1,4-beta-glucanase (cellulase), and two open reading frames (ORFs) with high levels of similarity to genes encoding biopolymer transporters (exbB to exbD).
Sequencing of cosmid pXIVA3 revealed an insert size of 38.1 kb and the presence of 22 ORFs. Four genes encoding putative agarases were designated aguC to aguF. Three of these genes were located in a 7.1-kb cluster (Fig. 2). Interestingly, aguD and aguE were identical to aguB and aguA, respectively. AguC was very similar to the β-agarase from Vibrio sp. The AguF protein was very similar to the protein encoded by a putative agarase gene, MSC115, in Microscilla sp. In addition to the four agarase genes present on pXIVA3, we identified one gene, uvs080 encoding a putative endo-1,3(4)-β-glucanase, which was similar to a corresponding gene in Clostridium thermocellum. Interestingly, several ORFs (uvs066 to uvs069) found on this cosmid did not exhibit homology to any of the genes annotated in the databases.
The genes homologous to β-agarase genes encoded on cosmid clone pIIIE5 were designated aguG to aguI, and all of them occurred in clusters. In fact, AguH was identical to AguB and AguD, and AguI was identical to AguC (Fig. 2 and Table 2). Additionally, we identified the amyB gene encoding an 1,4-α-glucan branching enzyme. Altogether, 25 kb of DNA sequences was determined for this cosmid clone.
Finally, another cluster of three agarase genes was identified on cosmid pXB11; these genes were designated aguJ, aguK, and aguL. The corresponding agarases were identical to other agarases identified previously (Fig. 2 and Table 2).
In summary, we identified 12 ORFs encoding potential and functional agarases. Most likely, several of these ORFs are the result of gene duplications. In fact, several of the functional and possible agarase genes encode identical proteins. This is indicated in Fig. 2. It is also very intriguing that nine of the agarase genes identified were located in clusters of three, as indicated in Fig. 2. The three clusters encode different agarases, which may have resulted from gene duplications and horizontal gene transfer, as suggested by the DNA sequences of the flanking regions. This hypothesis is supported by the observation that a 31-bp inverted repeat is located in the agarase clusters. The two repeated sequences were separated by 6,225 bp, and their positions are indicated in Fig. 2.
Subcloning and overexpression of the aguB gene as an N-terminally His-tagged protein resulted in production of a 52-kDa protein in E. coli BL21, as shown by Western blotting with a His-specific antibody (Fig. 1C). The estimated molecular mass of AguB (52 kDa) was in good agreement with the calculated molecular mass (49.1 kDa). Protein extracts from E. coli overproducing the aguB gene liquefied low-melting-point agar, in contrast to protein extracts derived from control strains (data not shown). Recombinant AguB was found to have a temperature optimum of 42°C, and no activity was detected at 55°C (data not shown). These findings clearly demonstrate that aguB and the homologues identified (aguD, aguH, and aguK) code for functional agarases. Further overexpression of the aguF gene indicated that this gene also encoded a fully functional agarase (data not shown). Additionally, tests with E. coli carrying cosmid pVIIC10 (Fig. 2 and Table 2) indicated that the genes encoding endoglucanse B (gnuB) and α-amylase (amyA) were both functional. A similar result was obtained for amyB encoded on cosmid pIIIE5 (data not shown).
Screening for DNA fragments containing lipolytic activities.
Cosmid clones encoding lipase activity were identified by transfer to agar plates containing triolein/tributyrin. Formation of clearing zones around the colonies after 4 to 5 days of incubation indicated that lipase or esterase genes were expressed. DNA was isolated and retransformed, and clones were reexamined on the same indicator plates to avoid isolation of false-positive clones. Finally, clone pXE1 was characterized by DNA sequencing and restriction analysis, which revealed an insert size of approximately 25 kb and a total of 18 ORFs. Two lipase genes appeared to be located in a cluster together with other hypothetical genes. The lipase amino acid sequences were very similar to the sequences of lipases from Pseudomonas sp. strain UB48 and Pseudomonas brassicacearum (Table 2). The lipase genes were located adjacent to genes encoding parts of a type I secretion system (aprE and aprF) (Fig. 2).
In addition, we identified on the same cosmid an ORF whose product was very similar to a stereoselective amidase from Rhodococcus sp. Further sequencing of randomly chosen cosmid clones resulted in identification of other genes presumably encoding interesting biocatalysts. Among them were two novel ORFs coding for pectate lyases, which were designated pelA and uvs119 (Fig. 2). Both of these genes were identified on cosmid pIIIF2, which harbored 28 kb of insert DNA. PelA was very similar to a pectate lyase from Pseudoalteromonas haloplanktis, and the second pectate lyase (Uvs119) was similar to a putative pectate lyase II from Bacillus halodurans (Table 2).
DISCUSSION
In this work we utilized a combined precultivation-metagenomics technology for isolation and comparative analysis of gene clusters from a microbial consortium. In general, cultivation has been successfully used to screen for single microbes or consortia with diverse catabolic capabilities, including the ability to degrade toluene (12) and phthalate (20). However, only a limited number of reports have described direct cloning from mixed laboratory cultures (8, 21) or enrichments (7) to isolate novel genes or operons useful for biotechnology.
The strategy applied here led to isolation of a microbial consortium consisting of several novel bacterial species which belong to the nonpathogenic gamma and beta subclasses of the Proteobacteria (Fig. 1B). Although many of the genes detected in this agarolytic consortium were surprisingly similar to genes of known microbes, we have no evidence indicating from which species the cosmids analyzed were derived.
In this work we isolated a significant number of genes with high potential for biotechnological applications. A total of 12 potential β-agarases were identified, and 6 of these enzymes were truly different (Fig. 2). Agar is a linear polymer of sulfated glucan composed of two regularly repeated galactose units alternatively linked by β-d-(1,4) and α-l-(1,3) linkages. The β-agarases cleave the β-d-(1,4) linkages, and the α-agarases cleave the α-l-(1,3) linkages. These enzymes are very interesting because agar is an additive in many foods and feed products, as well as an important compound for a large number of molecular biology applications.
It is very intriguing that most of the agarases identified in this study were in clusters, which was probably a result of gene duplications and gene transfer (Fig. 2). A similar observation has been made for the putative agarases located on a 101-kb plasmid of Microscilla sp. In the DNA sequence of pSD15 five ORFs were defined as ORFs that encode putative agarases, and several of them were very similar to each other. Interestingly, four of five agarases detected in pSD15 were members of clusters containing two enzymes (33).
In addition to the agarolytic genes, we identified several other genes which encode biocatalysts with high biotechnological potential (1, 11, 23, 32). These biocatalysts include two putative cellulases encoded by gnuB and uvs80 and the starch-degrading enzyme α-amylase encoded by amyA, as well as a 1,4-α-glucan branching enzyme encoded by amyB. Recently, we have shown that these genes encode functional enzymes (unpublished data). Furthermore, we have identified two putative pectate lyases encoded by pelA and uvs119 and also proteolytic enzymes with the ability to hydrolyze gellan gum (data not shown).
A cluster of lipase-encoding genes was identified (Table 2 and Fig. 2) together with the genes encoding a type I secretion system possibly involved in lipase secretion (6). The lipase genes were located in the 3′ direction of the aprE and aprF genes in a conserved gene organization, as previously identified in Pseudomonas fluorescens and P. brasicacearum. In both these microbes genes coding for alkaline proteases are linked to the lipase gene clusters (5, 6, 19). Such putative protease genes were missing on our cosmid clone pXE1. Lipases remain enzymatically active in organic solvents, they usually display exquisite chemo-, regio-, and stereoselectivities, they do not require cofactors, and they do not catalyze side reactions. These properties make lipases not only a very important class of enzymes for a variety of biotechnological applications but also the most widely used group of biocatalysts in organic chemistry (13-17).
Finally, it should be mentioned that in our study the frequency of isolating novel genes encoding potential biocatalysts was extremely high compared to the frequencies in other metagenome studies. In the other studies much larger numbers of clones were screened to identify the desired biocatalysts (3, 9, 10, 24, 28). Henne and coworkers screened 286,000 clones and obtained four clones carrying lipase genes (9). A similar number of clones (148,000 clones) were analyzed to identify two clones encoding possible Na+/H+ antiporter proteins (26). Although other workers analyzed smaller clone libraries, they still had to examine large numbers of clones (28). Rondon and coworkers (28) found clones expressing DNase (one clone), antibacterial activity (one clone), lipase (two clones), and amylase (eight clones) in a library consisting of 24,000 clones. Finally, Knietsch and coworkers (21) identified only eight functional clones when they screened more than 100,000 clones. In contrast, our results suggest that cocultivation in a mixed culture in combination with direct cloning allows rapid screening for and successful identification of multiple biocatalyst genes. Consequently, the technique which we used should prove to be a valuable tool for exploiting metagenomic biodiversity.
Acknowledgments
This work was supported by the BMBF-Kompetenznetzwerke (Genome Research on Bacteria for the Analysis of Biodiversity and Its Further Use for the Development of New Production Processes) and by the Fonds der chemischen Industrie.
REFERENCES
- 1.Alkorta, I., C. Garbisu, M. Llama, and J. Serra. 1998. Industrial applications of pectic enzymes: a review. Process Biochem. 33:21-28. [Google Scholar]
- 2.Amann, R. I., W. Ludwig, and K. H. Schleifer. 1995. Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol. Rev. 59:143-169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Beja, O., M. T. Suzuki, E. V. Koonin, L. Aravind, A. Hadd, L. P. Nguyen, R. Villacorta, M. Amjadi, C. Garrigues, S. B. Jovanovich, R. A. Feldman, and E. F. DeLong. 2000. Construction and analysis of bacterial artificial chromosome libraries from a marine microbial assemblage. Environ. Microbiol. 2:516-529. [DOI] [PubMed] [Google Scholar]
- 4.Caldwell, D. E., W. G. Korber, and J. R. Lawrence. 1997. Cultivation of microbial consortia and communities, p. 79-90. In J. H. Hurst, G. R. Knudsen, M. J. McInnerny, L. D. Stezenbach, and M. V. Walter (ed.), Manual of environmental microbiology. ASM Press, Washington, D.C.
- 5.Chabeaud, P., A. de Groot, W. Bitter, J. Tommassen, T. Heulin, and W. Achouak. 2001. Phase-variable expression of an operon encoding extracellular alkaline protease, a serine protease homolog, and lipase in Pseudomonas brassicacearum. J. Bacteriol. 183:2117-2120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Duong, F., C. Soscia, A. Lazdunski, and M. Murgier. 1994. The Pseudomonas fluorescens lipase has a C-terminal secretion signal and is secreted by a three-component bacterial ABC-exporter system. Mol. Microbiol. 11:1117-1126. [DOI] [PubMed] [Google Scholar]
- 7.Entcheva, P., W. Liebl, A. Johann, T. Hartsch, and W. R. Streit. 2001. Direct cloning from enrichment cultures, a reliable strategy for isolation of complete operons and genes from microbial consortia. Appl. Environ. Microbiol. 67:89-99. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Healy, F. G., R. M. Ray, H. C. Aldrich, A. C. Wilkie, L. O. Ingram, and K. T. Shanmugam. 1995. Direct isolation of functional genes encoding cellulases from the microbial consortia in a thermophilic, anaerobic digester maintained on lignocellulose. Appl. Microbiol. Biotechnol. 43:667-674. [DOI] [PubMed] [Google Scholar]
- 9.Henne, A., R. Daniel, R. A. Schmitz, and G. Gottschalk. 1999. Construction of environmental DNA libraries in Escherichia coli and screening for the presence of genes conferring utilization of 4-hydroxybutyrate. Appl. Environ. Microbiol. 65:3901-3907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Henne, A., R. A. Schmitz, M. Bomeke, G. Gottschalk, and R. Daniel. 2000. Screening of environmental DNA libraries for the presence of genes conferring lipolytic activity on Escherichia coli. Appl. Environ. Microbiol. 66:3113-3116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Henriksson, G., D. Akin, D. Slomczynski, and K.-E. L. Eriksson. 1999. Production of highly efficient enzymes for flax retting by Rhizomucor pusillus. J. Biotechnol. 68:115-123. [Google Scholar]
- 12.Hubert, C., Y. Shen, and G. Voordouw. 1999. Composition of toluene-degrading microbial communities from soil at different concentrations of toluene. Appl. Environ. Microbiol. 65:3064-3070. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Jaeger, K. E., B. W. Dijkstra, and M. T. Reetz. 1999. Bacterial biocatalysts: molecular biology, three-dimensional structures, and biotechnological applications of lipases. Annu. Rev. Microbiol. 53:315-351. [DOI] [PubMed] [Google Scholar]
- 14.Jaeger, K. E., and T. Eggert. 2002. Lipases for biotechnology. Curr. Opin. Biotechnol. 13:390-397. [DOI] [PubMed] [Google Scholar]
- 15.Jaeger, K. E., S. Ransac, B. W. Dijkstra, C. Colson, M. van Heuvel, and O. Misset. 1994. Bacterial lipases. FEMS. Microbiol. Rev. 15:29-63. [DOI] [PubMed] [Google Scholar]
- 16.Jaeger, K. E., and M. T. Reetz. 2000. Directed evolution of enantioselective enzymes for organic chemistry. Curr. Opin. Chem. Biol. 4:68-73. [DOI] [PubMed] [Google Scholar]
- 17.Jaeger, K. E., and M. T. Reetz. 1998. Microbial lipases form versatile tools for biotechnology. Trends Biotechnol. 16:396-403. [DOI] [PubMed] [Google Scholar]
- 18.Jukes, T. H., and C. R. Cantor. 1969. Evolution of protein molecules, p. 21-132. In H. N. Munro (ed.), Mammalian protein metabolism. Academic Press, New York, N.Y.
- 19.Kawai, E., A. Idei, H. Kumura, K. Shimazaki, H. Akatsuka, and K. Omori. 1999. The ABC-exporter genes involved in the lipase secretion are clustered with the genes for lipase, alkaline protease, and serine protease homologues in Pseudomonas fluorescens no. 33. Biochim. Biophys. Acta 1446:377-382. [DOI] [PubMed] [Google Scholar]
- 20.Kleerebezem, R., L. W. H. Pol, and G. Lettinga. 1999. Anaerobic degradation of phthalate isomers by methanogenic consortia. Appl. Environ. Microbiol. 65:1152-1160. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Knietsch, A., T. Waschkowitz, S. Bowien, A. Henne, and R. Daniel. 2003. Construction and screening of metagenomic libraries derived from enrichment cultures: generation of a gene bank for genes conferring alcohol oxidoreductase activity on Escherichia coli. Appl. Environ. Microbiol. 69:1408-1416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kok, R. G., V. M. Christoffels, B. Vosman, and K. J. Hellingwerf. 1993. Growth-phase-dependent expression of the lipolytic system of Acinetobacter calcoaceticus BD413: cloning of a gene encoding one of the esterases. J. Gen. Microbiol. 139:2329-2342. [DOI] [PubMed] [Google Scholar]
- 23.Lynd, L. R., P. J. Weimer, W. H. van Zyl, and I. S. Pretorius. 2002. Microbial cellulose utilization: fundamentals and biotechnology. Microbiol. Mol. Biol. Rev. 66:506-577. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.MacNeil, I. A., C. L. Tiong, C. Minor, P. R. August, T. H. Grossman, K. A. Loiacono, B. A. Lynch, T. Phillips, S. Narula, R. Sundaramoorthi, A. Tyler, T. Aldredge, H. Long, M. Gilman, D. Holt, and M. S. Osburne. 2001. Expression and isolation of antimicrobial small molecules from soil DNA libraries. J. Mol. Microbiol. Biotechnol. 3:301-308. [PubMed] [Google Scholar]
- 25.Maidak, B. L., J. R. Cole, T. G. Lilburn, C. T. Parker, Jr., P. R. Saxman, R. J. Farris, G. M. Garrity, G. J. Olsen, T. M. Schmidt, and J. M. Tiedje. 2001. The RDP-II (Ribosomal Database Project). Nucleic Acids Res. 29:173-174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Majernik, A., G. Gottschalk, and R. Daniel. 2001. Screening of environmental DNA libraries for the presence of genes conferring Na+ (Li+)/H+ antiporter activity on Escherichia coli: characterization of the recovered genes and the corresponding gene products. J. Bacteriol. 183:6645-6653. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Raasch, C., W. Streit, J. Schanzer, M. Bibel, U. Gosslar, and W. Liebl. 2000. Thermotoga maritima AglA, an extremely thermostable NAD+-, Mn2+-, and thiol-dependent alpha-glucosidase. Extremophiles 4:189-200. [DOI] [PubMed] [Google Scholar]
- 28.Rondon, M. R., P. R. August, A. D. Bettermann, S. F. Brady, T. H. Grossman, M. R. Liles, K. A. Loiacono, B. A. Lynch, I. A. MacNeil, C. Minor, C. L. Tiong, M. Gilman, M. S. Osburne, J. Clardy, J. Handelsman, and R. M. Goodman. 2000. Cloning the soil metagenome: a strategy for accessing the genetic and functional diversity of uncultured microorganisms. Appl. Environ. Microbiol. 66:2541-2547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Sambrook, J., E. F., Fritsch, and T. Maniatis. 1989. Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
- 30.Torsvik, V., and L. Ovreas. 2002. Microbial diversity and function in soil: from genes to ecosystems. Curr. Opin. Microbiol. 5:240-245. [DOI] [PubMed] [Google Scholar]
- 31.Tsai, Y. L., and B. H. Olson. 1992. Rapid method for separation of bacterial DNA from humic substances in sediments for polymerase chain reaction. Appl. Environ. Microbiol. 58:2292-2295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.van der Maarel, M. J., B. van der Veen, J. C. Uitdehaag, H. Leemhuis, and L. Dijkhuizen. 2002. Properties and applications of starch-converting enzymes of the alpha-amylase family. J. Biotechnol. 94:137-155. [DOI] [PubMed] [Google Scholar]
- 33.Zhong, Z., A. Toukdarian, D. Helinski, V. Knauf, S. Sykes, J. E. Wilkinson, C. O'Bryne, T. Shea, C. DeLoughery, and R. Caspi. 2001. Sequence analysis of a 101-kilobase plasmid required for agar degradation by a Microscilla isolate. Appl. Environ. Microbiol. 67:5771-5779. [DOI] [PMC free article] [PubMed] [Google Scholar]