Abstract
The phylogenetic relationship of chemoautotrophic, sulfur-oxidizing, ectosymbiotic bacteria growing on a marine nematode, a Laxus sp. (formerly a Catanema sp.), to known endosymbionts and free-living bacteria was determined. Comparative 16S rRNA sequencing was used to investigate the unculturable nematode epibionts, and rRNA-targeted oligonucleotide hybridization probes were used to identify the ectosymbionts in situ. Both analyses revealed a remarkably specific and stable symbiosis. Unique hybridization of a specific probe to the ectosymbionts indicated that only one species of bacteria was present and growing on the cuticle of the nematode. Distance and parsimony methods used to infer phylogenetic trees both placed the nematode ectosymbionts at the base of a branch containing chemoautotrophic, sulfur-oxidizing endosymbionts of three bivalve families and of the tube worm Riftia pachyptila. The most closely related free-living bacteria were chemoautotrophic sulfur oxidizers belonging to the genus Thiomicrospira. Furthermore, our results suggested that a second, only distantly related group of thioautotrophic endosymbionts has as its deepest branch surface-colonizing bacteria belonging to the genus Thiothrix, some of which are capable of sulfur-oxidizing chemoautotrophic growth.
Full text
PDF


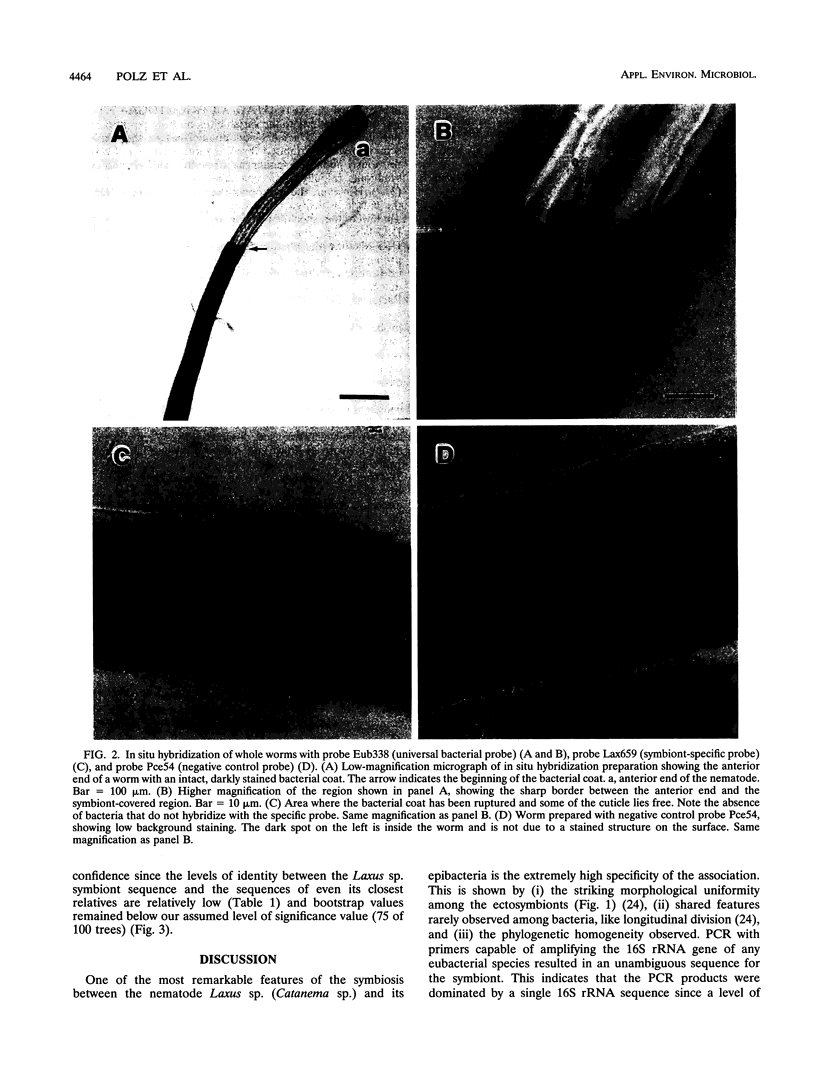



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Altschul S. F., Gish W., Miller W., Myers E. W., Lipman D. J. Basic local alignment search tool. J Mol Biol. 1990 Oct 5;215(3):403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- Amann R. I., Binder B. J., Olson R. J., Chisholm S. W., Devereux R., Stahl D. A. Combination of 16S rRNA-targeted oligonucleotide probes with flow cytometry for analyzing mixed microbial populations. Appl Environ Microbiol. 1990 Jun;56(6):1919–1925. doi: 10.1128/aem.56.6.1919-1925.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Amann R. I., Krumholz L., Stahl D. A. Fluorescent-oligonucleotide probing of whole cells for determinative, phylogenetic, and environmental studies in microbiology. J Bacteriol. 1990 Feb;172(2):762–770. doi: 10.1128/jb.172.2.762-770.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Distel D. L., Lane D. J., Olsen G. J., Giovannoni S. J., Pace B., Pace N. R., Stahl D. A., Felbeck H. Sulfur-oxidizing bacterial endosymbionts: analysis of phylogeny and specificity by 16S rRNA sequences. J Bacteriol. 1988 Jun;170(6):2506–2510. doi: 10.1128/jb.170.6.2506-2510.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Distel D. L., Wood A. P. Characterization of the gill symbiont of Thyasira flexuosa (Thyasiridae: Bivalvia) by use of polymerase chain reaction and 16S rRNA sequence analysis. J Bacteriol. 1992 Oct;174(19):6317–6320. doi: 10.1128/jb.174.19.6317-6320.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eisen J. A., Smith S. W., Cavanaugh C. M. Phylogenetic relationships of chemoautotrophic bacterial symbionts of Solemya velum say (Mollusca: Bivalvia) determined by 16S rRNA gene sequence analysis. J Bacteriol. 1992 May;174(10):3416–3421. doi: 10.1128/jb.174.10.3416-3421.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lane D. J., Harrison A. P., Jr, Stahl D., Pace B., Giovannoni S. J., Olsen G. J., Pace N. R. Evolutionary relationships among sulfur- and iron-oxidizing eubacteria. J Bacteriol. 1992 Jan;174(1):269–278. doi: 10.1128/jb.174.1.269-278.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larkin J. M., Henk M. C., Burton S. D. Occurrence of a Thiothrix sp. Attached to Mayfly Larvae and Presence of Parasitic Bacteria in the Thiothrix sp. Appl Environ Microbiol. 1990 Feb;56(2):357–361. doi: 10.1128/aem.56.2.357-361.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larsen N., Olsen G. J., Maidak B. L., McCaughey M. J., Overbeek R., Macke T. J., Marsh T. L., Woese C. R. The ribosomal database project. Nucleic Acids Res. 1993 Jul 1;21(13):3021–3023. doi: 10.1093/nar/21.13.3021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stahl D. A., Lane D. J., Olsen G. J., Pace N. R. Analysis of hydrothermal vent-associated symbionts by ribosomal RNA sequences. Science. 1984 Apr 27;224(4647):409–411. doi: 10.1126/science.224.4647.409. [DOI] [PubMed] [Google Scholar]
- Zarda B., Amann R., Wallner G., Schleifer K. H. Identification of single bacterial cells using digoxigenin-labelled, rRNA-targeted oligonucleotides. J Gen Microbiol. 1991 Dec;137(12):2823–2830. doi: 10.1099/00221287-137-12-2823. [DOI] [PubMed] [Google Scholar]
- Zharkikh A., Li W. H. Statistical properties of bootstrap estimation of phylogenetic variability from nucleotide sequences. I. Four taxa with a molecular clock. Mol Biol Evol. 1992 Nov;9(6):1119–1147. doi: 10.1093/oxfordjournals.molbev.a040782. [DOI] [PubMed] [Google Scholar]