Abstract
Glycosidase and lectins both bind sugars, but only the glycosidases have catalytic activity. The glycosidases occur among 91 evolved protein families and Family 18 is one of the two chitinases (EC 3, 2.1.14) families. Interestingly, lectins are also in this evolutionary group of Family 18 glycosidase proteins. The proteins belonging to the enzymatically inactive class are referred to as, chitolectins and have a binding site that is highly similar to the catalytic Family 18 enzymes. We present a comparison of the recently obtained structures of two Family 18 chitolectins, MGP40 (Mohanty, Singh et al., 2003) and HumGP39 (Fusetti, Pijning et al., 2003; Houston, Anneliese et al., 2003) with glycosidases active site. We compare the sequence and the structure of these two Family 18 proteins. The difference between the active and inactive protein is a glutamic acid which acts as the essential acid/base residue for chitin cleavage is replaced with leucine or glutamine. Furthermore, mechanism for the interaction between the chitolectin and oligosaccharides were proposed.
Keywords: Oligosaccharide, binding, mechanism, chitinases
Introduction
Chitinases (EC 3.2.1.14) hydrolyze chitin β(1 → 4) linkages in order to make available the large quantities of carbon and nitrogen elements potentially trapped in the biosphere as insoluble N-acetyl-glucosamine (GlcNAc) [1]. Henrissat and Davies [2] have classified glycosyl hydrolases into 91 evolutionary groups based on their homologous amino acid sequences. These enzyme families are further organized into clans that have common three-dimensional structures, but not necessarily common primary sequences. Glycosidase Family 18 and 19 encompass the chitin hydrolases. Of these, Family 18 proteins fall into two distinct classes. One class of Family 18 proteins (chitinases) are active, that is they bind and hydrolyze oligosaccharides, whereas the other class of proteins ("chitolectins") are highly similar in binding oligosaccharides but do not hydrolyze them. Through natural evolution the chitolectin members of Family 18 proteins [3,4] have lost their hydrolysis potential because the catalytic acid/base glutamic acid in the substrate binding groove has been substituted by non-acidic amino acids (e.g. Leu).
Currently, the physiological role of these non-enzymatic proteins has not been elucidated. The best evidence based on their unique times of physiological expression indicates that these proteins are involved in tissue remodeling. Most likely due to their structures, they function by binding oligosaccharides. Since these chitolectins all have a signal peptide, they are likely secreted whereby they can interact with extracellular matrix proteoglycans or cell surface sugars in glycolipids or glycoproteins. These proteins therefore do not function in a stand-alone mode, but instead function either as direct cofactors during molecular signaling processes or help to present oligosaccharide targets to other proteins.
MGP40
Recently, an x-ray structure for one member of the chitolectin class of proteins MGP40 (PDB ID: 1LJY) was determined [5]. The structure is consistent with the (β/α)8 barrel topology of the Family 18 glycosidase proteins [6–26]. MGP40 is an Asn-linked glycoprotein itself. Mohanty et al. in their study proposed that the single disaccharide that is covalently linked to the protein (at Asn39) forms hydrogen bonds with Arg84 and Ile40, influencing the backbone conformation of loop Val75-Phe85, which in turn alters the disposition of Trp78. Since Trp78 is an essential member of the binding site, its altered positioning leads to constriction of the binding site, thereby leading to the inability of the MGP40 to bind sugar. Therefore, they hypothesized that oligosaccharides cannot bind to the putative binding site. Finally, the authors point out that Arg84, which is hydrogen-bonded to the covalently bond sugar, is conserved in this particular class of chitolectins. The active chitinases in the Family 18 group which bind and cleave sugars possess a Pro in place of Arg84, which in turn leads to inability of hydrogen bonding to the covalent linked sugar. Subsequently, there is a “relaxed” backbone conformation of the Val75-Phe85 loop in the Family 18 chitinases and thereby no constriction of the binding site.
HUMGP39
Very recently x-ray crystal structures of another Family 18 chitolectin HumGP39 (PDB ID: 1HJX) was reported independently by two different groups [9, 27]. These structures are also consistent with the (β/α) 8 barrel topology of the Family 18 proteins. The HumGP39 protein structures show a disaccharide covalently linked to Asn39 of HumGP39. However, more importantly these structures depict in addition an oligosaccharide bound in the active site. Different crystal structures of this protein show complexes with di, tri, tetra, penta and hexasaccharide bound in the active site. Based on these studies and in contrast to the suggestions made by Mohanty et al., it appears that the chitolectin group of proteins from Family 18 can bind chitin oligosaccharides. We compare the sequence and the structure of these two Family 18 proteins to justify this point. We also propose a mechanism for the interaction between the chitolectin and oligosaccharides.
Results and discussion
The primary sequences of MGP40 and HumGP39 are highly identical (83% identity) (see Fig. 1). We found that since Arg84 is conserved between the two proteins, it is not the cause for the lack of oligosaccharide binding in MGP40 as proposed by Mohanty et al. [5]. Hence we compared the aromatic residues in both proteins, focusing on the Trp and Tyr residues present in the floor of the binding groove that are likely to interact strongly with the sugar molecules. These residues also are important for chitin oligosaccharide binding by the active Family 18 chitinases. The interactions between the protein aromatic residues and the sugars in a hexasaccharide are listed in Table 1.
Fig. 1.

Sequence alignment for Human GP39 (362 residues); Goat MGP40 (362 residues); and Human Chitotriosidase (366 residues) is shown. Identity (*), 181 residues, 49.05%; strongly similar (:), 78 residues, 21.14%; Weakly similar (.), 41 residues, 11.11%; Different 69 residues, 18.70%.
Table 1.
Aromatic residues interacting with the oligosaccharides
| Sugar in subsite | −6 | −5 | −4 | −3 | −2 | −1 | +1 | +2 | +3 |
|---|---|---|---|---|---|---|---|---|---|
| HumGP39 | W71 | Y34 | - | W31 | - | W352 | W99 | - | W212 |
| MGP40 | W50 | Y13 | - | W10 | - | W331 | W78 | - | W191 |
| HumCHI | W71 | Y34 | - | W31 | - | W358 | W99 | - | W218 |
All of the aromatic residues listed in Table 1 that define the binding groove are conserved between MGP40 and HumGP39, and even the human chitinase, chitotriosidase. Having ruled out Arg84 and the composition of binding groove aromatics as the cause for a lack of observed oligosaccharide binding by MGP40, we mapped out the full binding region of HumGP39 by identifying all residues within 4.5 Å of the sugar residues in the hexasaccharide-bound HumGP39 structure (PDB code: 1HJW), wherein the hexasaccharide binds from −4 to +2 subsites (see Fig 2).
Fig 2.

Residues within 4.5 Å of the sugar residues in the hexasaccharide-bound HumGP39 structure
The identified residues in HumGP39 were compared to the equivalent residues in MGP40. The residues are highly identical between the proteins near the binding site with only three differences observed between the two proteins (Table 2).
Table 2.
Differences between MGP40 and HumGP39 in the binding subside
| HumGP39 | Sugar in subsite | MGP40 |
|---|---|---|
| Thr 163 | +2 | Ala163 |
| Met 183 | −1, +1 | Leu183 |
| Thr 272 | −2 | Glu272 |
Of those three amino acids Thr 163 and Thr 272 are ~4 Å away from the +2 and the −2 sites respectively, and Met 183 is ~3 Å away from the −1 and +1 sugars. Hence the contribution from their interactions with the sugar to the overall interaction is very minimal. The majority of the favorable interaction between the protein and sugar is due to the interaction between sugar rings and the aromatic residues at −6, −5, −3, −1 +1 and +3 (Table 1).
Structural Comparison: Loop 78–85
Mohanty et al. proposed that the Trp 78 - Phe 85 loop in MGP40 is altered due to the hydrogen bond between Arg84 and the covalently linked sugar at Asn39. Hence, we compared the backbone configuration of the Trp 78 – Phe 85 loop in MGP40 with the equivalent residues in HumGP39 (Fig. 3). Upon superimposing the protein structures (PDB codes: 1LJY and 1HWX) we observed that Arg84 is in identical position in both proteins. Interestingly, we did not observe the proposed hydrogen bond between Arg84 and the covalent sugar in MGP40. The sugar is ~4 Å away from Arg84. The sequence comparison of the Trp 78 – Phe 85 (MGP40) loop is shown below.
Fig. 3.
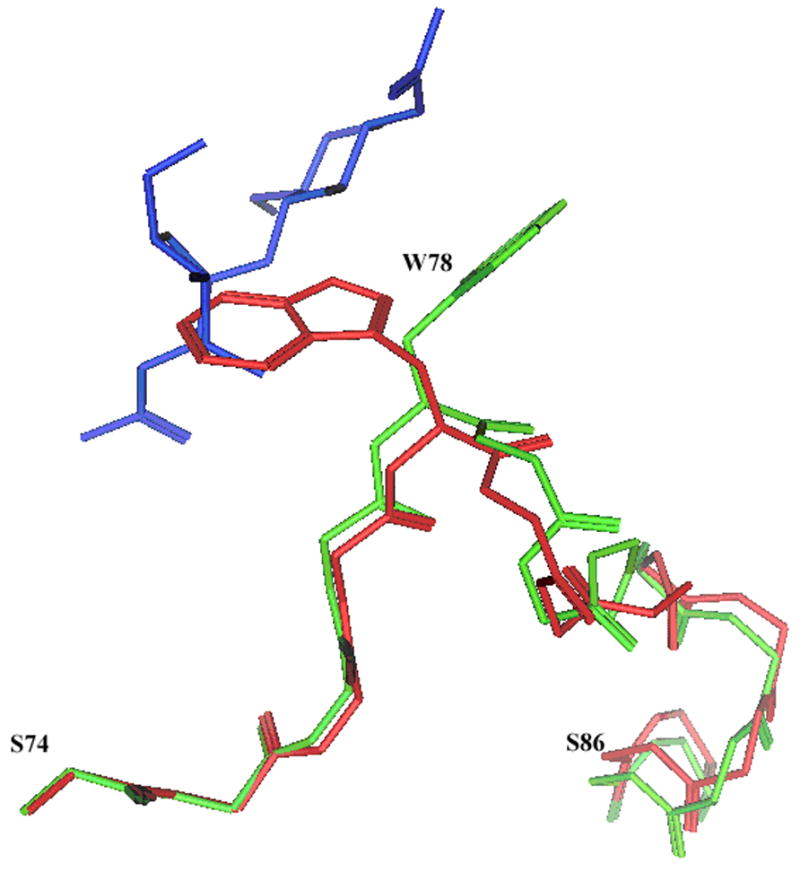
Backbone of the residues 74–86 of MGP40 (red) and equivalent residues 95–107 of HumGP39 (green) are shown. Sidechains of Trp 78 (MGP40) and Trp 99 (HumGP39) are also shown. The −1 +1 sugars found in the HumGP39 structure are shown in blue. The proteins and sugar are depicted in stick configuration.
| MGP40 | (78) WNFGPERF (85) |
| HumGP39 | (99) WNFGSQRF (107) |
The backbone conformation of Trp 78 – Phe 85 is very similar between HumGP39 and MGP40 (Fig. 3). The major difference observed in this loop is at position 82 where a Pro is present in MGP 40 and a Ser in HumGP39. This variation could potentially be the cause of slightly different backbone positioning observed in the two proteins. We also observed that Trp78/99 sidechains are in different positions in the two proteins. The slight difference in the backbone positioning should not contribute to this difference in their sidechains. We observed that the HumGP39 structures, when crystallized without any sugars (PDB code: 1HJX and 1NWR), have the Trp 99 sidechain conformation similar to that of Trp 78 in MGP40. Thus it appears that Trp 78/99 at the +1 position is the “gate” of the binding site. It constricts the binding site in absence of sugar and swings out to open up the binding site in the presence of sugar (Fig. 3).
We believe that there are no potential causes for the lack of observed oligosaccharide binding in MGP40. It appears that the MGP40 in this study was not crystallized in appropriate conditions and hence did not have sugar in the binding site. Careful crystallization and x-ray structure determination should allow sugars in the binding site of MGP40.
Structural Comparison: Binding Site +1
As mentioned above, the Trp at +1 position (Trp78 in MGP40 and Trp 99 in HumGP39) has two different conformations depending on the presence of the sugar. To further investigate the +1 binding site, we superimposed structures from different Family 18 proteins. We observe that the proteins with no sugar bound, MGP40 and two HumGP39 structures (pdb ids: 1LJY, 1NWR, 1HJX) [5,9,27] have the Trp in a different “pinching” orientation as opposed to the “stacking” orientation in structures with the sugar bound (Fig. 4). As seen in the figure the Trp in the “pinching” orientation (red) pinches the −1 binding site, whereas Trp in the stacking configuration (green) stacks with the +1 sugar to provide the favorable hydrophobic interaction. Hence, we propose that the Trp at +1 position functions as the “gate” for this large binding site. Whether the “key” is just the knocking of an oligosaccharide on this “gate” or involves a more sophisticated system of conformational change is yet to be determined. Fusetti et al. point out the rotation of the side chain of the Trp to make this parallel stacking interaction. Furthermore, as mentioned by the authors, this differing orientation of the +1 Trp is not observed in chitotriosidase (pdb: 1LG1 and 1LG2) [7]. Very recently Kumar et. al. also discussed the induce opening of the carbohydrate-binding grooves by large scale conformational changes in Trp78 [28]. Upon further investigation, we found that in Chitinase B (pdb: 1E15 and 1E6N) the +1 Trp has the same orientation in the apo- and holo form. Thus, there appears to be a clear distinction between the chitolectin and chitinase classes of Family 18 proteins; namely the +1 Trp. In chitolectins +1 Trp has distinct pinched and stacked conformations in the apo and the holo forms respectively, whereas in chitinases +1 Trp has the same orientation in the apo and the holo forms.
Fig. 4.
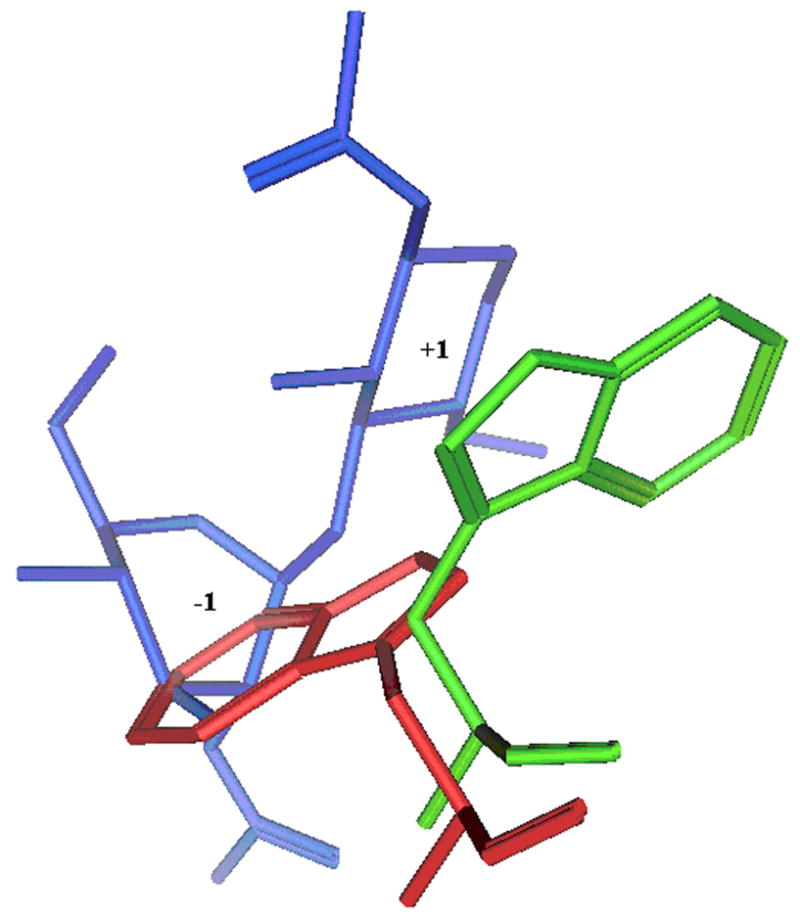
Trp99 of HumGP39 protein along with the binding sugars is shown in stick configuration. Trp99 of HumGP39 from a structure with no sugar bound (PDB code: 1HJX) is shown in red and with sugar bound (PDB code: 1HJW) is shown in green. The bound sugars at −1 and +1 positions are shown in blue. The proteins and sugar are shown in stick configuration
Structural Comparison: Binding Site +3, Conformation of Residues 209–213
Houston et al. proposed that Trp 212 (+3 binding site) and the neighboring residues Arg 213 and His 209 in HumGP39 are in different orientation depending upon the presence of the oligosaccharide in the binding site. They propose that there is a ligand (oligosaccharide) induced conformational change in this region. The conformation of this region is different in their x-ray structures in the presence and absence of sugar. However, upon further investigation, we determined that in one of the four protein chains with no sugar (PDB code: 1HJX – chain B) the conformation of the 209–213 region was similar to one with the sugar bound. Furthermore, His 209 and Arg 213 in MGP40 (no sugar bound) are also in similar position to that of sugar-bound HumGP39. Finally, recently published x-ray structures of GP39 [27] show no difference in the conformation of residues 209–213 in presence or absence of bound oligosaccharides. Thus, it would seem that binding of sugar likely does not influence the conformational change of His 209, Trp 212 and Arg 213 in the HumGP39 structures as proposed by Houston et al.
Comparison of HUMGP39 structures
As mentioned earlier, two groups have independently determined the structure of HumGP39 in the presence and absence of oligosaccharides. The tetrasaccharide and pentasaccharide have their sugars bound respectively in −2 to +2 and −3 to +2 configuration in both studies. Interestingly, hexasaccharide shows −4 to +2 binding in one study [9] and −3 to +3 binding in the second study [27]. W212 makes up the +3 binding subsite, and as described earlier it is similar between the two HumGP39 structures. Based on a) definite binding sites for tetrasaccharide (−2 to +2) and pentasaccharide (−3 to +2) and b) lack of a definite binding site for a hexasaccharide (−3 to +3 versus −4 to +2), it would appear that the energetic surface distal to the −3 and +2 subsites is shallow and not very well-defined. Fusetti et al. [27] also crystallized a disaccharide bound HumGP39. Interestingly, disaccharide appears to bind in the −5 −6 subsites rather than somewhere between −4 and +3 subsites. We further investigated the HumGP39 structures elucidated by Houston et al. We noticed that in tetrasaccharide-bound proteins there were also trisaccharides bound in the −5 −6 −7 subsites. Based on this information, we concur with Fusetti et al. that only larger (than tri) oligosaccharides bind in the traditional (−4 to +3) binding subsites, whereas the shorter oligosaccharides occupy the distal binding site.
Conclusion
In this study, we compare the structures of two Family 18 chitolectins, MGP40 and HumGP39. We propose that MGP40 is capable of binding oligosaccharides contrary to the authors’ conclusion. We also show that the conformation of residues 209–213 is not altered upon ligand binding as proposed by the authors. Based on the two HumGP39 structures, we concur that there are two distinct binding sites; a distal site for trisaccharides and the main site for tetrasaccharides and larger. Most importantly, we propose that the Trp in the +1 position functions as a “gate” to the main binding site that is in the “pinched” conformation when the oligosaccharide is not bound and is in the “stacked” conformation while interacting with the oligosaccharide. This equivalent conserved residue in Sm ChitinaseA and Sm ChitinaseB, both active Family 18 chitinases, never display a pinching conformation, only an open one.
Acknowledgments
This research was partially supported by the National Institutes of Health (Bethesda, MD, U.S.A.; grant no. GM-59471 to J.D.M. and N.N.A.), United States Public Health Service (Washington, DC, U.S.A.) and National Science Foundation EPSCoR (grant no. 91853 to N.N.A.).
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Keyhani NO, Roseman S. Physiological aspects of chitin catabolism in marine bacteria. Biochim Biophys Acta. 1999;1473:108–122. doi: 10.1016/s0304-4165(99)00172-5. [DOI] [PubMed] [Google Scholar]
- 2.Henrissat B, Davies GJ. Glycoside hydrolases and glycosyltransferases. Families, modules, and implications forgenomics. Plant Physiol. 2000;124:1515–1519. doi: 10.1104/pp.124.4.1515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Bakkers J, Kijne JW, Spaink HP. Function of chitin oligosaccharides in plant and animal development. EXS. 1999;87:71–83. doi: 10.1007/978-3-0348-8757-1_5. [DOI] [PubMed] [Google Scholar]
- 4.Bleau G, Massicotte F, Merlen Y, Boisvert C. Mammalian chitinase-like proteins. EXS. 1999;87:211–221. doi: 10.1007/978-3-0348-8757-1_15. [DOI] [PubMed] [Google Scholar]
- 5.Mohanty AK, Singh G, Paramasivam M, Saravanan K, Jabeen T, Sharma S, Yadav S, Kaur P, Kumar P, Srinivasan A, Singh TP. Crystal structure of a novel regulatory 40 kDa mammary gland protein (MGP-40) secreted during involution. J Biol Chem. 2003;278:14451–14460. doi: 10.1074/jbc.M208967200. [DOI] [PubMed] [Google Scholar]
- 6.Bortone K, Monzingo AF, Ernst S, Robertus JD. The structure of an allosamidin complex with the Coccidioides immitis chitinase defines a role for a second acid residue in substrate-assisted mechanism. J Mol Biol. 2002;320:293–302. doi: 10.1016/S0022-2836(02)00444-8. [DOI] [PubMed] [Google Scholar]
- 7.Fusetti F, Moeller HV, Houston D, Rozeboom HJ, Dijkstra BW, Boot RG, Aerts JM, Aalten DMV. Structure of human chiototriosidase. Implications for specific inhibitor design and function of mammalin chitnase-like lectins. J Biol Chem. 2002;277:25537–25544. doi: 10.1074/jbc.M201636200. [DOI] [PubMed] [Google Scholar]
- 8.Hollis T, Monzingo AF, Bortone K, Ernst S, Cox R, Robertus JD. The X-ray structure of a chitinase fromthe pathogenic fungs Coccidioides immitis. Protein Sci. 2000;9:544–551. doi: 10.1110/ps.9.3.544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Houston DR, Anneliese DR, Joanne CK, Aalten DMV. Structure and ligand-induced conformational change of the 39 kD glycoprotein from human articular chondrocytes. J Biol Chem. 2003;278:30206–30212. doi: 10.1074/jbc.M303371200. [DOI] [PubMed] [Google Scholar]
- 10.Houston DR, Eggleston I, Synstad B, Eijsink VG, Aalten DMV. The cyclic dipeptide C1–4[cyclo-(L-Arg-D-Pro)] inhibits family 18 chitinases by structural mimicry of a reaction intermediate. Biochem, J. 2002;368:23–27. doi: 10.1042/BJ20021034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Houston DR, Shiomi K, Arai N, Omura S, Peter MG, Turberg A, Synstad B, Eijsink VGH, Aalten DMFV. High-resolution structures of a chitinase complexed with natural product cyclopentapeptide inhibitors: mimicry of carbohydrate substrate. Proc Natl Acad Sci USA. 2002;99:9127–9132. doi: 10.1073/pnas.132060599. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Matsumoto T, Nonaka T, Hashimoto M, Watanabe T, Mitsui Y. Three-dimensional structure of the catalytic domain of chitinase A1 from Bacillus circulans WL-12 at a very high resolution. Proc Japan Acad. 1999;75:269–274. [Google Scholar]
- 13.Orikoshi H, Baba N, Nakayama S, Kashu H, Miyamoto K, Yasuda M, Inamori Y, Tsujibo H. Molecular analysis of the gene encoding a novel cold-adapted chitinase (ChiB) from a marine bacterium, Alteromonas sp. strain O-7. J Bacteriol. 2003;185:1153–1160. doi: 10.1128/JB.185.4.1153-1160.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Perrakis A, Tews I, Dauter Z, Oppenheim AB, Chet I, Wilson KS, Vorgias CE. Crystal structure of a bacterial chitinase at 2.3Å resolution. Structure. 1994;2:1169–1180. doi: 10.1016/s0969-2126(94)00119-7. [DOI] [PubMed] [Google Scholar]
- 15.Prag G, Papanikolau Y, Tavlas G, Vorgias CE, Petratos K, Oppenheim AB. Structures of chitobiase mutants complexed with the substrate Di-N-acetyl-D-glucosamine: the catalytic role of the conserved acidic pair, aspartate 539 and glutamate 540. J Mol Biol. 2000;300:611–617. doi: 10.1006/jmbi.2000.3906. [DOI] [PubMed] [Google Scholar]
- 16.Rao V, Cui T, Guan C, Roey PV. Mutations of endo-beta-N-acetylglucoseaminidase H active site residue Asp 130 and Glu 132: activities and conformations. Protein Sci. 1999;8:2338–2346. doi: 10.1110/ps.8.11.2338. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Rao V, Guan C, Roey PV. Crystal structure of endo-beta-N-acetylglucoseaminidase H at 1.9 A resolutio: active-site geometry and substrate recognition. Structure. 1995;3:449–457. doi: 10.1016/s0969-2126(01)00178-2. [DOI] [PubMed] [Google Scholar]
- 18.Sun YJ, Chang NC, Hung SI, Chou CC, Hsiao CD. The crystal structure of a novel mammalian lectin, Ym1, suggests a saccharide binding site. J Biol Chem. 2001;276:17507–17514. doi: 10.1074/jbc.M010416200. [DOI] [PubMed] [Google Scholar]
- 19.Terwisscha van Scheltinga AC, Armand S, Kalk KH, Isogai A, Henrissat B, Dijkstra BW. Stereochemistry of chitin hydrolysis by a plant chitinase/lysozyme and X-ray structure of a complex with allosamidin: evidence for substrate assisted catalysis. Biochemistry. 1995;34:15619–15623. doi: 10.1021/bi00048a003. [DOI] [PubMed] [Google Scholar]
- 20.Terwisscha van Scheltinga AC, Kalk KH, Beintema JJ, Dijkstra BW. Crystal structures of hevamine, a plant defence protein with chitinase and lysozyme activity, and its complex with inhibitor. Structure. 1994;2:1181–1189. doi: 10.1016/s0969-2126(94)00120-0. [DOI] [PubMed] [Google Scholar]
- 21.Vaaje-Kolstad G, Vasella A, Peter MG, Netter C, Houstan DR, Westereng B, Synstad B, Eijsink VG, van Aalten DM. Interactions of a family 18 chitinase with the designedinhibitor HM508, and its degradation product,chitobiono-delta-lactone. J Biol Chem. 2004;279:3612–3619. doi: 10.1074/jbc.M310057200. [DOI] [PubMed] [Google Scholar]
- 22.van Aalten DM, Komander D, Synstad B, Gaseidnes S, Peter MG, Eijsink VG. Structural insights into the catalytic mechanism of a family 18 exochitinase. Proc Natl Acad Sci USA. 2001;98:8979–8984. doi: 10.1073/pnas.151103798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Van Roey P, Rao V, Plummer TH, Jr, Tarentino AL. Crystal structure of endo-beta-N-acetylglucosaminidase F1. an alpha/beta-barrel enzyme adapted for a complex substrate. Biochemistry. 1994;33:13989–13996. doi: 10.1021/bi00251a005. [DOI] [PubMed] [Google Scholar]
- 24.Varela PF, LIera AS, Mariuzza RA, Tormo J. Crystal structure of imaginal disc growth factor-2. A member of a new family of growth-promoting glycoproteins from Drosophila melanogaster. J Biol Chem. 2002;277:13229–13236. doi: 10.1074/jbc.M110502200. [DOI] [PubMed] [Google Scholar]
- 25.Waddling CA, Plummer THJ, Tarentino AL, van Roey P. Structural basis for the substratespecificity of endo-beta-N-acetylglucoseaminidase F(3) Biochemistry. 2000;39:7878–7885. doi: 10.1021/bi0001731. [DOI] [PubMed] [Google Scholar]
- 26.Watanabe T, Ishibashi A, Ariga Y, Yashimoto M, Nikaidou N, Sugiyama J, Matsumoto T, Nonaka T. Trp122 and Trp134 on the surface of the catalytic domain are essential for crystalline chitine hydrolysis by Bacillus circulans chitinase A1. FEBS Lett. 2001;494:74–78. doi: 10.1016/s0014-5793(01)02317-1. [DOI] [PubMed] [Google Scholar]
- 27.Fusetti F, Pijning T, Kalk KH, Bos E, Dijkstra BW. Crystal structure and carbohydrate-binding properties of the human cartilage glycoprotein-39. J Biol Chem. 2003;278:37753–37760. doi: 10.1074/jbc.M303137200. [DOI] [PubMed] [Google Scholar]
- 28.Kumar J, Ethayathulla AS, Srivastava DB, Singh N, Sharma S, Kaur P, Srinivasan A, Singh TP. Carbohydrate-binding properties of goat secretory glycoprotein (SPG-40) and its functional implications: structures of the native glycoprotein and its four complexes with chitin-like oligosaccharides. Acta Cryst D. 2007;63:437–46. doi: 10.1107/S0907444907001631. [DOI] [PubMed] [Google Scholar]


