Abstract
Vibrio vulnificus is a ubiquitous inhabitant of the marine coastal environment, and an important pathogen of humans. We characterized a globally distributed sample of environmental isolates from a range of habitats and hosts and compared these with isolates recovered from cases of human infection. Multilocus sequence typing data using six housekeeping genes divided 63 of the 67 isolates into the two main lineages previously noted for this species, and this division was also confirmed using the 16S rRNA and open reading frame VV0401 markers. Lineage I was comprised exclusively of biotype 1 isolates, whereas lineage II contained biotype 1 and all biotype 2 isolates. Four isolates did not cluster within either lineage: two biotype 3 and two biotype 1 isolates. The proportion of isolates recovered from a clinical setting was noted to be higher in lineage I than in lineage II. Lineage I isolates were also associated with a 33-kb genomic island (region XII), one of three regions identified by genome comparisons as unique to the species. Region XII contained an arylsulfatase gene cluster, a sulfate reduction system, two chondroitinase genes, and an oligopeptide ABC transport system, all of which are absent from the majority of lineage II isolates. Arylsulfatases and the sulfate reduction system, along with performing a scavenging role, have been hypothesized to play a role in pathogenic processes in other bacteria. Our data suggest that lineage I may have a higher pathogenic potential and that region XII, along with other regions, may give isolates a selective advantage either in the human host or in the aquatic environment or both.
Vibrio vulnificus is a gram-negative aquatic bacterium that was first isolated by the U.S. Center for Disease Control and Prevention (CDC) in 1964 and recognized as a distinct species in 1976 (70). V. vulnificus is an obligate halophile, found in estuarine and marine coastal environments throughout the world, particularly in the warmer summer months (35, 36, 54, 73). V. vulnificus is found in association with zooplankton, crabs, and various filter feeders such as oysters and mussels (21, 27). Oysters harvested from the Gulf of Mexico during the summer months contain high numbers of this pathogen (28, 54). As well as comprising part of the normal microflora of coastal waters, V. vulnificus is a highly invasive pathogen of both fish and humans (23, 54). V. vulnificus causes significant economic losses in the eel aquaculture industry in Asia and Europe (4). V. vulnificus infections in humans are characterized by primary septicemia, wound infections and, to a lesser extent, gastrointestinal illness with mortality rates greater than 50% in susceptible individuals (42, 54). V. vulnificus is the leading cause of death associated with the consumption of contaminated seafood in the United States. Between 1989 and 2000, 274 cases involving oyster ingestion occurred in the United States, 52% of which were fatal (62). The infectious dose can be as low as 10 cells in a mouse model (68). Several putative virulence factors have been identified in V. vulnificus, in particular the capsule polysaccharide, an iron siderophore, and an rtxA (repeats in structural toxins) gene (VVA1030) which was recently shown to be essential for cytotoxicity (40, 43). However, these are found in both clinical and environmental isolates and no virulence determinants specifically associated with clinical isolates have been identified (23).
Three biotypes are recognized among V. vulnificus isolates based on phenotypic characteristics and host range criteria (30). Biotype 1 strains are indole positive, serologically heterogeneous, and pathogenic to humans (54). Biotype 2 strains are indole negative, have a homogeneous lipopolysaccharide O antigen named serovar E and are primarily considered pathogens of eels (vibriosis) and secondary pathogens of humans (6, 8). Biotype 3 strains are pathogenic to humans and appear to be geographically confined to Israel, where they caused an outbreak of systemic V. vulnificus infection among Israeli fish market workers in 1996 (9, 12). Atypical V. vulnificus isolates have been documented that are pathogenic for eels but are indole positive and have a different genotype than biotype 2 indole-negative, eel-pathogenic strains (7, 31). A reclassification of all biotype 2 and atypical strains to serovar E, a homogeneous lipopolysaccharide O serogroup, has been put forward (7).
A number of different molecular epidemiological tools have been used to study the genetic diversity of V. vulnificus, and many of these studies have focused on identifying genetic differences between clinical and environmental strains (5, 18, 32, 74). Buchrieser and coworkers used clamped homogeneous electric field gel electrophoresis to characterize V. vulnificus isolates from oysters (18). Pulsed-field gel electrophoresis and ribotyping have been used to assay diversity among clinical and environmental strains (18). Wong and colleagues examined isolates from Taiwan and the United States using pulsed-field gel electrophoresis and noted that clinical isolates from the United States were more homogeneous than isolates from Taiwan (79). Arias and colleagues noted significant diversity using ribotyping and randomly amplified polymorphic DNA PCR in 132 environmental, clinical, and eel-pathogenic V. vulnificus isolates recovered from different geographic regions (5). By using a typing method based on the differences in amplification and restriction analyses of the 16S rRNA sequence among clinical and environmental isolates, Nilsson and coworkers found most environmental isolates had a distinct A genotype and most clinical isolates had a B genotype, suggesting that the 16S rRNA locus might be a good indicator of virulence (52). Warner and Oliver (78) applied a randomly amplified polymorphic DNA PCR procedure to distinguish V. vulnificus from other Vibrio species as well as to compare clinical and environmental isolates. These authors identified a unique 178- to 200-bp segment present in all of the 39 clinical isolates but in only 3 of the 30 environmental isolates tested. Based on that study, they were able to develop a PCR assay that divides strains into two types, a clinical type (type C) and an environmental type (type E), according to their origins (62). Chatzidaki-Livanis and coworkers used a typing method based on differences of the group 1 capsule polysaccharide (CPS) operon that was previously reported and found that the A/E type associated with the CPS allele 2 and environmental isolates, whereas the B/C type associated with the CPS allele 1 and clinical isolates (19). Using repetitive extragenic palindromic PCR (rep-PCR), the study also found that some clinical isolates were more closely related to environmental isolates (mostly found in group III, profile A/E, and CPS allele 2) than to other clinical isolates (groups I, IV, and VII, profile B/C, and CPS allele 1).
Although molecular typing methods based on single gene loci may be used to distinguish strains, determining the precise evolutionary relationships between strains requires data from multiple gene loci. Gutacker et al. examined the relationships among 62 isolates with different geographic and host origins using multilocus enzyme electrophoresis and sequence analysis of two housekeeping genes and identified two distinct major clusters of isolates (24). Interestingly, they found that biotype 3 isolates grouped together but within different lineages depending on the method used (24). Single locus sequence analysis of gyrB sequences of V. vulnificus from oysters in India showed the presence of two distinct clonal groups (56). Bisharat and colleagues used multilocus sequence typing (MLST) analysis of 10 housekeeping genes to investigate the evolutionary origins of 62 biotype 3 isolates from a 1996 V. vulnificus outbreak in Israel (11). They showed that biotype 3 strains were genetically identical and distinct from the biotype 1 and biotype 2 isolates. From their analysis, they proposed that V. vulnificus biotype 3 contains a mosaic genome that evolved by the hybridization of the genomes from two distinct and independent populations (11). They also proposed that this hybridization event has led to heightened virulence compared to that of the existing biotype 1 strains present in Israel (11). In a more recent report, this group has suggested that higher water temperature might have impacted the ecology of the area and caused the emergence of the biotype 3 strains, as an effect of global climate change (57).
In this study, we examined the population genetic structure of V. vulnificus isolates based on MLST analysis of six housekeeping genes to determine whether phylogenetically closely related isolates share pathogenic potential. MLST analysis of 67 isolates with a worldwide distribution and from a range of hosts and habitats identified two distinct evolutionary lineages. We examined the genetic variation within each lineage and found that lineage I was genetically more diverse than lineage II. Lineage I was comprised of isolates that typed exclusively as B/C types and lineage II was comprised of isolates that typed as A/E by the molecular typing methods of Nilsson et al. (52) and Rosche et al. (62). Additionally, we identified a 33-kb region, termed region XII, that was present on chromosome II of both of the sequenced clinical isolates but absent from all other sequenced Vibrio species. We examined the occurrence and distribution of this region among our collection of isolates and found that it was present predominantly among isolates within lineage I. These data suggest that isolates of lineage I may have a higher pathogenic potential than those of lineage II, and we speculate that isolates that encode region XII, in addition to other regions, may have increased fitness in the aquatic environment or human host or both.
MATERIALS AND METHODS
Bacterial strains.
A total of 67 V. vulnificus isolates were examined in this study (Table 1). The 67 isolates were temporally (1980 to 2005) and geographically (Asia, Europe, and North America) widespread. Of the 67 isolates examined, 27 were recovered from clinical sources (wound infections and blood) and 40 from environmental sources (clam, oyster, mussel, eel, fish, seawater, and sea sediment). The collection contained isolates encompassing all three biotype groups identified in V. vulnificus. All V. vulnificus strains were grown in Luria-Bertani (LB) broth supplemented with 2% NaCl and stored at −70°C in LB broth with 20% (vol/vol) glycerol.
TABLE 1.
Strains used in this study
| Strain name | Source | Country of isolation | Yr of isolation | Biotype | Lineage | Genotypes |
|---|---|---|---|---|---|---|
| YJ002 | Clinical | Taiwan | 1993 | 1 | I | B, C |
| JJ068 | Clinical | Taiwan | 1993 | 1 | I | B, C |
| JJ072 | Clinical | Taiwan | 1993 | 1 | I | B, C |
| JJ067 | Clinical | Taiwan | 1993 | 1 | I | B, C |
| YJ016 | Clinical | Taiwan | 1993 | 1 | I | B, C |
| L-180 | Clinical | Japan | 1980 | 1 | I | B, C |
| N-87 | Clinical | Japan | 1987 | 1 | I | B, C |
| KH-03 | Clinical | Japan | 2003 | 1 | I | B, C |
| CDC9038-96 | Clinical | United States | 1996 | 1 | I | B, C |
| CDC9062-96 | Clinical | United States | 1996 | 1 | I | B, C |
| CDC9005-97 | Clinical | United States | 1997 | 1 | I | B, C |
| K2637 | Clinical | United States | 2005 | NKa | I | B, C |
| K2667 | Clinical | United States | 2005 | NK | I | B, C |
| MO6-24/O | Clinical | United States | NK | 1 | I | B, C |
| MO6 | Clinical | United States | NK | 1 | I | B, C |
| 85A667/O | Clinical | United States | NK | NK | I | B, C |
| C7184 | Clinical | United States | NK | 1 | I | B, C |
| SPRC 10143 | Clinical | United States | NK | 1 | I | B, C |
| CMCP6 | Clinical | NK | NK | 1 | I | B, C |
| LSU 1866 | Clinical | NK | NK | 1 | I | B, C |
| 6353/O | Clinical | United States | NK | NK | II | A, E |
| CDC9030-95 | Clinical | United States | 1995 | 1 | II | A, E |
| K2719 | Clinical | United States | 2005 | NK | II | A, E |
| IFVv8 | Clinical | United States | NK | NK | II | A, E |
| 313-98 | Clinical | Israel | NK | 3 | None | A, E |
| 11028 | Clinical | Israel | NK | 3 | None | A, E |
| CIP 81.90 | Clinical | France | 1980 | NK | II | A, E |
| MLT 362 | Seawater | United States | 1991 | 1 | I | B, C |
| MLT 364 | Seawater | United States | 1991 | 1 | I | B, C |
| SPRC 10215 | Seawater | United States | NK | NK | I | B, C |
| MLT 406 | Seawater | United States | 1991 | NK | II | A, E |
| CG62 | Seawater | Taiwan | 1993 | 1 | I | B, C |
| CG63 | Seawater | Taiwan | 1993 | 1 | I | B, C |
| CG123 | Seawater | Taiwan | 1993 | 1 | I | B, C |
| 300-1C1 | Seawater | NK | NK | NK | II | A, E |
| IFVv18 | Seawater | France | NK | NK | None | A, E |
| Env 1 | Environmental | United States | NK | 1 | II | A, E |
| MLT 365 | Environmental | United States | NK | NK | I | B, C |
| UNCC 1015 | Environmental | United States | 1981 | NK | I | B, C |
| UNCC 913 | Environmental | United States | 1981 | NK | I | B, C |
| 345/O | Environmental | United States | NK | NK | I | B, C |
| L-49 | Environmental | Japan | 1988 | 2 | II | A, E |
| 96-9-114s | Sediment | Denmark | NK | 1 | II | A, E |
| 99-779-DP-D2 | Oyster | United States | 1999 | 1 | II | A, E |
| 98-640-DP-B9 | Oyster | United States | 1998 | 1 | II | A, E |
| JY 1701 | Oyster | United States | 1999 | NK | II | A, E |
| JY 1305 | Oyster | United States | 1999 | NK | II | A, E |
| 99-509-DP-A6 | Oyster | United States | 1999 | 1 | II | A, E |
| 99-540-DP-B6 | Oyster | United States | 1998 | 1 | II | A, E |
| 99-609-DP-A4 | Oyster | United States | 1999 | 1 | II | A, E |
| SS108A-3A | Oyster | NK | NK | NK | II | A, E |
| CG27 | Oyster | Taiwan | 1993 | 1 | I | B, C |
| b60 | Oyster | India | NK | NK | I | B, C |
| b122 | Oyster | India | NK | NK | I | B, C |
| M79 | Eel | Spain | NK | 2 | II | A, E |
| E86 | Eel | Spain | 1990 | 2 | II | A, E |
| 90-2-11 | Eel | Norway | NK | 2 | II | A, E |
| ATCC 33149 | Eel | Japan | NK | 2 | II | A, E |
| NCIMB 2137 | Eel | Japan | NK | 2 | II | A, E |
| NCIMB 2136 | Eel | Japan | NK | 2 | II | A, E |
| IFVv10 | Mussel | France | NK | NK | II | A, E |
| IFVv11 | Mussel | France | NK | NK | II | A, E |
| 72M4 | Clam | India | NK | NK | I | B, C |
| 79M4 | Clam | India | NK | NK | I | B, C |
| 76M3 | Fish | India | NK | NK | I | B, C |
| 80M4 | Fish | India | NK | NK | I | B, C |
| G-83 | Fish | Korea | NK | 1 | None | A, C |
NK, not known.
Molecular analysis.
Chromosomal DNA was extracted from each of the V. vulnificus isolates using a genome DNA isolation kit from Bio 101. Briefly, a single colony of each isolate was inoculated into 3 ml LB broth and incubated overnight at 37°C with shaking at 150 rpm. The bacterial cells were pelleted at 3,000 rpm for 5 min, the supernatant was discarded, and the pellet was brought to a final volume of 1.85 ml in cell suspension solution. The cells were lysed and treated with RNase and protease. DNA was extracted with ethanol and resuspended in Tris-EDTA buffer (10 mM, pH 7.5).
PCR primers to amplify six chromosomal housekeeping genes, gyrB (open reading frame [ORF] VV0014), mdh (VV0467), rpoD (VV0561), groEL1 (VV3106), pyrC (VVA0407), and groEL2 (VVA1659), were designed based on the sequence of V. vulnificus YJ016 (20) (Table 2). The six genes encoded a range of proteins dispersed on chromosomes I and II: DNA gyrase subunit B (gyrB), malate dehydrogenase (mdh), RNA polymerase sigma factor (rpoD), and a chaperonin (groEL1) from chromosome I and a chaperonin (groEL2) and dihydroorotase (pyrC) from chromosome II. Gene fragments for each of the genes were amplified from chromosomal DNA isolated from all 67 V. vulnificus strains. PCRs were performed in a 50-μl reaction mixture as follows: an initial denaturation step at 96°C for 3 min, followed by 30 cycles of denaturation at 94°C for 30 s, 30 s of primer annealing at the temperatures shown in Table 2, and 1 min of primer extension at 72°C for each kilobase of product. PCR products were purified using a Jetquick PCR purification kit (GENOMED) or a QIAquick PCR purification kit (QIAGEN, Hilden, Germany) in accordance with the manufacturers’ instructions. After purification, an aliquot of 35 μl was used as a sequence template. The gyrB, mdh, rpoD, groEL1, pyrC, and groEL2 sequences were determined by MWG-Biotech based on the dye deoxy terminator method.
TABLE 2.
Primers designed for this study
| Primer target and designation | Sequence (5′-3′) | Ta (°C)a | Expected product (bp) |
|---|---|---|---|
| Housekeeping genes | |||
| VVgyrBF | CTATCGTCATGGTGTGC | 48 | 1,805 |
| VVgyrBR | TCTCACCCAAACCTTTG | ||
| VVmdhF | TGTTATTGGTGCCGCTG | 55 | 918 |
| VVmdhR | TACTTTACGAAGTCCACGC | ||
| VVrpoDF | CGTTTCGATCATGTGTACCG | 60 | 1,340 |
| VVrpoDR | ACGCCGAAGTAAATGACCAC | ||
| VVGroEL1bF | GCGTGGTATCGACAAAGCTG | 55.9 | 702 |
| VVGroEL1bR | TGACGAATTTGAGCAACACG | ||
| VVpyrC | AATGGCATTGAAGCAGGAAC | 60 | 858 |
| VVpyrC | GGACAGTGTTTGGCATGATG | ||
| VVGro2F | GACGCACGTCAAAAGATGCTG | 59 | 1,542 |
| VVGro2R | GTGATCGGTGATCATCGCTTC | ||
| Region XII | |||
| VVA1612F | ACC CTG ATC GTT GGC TAC TC | 57 | 1,284 |
| VVA1612R | AAG TGG TGC GTC TTT GAG CAG C | ||
| VVA1612F | ACC CTG ATC GTT GGC TAC TC | 61 | 1,305 |
| VVA1613R | GGA GCG GTG TGA TGG TGT TG | ||
| VVA1616F | ACA TAG TCA GAA GCG GAC G | 56.1 | 2,161 |
| VVA1618R | TTA GGC ACT GGA AAT GGG | ||
| VVA1618F | AAG GTA ATG CTC CAG TCG GG | 57 | 4,240 |
| VVA1623R | TGT GCT TCC TCT TTG TTG C | ||
| VVA1625F | CGG TCT GTG GTT TAT CG | 47 | 1,822 |
| VVA1625R | TCG TTT CCA GTC GTC AC | ||
| VVA1625bF | TCG TAG AAC TGT GGG TAA GG | 57.3 | 3,979 |
| VVA1627R | CAT AAA CTG GCG TGG TAC AG | ||
| VVA1630F | AAG TAA ATC CAG TGG CTA GG | 54.9 | 2,799 |
| VVA1632R | GGA AGG GCA AAT CCT TTC | ||
| VVA1634F | TGA CAC CCA ACC TAG ACC AC | 55 | 1,364 |
| VVA1634R | ATT GAT GCC AAC CTG AG | ||
| VVA1636F | TGT CCA CGA CTT GAA CAC G | 56 | 1,547 |
| VVA1637R | AAC ATC AAC CAG CGA GTC GAA C | ||
| VVA1637F | GCT TAT TGA CCA ACG TG | 52 | 1,631 |
| VVA1639R | CGG TTG AAA CTG ACG AG | ||
| VVA1612bF | TGT GGA GAG CGG CAA GAT CAA G | 61 | 1,200 |
| VVA1637R | AAC ATC AAC CAG CGA GTC GAA C |
Ta, annealing temperature.
Phylogenetic analysis.
The multiple sequence alignment program ClustalW was used to align nucleotide sequences for each gene (75). Nucleotide diversity was calculated as the average pairwise nucleotide difference per site. Rates of synonymous substitutions/synonymous site (KS) and nonsynonymous substitutions/nonsynonymous site (KN) were calculated by the methods of Nei and colleagues (50, 51). The substitution ratio was calculated, which determines the type and level of selection acting on genes. Purifying selection should result in KN/KS ratios that are less than 1.0, and diversifying selection should result in ratios higher than 1.0. Total nucleotide diversity (π) was measured for each gene among all strains, within strains from clinical sources and environmental sources, and for each gene within each of the two major lineages identified. To compare the evolutionary relationships among the V. vulnificus isolates, each gene sequence was used to reconstruct phylogenetic trees. Additionally, the sequences were concatenated, and sequence types (ST) were analyzed for all genes. Phylogenetic trees were constructed with MEGA 3.1 (39) by using the neighbor-joining method and Kimura two-parameter distance for all substitutions, and the inferred phylogenies were tested with 1,000 bootstrap replications. Kimura two-parameter distance takes into consideration that the rate of transitional nucleotide substitution (A↔ G, T↔ C) is often higher than the one found for transversional substitutions (A↔ T, A↔ C, T↔ G, C↔ G).
Tests for recombination.
Four statistical tests to identify homologous recombination within genes were used: Stephen's test, Sawyer's test, MaxChi test, and the PHI test (17, 63, 66, 69). Stephen's test detects significantly nonrandom clustering of polymorphic nucleotide sites, as a footprint of intragenic recombination (69). We used an implementation of this method developed by Thomas S. Whittam (http://www.shigatox.net/cgi-bin/stec/programs). Sawyer's test also examines intragenic recombination by considering only the distribution of synonymous sites in order to distinguish regions of sequence similarity due to recombination from regions of sequence similarity due to selection (63). This test is based on an analysis of whether some region(s) of a pair of sequences has more consecutive identical silent polymorphic sites in common than would be expected by chance and was applied using GENECONV 1.81 (http://www.math.wustl.edu/∼sawyer/geneconv/index.html). The maximum chi-square test (MaxChi) (66) was applied to the data set using MaxChi 4_2.c version 1.00 (http://www.lifesci.sussex.ac.uk/CSE/test/maxchi.php). This test consists of a statistical analysis for clustering of polymorphic sites and identifies the ends of segments of a mosaic allele. We also applied the PHI test (ϕw) (or pairwise homoplasy index), implemented by the SplitsTree 4 version 4.6 program (33), which distinguishes recurrent mutations (or homoplasies) from recombination in generating genotypic diversity.
Group-specific PCR.
The collection of 67 isolates was typed according to two methods (52, 62). Both methods were developed to differentiate between clinical and environmental isolates and are based on sequence differences found in chromosomal genes. The method of Nilsson and colleagues targets the 16S rRNA gene and separates the isolates into genotypes A and B, corresponding to environmental and clinical isolates, respectively (52). Universal primers (UFUL/URUL) were used to amplify the 16S rRNA gene (53), and PCR products were purified using a Jetquick PCR purification kit (GENOMED) in accordance with the manufacturer's instructions. After purification, an aliquot of 35 μl was used as a template. Restriction digestion was performed in a 50-μl reaction mix with the restriction enzyme AluI (New England Biolabs) for 2.5 h. Restriction products were run in a 2% low-melting-point agarose (Promega). Two patterns emerged: type A strains produced products of 248, 140, 62, and 42 bp, while type B strains produced products of 248, 202, and 42 bp. Rosche et al. targeted a hypothetical protein encoded by VV0401 to divide isolates into clinical C-type or environmental E-type strains. Primer pairs targeting the sequence differences at the VV0401 locus were used (62). Primer pair P1/P3 identified C-type strains and primer pair P2/P3 identified E-type strains.
Genome comparisons.
Initial interspecies whole-genome comparisons of V. vulnificus YJ016 with Vibrio parahaemolyticus RIMD2210633, Vibrio cholerae N16961, and Vibrio fischeri ES114 were generated using WebACT, and this analysis identified a number of chromosomal regions unique to V. vulnificus YJ016 (1). A similar analysis was performed with V. vulnificus strain CMCP6. Those regions unique to both V. vulnificus strains YJ016 and CMCP6 but absent from the other sequenced Vibrio species were further analyzed for aberrant base sequence composition. Compositional bias of dinucleotide frequency and the GC percentage of genomic regions unique to V. vulnificus were measured using deltarho, a web-based application which calculates the genomic dissimilarity values δ* (the average dinucleotide relative abundance difference) between input sequences (genomic regions) and the V. vulnificus genome sequence (http://deltarho.amc.uva.nl) (76). Three regions were identified on chromosome II with aberrant base composition: ORFs VVA0080 to VVA0186, VVA0301 to VVA0336, and VVA1613 to VVA1636. These regions were further analyzed for similarities to sequences in the databases using the BLAST algorithm (3).
PCR analysis of genomic region XII.
To determine whether region XII was present in all V. vulnificus isolates, PCR assays were performed on our collection of isolates. Primer pairs were designed to target regions within genomic region XII, flanking this region, or targeting the chromosomal insertion sites (Table 2). PCR was performed in a 25-μl reaction mixture as follows: an initial denaturation step at 96°C for 3 min followed by 30 cycles of denaturation at 94°C for 30 s, 30 s of primer pair annealing at the respective temperatures (Table 2), and 1 min of primer extension at 72°C per kilobase of product.
Southern hybridization.
Southern hybridization was carried out with DNA probes, generated by PCR from the reference strain YJ016, to confirm PCR negative results for region XII. Briefly, DNA from each isolate was digested with restriction enzymes (New England Biolabs), and the fragments were separated by electrophoresis in 0.6% Tris-acetate-EDTA agarose. DNA fragments were transferred to nylon membranes under positive pressure (Stratagene Posiblotter; Stratagene, La Jolla, CA). Probe DNAs were labeled using the ECL direct nucleic labeling system (Amersham Pharmacia Biotech), and positive hybridization was detected by the ECL chemiluminescent substrate.
Nucleotide sequence accession numbers.
The sequences of mdh, rpoD, gyrB, groEL1, groEL2, and pyrC were submitted to GenBank and given the following accession numbers: groEL1, EF642498 to EF642562; groEL2, EF642563 to EF642627; gyrB, EF642628 to EF642692; mdh, EF642693 to EF642757; pyrC, EF642758 to EF642822; and rpoD, EF642823 to EF642887.
RESULTS AND DISCUSSION
Nucleotide polymorphism and diversity.
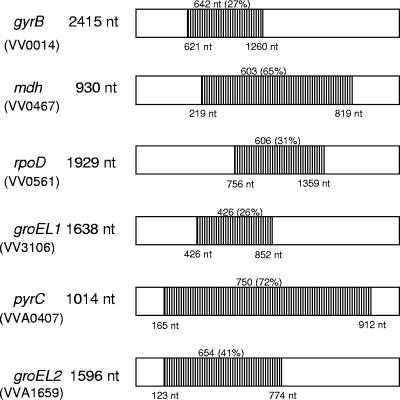
We examined a collection of V. vulnificus isolates with worldwide distribution recovered over a 25-year period from a wide variety of hosts and habitats (Table 1). Of the 67 V. vulnificus isolates analyzed, 27 were recovered from clinical settings and 40 from environmental sources. We sequenced gene fragments from four housekeeping genes, gyrB, mdh, rpoD, and groEL1, encoded on chromosome I and two housekeeping genes, pyrC and groEL2, encoded on chromosome II (Fig. 1). Comparison of GroEL1 with GroEL2 from V. vulnificus YJ016 showed 78% similarity.
FIG. 1.
Housekeeping genes used in this study. The gene fragments sequenced in this study are shown in gray.
The sequenced housekeeping gene fragments examined in this study ranged in size from 426 bp for groEL1 to 750 bp for pyrC (Fig. 1). The total nucleotide diversity among our sequenced V. vulnificus isolates was comparable to MLST data from a number of highly homogeneous pathogenic bacteria (2, 22, 67). We examined nucleotide polymorphism among the six housekeeping genes to determine differences in variation among genes and between chromosomes (Table 3). The number of polymorphic nucleotide sites for genes on chromosome I ranged from 33 for groEL1 (resulting in two amino acid replacement sites) to 54 polymorphic sites for the rpoD gene (resulting in seven amino acid replacements) (Table 3). Both pyrC and groEL2 examined on chromosome II had higher numbers of polymorphic sites compared to genes on chromosome I: 89 and 68 sites resulting in 12 and 10 amino acid replacement sites, respectively (Table 3). This is consistent with previous reports that show that chromosome I is more conserved than chromosome II (26, 61). The ratio KS/KN is an indication of the selective pressures acting on each of the housekeeping genes. We found that this ratio was far greater than 1 due to a predominance of synonymous change over nonsynonymous change. This indicates purifying selection acting against amino acid changes, as would be expected for functional conserved genes such as those involved in housekeeping functions. These results are broadly consistent with previous genetic analyses of MLST data (16, 22, 45, 58). The central region of rpoD is more variable than other parts of this gene. This region is also relatively variable between species, and this is thought to reflect functional redundancy (44).
TABLE 3.
Nucleotide polymorphism and diversity among six housekeeping genesa
| Gene | No. of alleles | No. of alleles with two or more isolates | Polymorphic sites
|
Ps (nt) | KS | KN | π | |
|---|---|---|---|---|---|---|---|---|
| nt | aa | |||||||
| gyrB | 33 | 10 | 42 | 1 | 34 | 8.0 × 10−2 ± 1.0 × 10−3 | 1.0 × 10−3 ± 1.0 × 10−3 | 1.8 × 10−2 ± 4 × 10−3 |
| mdh | 30 | 16 | 43 | 6 | 34 | 6.0 × 10−2 ± 1.2 × 10−2 | 1.6 × 10−3 ± 1.1 × 10−3 | 1.6 × 10−2 ± 3 × 10−3 |
| rpoD | 22 | 12 | 54 | 7 | 41 | 1.1 × 10−1 ± 2.3 × 10−2 | 6.4 × 10−3 ± 2.4 × 10−3 | 2.8 × 10−2 ± 5 × 10−3 |
| groEL1 | 27 | 10 | 33 | 2 | 20 | 7.0 × 10−2 ± 1.7 × 10−2 | 2.0 × 10−4 ± 1.0 × 10−4 | 1.6 × 10−2 ± 4 × 10−3 |
| pyrC | 42 | 9 | 89 | 12 | 73 | 1.4 × 10−1 ± 1.9 × 10−2 | 2.2 × 10−3 ± 7.0 × 10−4 | 3.2 × 10−2 ± 4 × 10−3 |
| groEL2 | 35 | 13 | 68 | 10 | 55 | 1.3 × 10−1 ± 2.1 × 10−2 | 3.6 × 10−3 ± 1.6 × 10−3 | 3.0 × 10−2 ± 4 × 10−3 |
| All | 54 | 8 | 329 | 38 | 257 | 1.0 × 10−1 ± 7.0 × 10−3 | 2.6 × 10−3 ± 5.0 × 10−4 | 2.4 × 10−2 ± 2 × 10−3 |
| Chr. I | 53 | 9 | 172 | 16 | 129 | 8.0 × 10−2 ± 9.0 × 10−3 | 2.0 × 10−3 ± 8.0 × 10−4 | 2.0 × 10−2 ± 2 × 10−3 |
| Chr. II | 48 | 11 | 157 | 22 | 128 | 1.3 × 10−1 ± 1.3 × 10−2 | 3.0 × 10−3 ± 8.0 × 10−4 | 3.1 × 10−2 ± 4 × 10−3 |
Chr., chromosome; nt, nucleotides; aa, amino acids; Ps, parsimonious sites.
MLST analysis.
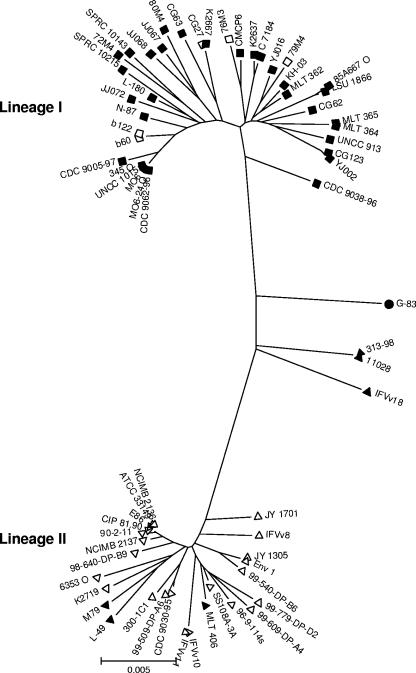
The number of sequence alleles for each gene sequenced among the 67 strains ranged from 22 for rpoD on chromosome I to 42 for pyrC on chromosome II (Table 3). Based on all six genes, 54 ST were identified among the 67 isolates. We constructed a neighbor-joining tree for all sites from the concatenated sequence of all six genes based on the Kimura two-parameter distance method (3,681 bp, 257 informative sites; Fig. 2). From the phylogenetic tree, two distinct evolutionary lineages can be identified, designated lineage I and II (Fig. 2). Both lineages have a star-shaped phylogeny, which is consistent with rapid clonal expansion after these lineages split from their most-recent common ancestor, and/or a history of frequent recombination.
FIG. 2.
Unrooted radial neighbor-joining tree (Kimura two-parameter distance; all substitution sites; bootstrap, 1,000; variable sites, 329/3,681) constructed by concatenating all six housekeeping genes. ▪, B/C type isolates containing region XII; □, B/C type isolates and absence of region XII; ▴, A/E type isolates containing region XII; ▵, A/E type isolates and absence of region XII; •, A/C isolate containing region XII.
The majority of isolates recovered from clinical sources are present in lineage I (n = 20), whereas isolates recovered from environmental sources are present within both lineage I (n = 17) and lineage II (n = 21). Lineage II contained only five isolates from clinical sources. One of these strains, K2719, was isolated from a wound infection from a victim of Hurricane Katrina and clustered with another clinical isolate from the United States (Fig. 2). Of the other two isolates recovered from victims of Hurricane Katrina in 2005 that were examined, strain K2667 shared identity at all six loci to strain CG27, an isolate from oysters from Taiwan, with both clustering in lineage I (Fig. 2). Strain K2637 clustered in lineage I with a clinical isolate from the United States. Within lineage II, all five isolates recovered from diseased eels clustered tightly together with a strain recovered from oysters in the United States (Fig. 2).
Four strains branched with neither lineage I nor II; strain G-83 was recovered from fish in Korea, strain IFVv18 was isolated from seawater in France in 2000, and two identical biotype 3 isolates were recovered in Israel, strains 11028 and 313-98 (Fig. 2). Bisharat and colleagues applied multilocus sequence analysis to a large collection of biotype 3 isolates and demonstrated a highly clonal population (10, 11). They have proposed that biotype 3 strains are a hybrid, virulent organism that acquired genes from two distinct and independent populations in a relatively recent genome hybridization event (11). However, it is difficult to explain how the complex mosaic structure observed in these strains could have been generated by a single event or why a reassortment of housekeeping genes could result in heightened virulence. By contrast, we favor the widely accepted mechanism found in many enteric pathogens that virulence is conferred upon particular strains by the acquisition of virulence factors that are either phage encoded or island encoded (14, 15, 25).
We next examined nucleotide diversity within each of the lineages and found lineage I (0.017) was more diverse than lineage II (0.008) (Table 4), but there was no clear difference in diversity when clinical and environmental isolates were directly compared (Table 5). Comparing nucleotide diversity with respect to geographical source revealed a striking homogeneity among the six isolates recovered from eels in Japan (n = 3), Spain (n = 2), and Norway (n = 1) (Table 5). This is similar to the results of Gutacker and colleagues, who identified a distinct eel-pathogenic clone when they examined diversity at two other housekeeping genes (recA and glnA) (24). In contrast, Hoi and colleagues found that V. vulnificus isolates recovered from diseased eels in Denmark were heterogeneous when analyzed by O serovar, capsule types, ribotyping, phage typing, and plasmid profiling (31).
TABLE 4.
Comparison of the lineages in the concatenated treea
| Lineage | No. of isolates | No. of alleles | Polymorphic sites
|
Ps (nt) | KS | KN | π | |
|---|---|---|---|---|---|---|---|---|
| nt | aa | |||||||
| I | 37 | 30 | 160 | 17 | 117 | 4 × 10−2 ± 5 × 10−3 | 7.0 × 10−4 ± 2 × 10−4 | 1.7 × 10−2 ± 1 × 10−3 |
| II | 26 | 21 | 128 | 18 | 73 | 3 × 10−2 ± 3 × 10−3 | 1.2 × 10−3 ± 3 × 10−4 | 8.0 × 10−3 ± 1 × 10−3 |
nt, nucleotides; aa, amino acids; Ps, parsimonious sites.
TABLE 5.
Comparison of nucleotide diversity among V. vulnificus isolates from clinical and environmental sources for the concatenated sequence
| Clusters | No. of isolates | No. of alleles | Polymorphic sitesa
|
π | |
|---|---|---|---|---|---|
| nt | aa | ||||
| Clinical isolates | 27 | 22 | 260 | 27 | 2.1 × 10−2 ± 1.3 × 10−3 |
| Environmental isolates | 40 | 33 | 294 | 31 | 2.4 × 10−2 ± 1.6 × 10−3 |
| Oyster, clam, mussel | 13 | 11 | 208 | 22 | 2.3 × 10−2 ± 6.0 × 10−4 |
| Seawater, sediment, other | 16 | 14 | 241 | 20 | 2.4 × 10−2 ± 2.0 × 10−3 |
| Fish (including eel) | 11 | 8 | 206 | 17 | 2.0 × 10−2 ± 1.4 × 10−3 |
| Eel only | 6 | 4 | 34 | 2 | 3.0 × 10−3 ± 5.0 × 10−4 |
nt, nucleotides; aa, amino acids.
Of the six genes sequenced, the rpoD gene is the most discriminatory between the two lineages, with a coefficient of differentiation of 0.839, but this division was broadly resolved for each of the genes examined (data not shown). However, some strain sequences within each lineage differed in their branching patterns for some genes. For example, the branching of sequences from strains G-83, IFVv18, 11028, and 313-98 diverges the most between gene trees. On the mdh and rpoD gene trees, sequences from strains G-83, 11028, and 313-98 all cluster within lineage II (bootstrap support, 57% and 76%, respectively) and strain IFVv18 clustered separately from both lineages I and II, as in the concatenated tree. On the groEL1 gene tree, 11028 and 313-98 clustered in lineage I (bootstrap 81%), whereas strain G-83 clustered with lineage II (bootstrap, 91%) and IFVv18 clustered separately. On the gyrB gene tree, G-83, IFVv18, 11028, and 313-98 branched separately from lineages I and II. In contrast, on the groEL2 gene tree from chromosome II, all four strains clustered with lineage I strains (bootstrap support, 100%), and on the pyrC gene tree, G-83 clustered within lineage I (bootstrap, 78%), whereas strains IFVv18, 11028, and 313-98 branched separately from both lineages (data not shown).
The phylogenetic inconsistencies suggest occasional horizontal transfer between the major lineages. We examined the extent of recombination using four statistical tests. The Stephen's test, which detects nonrandom clustering of polymorphic nucleotide sites, found evidence for recombination (polymorphic nucleotide sites nonrandomly distributed) in gyrB and pyrC. For gyrB, recombination was found in a 107-bp segment (five sites) partitioning the ST represented by the isolates G-83, L-49, MLT 406, MLT 362, 300-1C1, 6353/0, K2719, 99-779-DP-D2, 98-640-DP-B9, SS108A-3A, 99-540-DP-B6, 96-9-114s, M79, 90-2-11, IFVv8, and IFVv10 from all other ST (data not shown). For pyrC, recombination was found in a 143-bp segment (five sites) partitioning the ST represented by the isolates L-49, MLT 406, 300-1C1, 6353/0, K2719, 99-779-DP-D2, 98-640-DP-B9, JY1701, SS108A-3A, 99-540-DP-B6, 99-609-DP-A4, 96-9-114s, Env1, M79, 90-2-11, and IFVv10 from all other ST (data not shown). Applying the Sawyer's test to all six genes gave no evidence of recombination. The MaxChi test performs a statistical analysis for clustering of polymorphic sites and found evidence for recombination in four housekeeping genes, gyrB, rpoD, pyrC, and groEL2 (P ≤ 0.05). The Sawyer's and MaxChi tests tend to underestimate the presence of recombination under strong population growth (star-like topology), which is consistent with our results. Moreover, MaxChi may falsely infer the presence of recombination under a simple model of mutation rate correlation (17). Therefore, we employed the PHI test implemented by SplitsTree 4, which distinguishes homoplasies (or convergent mutations) from recombination in generating genotypic diversity. The test found statistical evidence for recombination in the genes gyrB, mdh, groEL1, and pyrC. Interestingly, these results did not support the previous tests used, which might be explained by the low level of phylogenetic signal in our data. As Posada and coworkers have suggested, a minimum nucleotide diversity of 5% is required to obtain substantial statistical power, and in our data set our values range from only 1.6% to 3.2% (59). Perez-Losada and coworkers (58) analyzed MLST data available in the databases for several species and found no evidence of recombination for V. vulnificus. From our study of V. vulnificus, the low phylogenetic signal in our data set did not give strong statistical support for recombination but single-gene analysis did suggest that there is some recombination occurring in this microorganism.
Group-specific PCR.
Two more previously described PCR-based genotyping methods were performed on these isolates in order to compare these methods to the MLST method. These approaches have been successfully used to distinguish clinical and environmental isolates and are based on the 16S rRNA locus, which differentiates isolates into genotypes A or B (52), and a hypothetical protein encoded by VV0401, which differentiates isolates into types C or E (62). Thirty-seven isolates had a B/C type, 29 had the A/E genotype, and 1 isolate G-83 had the A-C type (Table 1). All B/C type isolates clustered within lineage I, including 20 of the 27 isolates from clinical sources (Fig. 3). Lineage II is comprised of all isolates that were of the A/E type, which included only five isolates recovered from clinical sources: four from the United States and a single isolate from France. Of the 27 clinical isolates, 20 were B type and 7 were A type. In contrast, of the 40 environmental isolates, 18 were of the B type and 22 of the A type according to the method of Nilsson et al. (52), confirming that the B type is significantly associated with virulence (Table 1) (two-by-two chi-square tests [20, 7, 18, 22]; P = 0.018). Type A and type B isolates showed a nearly perfect correlation with the E and C type, respectively, using the Rosche et al. method, suggesting that E/C genotypes can similarly distinguish between clinical and environmental strains.
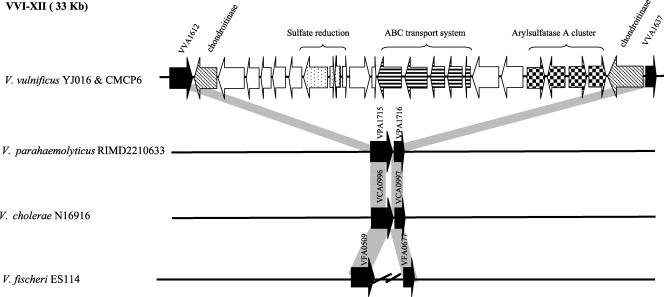
FIG. 3.
BLAST analysis of each ORF present within region XII and flanking core ORFs. Horizontal arrows represent annotated ORFs and ORF orientation. Black arrows represent core ORFs flanking region XII. ORF number refers to the V. vulnificus YJ016 genome. Homologues of core chromosomal ORFs are adjacent to one another in the V. parahaemolyticus RIMD2210633 and V. cholerae N16961 genomes, indicating that this region is absent from these species. In V. fischeri ES114 the 5′ and 3′ core genes are dispersed on the genome.
As noted above, the MLST data are similarly associated with virulence and show a perfect concordance with the previously described genotyping assays; all isolates in lineage I were of the B/C type, and all isolates in lineage II were of the A/E type (Fig. 2). Three of the four isolates that did not cluster with either of the two lineages were A/E type strains, the exception being strain G-83, which was A/C type (Fig. 2).
A study by Vickery and coworkers using a real-time PCR assay for the rapid determination of 16S rRNA genotype found that some isolates contained both types of the 16S rRNA gene (A and B) (77). The genomes of V. vulnificus YJ016 and CMCP6 contain nine rRNA loci, eight on chromosome I and one on chromosome II, all of them type B. In our study, we applied conventional PCR to examine our isolates. The 12 isolates in common to both studies produced the same results, except for CDC9030-95 and 313-98, which in our study are type A and in their study by real-time PCR were type AB. This suggests that our results present the most predominant form of 16S rRNA type found in the isolates. In addition, our study examined some of the same strains previously used by Chatzidaki-Livanis and coworkers in their analysis of REP sequences (19). Our MLST data show a correlation of lineage I with rep-PCR profiles I, IV, and VII and lineage II with rep-PCR profile III. These data indicate more diversity in rep-PCR profiles within lineages.
Our results suggest that a large number of strains from environmental sources have the potential to be virulent and cause disease, i.e., become “clinical.” It is estimated that between 2 and 4 million people in the United States are at risk from infection with V. vulnificus, and annually a large number of people consume raw oysters which contain a significant number of “clinical” isolates (B/C type). Yet the number of cases is limited; between 1989 and 2000, 274 cases involving oyster ingestion occurred in the United States, of which 142 (52%) were fatal (62). This argues heavily in favor of host susceptibility as a key factor in developing disease. This has also been suggested by Starks and coworkers, who examined differences in the abilities of clinical and environmental V. vulnificus isolates to replicate in the host (68).
Identification and distribution of genomic region XII.
Genome comparisons of V. vulnificus YJ016 and CMCP6 versus V. parahaemolyticus RIMD2210633, V. cholerae N16961, and V. fischeri ES114 identified, among others, three regions on chromosome II (ORFs VVA0080 to VVA0186, VVA0301 to VVA0336, and VVA1613 to VVA1636) present in both YJ016 and CMCP6 and absent from the other Vibrio genomes. These three regions did not conform to the definition or features of genomic islands identified previously in V. vulnificus or other gram-negative bacteria, such as a GC content lower than the whole genome, the presence of integrase genes, or flanking repeat sequences (55, 60). ORFs VVA0301 to VVA0336 encompass a 53-kb region that encodes a phospholipase/hemolysin, an RTX toxin gene cluster, a hydrolase, RelB, a plasmid-stabilizing system, a number of transcriptional regulators, VieS (a sensor kinase), as well as a number of hypothetical proteins. Hydrolases are invasion-associated proteins and have been implicated in pathogenicity in other bacteria. In V. cholerae, RTX has been shown to play a role in V. cholerae cytotoxicity and appears to affect regulators of host cell actin polymerization, leading to loss of tight junction integrity (64). RTX toxin has been found in all sequenced Vibrio species to date. In V. vulnificus YJ016, three rtxA homologues are annotated VV1546, VVA1030, and VVA0331 (20). The rtxA1 gene (VVA1030) has recently been shown to be essential for cytotoxicity of V. vulnificus in vitro and in mice (40, 43). Unlike rtxA in V. cholerae, results show that rtxA1 from V. vulnificus is able to disrupt cell membrane integrity (40, 43). Region VVA0080 to VVA0186 is a 124-kb region which encodes a large number of transport systems as well as amino acid metabolism genes, maltoporins, thermostable hemolysin, ferredoxin, flavodoxin, and oxidoreductase. Although most of these ORFs are also present in CMCP6, the region does contain some deletions in this strain (data not shown). The region has an overall GC content of 49%. The third region encompassing ORFs VVA1613 to VVA1636, which we focused on in this study, is a 33-kb region that showed an aberrant GC content of 50% compared to 47% found for the entire genome of V. vulnificus. This region, which we named region XII, encompasses 23 ORFs encoding two chondroitinases, a sulfate reduction system, an oligopeptide ABC transport system, an arylsulfatase A gene cluster, and a methyl-accepting chemotaxis protein as well as a number of hypothetical proteins (Table 6). We did not identify a phage-like integrase or direct repeats associated with the region, which are the usual signatures of genomic islands (55, 60). To examine the distribution of this region among our collection of V. vulnificus isolates, we performed PCR analysis using four primers pairs within region XII (VVA1616F/VVA1618R, VVA1625F/VVA1625R, VVA1634F/VVA1634R, and VVA1630F/VVA1632R) (Fig. 4B). PCR analysis of DNA from our 67 V. vulnificus isolates gave an expected 2.1-kb product with primer pair VVA1616F/VVA1618R from 39 of the isolates. Similarly, PCR analysis with primer pair VVA1625F/VVA1625R gave an expected 1.8-kb PCR band from the same 39 isolates (a subset of these strains is shown in Fig. 4A). The primer pairs VVA1618F/VVA1623R and VVA1625bF/VVA1627R (shown in Fig. 4B) overlap with the previous two primer sets tested and were used to ensure that the regions are contiguous with each other. These two primer pairs were used in PCR assays of the 39 positive isolates examined above, all of which gave the expected size products, indicating that region XII is contiguous. Next, we examined the chromosomal insertion site for region XII in all 67 isolates using two primer pairs, VVA1612bF/VVA1613R targeting the 5′ insertion site and VVA1636F/VVA1637R targeting the 3′ insertion site (shown in Fig. 4B). We found that the 39 isolates that gave positive PCR results for the presence of region XII also give positive PCR bands of the expected sizes with the flanking primer pairs. Figure 4A shows the results for a subset of isolates using the primer pair VVA1612F/VVA1613R; a PCR band of 1.3 kb was obtained only from the region XII-positive isolates. The core chromosomal flanking genes, VVA1612 in the 5′ region and VVA1637 to VVA1639 in the 3′ region, were examined to determine their presence among all 67 isolates (Fig. 4B). As expected, primer pairs targeting these genes gave a PCR band for all isolates. Results of PCR assays for a subset of isolates with both of these primer pairs are shown in Fig. 4C. V. vulnificus isolates that were negative for the presence of region XII were further examined to determine whether VVA1612 is adjacent to VVA1637 in these isolates. The primer pair VVA1612F/VVA1637R, designed within the core 5′ and 3′ genes, was used in PCR assays of all 28 region XII-negative isolates. An approximately 1.2-kb PCR product was obtained from all isolates tested, demonstrating that VVA1612 and VVA1637 are adjacent to one another and that no novel DNA is present (Fig. 4D).
TABLE 6.
BLAST results for the ORFs within region XII (VVA1613 to VVA1636)
| ORF | Length (aa)a | Product | Organism with homologous protein | ORF | % Nucleotide identity | E value |
|---|---|---|---|---|---|---|
| VVA1613 | 683 | Chondroitinase AC lyase | Pedobacter heparinus | 33 | 4e−75 | |
| VVA1614 | 732 | Hypothetical protein | Ruminococcus torques ATCC 27756 | RUMTOR_02141 | 40 | 5e−153 |
| VVA1615 | 394 | Hypothetical protein | Enterobacter sp. strain 638 | Ent638_3433 | 35 | 5e−61 |
| VVA1616 | 253 | Dehydrogenase | Vibrio parahaemolyticus RIMD 2210633 | VPA0077 | 76 | 1e−103 |
| VVA1617 | 376 | Galactose-1-phosphate uridylyltransferase | Salmonella enterica serovar Typhimurium LT2 | STM0775 | 67 | 2e−138 |
| VVA1618 | 338 | UDP-glucose 4-epimerase | Vibrionales bacterium SWAT-3 | VSWAT3_19611 | 64 | 1e−129 |
| VVA1619 | 80 | Hypothetical protein | None | |||
| VVA1620 | 542 | Sulfate permease | Magnetococcus sp. strain MC-1 | Mmc1_1638 | 43 | 2e−117 |
| VVA1621 | 59 | Hypothetical protein | None | |||
| VVA1622 | 49 | Hypothetical protein | None | |||
| VVA1623 | 479 | Hypothetical protein | Chromobacterium violaceum ATCC 12472 | CV0263 | 36 | 1e−58 |
| VVA1624 | 65 | CBS-domain-containing membrane protein | Vibrio cholerae 2740-80 | VC274080_A0374 | 77 | 8e−14 |
| VVA1625 | 648 | Methyl-accepting chemotaxis protein | Aeromonas hydrophila subsp. hydrophila ATCC 7966 | AHA_2600 | 25 | 5e−40 |
| VVA1626 | 566 | ABC-type transport system, ATPase component | Rhizobium leguminosarum bv. viciae 3841 | RL3798 | 51 | 3e−148 |
| VVA1627 | 386 | Oligopeptide ABC transporter, permease protein | Thermotoga petrophila RKU-1 | Tpet_1684 | 55 | 4e−101 |
| VVA1628 | 335 | Oligopeptide ABC transporter, permease protein | Rhizobium leguminosarum bv. viciae 3841 | pRL100277 | 51 | 3e−78 |
| VVA1629 | 56 | Hypothetical protein | None | |||
| VVA1630 | 639 | ABC-type dipeptide transport system, periplasmic component | Rhizobium leguminosarum bv. viciae 3841 | pRL100276 | 37 | 1e−106 |
| VVA1631 | 573 | Conserved hypothetical protein | Pichia stipitis CBS 6054 | PICST_57213 | 32 | 4e−72 |
| VVA1632 | 486 | Arylsulfatase A | Haemophilus influenzae PittHH | CGSHiHH_09635 | 55 | 4e−169 |
| VVA1633 | 403 | Arylsulfatase regulator | Escherichia coli | atsB | 44 | 6e−97 |
| VVA1634 | 497 | Arylsulfatase A | Bacteroides thetaiotaomicron VPI-5482 | BT3349 | 49 | 1e−137 |
| VVA1635 | 488 | Arylsulfatase A | Algoriphagus sp. strain PR1 | ALPR1_08823 | 54 | 5e−151 |
| VVA1636 | 858 | Chondroitinase AC lyase | Pedobacter heparinus | 32 | 5e−85 |
aa, amino acids.
FIG. 4.
PCR results obtained for the analysis of region XII encompassing ORFs VVA1613 to VVA1636 for a subset of isolates. (A) PCR results obtained for a subset of isolates obtained for the primer pairs VVA1212F/VVA1213R, VVA1618F/VVA1623R, VVA1625F/VVA1625R, and VVA1625bF/VVA1627R targeting the 5′ insertion site and ORFs within the island. PCR results were positive for the clinical isolates and negative for the environmental isolates. Note that the primer pairs VVA1618F/VVA1623R and VVA1625bF/VVA1627R were only used for strains that were previously shown to be positive for region XII. (B) Location of the primer pairs used in this study. Region XII is shown in white. Large black arrows represent the core flanking chromosomal ORFs. Small black arrows refer to the primer pairs used within region XII. (C) A subset of strains analyzed with the primer pairs VVA1612F/VVA1612R and VVA1637F/VVA1639R targeting the 5′ and the 3′ core flanking ORFs, respectively. All isolates were positive for these primers pairs. (D) A subset of strains analyzed with the primer pair VVA1612F/VVA1637R designed within the core 5′ and 3′ genes, demonstrating that the genes are adjacent to one another. The molecular ladder used was a 1-kb DNA ladder (New England Biolabs).
In summary, our results of the distribution of region XII show that of the 37 isolates that were of the B/C type (lineage I), 32 contained region XII whereas only 6 of the 29 A/E type strains contained this region (only 3 of the 6 strains belonged to lineage II). Region XII is therefore significantly associated with the more virulent B/C (lineage I) genotype (two-by-two chi-square tests [32, 5, 6, 23]; P < 0.0001). The single A/C type strain also contained region XII. Of the five B/C isolates (CG27, B60, B122, 76M3, and 79M4) within lineage I that did not contain region XII, all were from filter feeders, with four strains originating from India and one from Taiwan. Of the three isolates (L-49, MLT 406, and M79) that contained region XII from lineage II, all were environmental isolates: two were biotype 2 isolates from Spain and Japan, and one was from seawater isolated in the United States in 1991. The two A/E type isolates that contained region XII were both biotype 3 Israeli strains, known to be associated with a recent V. vulnificus outbreak (9). Whether region XII has been acquired by lineage I isolates or alternatively deleted from lineage II isolates is unknown. However, one possible mechanism of acquisition could be transformation. The Schoolnik group has shown that in the presence of chitin, an abundant polymer in the aquatic environment, V. cholerae is competent for the uptake of DNA (13, 46, 47). A similar mechanism for the acquisition of novel regions could operate in V. vulnificus.
In conclusion, two major patterns were found among our collection of V. vulnificus isolates. The lineage I isolates were all of the B/C type, and the majority of these strains contained region XII. Lineage II isolates were all of the A/E type, and the majority of these did not encode region XII (three strains were the exception). The association of region XII with the clinical and B/C type isolates suggests that this region may play a role in virulence either directly or indirectly by increasing the fitness of the strains that contain the region. The identification of two additional regions, ORFs VVA0080 to VVA0186 and ORFs VVA0301 to VVA0336, unique to V. vulnificus is now under investigation to determine whether these regions are present in all V. vulnificus isolates or confined to a subset of isolates associated exclusively with a particular lineage.
Region XII, among other genes, encodes two putative chondroitinase AC lyases (VVA1613 and VVA1636) and sulfate permease as well as an arylsulfatase A gene cluster (VVA1632 to VVA1635), which consists of an arylsulfatase hydrolase, a putative regulator, a sulfatase, and an arylsulfatase A precursor. Chondroitinase is an enzyme that degrades chondroitin sulfate proteoglycans, and this enzyme is believed to play a role in the pathogenicity of bacteria that cause oral infections (71, 72). In Porphyromonas gingivalis, it has been speculated that chondroitinase may be involved in the initial permeation of the gingival epithelium, permitting the entrance of additional microbial virulence factors (65). Jacobs and Stobberingh found that chondroitin sulfatase activity tended to be present more frequently in Streptococcus intermedius and Streptococcus constellatus than in Streptococcus anginosus and was associated with infection-related strains (34).
Arylsulfatases are distributed in a wide range of organisms from mammals to bacteria. In fact, the digestive glands of various molluscs (oysters, clams, and mussels) are found to be a rich source of arylsulfatases (38). Arylsulfatases hydrolyze arylsulfate ester bonds to the corresponding alcohol (or amine) and free sulfate. Sulfur is an essential element for bacterial growth and survival. In bacteria, sulfatase genes are expressed under conditions of sulfur starvation, where the enzymes function in sulfate scavenging (37). Mutations in sulfur metabolism have been shown to be attenuating in Brucella melitensis (41). Mougous and colleagues hypothesized that sulfatases may be involved in novel host-pathogen interaction and speculated that mycobacterial sulfatases may act on extracellular glycosaminoglycans, resulting in the remodeling of extracellular sulfoforms and thereby modulating bacterial adhesion (48). In enteric bacteria, arylsulfatase synthesis is regulated by sulfur supply and by monoamine compounds such as tyramine, dopamine, or norepinephrine (37). Murooka and coworkers have proposed that the monoamine regulon in enteric bacteria is related to the utilization of dopamine sulfate, a compound transported in the human body (49). Hoffman and coworkers identified in Escherichia coli K1 aslA, which encodes arylsulfatase, part of a group of factors that play a role in the E. coli invasion of the blood-brain barrier (29).
In clinical V. vulnificus isolates, the presence of region XII, encoding sulfur accumulation and metabolism functions, suggests that this region may have an important scavenging function removing sulfate groups from exogenous substrates, providing sulfur and carbon sources, which could facilitate survival in the human host where free sulfur is limited. The role of sulfur in V. vulnificus physiology is now under investigation.
Acknowledgments
We thank those who kindly provided us with some of the V. vulnificus strains used in this study (H. C. Wong, S. Miyoshi, A. C. Wright, C. A. Bopp, I. Karunasagar, and D. Hervio-Heath). We also thank the anonymous reviewers for their comments and suggestions that greatly improved the manuscript.
This study was supported in part by Science Foundation Ireland and University of Delaware research grants.
Footnotes
Published ahead of print on 6 July 2007.
REFERENCES
- 1.Abbott, J. C., D. M. Aanensen, K. Rutherford, S. Butcher, and B. G. Spratt. 2005. WebACT—an online companion for the Artemis comparison tool. Bioinformatics 21:3665-3666. [DOI] [PubMed] [Google Scholar]
- 2.Achtman, M., K. Zurth, G. Morelli, G. Torrea, A. Guiyoule, and E. Carniel. 1999. Yersinia pestis, the cause of plague, is a recently emerged clone of Yersinia pseudotuberculosis. Proc. Natl. Acad. Sci. USA 96:14043-14048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Altschul, S. F., T. L. Madden, A. A. Schaffer, J. Zhang, Z. Zhang, W. Miller, and D. J. Lipman. 1997. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25:3389-3402. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Amaro, C., E. G. Biosca, C. Esteve, B. Fouz, and A. E. Toranzo. 1992. Comparative study of phenotypic and virulence properties in Vibrio vulnificus biotype 1 and 2 obtained from a European eel farm experiencing mortalities. Dis. Aquat. Organ. 13:29-35. [Google Scholar]
- 5.Arias, C. R., M. J. Pujalte, E. Garay, and R. Aznar. 1998. Genetic relatedness among environmental, clinical, and diseased-eel Vibrio vulnificus isolates from different geographic regions by ribotyping and randomly amplified polymorphic DNA PCR. Appl. Environ. Microbiol. 64:3403-3410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Arias, C. R., L. Verdonck, J. Swings, E. Garay, and R. Aznar. 1997. Intraspecific differentiation of Vibrio vulnificus biotypes by amplified fragment length polymorphism and ribotyping. Appl. Environ. Microbiol. 63:2600-2606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Biosca, E. G., C. Amaro, J. L. Larsen, and K. Pedersen. 1997. Phenotypic and genotypic characterization of Vibrio vulnificus: proposal for the substitution of the subspecific taxon biotype for serovar. Appl. Environ. Microbiol. 63:1460-1466. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Biosca, E. G., R. M. Collado, J. D. Oliver, and C. Amaro. 1999. Comparative study of biological properties and electrophoretic characteristics of lipopolysaccharide from eel-virulent and eel-avirulent Vibrio vulnificus strains. Appl. Environ. Microbiol. 65:856-858. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Bisharat, N., V. Agmon, R. Finkelstein, R. Raz, G. Ben-Dror, L. Lerner, S. Soboh, R. Colodner, D. N. Cameron, D. L. Wykstra, D. L. Swerdlow, and J. J. Farmer III. 1999. Clinical, epidemiological, and microbiological features of Vibrio vulnificus biogroup 3 causing outbreaks of wound infection and bacteraemia in Israel. Lancet 354:1421-1424. [DOI] [PubMed] [Google Scholar]
- 10.Bisharat, N., C. Amaro, B. Fouz, A. Llorens, and D. I. Cohen. 2007. Serological and molecular characteristics of Vibrio vulnificus biotype 3: evidence for high clonality. Microbiology 153:847-856. [DOI] [PubMed] [Google Scholar]
- 11.Bisharat, N., D. I. Cohen, R. M. Harding, D. Falush, D. W. Crook, T. Peto, and M. C. Maiden. 2005. Hybrid Vibrio vulnificus. Emerg. Infect. Dis. 11:30-35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Bisharat, N., and R. Raz. 1996. Vibrio infection in Israel due to changes in fish marketing. Lancet 348:1585-1586. [DOI] [PubMed] [Google Scholar]
- 13.Blokesch, M., and G. K. Schoolnik. 2007. Serogroup conversion of Vibrio cholerae in aquatic reservoirs. PLoS Pathog. 3:e81. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Boyd, E. F. 2005. Bacteriophages and bacterial virulence, p. 223-266. In E. Kutter and A. Sulakvelidze (ed.), Bacteriophages: biology and applications. CRC Press, Boca Raton, Florida.
- 15.Boyd, E. F., and H. Brussow. 2002. Common themes among bacteriophage-encoded virulence factors and diversity among the bacteriophages involved. Trends Microbiol. 10:521-529. [DOI] [PubMed] [Google Scholar]
- 16.Boyd, E. F., J. Li, H. Ochman, and R. K. Selander. 1997. Comparative genetics of the inv-spa invasion gene complex of Salmonella enterica. J. Bacteriol. 179:1985-1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Bruen, T. C., H. Philippe, and D. Bryant. 2006. A simple and robust statistical test for detecting the presence of recombination. Genetics 172:2665-2681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Buchrieser, C., V. V. Gangar, R. L. Murphree, M. L. Tamplin, and C. W. Kaspar. 1995. Multiple Vibrio vulnificus strains in oysters as demonstrated by clamped homogeneous electric field gel electrophoresis. Appl. Environ. Microbiol. 61:1163-1168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Chatzidaki-Livanis, M., M. A. Hubbard, K. Gordon, V. J. Harwood, and A. C. Wright. 2006. Genetic distinctions among clinical and environmental strains of Vibrio vulnificus. Appl. Environ. Microbiol. 72:6136-6141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Chen, C. Y., K. M. Wu, Y. C. Chang, C. H. Chang, H. C. Tsai, T. L. Liao, Y. M. Liu, H. J. Chen, A. B. Shen, J. C. Li, T. L. Su, C. P. Shao, C. T. Lee, L. I. Hor, and S. F. Tsai. 2003. Comparative genome analysis of Vibrio vulnificus, a marine pathogen. Genome Res. 13:2577-2587. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.DePaola, A., G. M. Capers, and D. Alexander. 1994. Densities of Vibrio vulnificus in the intestines of fish from the U.S. Gulf Coast. Appl. Environ. Microbiol. 60:984-988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Feil, E. J., J. E. Cooper, H. Grundmann, D. A. Robinson, M. C. Enright, T. Berendt, S. J. Peacock, J. M. Smith, M. Murphy, B. G. Spratt, C. E. Moore, and N. P. J. Day. 2003. How clonal is Staphylococcus aureus? J. Bacteriol. 185:3307-3316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Gulig, P. A., K. L. Bourdage, and A. M. Starks. 2005. Molecular pathogenesis of Vibrio vulnificus. J. Microbiol. 43:118-131. [PubMed] [Google Scholar]
- 24.Gutacker, M., N. Conza, C. Benagli, A. Pedroli, M. V. Bernasconi, L. Permin, R. Aznar, and J. C. Piffaretti. 2003. Population genetics of Vibrio vulnificus: identification of two divisions and a distinct eel-pathogenic clone. Appl. Environ. Microbiol. 69:3203-3212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Hacker, J., and E. Carniel. 2001. Ecological fitness, genomic islands and bacterial pathogenicity. A Darwinian view of the evolution of microbes. EMBO Rep. 2:376-381. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Hallin, P. F., and D. W. Ussery. 2004. CBS genome atlas database: a dynamic storage for bioinformatic results and sequence data. Bioinformatics 20:3682-3686. [DOI] [PubMed] [Google Scholar]
- 27.Heidelberg, J. F., K. B. Heidelberg, and R. R. Colwell. 2002. Bacteria of the γ-subclass Proteobacteria associated with zooplankton in Chesapeake Bay. Appl. Environ. Microbiol. 68:5498-5507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Hlady, W. G., and K. C. Klontz. 1996. The epidemiology of Vibrio infections in Florida, 1981-1993. J. Infect. Dis. 173:1176-1183. [DOI] [PubMed] [Google Scholar]
- 29.Hoffman, J. A., J. L. Badger, Y. Zhang, S.-H. Huang, and K. S. Kim. 2000. Escherichia coli K1 aslA contributes to invasion of brain microvascular endothelial cells in vitro and in vivo. Infect. Immun. 68:5062-5067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Hoi, L., A. Dalsgaard, J. L. Larsen, J. M. Warner, and J. D. Oliver. 1997. Comparison of ribotyping and randomly amplified polymorphic DNA PCR for characterization of Vibrio vulnificus. Appl. Environ. Microbiol. 63:1674-1678. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Hoi, L., J. L. Larsen, I. Dalsgaard, and A. Dalsgaard. 1998. Occurrence of Vibrio vulnificus biotypes in Danish marine environments. Appl. Environ. Microbiol. 64:7-13. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Hor, L. I., C. T. Gao, and L. Wan. 1995. Isolation and characterization of Vibrio vulnificus inhabiting the marine environment of the southwestern area of Taiwan. J. Biomed. Sci. 2:384-389. [DOI] [PubMed] [Google Scholar]
- 33.Huson, D. H., and D. Bryant. 2006. Application of phylogenetic networks in evolutionary studies. Mol. Biol. Evol. 23:254-267. [DOI] [PubMed] [Google Scholar]
- 34.Jacobs, J. A., and E. E. Stobberingh 1995. Hydrolytic enzymes of Streptococcus anginosus, Streptococcus constellatus and Streptococcus intermedius in relation to infection. Eur. J. Clin. Microbiol. Infect. Dis. 14:818-820. [DOI] [PubMed] [Google Scholar]
- 35.Kaysner, C. A., C. Abeyta, Jr., M. M. Wekell, A. DePaola, Jr., R. F. Stott, and J. M. Leitch. 1987. Virulent strains of Vibrio vulnificus isolated from estuaries of the United States West Coast. Appl. Environ. Microbiol. 53:1349-1351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Kaysner, C. A., M. L. Tamplin, M. M. Wekell, R. F. Stott, and K. G. Colburn. 1989. Survival of Vibrio vulnificus in shellstock and shucked oysters (Crassostrea gigas and Crassostrea virginica) and effects of isolation medium on recovery. Appl. Environ. Microbiol. 55:3072-3079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kertesz, M. A. 2000. Riding the sulfur cycle—metabolism of sulfonates and sulfate esters in gram-negative bacteria. FEMS Microbiol. Rev. 24:135-175. [DOI] [PubMed] [Google Scholar]
- 38.Kim, D.-E., K.-H. Kim, Y.-J. Bae, J.-H. Lee, Y.-H. Jang, and S.-W. Nam. 2005. Purification and characterization of the recombinant arylsulfatase cloned from Pseudoalteromonas carrageenovora. Protein Expr. Purif. 39:107-115. [DOI] [PubMed] [Google Scholar]
- 39.Kumar, S., K. Tamura, and M. Nei. 2004. MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief. Bioinform. 5:150-163. [DOI] [PubMed] [Google Scholar]
- 40.Lee, J. H., M. W. Kim, B. S. Kim, S. M. Kim, B. C. Lee, T. S. Kim, and S. H. Choi. 2007. Identification and characterization of the Vibrio vulnificus rtxA essential for cytotoxicity in vitro and virulence in mice. J. Microbiol. 45:146-152. [PubMed] [Google Scholar]
- 41.Lestrate, P., R. M. Delrue, I. Danese, C. Didembourg, B. Taminiau, P. Mertens, X. De Bolle, A. Tibor, C. M. Tang, and J. J. Letesson. 2000. Identification and characterization of in vivo attenuated mutants of Brucella melitensis. Mol. Microbiol. 38:543-551. [DOI] [PubMed] [Google Scholar]
- 42.Linkous, D. A., and J. D. Oliver. 1999. Pathogenesis of Vibrio vulnificus. FEMS Microbiol. Lett. 174:207-214. [DOI] [PubMed] [Google Scholar]
- 43.Liu, M., A. F. Alice, H. Naka, and J. H. Crosa. 2007. The HlyU protein is a positive regulator of rtxA1, a gene responsible for cytotoxicity and virulence in the human pathogen Vibrio vulnificus. Infect. Immun. 75:3282-3289. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Malhotra, A., E. Severinova, and S. A. Darst. 1996. Crystal structure of a σ70 subunit fragment from E. coli RNA polymerase. Cell 87:127-136. [DOI] [PubMed] [Google Scholar]
- 45.Meats, E., E. J. Feil, S. Stringer, A. J. Cody, R. Goldstein, J. S. Kroll, T. Popovic, and B. G. Spratt. 2003. Characterization of encapsulated and noncapsulated Haemophilus influenzae and determination of phylogenetic relationships by multilocus sequence typing. J. Clin. Microbiol. 41:1623-1636. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Meibom, K. L., M. Blokesch, N. A. Dolganov, C.-Y. Wu, and G. K. Schoolnik. 2005. Chitin induces natural competence in Vibrio cholerae. Science 310:1824-1827. [DOI] [PubMed] [Google Scholar]
- 47.Miller, M. C., D. P. Keymer, A. Avelar, A. B. Boehm, and G. K. Schoolnik. 2007. Detection and transformation of genome segments that differ within a coastal population of Vibrio cholerae strains. Appl. Environ. Microbiol. 73:3695-3704. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Mougous, J. D., R. E. Green, S. J. Williams, S. E. Brenner, and C. R. Bertozzi. 2002. Sulfotransferases and sulfatases in mycobacteria. Chem. Biol. 9:767-776. [DOI] [PubMed] [Google Scholar]
- 49.Murooka, Y., H. Azakami, and M. Yamashita. 1996. The monoamine regulon including syntheses of arylsulfatase and monoamine oxidase in bacteria. Biosci. Biotechnol. Biochem. 60:935-941. [DOI] [PubMed] [Google Scholar]
- 50.Nei, M., and T. Gojobori. 1986. Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol. Biol. Evol. 3:418-426. [DOI] [PubMed] [Google Scholar]
- 51.Nei, M., and L. Jin. 1989. Variances of the average numbers of nucleotide substitutions within and between populations. Mol. Biol. Evol. 6:290-300. [DOI] [PubMed] [Google Scholar]
- 52.Nilsson, W. B., R. N. Paranjype, A. DePaola, and M. S. Strom. 2003. Sequence polymorphism of the 16S rRNA gene of Vibrio vulnificus is a possible indicator of strain virulence. J. Clin. Microbiol. 41:442-446. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Nilsson, W. B., and M. S. Strom. 2002. Detection and identification of bacterial pathogens of fish in kidney tissue using terminal restriction fragment length polymorphism (T-RFLP) analysis of 16S rRNA genes. Dis. Aquat. Organ. 48:175-185. [DOI] [PubMed] [Google Scholar]
- 54.Oliver, J. D. 2006. Vibrio vulnificus, p. 349-366. In F. L. Thompson, B. Austin, and J. Swings (ed.), The biology of vibrios. ASM Press, Washington, DC.
- 55.O'Shea, Y. A., S. Finnan, F. J. Reen, J. P. Morrissey, F. O'Gara, and E. F. Boyd. 2004. The Vibrio seventh pandemic island-II is a 26.9 kb genomic island present in Vibrio cholerae El Tor and O139 serogroup isolates that shows homology to a 43.4 kb genomic island in V. vulnificus. Microbiology 150:4053-4063. [DOI] [PubMed] [Google Scholar]
- 56.Parvathi, A., H. S. Kumar, I. Karunasagar, and I. Karunasagar. 2005. Study of the occurrence of Vibrio vulnificus in oysters in India by polymerase chain reaction (PCR) and heterogeneity among V. vulnificus by randomly amplified polymorphic DNA PCR and gyrB sequence analysis. Environ. Microbiol. 7:995-1002. [DOI] [PubMed] [Google Scholar]
- 57.Paz, S., N. Bisharat, E. Paz, O. Kidar, and D. Cohen. 2007. Climate change and the emergence of Vibrio vulnificus disease in Israel. Environ. Res. 103:390-396. [DOI] [PubMed] [Google Scholar]
- 58.Perez-Losada, M., E. B. Browne, A. Madsen, T. Wirth, R. P. Viscidi, and K. A. Crandall. 2006. Population genetics of microbial pathogens estimated from multilocus sequence typing (MLST) data. Infect. Genet. Evol. 6:97-112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Posada, D., K. A. Crandall, and E. C. Holmes. 2002. Recombination in evolutionary genomics. Annu. Rev. Genet. 36:75-97. [DOI] [PubMed] [Google Scholar]
- 60.Quirke, A. M., F. J. Reen, M. J. Claesson, and E. F. Boyd. 2006. Genomic island identification in Vibrio vulnificus reveals significant genome plasticity in this human pathogen. Bioinformatics 22:905-910. [DOI] [PubMed] [Google Scholar]
- 61.Reen, F. J., S. Almagro-Moreno, D. Ussery, and E. F. Boyd. 2006. The genomic code: inferring Vibrionaceae niche specialization. Nat. Rev. Microbiol. 4:697-704. [DOI] [PubMed] [Google Scholar]
- 62.Rosche, T., Y. Yano, and J. D. Oliver. 2005. A rapid and simple PCR analysis indicates there are two subgroups of Vibrio vulnificus which correlate with clinical or environmental isolation. Microbiol. Immunol. 49:381-389. [DOI] [PubMed] [Google Scholar]
- 63.Sawyer, S. 1989. Statistical tests for detecting gene conversion. Mol. Biol. Evol. 6:526-538. [DOI] [PubMed] [Google Scholar]
- 64.Sheahan, K.-L., C. L. Cordero, and K. J. Fullner Satchell. 2004. Identification of a domain within the multifunctional Vibrio cholerae RTX toxin that covalently cross-links actin. Proc. Natl. Acad. Sci. USA 101:9798-9803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Smith, A. J., J. Greenman, and G. Embery. 1997. Detection and possible biological role of chondroitinase and heparitinase enzymes produced by Porphyromonas gingivalis W50. J. Periodontal Res. 32:1-8. [DOI] [PubMed] [Google Scholar]
- 66.Smith, J. M. 1992. Analyzing the mosaic structure of genes. J. Mol. Evol. 34:126-129. [DOI] [PubMed] [Google Scholar]
- 67.Sreevatsan, S., X. Pan, K. E. Stockbauer, N. D. Connell, B. N. Kreiswirth, T. S. Whittam, and J. M. Musser. 1997. Restricted structural gene polymorphism in the Mycobacterium tuberculosis complex indicates evolutionarily recent global dissemination. Proc. Natl. Acad. Sci. USA 94:9869-9874. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Starks, A. M., T. R. Schoeb, M. L. Tamplin, S. Parveen, T. J. Doyle, P. E. Bomeisl, G. M. Escudero, and P. A. Gulig. 2000. Pathogenesis of infection by clinical and environmental strains of Vibrio vulnificus in iron-dextran-treated mice. Infect. Immun. 68:5785-5793. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Stephens, J. C. 1985. Statistical methods of DNA sequence analysis: detection of intragenic recombination or gene conversion. Mol. Biol. Evol. 2:539-556. [DOI] [PubMed] [Google Scholar]
- 70.Strom, M. S., and R. N. Paranjpye. 2000. Epidemiology and pathogenesis of Vibrio vulnificus. Microbes Infect. 2:177-188. [DOI] [PubMed] [Google Scholar]
- 71.Takeuchi, H., Y. Hori, J. Kanehisa, A. Tani, R. Takay, and H. Sagawa. 1982. Bacterial chondroitinase ABC and hyaluronidase in human dental plaque and inflamed gingiva. J. Osaka Dent. Univ. 16:183-187. [PubMed] [Google Scholar]
- 72.Tam, Y. C., R. F. Harvey, and E. C. S. Chan. 1982. Chondroitin sulfatase-producing and hyaluronidase-producing oral bacteria associated with periodontal disease. J. Can. Dent. Assoc. 48:115-120. [PubMed] [Google Scholar]
- 73.Tamplin, M., G. E. Rodrick, N. J. Blake, and T. Cuba. 1982. Isolation and characterization of Vibrio vulnificus from two Florida estuaries. Appl. Environ. Microbiol. 44:1466-1470. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 74.Tamplin, M. L., J. K. Jackson, C. Buchrieser, R. L. Murphree, K. M. Portier, V. Gangar, L. G. Miller, and C. W. Kaspar. 1996. Pulsed-field gel electrophoresis and ribotype profiles of clinical and environmental Vibrio vulnificus isolates. Appl. Environ. Microbiol. 62:3572-3580. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Thompson, J. D., D. G. Higgins, and T. J. Gibson. 1994. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22:4673-4680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 76.van Passel, M. W. J., A. C. M. Luyf, A. H. C. van Kampen, A. Bart, and A. van der Ende. 2005. Deltarho-web, an online tool to assess composition similarity of individual nucleic acid sequences. Bioinformatics 21:3053-3055. [DOI] [PubMed] [Google Scholar]
- 77.Vickery, M. C. L., W. B. Nilsson, M. S. Strom, J. L. Nordstrom, and A. DePaola. 2007. A real-time PCR assay for the rapid determination of 16S rRNA genotype in Vibrio vulnificus. J. Microbiol. Methods 68:376-384. [DOI] [PubMed] [Google Scholar]
- 78.Warner, J. M., and J. D. Oliver. 1999. Randomly amplified polymorphic DNA analysis of clinical and environmental isolates of Vibrio vulnificus and other Vibrio species. Appl. Environ. Microbiol. 65:1141-1144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 79.Wong, H. C., S. Y. Chen, M. Y. Chen, J. D. Oliver, L. I. Hor, and W. C. Tsai. 2004. Pulsed-field gel electrophoresis analysis of Vibrio vulnificus strains isolated from Taiwan and the United States. Appl. Environ. Microbiol. 70:5153-5158. [DOI] [PMC free article] [PubMed] [Google Scholar]