Abstract
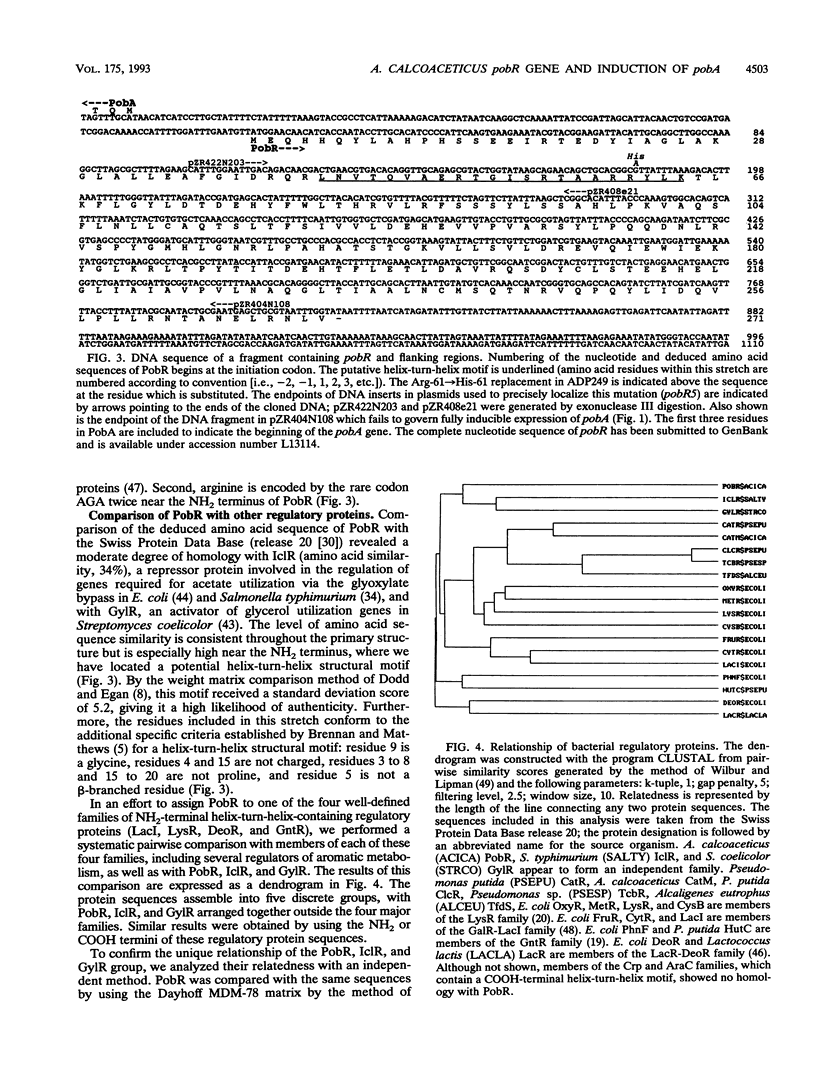
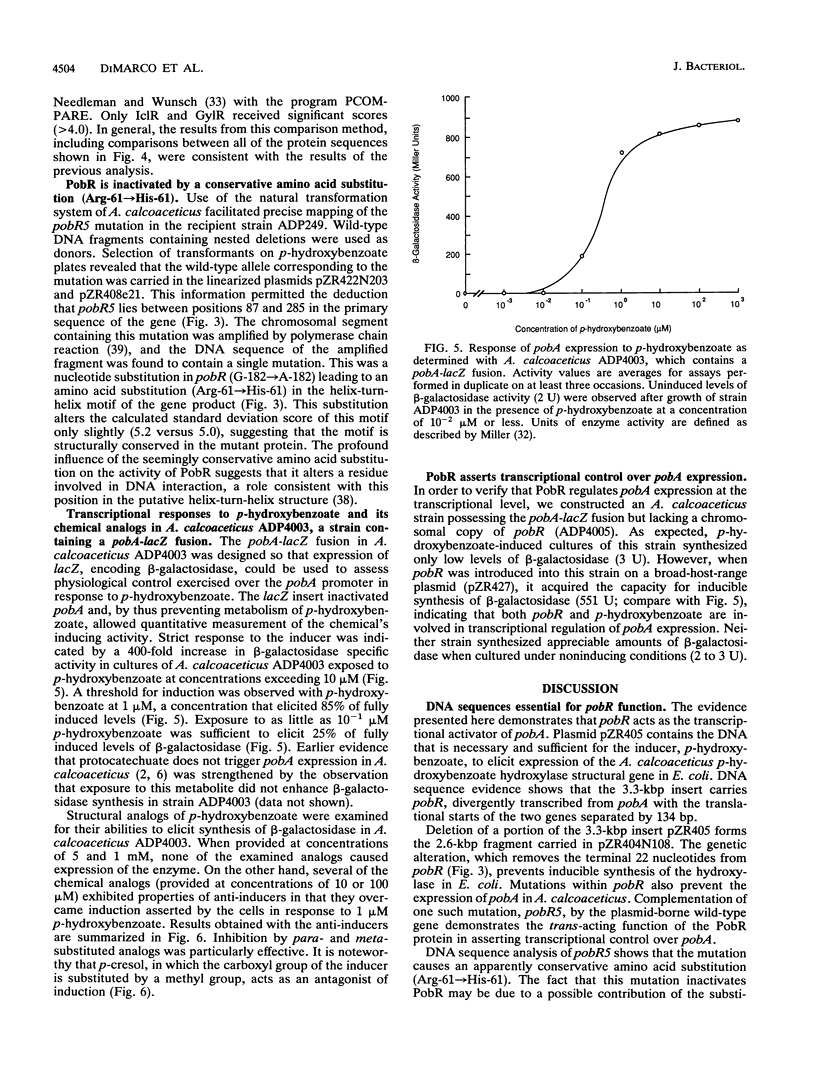
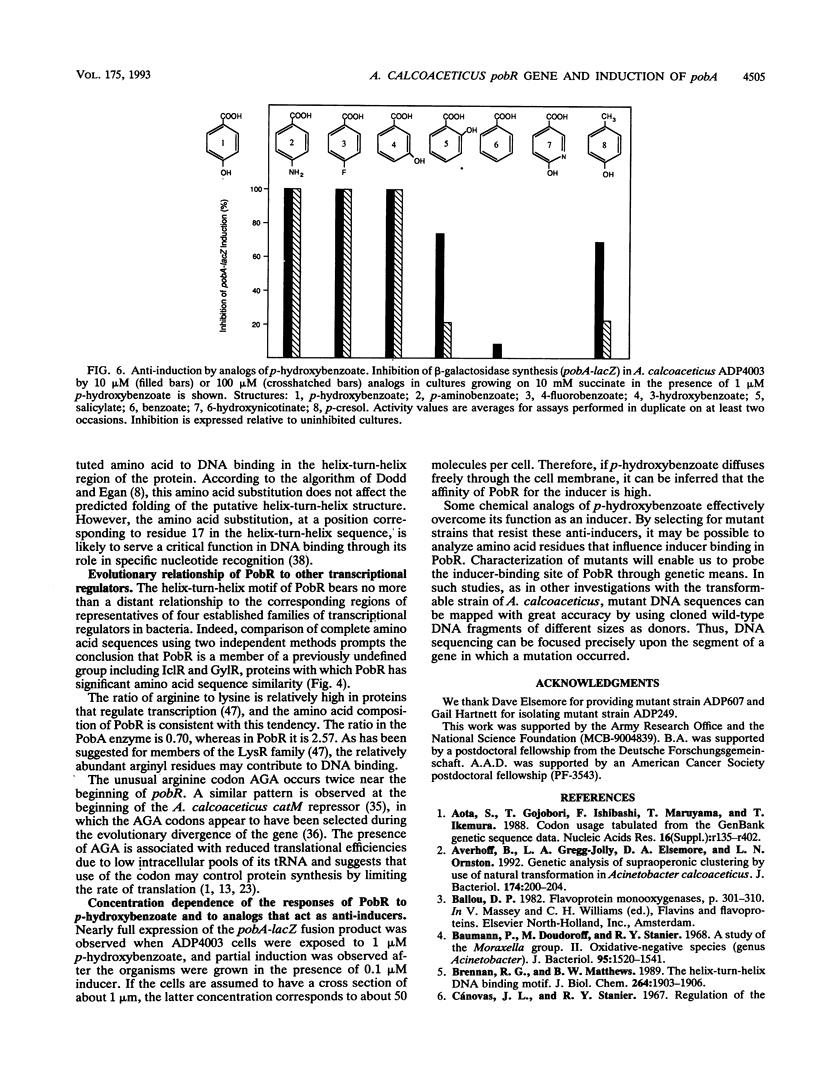
We have identified pobR, a gene encoding a transcriptional activator that regulates expression of pobA, the structural gene for p-hydroxybenzoate hydroxylase (PobA) in Acinetobacter calcoaceticus ADP1. Inducible expression of cloned pobA in Escherichia coli depended upon the presence of a functional pobR gene, and mutations within pobR prevented pobA expression in A. calcoaceticus. A pobA-lacZ operon fusion was used to demonstrate that pobA expression in A. calcoaceticus is enhanced up to 400-fold by the inducer p-hydroxybenzoate. Inducer concentrations as low as 10(-7) M were sufficient to elicit partial induction. Some structurally related analogs of p-hydroxybenzoate, unable to cause induction by themselves, were effective anti-inducers. The nucleotide sequence of pobR was determined, and the activator gene was shown to be transcribed divergently from pobA; the genes are separated by 134 DNA base pairs. The deduced amino acid sequence yielded a polypeptide of M(r) = 30,764. Analysis of this sequence revealed at the NH2 terminus a stretch of residues with high potential for forming a helix-turn-helix structure that could serve as a DNA-binding domain. A conservative amino acid substitution (Arg-61-->His-61) in this region inactivated PobR. The primary structure of PobR appears to be evolutionarily distinct from the four major families of NH2-terminal helix-turn-helix containing bacterial regulatory proteins that have been identified thus far.
Full text
PDF





Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aota S., Gojobori T., Ishibashi F., Maruyama T., Ikemura T. Codon usage tabulated from the GenBank Genetic Sequence Data. Nucleic Acids Res. 1988;16 (Suppl):r315–r402. doi: 10.1093/nar/16.suppl.r315. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Averhoff B., Gregg-Jolly L., Elsemore D., Ornston L. N. Genetic analysis of supraoperonic clustering by use of natural transformation in Acinetobacter calcoaceticus. J Bacteriol. 1992 Jan;174(1):200–204. doi: 10.1128/jb.174.1.200-204.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baumann P., Doudoroff M., Stanier R. Y. A study of the Moraxella group. II. Oxidative-negative species (genus Acinetobacter). J Bacteriol. 1968 May;95(5):1520–1541. doi: 10.1128/jb.95.5.1520-1541.1968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brennan R. G., Matthews B. W. The helix-turn-helix DNA binding motif. J Biol Chem. 1989 Feb 5;264(4):1903–1906. [PubMed] [Google Scholar]
- DiMarco A. A., Averhoff B. A., Kim E. E., Ornston L. N. Evolutionary divergence of pobA, the structural gene encoding p-hydroxybenzoate hydroxylase in an Acinetobacter calcoaceticus strain well-suited for genetic analysis. Gene. 1993 Mar 15;125(1):25–33. doi: 10.1016/0378-1119(93)90741-k. [DOI] [PubMed] [Google Scholar]
- Dodd I. B., Egan J. B. Improved detection of helix-turn-helix DNA-binding motifs in protein sequences. Nucleic Acids Res. 1990 Sep 11;18(17):5019–5026. doi: 10.1093/nar/18.17.5019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doten R. C., Ngai K. L., Mitchell D. J., Ornston L. N. Cloning and genetic organization of the pca gene cluster from Acinetobacter calcoaceticus. J Bacteriol. 1987 Jul;169(7):3168–3174. doi: 10.1128/jb.169.7.3168-3174.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Entsch B., Nan Y., Weaich K., Scott K. F. Sequence and organization of pobA, the gene coding for p-hydroxybenzoate hydroxylase, an inducible enzyme from Pseudomonas aeruginosa. Gene. 1988 Nov 30;71(2):279–291. doi: 10.1016/0378-1119(88)90044-3. [DOI] [PubMed] [Google Scholar]
- Garcia G. M., Mar P. K., Mullin D. A., Walker J. R., Prather N. E. The E. coli dnaY gene encodes an arginine transfer RNA. Cell. 1986 May 9;45(3):453–459. doi: 10.1016/0092-8674(86)90331-4. [DOI] [PubMed] [Google Scholar]
- Gregg-Jolly L. A., Ornston L. N. Recovery of DNA from the Acinetobacter calcoaceticus chromosome by gap repair. J Bacteriol. 1990 Oct;172(10):6169–6172. doi: 10.1128/jb.172.10.6169-6172.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hartnett C., Neidle E. L., Ngai K. L., Ornston L. N. DNA sequences of genes encoding Acinetobacter calcoaceticus protocatechuate 3,4-dioxygenase: evidence indicating shuffling of genes and of DNA sequences within genes during their evolutionary divergence. J Bacteriol. 1990 Feb;172(2):956–966. doi: 10.1128/jb.172.2.956-966.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hartnett G. B., Averhoff B., Ornston L. N. Selection of Acinetobacter calcoaceticus mutants deficient in the p-hydroxybenzoate hydroxylase gene (pobA), a member of a supraoperonic cluster. J Bacteriol. 1990 Oct;172(10):6160–6161. doi: 10.1128/jb.172.10.6160-6161.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Haydon D. J., Guest J. R. A new family of bacterial regulatory proteins. FEMS Microbiol Lett. 1991 Apr 15;63(2-3):291–295. doi: 10.1016/0378-1097(91)90101-f. [DOI] [PubMed] [Google Scholar]
- Henikoff S., Haughn G. W., Calvo J. M., Wallace J. C. A large family of bacterial activator proteins. Proc Natl Acad Sci U S A. 1988 Sep;85(18):6602–6606. doi: 10.1073/pnas.85.18.6602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holmes D. S., Quigley M. A rapid boiling method for the preparation of bacterial plasmids. Anal Biochem. 1981 Jun;114(1):193–197. doi: 10.1016/0003-2697(81)90473-5. [DOI] [PubMed] [Google Scholar]
- Howell L. G., Spector T., Massey V. Purification and properties of p-hydroxybenzoate hydroxylase from Pseudomonas fluorescens. J Biol Chem. 1972 Jul 10;247(13):4340–4350. [PubMed] [Google Scholar]
- Ikemura T. Correlation between the abundance of Escherichia coli transfer RNAs and the occurrence of the respective codons in its protein genes. J Mol Biol. 1981 Feb 15;146(1):1–21. doi: 10.1016/0022-2836(81)90363-6. [DOI] [PubMed] [Google Scholar]
- Juni E. Genetics and physiology of Acinetobacter. Annu Rev Microbiol. 1978;32:349–371. doi: 10.1146/annurev.mi.32.100178.002025. [DOI] [PubMed] [Google Scholar]
- Juni E., Janik A. Transformation of Acinetobacter calco-aceticus (Bacterium anitratum). J Bacteriol. 1969 Apr;98(1):281–288. doi: 10.1128/jb.98.1.281-288.1969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keen N. T., Tamaki S., Kobayashi D., Trollinger D. Improved broad-host-range plasmids for DNA cloning in gram-negative bacteria. Gene. 1988 Oct 15;70(1):191–197. doi: 10.1016/0378-1119(88)90117-5. [DOI] [PubMed] [Google Scholar]
- Kokotek W., Lotz W. Construction of a lacZ-kanamycin-resistance cassette, useful for site-directed mutagenesis and as a promoter probe. Gene. 1989 Dec 14;84(2):467–471. doi: 10.1016/0378-1119(89)90522-2. [DOI] [PubMed] [Google Scholar]
- LOWRY O. H., ROSEBROUGH N. J., FARR A. L., RANDALL R. J. Protein measurement with the Folin phenol reagent. J Biol Chem. 1951 Nov;193(1):265–275. [PubMed] [Google Scholar]
- Lipman D. J., Pearson W. R. Rapid and sensitive protein similarity searches. Science. 1985 Mar 22;227(4693):1435–1441. doi: 10.1126/science.2983426. [DOI] [PubMed] [Google Scholar]
- Needleman S. B., Wunsch C. D. A general method applicable to the search for similarities in the amino acid sequence of two proteins. J Mol Biol. 1970 Mar;48(3):443–453. doi: 10.1016/0022-2836(70)90057-4. [DOI] [PubMed] [Google Scholar]
- Neidle E. L., Hartnett C., Ornston L. N. Characterization of Acinetobacter calcoaceticus catM, a repressor gene homologous in sequence to transcriptional activator genes. J Bacteriol. 1989 Oct;171(10):5410–5421. doi: 10.1128/jb.171.10.5410-5421.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nègre D., Cortay J. C., Old I. G., Galinier A., Richaud C., Saint Girons I., Cozzone A. J. Overproduction and characterization of the iclR gene product of Escherichia coli K-12 and comparison with that of Salmonella typhimurium LT2. Gene. 1991 Jan 2;97(1):29–37. doi: 10.1016/0378-1119(91)90006-w. [DOI] [PubMed] [Google Scholar]
- Ornston L. N., Stanier R. Y. The conversion of catechol and protocatechuate to beta-ketoadipate by Pseudomonas putida. J Biol Chem. 1966 Aug 25;241(16):3776–3786. [PubMed] [Google Scholar]
- Pabo C. O., Sauer R. T. Protein-DNA recognition. Annu Rev Biochem. 1984;53:293–321. doi: 10.1146/annurev.bi.53.070184.001453. [DOI] [PubMed] [Google Scholar]
- Saiki R. K., Gelfand D. H., Stoffel S., Scharf S. J., Higuchi R., Horn G. T., Mullis K. B., Erlich H. A. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science. 1988 Jan 29;239(4839):487–491. doi: 10.1126/science.2448875. [DOI] [PubMed] [Google Scholar]
- Schreuder H. A., Prick P. A., Wierenga R. K., Vriend G., Wilson K. S., Hol W. G., Drenth J. Crystal structure of the p-hydroxybenzoate hydroxylase-substrate complex refined at 1.9 A resolution. Analysis of the enzyme-substrate and enzyme-product complexes. J Mol Biol. 1989 Aug 20;208(4):679–696. doi: 10.1016/0022-2836(89)90158-7. [DOI] [PubMed] [Google Scholar]
- Simon R., O'Connell M., Labes M., Pühler A. Plasmid vectors for the genetic analysis and manipulation of rhizobia and other gram-negative bacteria. Methods Enzymol. 1986;118:640–659. doi: 10.1016/0076-6879(86)18106-7. [DOI] [PubMed] [Google Scholar]
- Smith C. P., Chater K. F. Structure and regulation of controlling sequences for the Streptomyces coelicolor glycerol operon. J Mol Biol. 1988 Dec 5;204(3):569–580. doi: 10.1016/0022-2836(88)90356-7. [DOI] [PubMed] [Google Scholar]
- Sunnarborg A., Klumpp D., Chung T., LaPorte D. C. Regulation of the glyoxylate bypass operon: cloning and characterization of iclR. J Bacteriol. 1990 May;172(5):2642–2649. doi: 10.1128/jb.172.5.2642-2649.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Viale A. M., Kobayashi H., Akazawa T., Henikoff S. rbcR [correction of rcbR], a gene coding for a member of the LysR family of transcriptional regulators, is located upstream of the expressed set of ribulose 1,5-bisphosphate carboxylase/oxygenase genes in the photosynthetic bacterium Chromatium vinosum. J Bacteriol. 1991 Aug;173(16):5224–5229. doi: 10.1128/jb.173.16.5224-5229.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weickert M. J., Adhya S. A family of bacterial regulators homologous to Gal and Lac repressors. J Biol Chem. 1992 Aug 5;267(22):15869–15874. [PubMed] [Google Scholar]
- Wilbur W. J., Lipman D. J. Rapid similarity searches of nucleic acid and protein data banks. Proc Natl Acad Sci U S A. 1983 Feb;80(3):726–730. doi: 10.1073/pnas.80.3.726. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]
- van Berkel W., Westphal A., Eschrich K., Eppink M., de Kok A. Substitution of Arg214 at the substrate-binding site of p-hydroxybenzoate hydroxylase from Pseudomonas fluorescens. Eur J Biochem. 1992 Dec 1;210(2):411–419. doi: 10.1111/j.1432-1033.1992.tb17436.x. [DOI] [PubMed] [Google Scholar]
- van Rooijen R. J., de Vos W. M. Molecular cloning, transcriptional analysis, and nucleotide sequence of lacR, a gene encoding the repressor of the lactose phosphotransferase system of Lactococcus lactis. J Biol Chem. 1990 Oct 25;265(30):18499–18503. [PubMed] [Google Scholar]



