Abstract
Lethal alleles of the Drosophila k43 gene result in small or missing imaginal discs, greatly reduced mitotic index, and fragmented and abnormally condensed chromosomes. A female-sterile allele of k43 specifically reduces chorion gene amplification in ovarian follicle cells. k43 was cloned by chromosomal walking, and the identification of the k43 gene was confirmed by phenotypic rescue and sequence analysis of mutant alleles. The sequence analyses reveal that the k43 gene encodes the Drosophila homolog of the yeast origin recognition complex subunit 2 (Orc2p), a protein required for replication origin function and transcriptional silencing in yeast. These results suggest an evolutionarily conserved role for Orc2p in eukaryotic chromosomal DNA replication.
The Drosophila larva is composed primarily of large polyploid (polytene) cells, which do not divide but increase in size and ploidy during larval development. Most of the larval polytene cells are destroyed during morphogenesis. Also present in the larva are groups of rapidly dividing diploid cells, many in discrete structures called imaginal discs, which will give rise to the structures of the adult fly. The Drosophila k43 gene was first identified as a late-larval lethal mutation with small or missing imaginal discs (1, 2). Subsequent analyses revealed that k43 was required for normal mitosis in the dividing diploid cells of the larva. Diploid cells in k43 larvae had greatly reduced mitotic index, fragmented chromosomes, and abnormal chromosome condensation (3, 4). k43 appears to be a member of a large class of late-larval lethal Drosophila cell cycle genes required for normal mitosis (4). The mutant animals are thought to survive to larval stages due to maternal supply of products sufficient for embryonic mitoses.
The k43 gene was independently identified in the Nusslein-Volhard laboratory by a single female-sterile mutant allele causing abnormally thin and fragile chorions (eggshells). The chorion defect was found to be due to greatly reduced levels of chorion gene amplification during oogenesis (5). During Drosophila oogenesis the chorion gene loci are normally amplified ≈80-fold through repeated initiation of DNA replication at one or a small number of origins located within each of the two chorion gene clusters (6). Chorion gene amplification allows the follicle cells to produce the large amounts of protein required in a short period of time for normal chorion synthesis. Constructs derived from the chorion gene loci will amplify with the correct tissue and temporal specificity when reintroduced into the genome by P element-mediated germ-line transformation (7, 8). This has allowed analysis of cis-sequence requirements for amplification. A 310-bp cis-acting amplification control element, 3rd chromosome (ACE3) (9, 10) is required for high levels of amplification initiating at the nearby major origin of replication, called Ori-β. Ori-β was identified by two-dimensional gel analysis of DNA replication intermediates isolated from the ovarian follicle cells (11, 12). Based upon its mutant phenotypes, the k43 gene was predicted to be a component of the general DNA replication machinery, required for both diploid cell chromosome replication and the function of the chorion gene DNA replication origins (6).
For budding yeast, analysis of DNA replication origin function has advanced rapidly, and has allowed the identification of an origin recognition complex (ORC). The ORC is a complex of six polypeptides (Orc1p-Orc6p), which bind to yeast origins in vitro and in vivo (13–15). The phenotype of mutations in the orc2 gene demonstrates the requirement for Orc2p in DNA replication origin function and in transcriptional silencing of the yeast mating-type loci (16, 17). Similar results have been obtained by analysis of mutations in other ORC subunit genes (18). A Drosophila homolog of the yeast Orc2p subunit was previously identified by sequence homology (19); however, the genomic location and mutant phenotype of the Drosophila gene were unknown.
Here we report the cloning of the Drosophila k43 gene by chromosomal walking. Identification of the k43 gene was confirmed by phenotypic rescue using germ-line transformation, and by sequencing of mutant k43 alleles. The k43 gene was found to encode the Drosophila homolog of the yeast Orc2p subunit. These results suggest an evolutionarily conserved role for Orc2p and the ORC in eukaryotic chromosomal DNA replication.
MATERIALS AND METHODS
Generation of New k43 Alleles.
red e Drosophila males were mutagenized by γ-irradiation (4,000 rads), and crossed to +/TM3 virgin females. Approximately 9,000 red e/TM3 male progeny were crossed individually to k43fs293/TM3 virgin females. red e/k43fs293 female progeny from each cross were scored for fertility. New k43 alleles were recovered from red e/TM3 male siblings, balanced lines were derived, and the homozygous phenotype of the new mutations was examined. The new k43 alleles were characterized for cytologically visible lesions by examination of polytene chromosomes isolated from larval salivary gland cells. Results are summarized in Table 1.
Table 1.
Summary of k43 mutant alleles
| Allele | Mutagen | Cytology | Phenotype | Ref(s). |
|---|---|---|---|---|
| k43fs293 | EMS | Normal | Female-sterile | 5 |
| k431 | EMS | Normal | Lethal | 1–4 |
| k43γ3 | γ-rays | Normal | Lethal | This work |
| k43γ4 | γ-rays | Normal | Lethal | This work |
| k43γ5 | γ-rays | Df 87E2-88A4,6 | Lethal | This work |
| k43γ7 | γ-rays | Df 88A2-88A4 | Lethal | This work |
| Df redP52 | x-rays | Df 88A2,4-88B1,2 | Lethal | 20 |
Chromosome Walking.
The chromosomal walk was initiated with an anonymous cDNA clone, which mapped to the 88B region (clone 88b.1; R.K. and A.C.S., unpublished data). The clone was used as a probe to recover cosmids from a library of Drosophila Canton S genomic DNA inserted into the cosPneo vector (21, 22). Overlapping cosmid clones were obtained spanning approximately 400 kb. The cosmid clones were aligned to several chromosomal deficiency breakpoints by Southern analysis and by in situ hybridization to polytene chromosomes. Results are summarized in Fig. 1.
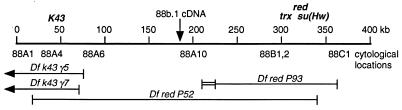
Figure 1.
Diagram of DNA cloned in the chromosomal walk. The 400-kb chromosomal walk is diagrammed. The position of the 88b.1 cDNA used to initiate the walk is indicated with an arrow. The numbers below the line indicate cytological locations determined by in situ hybridization of probes from the walk to polytene chromosomes. The extent of the chromosomal deficiencies are indicated, as are the approximate locations of the k43, red, trx, and su(Hw) genes. The region of overlap between Df k43γ5 and Df redP52 is approximately 50 kb, and the leftmost 20 kb is presented in greater detail in Fig. 2.
Northern Analysis.
Total RNA was isolated from dissected Drosophila ovaries, fractionated on 1.0% agarose/formaldehyde gels, and transferred to GeneScreen membranes (DuPont/NEN). DNA probes were 32P-labeled by random oligomer priming, using the Prime-It II DNA labeling kit (Stratagene). Hybridization signals were visualized by autoradiography.
cDNA Cloning.
Multiple k43 cDNAs were isolated from a commercially available adult Drosophila Canton S cDNA library, cloned in the lambda gt10 vector (CLONTECH). The probe was the 1.5-kb ApaI to EcoRI genomic fragment, designated probe No. 4 in Fig. 2. The largest cDNA was chosen for sequencing.
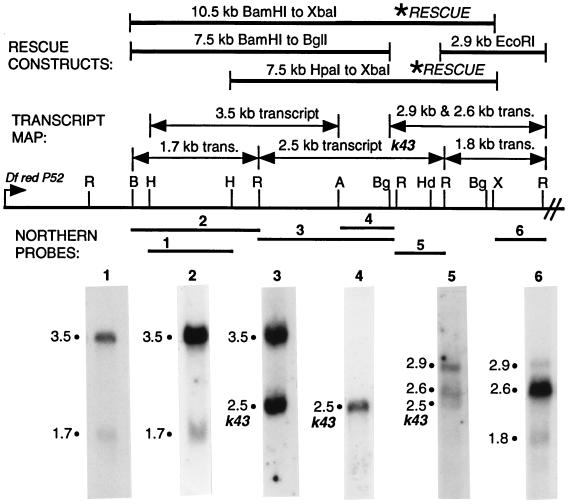
Figure 2.
Restriction map and transcript map of the k43 region. Six ovarian transcripts were identified within the leftmost 16 kb of interval of overlap between Df k43γ5 and Df redP52, as shown. The regions of DNA that hybridize to the various transcripts are summarized as the transcript map, as shown. The four indicated rescue fragments were introduced into the Drosophila genome by P element-mediated transformation. The 10.5-kb BamHI to XbaI fragment and the 7.5-kb HpaI to XbaI fragments, indicated by asterisks, were able to completely rescue the k43 lethal- and female-sterile phenotypes. Multiple independent transformant lines were tested for each construct. As demonstrated by the Northern analysis, only the 2.5-kb transcript is contained entirely within the fragments able to rescue, indicating that this transcript is k43. H, HpaI; R, EcoRI; B, BamHI; A, ApaI; Bg, BglII; Hd, HindIII; X, XbaI.
Measurement of Chorion Gene Amplification.
Amplification of chorion genes was assayed and quantitated as previously described (23).
Electron Microscopy.
Scanning electron microscopy of Drosophila eggs was carried out at the University of Southern California Center for Electron Microscopy and Microanalysis, using a Cambridge 360 SEM. Samples were prepared using standard methods, except that critical point drying was replaced by a 15-min treatment with hexamethyldisilazane (24).
DNA Sequencing.
The k43 cDNA was subcloned into pBluescript (Stratagene) and sequenced using dideoxy sequencing method. For mutant k43 alleles, overlapping segments of the k43 coding region were amplified by PCR from homozygous k43 mutant Drosophila genomic DNA, and subcloned. Dideoxy sequencing was carried out using the Sequenase version 2.0 DNA sequencing kit (United States Biochemical). For k43γ4, six independent clones, three each from two independent PCR reactions, were individually sequenced. For both k43fs293 and its control chromosome, an equimolar pool of six independent clones, three each from two independent PCR reactions, was sequenced. The control chromosome was from a line derived from the same screen in which k43fs293 was isolated, line fs(3)272 (5), which is wild type for k43.
P Element-Mediated Transformation.
Four different restriction fragments from the genomic walk (indicated in Fig. 2) were cloned into the pY.E.S. transformation vector (25), as follows: The 10.5-kb BamHI to XbaI genomic restriction fragment from the walk (Fig. 2) was first subcloned into the unique BamHI–XbaI sites of vector pBS2N, to generate construct pBS2N-BX10.5. The pBS2N vector used is pBluescript II KS+ (Stratagene), in which the unique KpnI site has been converted to a NotI site (gift of L. R. Bell, University of Southern California). Two smaller derivatives of pBS2N-BX10.5 were generated, as follows: pBS2N-BX10.5 was restriction digested with BglII and XbaI, treated with S1 nuclease and religated to generate pBS2N-BB7.5, containing the 7.5-kb BamHI to BglII genomic fragment. pBS2N-BX10.5 was also restriction-digested with BamHI and HpaI, treated with S1 nuclease, and religated to generate construct pBS2N-HX7.5, containing the 7.5-kb HpaI to XbaI genomic fragment. The inserts from each of these three constructs were excised by digestion with NotI, and cloned into the unique NotI site of transformation vector pY.E.S. To generate the 2.9-kb EcoRI rescue construct (Fig. 2), this genomic fragment was first cloned into the unique EcoRI site of pBS2N. The insert was then excised by digestion with NotI and cloned into the unique NotI site of pY.E.S. Multiple germ-line transformants for each of the four rescue constructs were generated using standard methods (26), using the y-ac-w1118 recipient strain (25). Multiple independent insertions for each construct were crossed into k43 lethal- and female-sterile mutant backgrounds, and assayed for phenotypic rescue.
RESULTS
To facilitate cloning by chromosomal walking, new γ-ray induced alleles of k43 were generated by screening for failure to complement the female-sterile k43fs293 allele. Four new k43 alleles were generated (Table 1). Analysis of polytene chromosomes in larval salivary gland cell squashes revealed that one new allele (k43γ5) was a chromosomal deficiency extending from cytological location 87E2 to 88A4,6. A 400-kb genomic walk was initiated from a cDNA clone at 88B to clone the 50-kb interval between the breakpoints of k43γ5 and DfredP52, which defines the region containing k43 as determined by complementation analyses (summarized in Fig. 1). The locations of the chromosomal breakpoints within the walk were determined by Southern analyses and by in situ hybridization of probes from the walk to polytene chromosomes (summarized in Fig. 1). The 50-kb interval containing k43 was restriction-mapped, and transcripts within the region were identified by hybridization of various radiolabeled restriction fragments with Northern blots of ovary RNA. Six transcripts were identified, all of which were located in the leftmost 16 kb of the 50-kb interval containing the k43 gene (Fig. 2). To identify which transcript might correspond to k43, restriction fragments containing one or more transcripts were tested for their ability to rescue the k43 phenotypes by P element-mediated germ-line transformation (summarized in Fig. 2). The smallest fragment tested that was able to rescue the k43 lethal- and female-sterile phenotypes was the 7.5-kb HindIII–XbaI fragment (indicated with an asterisk in Fig. 2). Northern analysis demonstrated that only the 2.5-kb transcript was contained entirely within this fragment (Fig. 2), thus indicating that the 2.5-kb transcript is k43. Consistent with its ability to rescue female fertility, the 7.5-kb HindIII to XbaI rescue fragment was able to restore normal levels of chorion gene amplification to the k43fs293 mutant (Fig. 3), and was able to restore normal chorion morphology (Fig. 4). A nearly full-length cDNA corresponding to the 2.5-kb transcript was cloned and sequenced, and found to contain the entire coding region of the Drosophila homolog of yeast orc2. This gene had been previously cloned by virtue of sequence homology to the yeast gene and called dmOrc2 (19). To confirm the identification of k43, the coding region was PCR-amplified from genomic DNA isolated from larvae homozygous for the k43γ4 lethal allele. DNA sequence analysis revealed a substitution of CT for G at nucleotide position 121, as indicated by underlining in Fig. 5. The numbering is according to the published sequence (19), where +1 is the first nucleotide of the translation initiation codon. The lethal mutation in k43γ4 thus causes a frameshift of the encoded protein beginning at amino acid residue 41. The female-sterile k43fs293 allele was likewise PCR-amplified from genomic DNA isolated from adult flies homozygous for the mutation. As a control, the coding region was amplified from adult flies homozygous for the same third chromosome used in the screen in which k43fs293 was isolated. k43fs293 differed from the control at two positions, which caused a change in the predicted amino acid sequence. The first mutation was nucleotide 961, where a C was substituted for A, resulting in a threonine-321 → proline mutation in the encoded protein. It should be pointed out that both the human and mouse orc2 genes encode a proline at this position (27, 28), suggesting the possibility that this change may not be detrimental to the function of the protein. The second mutation in k43fs293 is at nucleotide 1003, where A is substituted for G, resulting in a glutamate-335 → lysine mutation in the encoded protein.
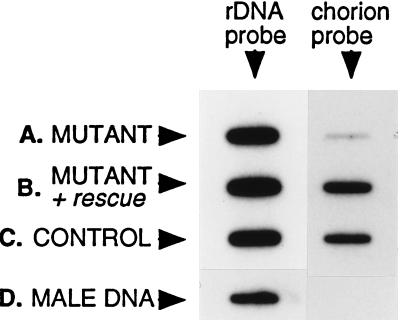
Figure 3.
Phenotypic rescue of the chorion gene amplification defect in k43fs293. Chorion gene amplification was quantitated by isolating DNA from stage 13 egg chambers, and transferring equal portions to identical slot blots. One membrane was hybridized with an rDNA probe, as indicated, which serves as a nonamplifying control for DNA concentration. The second membrane was hybridized with a chorion probe fragment from the third chromosome chorion gene locus, as indicated. (A) Mutant: y ac w; k43fs293. (B) Mutant + rescue: the transgenic 7.5-kb HpaI to XbaI rescue fragment in the k43fs293 background, y ac w; p[yellow+; HX7.5]/+; k43fs293. (C) Control: flies heterozygous for the recessive k43fs293 mutation, y ac w; k43fs293/+. (D) Male DNA: DNA isolated from adult male Drosophila as a nonamplifying control. Amplification was quantitated in this and additional experiments. Amplification in the mutant (A) was ≤7% of control. Amplification in the rescue strain (B) was equal to control.
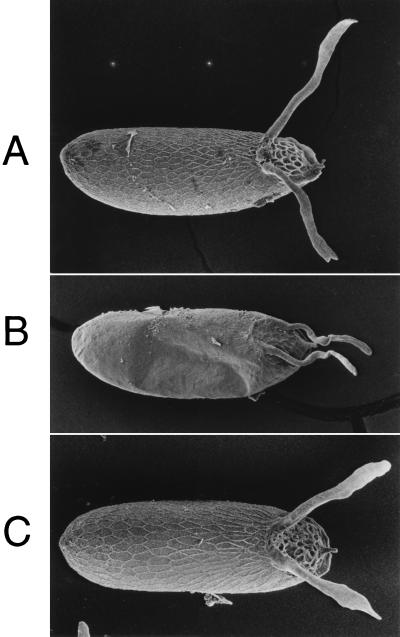
Figure 4.
Phenotypic rescue of the chorion defects in k43fs293. Laid eggs were analyzed by scanning electron microscopy for chorion morphology and are presented in a dorsal view, with anterior to the right. The two large chorion dorsal appendages are apparent at the dorsal anterior (right). (A) Wild-type Oregon R strain. (B) Mutant y ac w; k43fs293. (C) Mutant + rescue y ac w; p[yellow+; HX7.5]/+; k43fs293. The honeycomb-like pattern on the egg surface is the outline of chorion material deposited by each individual follicle cell. Note that in the mutant (B) the amount of chorion material present in the dorsal appendages and surrounding the egg is greatly reduced, and the mutant egg is slightly collapsed. All of these defects are corrected in the rescue strain (C).
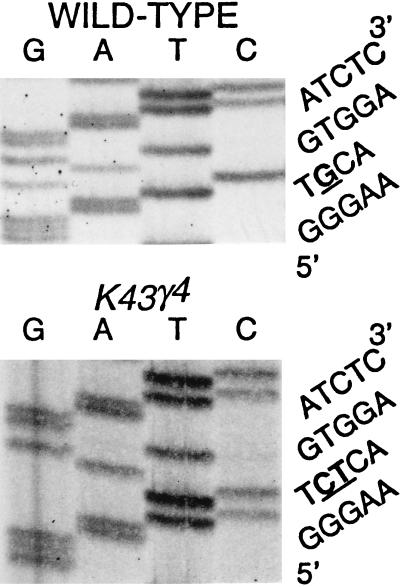
Figure 5.
Sequence analysis of the k43γ4 lethal mutant allele. The k43/dmOrc2p coding region was sequenced in the control (wild type) and k43γ4 mutant backgrounds, as indicated. The sequence of the region from nucleotide 115 to 133 is presented (5′-GGGAATGCAGTGGAATCTC-3′). The k43γ4 allele contains a substitution of CT for the G at position 121, as indicated by underlining. This mutation thus causes a frameshift of the k43γ4-encoded protein beginning at amino acid residue 41.
DISCUSSION
Chorion gene amplification results from the repeated firing of an origin or a small number of origins located within the two chromosomal chorion gene clusters. This process is likely to require at least some of the general DNA replication factors involved in the function of other chromosomal DNA replication origins, at other stages of the life cycle. In addition, there must be amplification-specific activities that limit the process to the follicle cells, and to only the DNA replication origins associated with the chorion gene clusters. k43 gene function is required for normal chorion gene amplification. The recessive female-sterile allele k43fs293 greatly reduces chorion gene amplification, resulting in underproduction of chorion proteins, and thus defective chorions (5). The null phenotype of k43 is late-larval lethality, with defects limited to dividing diploid cells. Homozygous larvae exhibit small or missing imaginal discs, and low mitotic index. The few mitotic cells present exhibit fragmented chromosomes and irregular chromatin condensation (3, 4). Taken together, the k43 mutant phenotypes suggest a role for k43 in chromosomal DNA replication. The determination that k43 encodes the Drosophila homolog of yeast orc2 is consistent with a role in chromosomal DNA replication origin function. In yeast, Orc2p is part of the ORC that binds to both active and inactive chromosomal origins throughout the cell cycle (13, 14). The structure and/or subunit composition of the yeast ORC is altered in correspondence with origin activation (15). Temperature-sensitive mutations in yeast orc2 reduce or eliminate origin firing, and cause increased loss of extrachromosomal plasmids (16, 17, 29–31). Thus for yeast the genetic analyses indicate that the ORC and Orc2p in particular are required for the activity of most or all DNA replication origins. The finding that k43 encodes the Drosophila homolog of yeast Orc2p confirms the hypothesis that Drosophila chorion gene amplification uses at least some of the general DNA replication machinery. These results also demonstrate an evolutionarily conserved role for Orc2p in DNA replication in eukaryotes. This is consistent with recent biochemical evidence that Xenopus homologs of Orc2p and the ORC are required for DNA replication in Xenopus egg extracts (32–34).
In addition to being required for DNA replication, functional yeast DNA replication origins and yeast orc2 both are required for transcriptional silencing at the mating type locus (16, 17, 29–31). Transcriptional silencing requires additional components, some of which are known to be involved in chromatin structure, such as histone H4 (35). It is hypothesized that a functional DNA replication origin and the ORC are required, along with these other factors, to establish a particular chromatin structure that causes silencing at the mating-type locus. The diploid cells of Drosophila k43 lethal mutants also have abnormal chromatin structure, in particular irregular chromatin condensation in mitotic cells (4). The defects in chromatin structure in k43 lethal mutants may be a secondary result of defects in DNA replication, or they may indicate a direct role for Orc2p and the ORC in chromatin structure in Drosophila as well.
The mutant phenotypes suggest that k43 is required for DNA replication in Drosophila, at least in diploid cells and during chorion gene amplification. However, in k43 null mutants the embryonic mitoses are normal, and the animals survive to late-larval stages. The ability of the k43 lethal mutants to survive to late-larval stages is likely due to maternal supply of the k43 gene product, dmOrc2p. The rapid embryonic mitoses in Drosophila appear in general to be supported by maternal products for mitotic cell functions. For example, the phenotype for many mutations affecting the mitotic spindle is late-larval lethality (4). The majority of larval tissues are not diploid, but consist of polyploid (polytene) cells, which continue to increase in size and ploidy throughout larval development. In lethal k43 mutant larvae the polytene tissues are normal, indicating that either maternal dmOrc2p is also sufficient to support polytene cell DNA replication, or that dmOrc2p is not required for polytene cell DNA replication. Further experiments will be required to distinguish between these two possibilities.
An important question is why the k43fs293 female-sterile mutation affects only chorion gene amplification. One possibility is that k43fs293 is a hypomorphic (partial loss of function) mutation, which decreases the ability of the dmOrc2p to function for all DNA replication. This model would require that chorion gene amplification has an especially stringent requirement for dmOrc2p activity, because development and viability are normal in k43fs293. Moreover, in k43fs293 mutant females the follicle cells appear to achieve their normal level of polyploidy (≈16×), yet these same follicle cells fail to support the relatively tiny amount of DNA replication required for chorion gene amplification, which begins even before polyploidization has finished (6). It previously has been determined that the rate of replication fork movement during chorion gene amplification is normal in the k43fs293 mutant, and that the defect in amplification is due to an ≈2-fold decrease in the rate of initiation (36). It may be that chromosomal DNA replication is normal in k43fs293 because the multiple, possibly partially redundant, origins located along the chromosomes are able to tolerate a 2-fold decrease in the rate of initiation without exhibiting a detectable phenotype. In contrast, chorion gene amplification uses a single major origin (Ori-β), with much smaller contributions from 1–3 nearby minor origins (11, 12). Because chorion gene amplification is dependent upon the repeated firing of a single major origin, a 2-fold decrease in the rate of initiation has a profound effect.
An alternative possibility to explain the amplification-specific phenotype of k43fs293 is that the k43fs293 mutation disrupts a domain or function of the dmOrc2p protein that is specific for amplification, such as a site on dmOrc2p that may contact an amplification-specific factor. Consistent with this possibility is the fact that when k43fs293 is placed opposite a chromosomal deficiency for the region (null mutation), a situation that usually increases the severity of hypomorphic phenotypes, no larval-lethality or defects in follicle cell polytene chromosomes are observed. If this second model should prove to be correct, the k43fs293 mutation may be a useful negative control for identifying amplification-specific factors that might contact dmOrc2p, using procedures such as the yeast two-hybrid screen (37).
Finally, it is of interest that there are two positions at which the k43fs293 amino acid sequence differs from both the published sequence (19) and the control chromosome used in this study: a Thr-321 → Pro mutation and a Glu-335 → Lys mutation. It is not possible at this time to determine whether one or both of these changes is responsible for the k43fs293 phenotype. However, the fact that both the human and mouse orc2 genes encode a proline at the position corresponding to residue 321 (27, 28) suggests that the Glu-335 → Lys mutation is more likely to be the relevant alteration.
Acknowledgments
We thank Lucy Lu for providing the data for Fig. 3, Erik T. Bieschke for generating the transgenic flies, and Alicia K. Thompson for assistance with electron microscopy. This research was supported by grants from the U.S. Department of Health and Human Services to A.C.S. (GM27875) and J.T (GM48449).
ABBREVIATION
- ORC
origin recognition complex
References
- 1.Shearn A, Rice T, Garen A, Gehring W. Proc Natl Acad Sci USA. 1971;68:2594–2598. doi: 10.1073/pnas.68.10.2594. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Shearn A, Garen A. Proc Natl Acad Sci USA. 1974;71:1393–1397. doi: 10.1073/pnas.71.4.1393. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Szabad J, Bryant P J. Dev Biol. 1982;93:240–256. doi: 10.1016/0012-1606(82)90256-1. [DOI] [PubMed] [Google Scholar]
- 4.Gatti M, Baker B S. Genes Dev. 1989;3:438–453. doi: 10.1101/gad.3.4.438. [DOI] [PubMed] [Google Scholar]
- 5.Snyder P B, Galanopoulos V K, Kafatos F C. Proc Natl Acad Sci USA. 1986;83:3341–3345. doi: 10.1073/pnas.83.10.3341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Orr-Weaver T L. BioEssays. 1991;13:97–105. doi: 10.1002/bies.950130302. [DOI] [PubMed] [Google Scholar]
- 7.deCicco D V, Spradling A C. Cell. 1984;38:45–54. doi: 10.1016/0092-8674(84)90525-7. [DOI] [PubMed] [Google Scholar]
- 8.Lu, L. & Tower, J. (1997) Mol. Cell. Biol. 17, in press. [DOI] [PMC free article] [PubMed]
- 9.Orr-Weaver T L, Johnston C G, Spradling A C. EMBO J. 1989;8:4153–4162. doi: 10.1002/j.1460-2075.1989.tb08600.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Swimmer S, Delidakis C, Kafatos F C. Proc Natl Acad Sci USA. 1989;86:8823–8827. doi: 10.1073/pnas.86.22.8823. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Delidakis C, Kafatos F C. EMBO J. 1989;8:891–901. doi: 10.1002/j.1460-2075.1989.tb03450.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Heck M M S, Spradling A C. J Cell Biol. 1990;110:903–914. doi: 10.1083/jcb.110.4.903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Bell S P, Stillman B. Nature (London) 1992;357:128–134. doi: 10.1038/357128a0. [DOI] [PubMed] [Google Scholar]
- 14.Diffley J F X, Cocker J H. Nature (London) 1992;357:169–172. doi: 10.1038/357169a0. [DOI] [PubMed] [Google Scholar]
- 15.Diffley J F X, Cocker J H, Dowell S J, Rowley A. Cell. 1994;78:303–316. doi: 10.1016/0092-8674(94)90299-2. [DOI] [PubMed] [Google Scholar]
- 16.Bell S P, Kobayashi R, Stillman B. Science. 1993;262:1844–1849. doi: 10.1126/science.8266072. [DOI] [PubMed] [Google Scholar]
- 17.Foss M, McNally F J, Laurenson P, Rine J. Science. 1993;262:1838–1844. doi: 10.1126/science.8266071. [DOI] [PubMed] [Google Scholar]
- 18.Diffley J F X. Genes Dev. 1996;10:2819–2830. doi: 10.1101/gad.10.22.2819. [DOI] [PubMed] [Google Scholar]
- 19.Gossen M, Pak D T, Hansen J K, Acharya J K, Botchan M. Science. 1995;270:1674–1677. doi: 10.1126/science.270.5242.1674. [DOI] [PubMed] [Google Scholar]
- 20.Lindsley D L, Zimm G G. The Genome of Drosophila melanogaster. San Diego: Academic; 1992. p. 872. [Google Scholar]
- 21.Cooley L, Kelley R, Spradling A C. Science. 1988;239:1121–1128. doi: 10.1126/science.2830671. [DOI] [PubMed] [Google Scholar]
- 22.Steller H, Pirrotta V. EMBO J. 1985;4:167–172. doi: 10.1002/j.1460-2075.1985.tb02332.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Underwood E M, Briot A S, Doll K Z, Ludwiczak R L, Otteson D C, Tower J, Vessey K B, Yu K. Genetics. 1990;126:639–650. doi: 10.1093/genetics/126.3.639. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Adams J L, Battjes C J, Buthala D A. In: Proceedings of the 45th Annual Meeting of the Electron Microscopy Society of America. Baily G W, editor. San Francisco: San Francisco Press; 1987. p. 956. [Google Scholar]
- 25.Patton J S, Gomes X V, Geyer P K. Nucleic Acids Res. 1992;20:5859–5860. doi: 10.1093/nar/20.21.5859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Rubin G M, Spradling A C. Science. 1982;218:348–353. doi: 10.1126/science.6289436. [DOI] [PubMed] [Google Scholar]
- 27.Takahara K, Bong M, Brevard R, Eddy R L, Haley L L, Sait S S, Shows T B, Hoffman G G, Greenspan D S. Genomics. 1996;31:119–122. doi: 10.1006/geno.1996.0018. [DOI] [PubMed] [Google Scholar]
- 28.Gavin K A, Hidaka M, Stillman B. Science. 1995;270:1667–1671. doi: 10.1126/science.270.5242.1667. [DOI] [PubMed] [Google Scholar]
- 29.Micklem G, Rowley A, Harwood J, Nasmyth K, Diffley J F X. Nature (London) 1993;366:87–89. doi: 10.1038/366087a0. [DOI] [PubMed] [Google Scholar]
- 30.Fox C A, Loo S, Dillin A, Rine J. Genes Dev. 1995;9:911–924. doi: 10.1101/gad.9.8.911. [DOI] [PubMed] [Google Scholar]
- 31.Liang C, Weinreich M, Stillman B. Cell. 1995;81:667–676. doi: 10.1016/0092-8674(95)90528-6. [DOI] [PubMed] [Google Scholar]
- 32.Carpenter P B, Mueller P R, Dunphy W G. Nature (London) 1996;379:357–360. doi: 10.1038/379357a0. [DOI] [PubMed] [Google Scholar]
- 33.Rowles A, Chong J P J, Brown L, Howell M, Evan G I, Blow J J. Cell. 1996;87:287–296. doi: 10.1016/s0092-8674(00)81346-x. [DOI] [PubMed] [Google Scholar]
- 34.Romanowski P, Madine M A, Rowles A, Blow J J, Laskey R A. Curr Biol. 1996;6:1416–1425. doi: 10.1016/s0960-9822(96)00746-4. [DOI] [PubMed] [Google Scholar]
- 35.Kayne P S, Kim U-J, Han M, Mullen J R, Yoshizaki F, Grunstein M. Cell. 1988;55:27–39. doi: 10.1016/0092-8674(88)90006-2. [DOI] [PubMed] [Google Scholar]
- 36.Leys E, Spradling A C. Cancer Cells 6/Eukaryotic DNA Replication. Plainview, NY: Cold Spring Harbor Lab. Press; 1988. pp. 305–309. [Google Scholar]
- 37.Hardy C F J. Mol Cell Biol. 1996;16:1832–1841. doi: 10.1128/mcb.16.4.1832. [DOI] [PMC free article] [PubMed] [Google Scholar]