Abstract
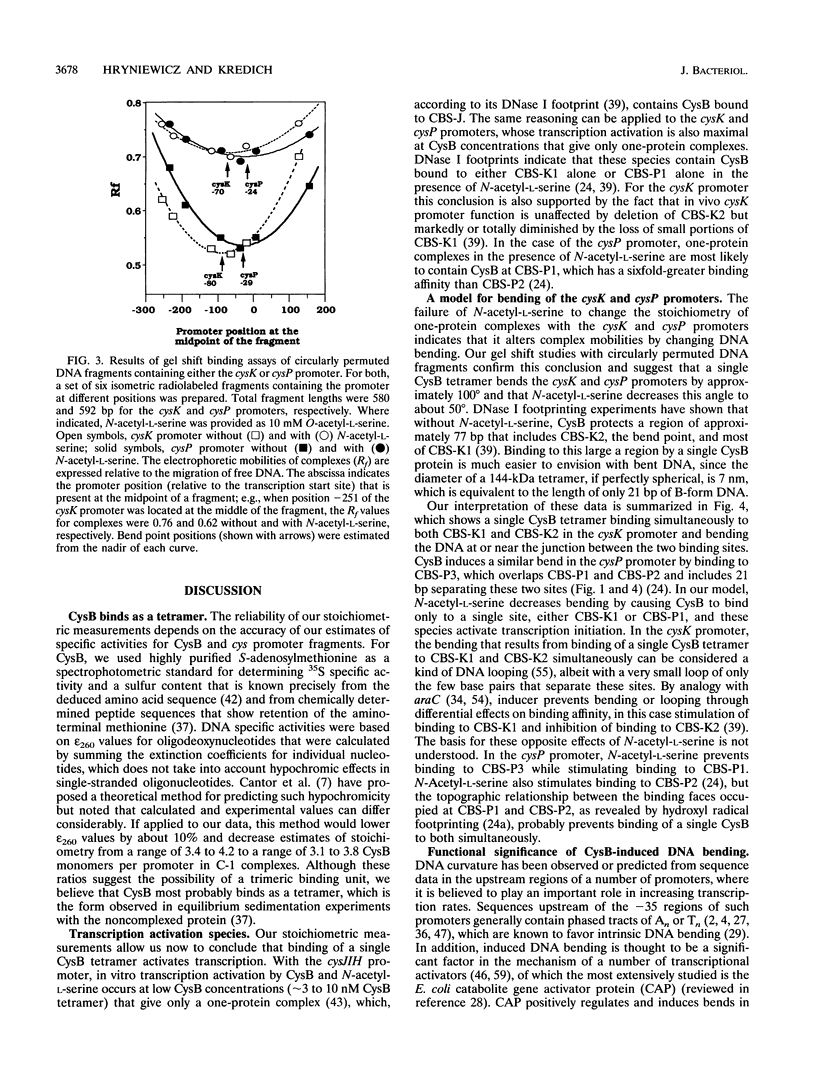
CysB is a member of the LysR family of transcriptional activators and regulates genes of the cysteine regulon in Salmonella typhimurium and Escherichia coli. CysB binds to specific sites just upstream of the -35 regions of the cysJIH, cysK, and cysP promoters, where, in the presence of N-acetyl-L-serine, it stimulates transcription initiation. The cysK and cysP promoters contain additional binding sites, and we have proposed that CysB bends these promoters by binding to adjacent sites. N-Acetyl-L-serine is thought to decrease the magnitude of such bending. Since stoichiometric data bearing on this model have been lacking, we analyzed complexes in gel mobility shift experiments with 35S-labeled CysB and 32P-labeled promoter fragments. CysB was found to bind as a tetramer, and N-acetyl-L-serine increased the electrophoretic mobilities of one-protein complexes of the multibinding site cysK and cysP promoters without changing their stoichiometry, indicating that a single CysB tetramer can bend these promoters and that N-acetyl-L-serine diminishes such bending. Bend angles for both promoters were calculated to be 100 and 50 degrees in the absence and presence of N-acetyl-L-serine. N-Acetyl-L-serine affected neither the stoichiometry nor the electrophoretic mobility of cysJIH promoter complexes, which are not known to contain bent DNA. DNA bending may be a mechanism for sequestering CysB at certain promoter sites by increasing their affinity for this protein in the absence of N-acetyl-L-serine.
Full text
PDF


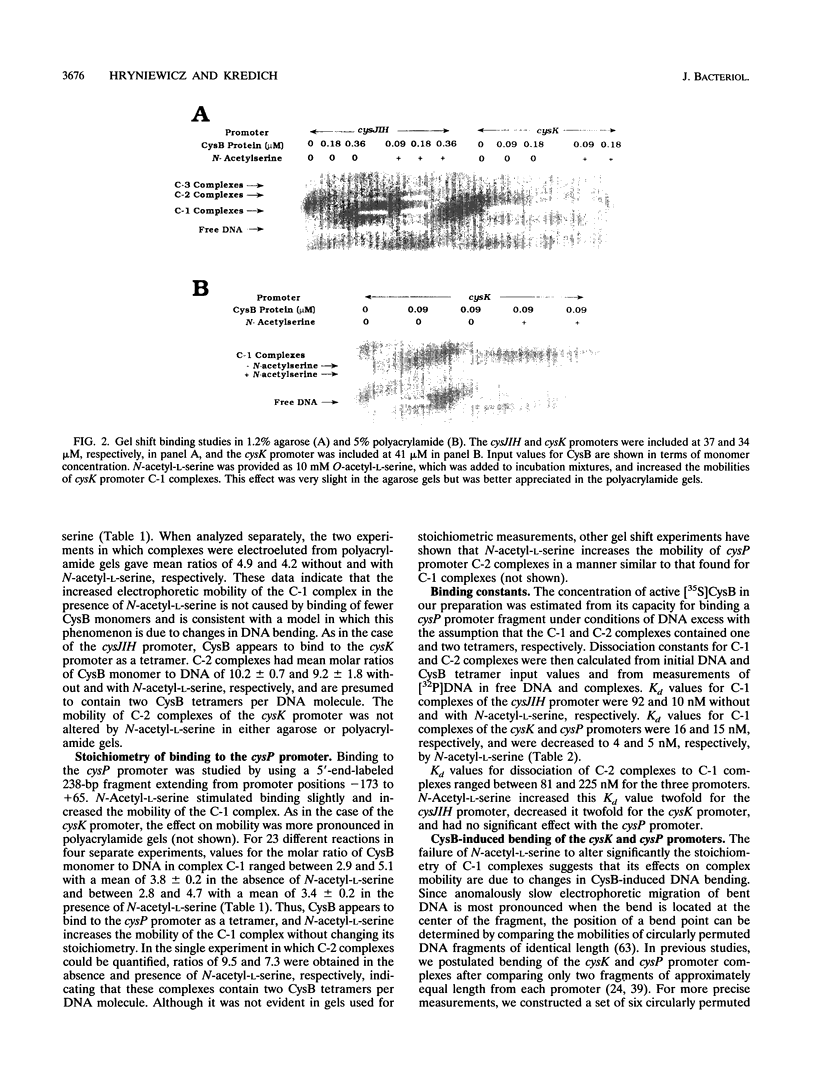






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- BOCK R. M., LING N. S., MORELL S. A., LIPTON S. H. Ultraviolet absorption spectra of adenosine-5'-triphosphate and related 5'-ribonucleotides. Arch Biochem Biophys. 1956 Jun;62(2):253–264. doi: 10.1016/0003-9861(56)90123-0. [DOI] [PubMed] [Google Scholar]
- Banner C. D., Moran C. P., Jr, Losick R. Deletion analysis of a complex promoter for a developmentally regulated gene from Bacillus subtilis. J Mol Biol. 1983 Aug 5;168(2):351–365. doi: 10.1016/s0022-2836(83)80023-0. [DOI] [PubMed] [Google Scholar]
- Bertrand-Burggraf E., Dunand J., Fuchs R. P., Lefèvre J. F. Kinetic studies of the modulation of ada promoter activity by upstream elements. EMBO J. 1990 Jul;9(7):2265–2271. doi: 10.1002/j.1460-2075.1990.tb07397.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bossi L., Smith D. M. Conformational change in the DNA associated with an unusual promoter mutation in a tRNA operon of Salmonella. Cell. 1984 Dec;39(3 Pt 2):643–652. doi: 10.1016/0092-8674(84)90471-9. [DOI] [PubMed] [Google Scholar]
- Bracco L., Kotlarz D., Kolb A., Diekmann S., Buc H. Synthetic curved DNA sequences can act as transcriptional activators in Escherichia coli. EMBO J. 1989 Dec 20;8(13):4289–4296. doi: 10.1002/j.1460-2075.1989.tb08615.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Byrne C. R., Monroe R. S., Ward K. A., Kredich N. M. DNA sequences of the cysK regions of Salmonella typhimurium and Escherichia coli and linkage of the cysK regions to ptsH. J Bacteriol. 1988 Jul;170(7):3150–3157. doi: 10.1128/jb.170.7.3150-3157.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cantor C. R., Warshaw M. M., Shapiro H. Oligonucleotide interactions. 3. Circular dichroism studies of the conformation of deoxyoligonucleotides. Biopolymers. 1970;9(9):1059–1077. doi: 10.1002/bip.1970.360090909. [DOI] [PubMed] [Google Scholar]
- Chang M., Crawford I. P. In vitro determination of the effect of indoleglycerol phosphate on the interaction of purified TrpI protein with its DNA-binding sites. J Bacteriol. 1991 Mar;173(5):1590–1597. doi: 10.1128/jb.173.5.1590-1597.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deuschle U., Kammerer W., Gentz R., Bujard H. Promoters of Escherichia coli: a hierarchy of in vivo strength indicates alternate structures. EMBO J. 1986 Nov;5(11):2987–2994. doi: 10.1002/j.1460-2075.1986.tb04596.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eschenlauer A. C., Reznikoff W. S. Escherichia coli catabolite gene activator protein mutants defective in positive control of lac operon transcription. J Bacteriol. 1991 Aug;173(16):5024–5029. doi: 10.1128/jb.173.16.5024-5029.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fisher R. F., Long S. R. DNA footprint analysis of the transcriptional activator proteins NodD1 and NodD3 on inducible nod gene promoters. J Bacteriol. 1989 Oct;171(10):5492–5502. doi: 10.1128/jb.171.10.5492-5502.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fisher R. F., Long S. R. Interactions of NodD at the nod Box: NodD binds to two distinct sites on the same face of the helix and induces a bend in the DNA. J Mol Biol. 1993 Oct 5;233(3):336–348. doi: 10.1006/jmbi.1993.1515. [DOI] [PubMed] [Google Scholar]
- Fried M., Crothers D. M. Equilibria and kinetics of lac repressor-operator interactions by polyacrylamide gel electrophoresis. Nucleic Acids Res. 1981 Dec 11;9(23):6505–6525. doi: 10.1093/nar/9.23.6505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Frisby D., Zuber P. Analysis of the upstream activating sequence and site of carbon and nitrogen source repression in the promoter of an early-induced sporulation gene of Bacillus subtilis. J Bacteriol. 1991 Dec;173(23):7557–7564. doi: 10.1128/jb.173.23.7557-7564.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Galas D. J., Eggert M., Waterman M. S. Rigorous pattern-recognition methods for DNA sequences. Analysis of promoter sequences from Escherichia coli. J Mol Biol. 1985 Nov 5;186(1):117–128. doi: 10.1016/0022-2836(85)90262-1. [DOI] [PubMed] [Google Scholar]
- Gao J., Gussin G. N. Mutations in TrpI binding site II that differentially affect activation of the trpBA promoter of Pseudomonas aeruginosa. EMBO J. 1991 Dec;10(13):4137–4144. doi: 10.1002/j.1460-2075.1991.tb04991.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garner M. M., Revzin A. A gel electrophoresis method for quantifying the binding of proteins to specific DNA regions: application to components of the Escherichia coli lactose operon regulatory system. Nucleic Acids Res. 1981 Jul 10;9(13):3047–3060. doi: 10.1093/nar/9.13.3047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gartenberg M. R., Crothers D. M. Synthetic DNA bending sequences increase the rate of in vitro transcription initiation at the Escherichia coli lac promoter. J Mol Biol. 1991 May 20;219(2):217–230. doi: 10.1016/0022-2836(91)90563-l. [DOI] [PubMed] [Google Scholar]
- German D. C., Bloch C. A., Kredich N. M. Measurements of S-adenosylmethionine and L-homocysteine metabolism in cultured human lymphoid cells. J Biol Chem. 1983 Sep 25;258(18):10997–11003. [PubMed] [Google Scholar]
- Goethals K., Van Montagu M., Holsters M. Conserved motifs in a divergent nod box of Azorhizobium caulinodans ORS571 reveal a common structure in promoters regulated by LysR-type proteins. Proc Natl Acad Sci U S A. 1992 Mar 1;89(5):1646–1650. doi: 10.1073/pnas.89.5.1646. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gourse R. L., de Boer H. A., Nomura M. DNA determinants of rRNA synthesis in E. coli: growth rate dependent regulation, feedback inhibition, upstream activation, antitermination. Cell. 1986 Jan 17;44(1):197–205. doi: 10.1016/0092-8674(86)90498-8. [DOI] [PubMed] [Google Scholar]
- Habeeb L. F., Wang L., Winans S. C. Transcription of the octopine catabolism operon of the Agrobacterium tumor-inducing plasmid pTiA6 is activated by a LysR-type regulatory protein. Mol Plant Microbe Interact. 1991 Jul-Aug;4(4):379–385. doi: 10.1094/mpmi-4-379. [DOI] [PubMed] [Google Scholar]
- Henikoff S., Haughn G. W., Calvo J. M., Wallace J. C. A large family of bacterial activator proteins. Proc Natl Acad Sci U S A. 1988 Sep;85(18):6602–6606. doi: 10.1073/pnas.85.18.6602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hryniewicz M. M., Kredich N. M. The cysP promoter of Salmonella typhimurium: characterization of two binding sites for CysB protein, studies of in vivo transcription initiation, and demonstration of the anti-inducer effects of thiosulfate. J Bacteriol. 1991 Sep;173(18):5876–5886. doi: 10.1128/jb.173.18.5876-5886.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang J. Z., Schell M. A. In vivo interactions of the NahR transcriptional activator with its target sequences. Inducer-mediated changes resulting in transcription activation. J Biol Chem. 1991 Jun 15;266(17):10830–10838. [PubMed] [Google Scholar]
- Igarashi K., Ishihama A. Bipartite functional map of the E. coli RNA polymerase alpha subunit: involvement of the C-terminal region in transcription activation by cAMP-CRP. Cell. 1991 Jun 14;65(6):1015–1022. doi: 10.1016/0092-8674(91)90553-b. [DOI] [PubMed] [Google Scholar]
- Jo Y. L., Nara F., Ichihara S., Mizuno T., Mizushima S. Purification and characterization of the OmpR protein, a positive regulator involved in osmoregulatory expression of the ompF and ompC genes in Escherichia coli. J Biol Chem. 1986 Nov 15;261(32):15252–15256. [PubMed] [Google Scholar]
- Kolb A., Busby S., Buc H., Garges S., Adhya S. Transcriptional regulation by cAMP and its receptor protein. Annu Rev Biochem. 1993;62:749–795. doi: 10.1146/annurev.bi.62.070193.003533. [DOI] [PubMed] [Google Scholar]
- Koo H. S., Wu H. M., Crothers D. M. DNA bending at adenine . thymine tracts. Nature. 1986 Apr 10;320(6062):501–506. doi: 10.1038/320501a0. [DOI] [PubMed] [Google Scholar]
- Kotb M., Kredich N. M. S-Adenosylmethionine synthetase from human lymphocytes. Purification and characterization. J Biol Chem. 1985 Apr 10;260(7):3923–3930. [PubMed] [Google Scholar]
- Kredich N. M. The molecular basis for positive regulation of cys promoters in Salmonella typhimurium and Escherichia coli. Mol Microbiol. 1992 Oct;6(19):2747–2753. doi: 10.1111/j.1365-2958.1992.tb01453.x. [DOI] [PubMed] [Google Scholar]
- Lindquist S., Lindberg F., Normark S. Binding of the Citrobacter freundii AmpR regulator to a single DNA site provides both autoregulation and activation of the inducible ampC beta-lactamase gene. J Bacteriol. 1989 Jul;171(7):3746–3753. doi: 10.1128/jb.171.7.3746-3753.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lobell R. B., Schleif R. F. DNA looping and unlooping by AraC protein. Science. 1990 Oct 26;250(4980):528–532. doi: 10.1126/science.2237403. [DOI] [PubMed] [Google Scholar]
- Markham G. D., Hafner E. W., Tabor C. W., Tabor H. S-Adenosylmethionine synthetase from Escherichia coli. J Biol Chem. 1980 Oct 10;255(19):9082–9092. [PubMed] [Google Scholar]
- McAllister C. F., Achberger E. C. Effect of polyadenine-containing curved DNA on promoter utilization in Bacillus subtilis. J Biol Chem. 1988 Aug 25;263(24):11743–11749. [PubMed] [Google Scholar]
- Miller B. E., Kredich N. M. Purification of the cysB protein from Salmonella typhimurium. J Biol Chem. 1987 May 5;262(13):6006–6009. [PubMed] [Google Scholar]
- Monroe R. S., Kredich N. M. Isolation of Salmonella typhimurium cys genes by transduction with a library of recombinant plasmids packaged in bacteriophage P22HT capsids. J Bacteriol. 1988 Jan;170(1):42–47. doi: 10.1128/jb.170.1.42-47.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Monroe R. S., Ostrowski J., Hryniewicz M. M., Kredich N. M. In vitro interactions of CysB protein with the cysK and cysJIH promoter regions of Salmonella typhimurium. J Bacteriol. 1990 Dec;172(12):6919–6929. doi: 10.1128/jb.172.12.6919-6929.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Newlands J. T., Josaitis C. A., Ross W., Gourse R. L. Both fis-dependent and factor-independent upstream activation of the rrnB P1 promoter are face of the helix dependent. Nucleic Acids Res. 1992 Feb 25;20(4):719–726. doi: 10.1093/nar/20.4.719. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Newlands J. T., Ross W., Gosink K. K., Gourse R. L. Factor-independent activation of Escherichia coli rRNA transcription. II. characterization of complexes of rrnB P1 promoters containing or lacking the upstream activator region with Escherichia coli RNA polymerase. J Mol Biol. 1991 Aug 5;220(3):569–583. doi: 10.1016/0022-2836(91)90101-b. [DOI] [PubMed] [Google Scholar]
- Ostrowski J., Jagura-Burdzy G., Kredich N. M. DNA sequences of the cysB regions of Salmonella typhimurium and Escherichia coli. J Biol Chem. 1987 May 5;262(13):5999–6005. [PubMed] [Google Scholar]
- Ostrowski J., Kredich N. M. In vitro interactions of CysB protein with the cysJIH promoter of Salmonella typhimurium: inhibitory effects of sulfide. J Bacteriol. 1990 Feb;172(2):779–785. doi: 10.1128/jb.172.2.779-785.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ostrowski J., Kredich N. M. Molecular characterization of the cysJIH promoters of Salmonella typhimurium and Escherichia coli: regulation by cysB protein and N-acetyl-L-serine. J Bacteriol. 1989 Jan;171(1):130–140. doi: 10.1128/jb.171.1.130-140.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ostrowski J., Kredich N. M. Negative autoregulation of cysB in Salmonella typhimurium: in vitro interactions of CysB protein with the cysB promoter. J Bacteriol. 1991 Apr;173(7):2212–2218. doi: 10.1128/jb.173.7.2212-2218.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Plaskon R. R., Wartell R. M. Sequence distributions associated with DNA curvature are found upstream of strong E. coli promoters. Nucleic Acids Res. 1987 Jan 26;15(2):785–796. doi: 10.1093/nar/15.2.785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Prentki P., Pham M. H., Galas D. J. Plasmid permutation vectors to monitor DNA bending. Nucleic Acids Res. 1987 Dec 10;15(23):10060–10060. doi: 10.1093/nar/15.23.10060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pérez-Martín J., Espinosa M. Protein-induced bending as a transcriptional switch. Science. 1993 May 7;260(5109):805–807. doi: 10.1126/science.8387228. [DOI] [PubMed] [Google Scholar]
- Revzin A. Gel electrophoresis assays for DNA-protein interactions. Biotechniques. 1989 Apr;7(4):346–355. [PubMed] [Google Scholar]
- Ross W., Gosink K. K., Salomon J., Igarashi K., Zou C., Ishihama A., Severinov K., Gourse R. L. A third recognition element in bacterial promoters: DNA binding by the alpha subunit of RNA polymerase. Science. 1993 Nov 26;262(5138):1407–1413. doi: 10.1126/science.8248780. [DOI] [PubMed] [Google Scholar]
- Schell M. A. Molecular biology of the LysR family of transcriptional regulators. Annu Rev Microbiol. 1993;47:597–626. doi: 10.1146/annurev.mi.47.100193.003121. [DOI] [PubMed] [Google Scholar]
- Schell M. A., Poser E. F. Demonstration, characterization, and mutational analysis of NahR protein binding to nah and sal promoters. J Bacteriol. 1989 Feb;171(2):837–846. doi: 10.1128/jb.171.2.837-846.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schleif R. DNA looping. Annu Rev Biochem. 1992;61:199–223. doi: 10.1146/annurev.bi.61.070192.001215. [DOI] [PubMed] [Google Scholar]
- Tartaglia L. A., Gimeno C. J., Storz G., Ames B. N. Multidegenerate DNA recognition by the OxyR transcriptional regulator. J Biol Chem. 1992 Jan 25;267(3):2038–2045. [PubMed] [Google Scholar]
- Tartaglia L. A., Storz G., Ames B. N. Identification and molecular analysis of oxyR-regulated promoters important for the bacterial adaptation to oxidative stress. J Mol Biol. 1989 Dec 20;210(4):709–719. doi: 10.1016/0022-2836(89)90104-6. [DOI] [PubMed] [Google Scholar]
- Thompson J. F., Landy A. Empirical estimation of protein-induced DNA bending angles: applications to lambda site-specific recombination complexes. Nucleic Acids Res. 1988 Oct 25;16(20):9687–9705. doi: 10.1093/nar/16.20.9687. [DOI] [PMC free article] [PubMed] [Google Scholar]
- VOGEL H. J., BONNER D. M. Acetylornithinase of Escherichia coli: partial purification and some properties. J Biol Chem. 1956 Jan;218(1):97–106. [PubMed] [Google Scholar]
- Wang S. P., Stacey G. Studies of the Bradyrhizobium japonicum nodD1 promoter: a repeated structure for the nod box. J Bacteriol. 1991 Jun;173(11):3356–3365. doi: 10.1128/jb.173.11.3356-3365.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu H. M., Crothers D. M. The locus of sequence-directed and protein-induced DNA bending. Nature. 1984 Apr 5;308(5959):509–513. doi: 10.1038/308509a0. [DOI] [PubMed] [Google Scholar]
- Zhou Y., Zhang X., Ebright R. H. Identification of the activating region of catabolite gene activator protein (CAP): isolation and characterization of mutants of CAP specifically defective in transcription activation. Proc Natl Acad Sci U S A. 1993 Jul 1;90(13):6081–6085. doi: 10.1073/pnas.90.13.6081. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Vliet P. C., Verrijzer C. P. Bending of DNA by transcription factors. Bioessays. 1993 Jan;15(1):25–32. doi: 10.1002/bies.950150105. [DOI] [PubMed] [Google Scholar]




