Abstract
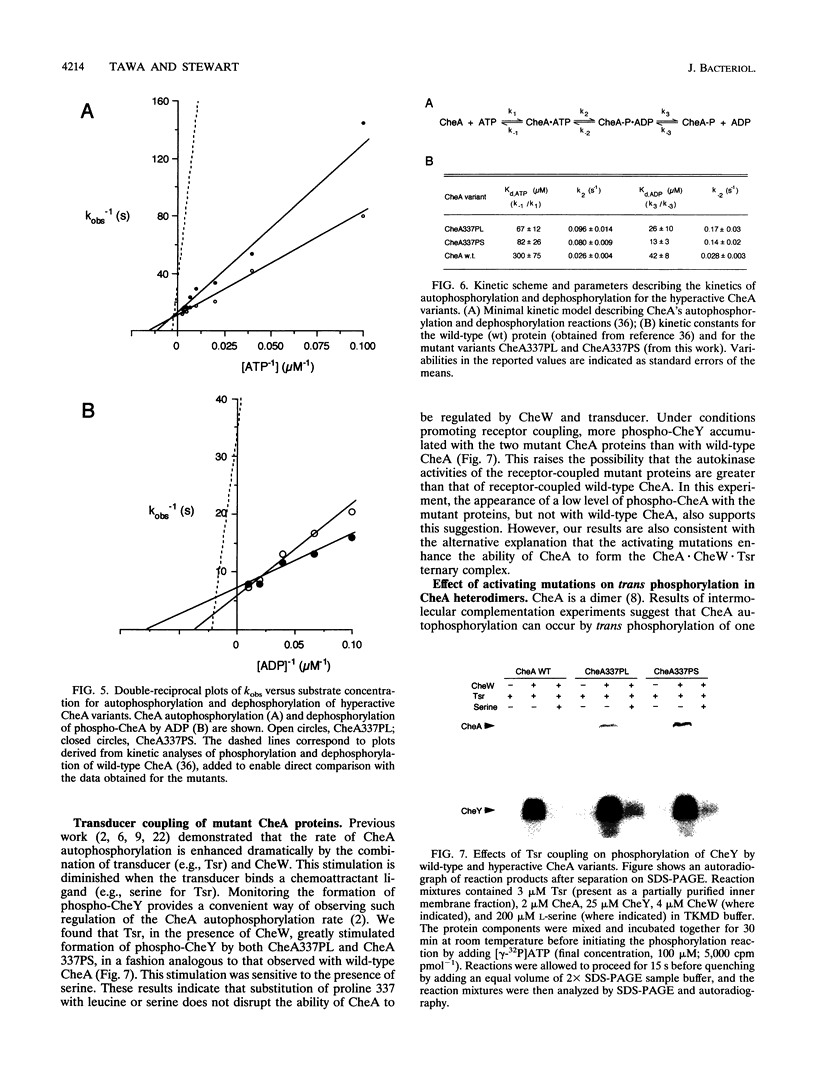
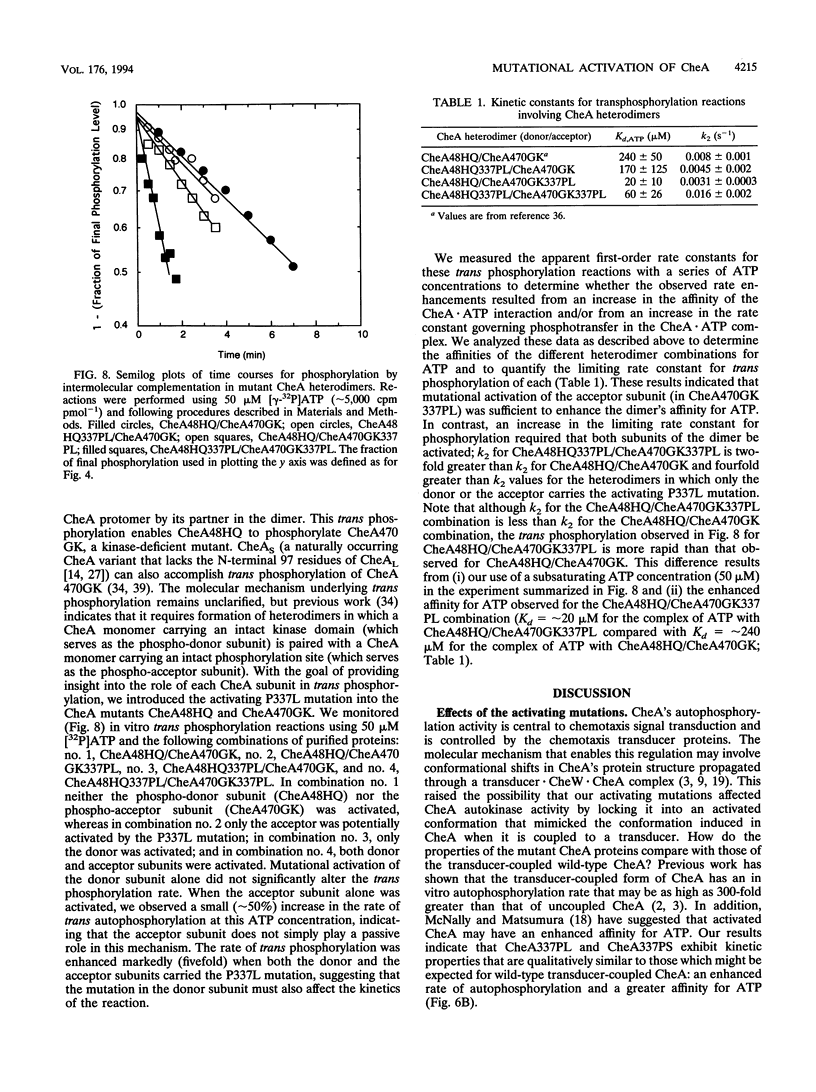
In Escherichia coli and Salmonella typhimurium, appropriate changes of cell swimming patterns are mediated by CheA, an autophosphorylating histidine protein kinase whose activity is regulated by receptor/transducer proteins. The molecular mechanism underlying this regulation remains unelucidated but may involve CheA shifting between high-activity and low-activity conformations. We devised an in vivo screen to search for potential hyperkinase variants of CheA and used this screen to identify two cheA point mutations that cause the CheA protein to have elevated autokinase activity. Each point mutation resulted in alteration of proline 337. In vitro, CheA337PL and CheA337PS autophosphorylated significantly more rapidly than did wild-type CheA. This rate enhancement reflected the higher affinities of the mutant proteins for ATP and an increased rate constant for acquisition by CheA of the gamma-phosphoryl group of ATP within a kinetically defined CheA.ATP complex. In addition, the mutant proteins reacted with ADP more rapidly than did wild-type CheA. We considered the possibility that the mutations served to lock CheA into an activated signaling conformation; however, we found that both mutant proteins were regulated in a normal fashion by the transducer Tsr in the presence of CheW. We exploited the activated properties of one of these mutants to investigate whether the CheA subunits within a CheA dimer make equivalent contributions to the mechanism of trans phosphorylation. Our results indicate that CheA trans phosphorylation may involve active-site residues that are located both in cis and in trans to the autophosphorylation site and that the two protomers of a CheA dimer make nonequivalent contributions in determining the affinity of the ATP-binding site(s) of CheA.
Full text
PDF




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ames P., Parkinson J. S. Transmembrane signaling by bacterial chemoreceptors: E. coli transducers with locked signal output. Cell. 1988 Dec 2;55(5):817–826. doi: 10.1016/0092-8674(88)90137-7. [DOI] [PubMed] [Google Scholar]
- Borkovich K. A., Kaplan N., Hess J. F., Simon M. I. Transmembrane signal transduction in bacterial chemotaxis involves ligand-dependent activation of phosphate group transfer. Proc Natl Acad Sci U S A. 1989 Feb;86(4):1208–1212. doi: 10.1073/pnas.86.4.1208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Borkovich K. A., Simon M. I. The dynamics of protein phosphorylation in bacterial chemotaxis. Cell. 1990 Dec 21;63(6):1339–1348. doi: 10.1016/0092-8674(90)90429-i. [DOI] [PubMed] [Google Scholar]
- Bourret R. B., Borkovich K. A., Simon M. I. Signal transduction pathways involving protein phosphorylation in prokaryotes. Annu Rev Biochem. 1991;60:401–441. doi: 10.1146/annurev.bi.60.070191.002153. [DOI] [PubMed] [Google Scholar]
- Bourret R. B., Davagnino J., Simon M. I. The carboxy-terminal portion of the CheA kinase mediates regulation of autophosphorylation by transducer and CheW. J Bacteriol. 1993 Apr;175(7):2097–2101. doi: 10.1128/jb.175.7.2097-2101.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bourret R. B., Hess J. F., Simon M. I. Conserved aspartate residues and phosphorylation in signal transduction by the chemotaxis protein CheY. Proc Natl Acad Sci U S A. 1990 Jan;87(1):41–45. doi: 10.1073/pnas.87.1.41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fuhrer D. K., Ordal G. W. Bacillus subtilis CheN, a homolog of CheA, the central regulator of chemotaxis in Escherichia coli. J Bacteriol. 1991 Dec;173(23):7443–7448. doi: 10.1128/jb.173.23.7443-7448.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gegner J. A., Dahlquist F. W. Signal transduction in bacteria: CheW forms a reversible complex with the protein kinase CheA. Proc Natl Acad Sci U S A. 1991 Feb 1;88(3):750–754. doi: 10.1073/pnas.88.3.750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gegner J. A., Graham D. R., Roth A. F., Dahlquist F. W. Assembly of an MCP receptor, CheW, and kinase CheA complex in the bacterial chemotaxis signal transduction pathway. Cell. 1992 Sep 18;70(6):975–982. doi: 10.1016/0092-8674(92)90247-a. [DOI] [PubMed] [Google Scholar]
- Hess J. F., Bourret R. B., Simon M. I. Histidine phosphorylation and phosphoryl group transfer in bacterial chemotaxis. Nature. 1988 Nov 10;336(6195):139–143. doi: 10.1038/336139a0. [DOI] [PubMed] [Google Scholar]
- Hess J. F., Oosawa K., Kaplan N., Simon M. I. Phosphorylation of three proteins in the signaling pathway of bacterial chemotaxis. Cell. 1988 Apr 8;53(1):79–87. doi: 10.1016/0092-8674(88)90489-8. [DOI] [PubMed] [Google Scholar]
- Hess J. F., Oosawa K., Matsumura P., Simon M. I. Protein phosphorylation is involved in bacterial chemotaxis. Proc Natl Acad Sci U S A. 1987 Nov;84(21):7609–7613. doi: 10.1073/pnas.84.21.7609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kalman L. V., Gunsalus R. P. Nitrate- and molybdenum-independent signal transduction mutations in narX that alter regulation of anaerobic respiratory genes in Escherichia coli. J Bacteriol. 1990 Dec;172(12):7049–7056. doi: 10.1128/jb.172.12.7049-7056.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kofoid E. C., Parkinson J. S. Tandem translation starts in the cheA locus of Escherichia coli. J Bacteriol. 1991 Mar;173(6):2116–2119. doi: 10.1128/jb.173.6.2116-2119.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larsen S. H., Reader R. W., Kort E. N., Tso W. W., Adler J. Change in direction of flagellar rotation is the basis of the chemotactic response in Escherichia coli. Nature. 1974 May 3;249(452):74–77. doi: 10.1038/249074a0. [DOI] [PubMed] [Google Scholar]
- Liu J. D., Parkinson J. S. Role of CheW protein in coupling membrane receptors to the intracellular signaling system of bacterial chemotaxis. Proc Natl Acad Sci U S A. 1989 Nov;86(22):8703–8707. doi: 10.1073/pnas.86.22.8703. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lupas A., Stock J. Phosphorylation of an N-terminal regulatory domain activates the CheB methylesterase in bacterial chemotaxis. J Biol Chem. 1989 Oct 15;264(29):17337–17342. [PubMed] [Google Scholar]
- McNally D. F., Matsumura P. Bacterial chemotaxis signaling complexes: formation of a CheA/CheW complex enhances autophosphorylation and affinity for CheY. Proc Natl Acad Sci U S A. 1991 Jul 15;88(14):6269–6273. doi: 10.1073/pnas.88.14.6269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Milburn M. V., Privé G. G., Milligan D. L., Scott W. G., Yeh J., Jancarik J., Koshland D. E., Jr, Kim S. H. Three-dimensional structures of the ligand-binding domain of the bacterial aspartate receptor with and without a ligand. Science. 1991 Nov 29;254(5036):1342–1347. doi: 10.1126/science.1660187. [DOI] [PubMed] [Google Scholar]
- Miller J. F., Johnson S. A., Black W. J., Beattie D. T., Mekalanos J. J., Falkow S. Constitutive sensory transduction mutations in the Bordetella pertussis bvgS gene. J Bacteriol. 1992 Feb;174(3):970–979. doi: 10.1128/jb.174.3.970-979.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Milligan D. L., Koshland D. E., Jr Intrasubunit signal transduction by the aspartate chemoreceptor. Science. 1991 Dec 13;254(5038):1651–1654. doi: 10.1126/science.1661030. [DOI] [PubMed] [Google Scholar]
- Ninfa E. G., Stock A., Mowbray S., Stock J. Reconstitution of the bacterial chemotaxis signal transduction system from purified components. J Biol Chem. 1991 May 25;266(15):9764–9770. [PubMed] [Google Scholar]
- Oosawa K., Hess J. F., Simon M. I. Mutants defective in bacterial chemotaxis show modified protein phosphorylation. Cell. 1988 Apr 8;53(1):89–96. doi: 10.1016/0092-8674(88)90490-4. [DOI] [PubMed] [Google Scholar]
- Parkinson J. S., Kofoid E. C. Communication modules in bacterial signaling proteins. Annu Rev Genet. 1992;26:71–112. doi: 10.1146/annurev.ge.26.120192.000443. [DOI] [PubMed] [Google Scholar]
- Parkinson J. S. cheA, cheB, and cheC genes of Escherichia coli and their role in chemotaxis. J Bacteriol. 1976 May;126(2):758–770. doi: 10.1128/jb.126.2.758-770.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith R. A., Parkinson J. S. Overlapping genes at the cheA locus of Escherichia coli. Proc Natl Acad Sci U S A. 1980 Sep;77(9):5370–5374. doi: 10.1073/pnas.77.9.5370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soderling T. R. Protein kinases. Regulation by autoinhibitory domains. J Biol Chem. 1990 Feb 5;265(4):1823–1826. [PubMed] [Google Scholar]
- Stewart R. C. Activating and inhibitory mutations in the regulatory domain of CheB, the methylesterase in bacterial chemotaxis. J Biol Chem. 1993 Jan 25;268(3):1921–1930. [PubMed] [Google Scholar]
- Stewart R. C., Roth A. F., Dahlquist F. W. Mutations that affect control of the methylesterase activity of CheB, a component of the chemotaxis adaptation system in Escherichia coli. J Bacteriol. 1990 Jun;172(6):3388–3399. doi: 10.1128/jb.172.6.3388-3399.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stock J. B., Lukat G. S., Stock A. M. Bacterial chemotaxis and the molecular logic of intracellular signal transduction networks. Annu Rev Biophys Biophys Chem. 1991;20:109–136. doi: 10.1146/annurev.bb.20.060191.000545. [DOI] [PubMed] [Google Scholar]
- Stock J. B., Ninfa A. J., Stock A. M. Protein phosphorylation and regulation of adaptive responses in bacteria. Microbiol Rev. 1989 Dec;53(4):450–490. doi: 10.1128/mr.53.4.450-490.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stock J. B., Stock A. M., Mottonen J. M. Signal transduction in bacteria. Nature. 1990 Mar 29;344(6265):395–400. doi: 10.1038/344395a0. [DOI] [PubMed] [Google Scholar]
- Swanson R. V., Bourret R. B., Simon M. I. Intermolecular complementation of the kinase activity of CheA. Mol Microbiol. 1993 May;8(3):435–441. doi: 10.1111/j.1365-2958.1993.tb01588.x. [DOI] [PubMed] [Google Scholar]
- Swanson R. V., Schuster S. C., Simon M. I. Expression of CheA fragments which define domains encoding kinase, phosphotransfer, and CheY binding activities. Biochemistry. 1993 Aug 3;32(30):7623–7629. doi: 10.1021/bi00081a004. [DOI] [PubMed] [Google Scholar]
- Welch M., Oosawa K., Aizawa S., Eisenbach M. Phosphorylation-dependent binding of a signal molecule to the flagellar switch of bacteria. Proc Natl Acad Sci U S A. 1993 Oct 1;90(19):8787–8791. doi: 10.1073/pnas.90.19.8787. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolfe A. J., Berg H. C. Migration of bacteria in semisolid agar. Proc Natl Acad Sci U S A. 1989 Sep;86(18):6973–6977. doi: 10.1073/pnas.86.18.6973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolfe A. J., Stewart R. C. The short form of the CheA protein restores kinase activity and chemotactic ability to kinase-deficient mutants. Proc Natl Acad Sci U S A. 1993 Feb 15;90(4):1518–1522. doi: 10.1073/pnas.90.4.1518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wylie D., Stock A., Wong C. Y., Stock J. Sensory transduction in bacterial chemotaxis involves phosphotransfer between Che proteins. Biochem Biophys Res Commun. 1988 Mar 15;151(2):891–896. doi: 10.1016/s0006-291x(88)80365-6. [DOI] [PubMed] [Google Scholar]
- Yamaguchi S., Aizawa S., Kihara M., Isomura M., Jones C. J., Macnab R. M. Genetic evidence for a switching and energy-transducing complex in the flagellar motor of Salmonella typhimurium. J Bacteriol. 1986 Dec;168(3):1172–1179. doi: 10.1128/jb.168.3.1172-1179.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]