Abstract
The differentiating bacterium Caulobacter crescentus has been studied extensively to understand how a relatively simple life form can govern the timing of expression of genes needed for the production of stage-specific structures. In this study, a clone containing the 5.3-kb flaP region was shown to contain the flgI, cheL, and flbY genes arranged in an operon with transcription proceeding from flgI to flbY. The predicted flgI polypeptide shows remarkable identity (44%) to the flagellar basal body P-ring protein encoded by the flgI gene of Salmonella typhimurium. flgI mutations case a reduction in the levels of flagellin production and the overproduction of the hook proteins. Therefore, the flgI-encoded P-ring protein is required for normal flagellin and hook protein synthesis, suggesting that basal body assembly may play a role in the regulation of flagellar gene expression. The flbY gene probably is a basal body component as well, since flbY mutants have flagellin and hook protein synthesis patterns similar to those exhibited by other basal body mutants. The smaller cheL gene complements a mutant that is unable to respond to chemotactic signals despite possessing a functional flagellum. This is the first example of an operon containing both flagellar and chemotaxis genes in C. crescentus.
Full text
PDF



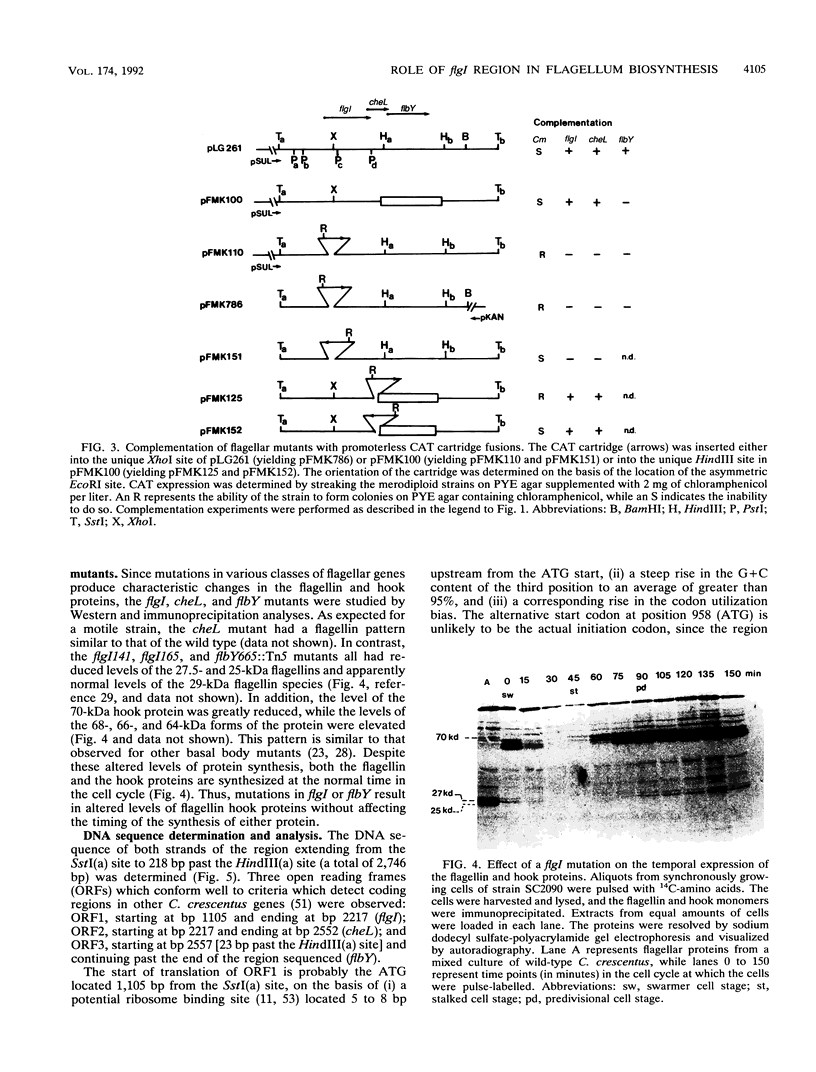




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bagdasarian M., Lurz R., Rückert B., Franklin F. C., Bagdasarian M. M., Frey J., Timmis K. N. Specific-purpose plasmid cloning vectors. II. Broad host range, high copy number, RSF1010-derived vectors, and a host-vector system for gene cloning in Pseudomonas. Gene. 1981 Dec;16(1-3):237–247. doi: 10.1016/0378-1119(81)90080-9. [DOI] [PubMed] [Google Scholar]
- Chen L. S., Mullin D., Newton A. Identification, nucleotide sequence, and control of developmentally regulated promoters in the hook operon region of Caulobacter crescentus. Proc Natl Acad Sci U S A. 1986 May;83(9):2860–2864. doi: 10.1073/pnas.83.9.2860. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheung K. K., Newton A. Patterns of protein synthesis during development in Caulobacter crescentus. Dev Biol. 1977 Apr;56(2):417–425. doi: 10.1016/0012-1606(77)90281-0. [DOI] [PubMed] [Google Scholar]
- Dente L., Cesareni G., Cortese R. pEMBL: a new family of single stranded plasmids. Nucleic Acids Res. 1983 Mar 25;11(6):1645–1655. doi: 10.1093/nar/11.6.1645. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deretic V., Chandrasekharappa S., Gill J. F., Chatterjee D. K., Chakrabarty A. M. A set of cassettes and improved vectors for genetic and biochemical characterization of Pseudomonas genes. Gene. 1987;57(1):61–72. doi: 10.1016/0378-1119(87)90177-6. [DOI] [PubMed] [Google Scholar]
- Driks A., Bryan R., Shapiro L., DeRosier D. J. The organization of the Caulobacter crescentus flagellar filament. J Mol Biol. 1989 Apr 20;206(4):627–636. doi: 10.1016/0022-2836(89)90571-8. [DOI] [PubMed] [Google Scholar]
- Ely B., Croft R. H., Gerardot C. J. Genetic mapping of genes required for motility in Caulobacter crescentus. Genetics. 1984 Nov;108(3):523–532. doi: 10.1093/genetics/108.3.523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ely B. DNA sequence of the 3' end of the Caulobacter crescentus 16S rRNA gene. Nucleic Acids Res. 1992 Mar 25;20(6):1423–1423. doi: 10.1093/nar/20.6.1423. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ely B., Ely T. W. Use of pulsed field gel electrophoresis and transposon mutagenesis to estimate the minimal number of genes required for motility in Caulobacter crescentus. Genetics. 1989 Dec;123(4):649–654. doi: 10.1093/genetics/123.4.649. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ely B. Transfer of drug resistance factors to the dimorphic bacterium Caulobacter crescentus. Genetics. 1979 Mar;91(3):371–380. doi: 10.1093/genetics/91.3.371. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Evinger M., Agabian N. Envelope-associated nucleoid from Caulobacter crescentus stalked and swarmer cells. J Bacteriol. 1977 Oct;132(1):294–301. doi: 10.1128/jb.132.1.294-301.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Figurski D. H., Helinski D. R. Replication of an origin-containing derivative of plasmid RK2 dependent on a plasmid function provided in trans. Proc Natl Acad Sci U S A. 1979 Apr;76(4):1648–1652. doi: 10.1073/pnas.76.4.1648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fukuda A., Koyasu S., Okada Y. Characterization of two flagella-related proteins from Caulobacter crescentus. FEBS Lett. 1978 Nov 1;95(1):70–75. doi: 10.1016/0014-5793(78)80054-4. [DOI] [PubMed] [Google Scholar]
- Gill P. R., Agabian N. The nucleotide sequence of the Mr = 28,500 flagellin gene of Caulobacter crescentus. J Biol Chem. 1983 Jun 25;258(12):7395–7401. [PubMed] [Google Scholar]
- Gober J. W., Champer R., Reuter S., Shapiro L. Expression of positional information during cell differentiation of Caulobacter. Cell. 1991 Jan 25;64(2):381–391. doi: 10.1016/0092-8674(91)90646-g. [DOI] [PubMed] [Google Scholar]
- Gober J. W., Shapiro L. Integration host factor is required for the activation of developmentally regulated genes in Caulobacter. Genes Dev. 1990 Sep;4(9):1494–1504. doi: 10.1101/gad.4.9.1494. [DOI] [PubMed] [Google Scholar]
- Gomes S. L., Shapiro L. Differential expression and positioning of chemotaxis methylation proteins in Caulobacter. J Mol Biol. 1984 Sep 25;178(3):551–568. doi: 10.1016/0022-2836(84)90238-9. [DOI] [PubMed] [Google Scholar]
- Hahnenberger K. M., Shapiro L. Identification of a gene cluster involved in flagellar basal body biogenesis in Caulobacter crescentus. J Mol Biol. 1987 Mar 5;194(1):91–103. doi: 10.1016/0022-2836(87)90718-2. [DOI] [PubMed] [Google Scholar]
- Henikoff S. Unidirectional digestion with exonuclease III creates targeted breakpoints for DNA sequencing. Gene. 1984 Jun;28(3):351–359. doi: 10.1016/0378-1119(84)90153-7. [DOI] [PubMed] [Google Scholar]
- Homma M., DeRosier D. J., Macnab R. M. Flagellar hook and hook-associated proteins of Salmonella typhimurium and their relationship to other axial components of the flagellum. J Mol Biol. 1990 Jun 20;213(4):819–832. doi: 10.1016/S0022-2836(05)80266-9. [DOI] [PubMed] [Google Scholar]
- Johnson R. C., Ely B. Analysis of nonmotile mutants of the dimorphic bacterium Caulobacter crescentus. J Bacteriol. 1979 Jan;137(1):627–634. doi: 10.1128/jb.137.1.627-634.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson R. C., Ely B. Isolation of spontaneously derived mutants of Caulobacter crescentus. Genetics. 1977 May;86(1):25–32. doi: 10.1093/genetics/86.1.25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson R. C., Ferber D. M., Ely B. Synthesis and assembly of flagellar components by Caulobacter crescentus motility mutants. J Bacteriol. 1983 Jun;154(3):1137–1144. doi: 10.1128/jb.154.3.1137-1144.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson R. C., Walsh M. P., Ely B., Shapiro L. Flagellar hook and basal complex of Caulobacter crescentus. J Bacteriol. 1979 Jun;138(3):984–989. doi: 10.1128/jb.138.3.984-989.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones C. J., Homma M., Macnab R. M. L-, P-, and M-ring proteins of the flagellar basal body of Salmonella typhimurium: gene sequences and deduced protein sequences. J Bacteriol. 1989 Jul;171(7):3890–3900. doi: 10.1128/jb.171.7.3890-3900.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khambaty F. M., Ely B. A method for the determination of sequence of high G+C% DNA by the sequential application of sequenase and Taq polymerase. Biotechniques. 1990 Dec;9(6):714–716. [PubMed] [Google Scholar]
- Kihara M., Homma M., Kutsukake K., Macnab R. M. Flagellar switch of Salmonella typhimurium: gene sequences and deduced protein sequences. J Bacteriol. 1989 Jun;171(6):3247–3257. doi: 10.1128/jb.171.6.3247-3257.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuwajima G., Kawagishi I., Homma M., Asaka J., Kondo E., Macnab R. M. Export of an N-terminal fragment of Escherichia coli flagellin by a flagellum-specific pathway. Proc Natl Acad Sci U S A. 1989 Jul;86(13):4953–4957. doi: 10.1073/pnas.86.13.4953. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kyte J., Doolittle R. F. A simple method for displaying the hydropathic character of a protein. J Mol Biol. 1982 May 5;157(1):105–132. doi: 10.1016/0022-2836(82)90515-0. [DOI] [PubMed] [Google Scholar]
- Lagenaur C., Agabian N. Physical characterization of Caulobacter crescentus flagella. J Bacteriol. 1976 Oct;128(1):435–444. doi: 10.1128/jb.128.1.435-444.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Malakooti J., Komeda Y., Matsumura P. DNA sequence analysis, gene product identification, and localization of flagellar motor components of Escherichia coli. J Bacteriol. 1989 May;171(5):2728–2734. doi: 10.1128/jb.171.5.2728-2734.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Milhausen M., Agabian N. Caulobacter flagellin mRNA segregates asymmetrically at cell division. Nature. 1983 Apr 14;302(5909):630–632. doi: 10.1038/302630a0. [DOI] [PubMed] [Google Scholar]
- Mullin D., Minnich S., Chen L. S., Newton A. A set of positively regulated flagellar gene promoters in Caulobacter crescentus with sequence homology to the nif gene promoters of Klebsiella pneumoniae. J Mol Biol. 1987 Jun 20;195(4):939–943. doi: 10.1016/0022-2836(87)90497-9. [DOI] [PubMed] [Google Scholar]
- Norrander J., Kempe T., Messing J. Construction of improved M13 vectors using oligodeoxynucleotide-directed mutagenesis. Gene. 1983 Dec;26(1):101–106. doi: 10.1016/0378-1119(83)90040-9. [DOI] [PubMed] [Google Scholar]
- Ohta N., Chen L. S., Newton A. Isolation and expression of cloned hook protein gene from Caulobacter crescentus. Proc Natl Acad Sci U S A. 1982 Aug;79(16):4863–4867. doi: 10.1073/pnas.79.16.4863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohta N., Mullin D. A., Tarleton J., Ely B., Newton A. Identification, distribution, and sequence analysis of new insertion elements in Caulobacter crescentus. J Bacteriol. 1990 Jan;172(1):236–242. doi: 10.1128/jb.172.1.236-242.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oliver D. Protein secretion in Escherichia coli. Annu Rev Microbiol. 1985;39:615–648. doi: 10.1146/annurev.mi.39.100185.003151. [DOI] [PubMed] [Google Scholar]
- Osley M. A., Sheffery M., Newton A. Regulation of flagellin synthesis in the cell cycle of caulobacter: dependence on DNA replication. Cell. 1977 Oct;12(2):393–400. doi: 10.1016/0092-8674(77)90115-5. [DOI] [PubMed] [Google Scholar]
- POINDEXTER J. S. BIOLOGICAL PROPERTIES AND CLASSIFICATION OF THE CAULOBACTER GROUP. Bacteriol Rev. 1964 Sep;28:231–295. doi: 10.1128/br.28.3.231-295.1964. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ross C. M., Winkler M. E. Structure of the Caulobacter crescentus trpFBA operon. J Bacteriol. 1988 Feb;170(2):757–768. doi: 10.1128/jb.170.2.757-768.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schoenlein P. V., Gallman L. M., Ely B. Use of transmissible plasmids as cloning vectors in Caulobacter crescentus. Gene. 1988 Oct 30;70(2):321–329. doi: 10.1016/0378-1119(88)90204-1. [DOI] [PubMed] [Google Scholar]
- Schoenlein P. V., Gallman L. S., Winkler M. E., Ely B. Nucleotide sequence of the Caulobacter crescentus flaF and flbT genes and an analysis of codon usage in organisms with G + C-rich genomes. Gene. 1990 Sep 1;93(1):17–25. doi: 10.1016/0378-1119(90)90130-j. [DOI] [PubMed] [Google Scholar]
- Sheffery M., Newton A. Reconstitution and purification of flagellar filaments from Caulobacter crescentus. J Bacteriol. 1977 Dec;132(3):1027–1030. doi: 10.1128/jb.132.3.1027-1030.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shine J., Dalgarno L. Determinant of cistron specificity in bacterial ribosomes. Nature. 1975 Mar 6;254(5495):34–38. doi: 10.1038/254034a0. [DOI] [PubMed] [Google Scholar]
- Sommer J. M., Newton A. Sequential regulation of developmental events during polar morphogenesis in Caulobacter crescentus: assembly of pili on swarmer cells requires cell separation. J Bacteriol. 1988 Jan;170(1):409–415. doi: 10.1128/jb.170.1.409-415.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stallmeyer M. J., Hahnenberger K. M., Sosinsky G. E., Shapiro L., DeRosier D. J. Image reconstruction of the flagellar basal body of Caulobacter crescentus. J Mol Biol. 1989 Feb 5;205(3):511–518. doi: 10.1016/0022-2836(89)90222-2. [DOI] [PubMed] [Google Scholar]
- VOGEL H. J., BONNER D. M. Acetylornithinase of Escherichia coli: partial purification and some properties. J Biol Chem. 1956 Jan;218(1):97–106. [PubMed] [Google Scholar]




