Abstract
We have investigated the organization and expression of the Caulobacter crescentus flbF gene because it occupies a high level in the flagellar gene regulatory hierarchy. The nucleotide sequence comprising the 3' end of the flaO operon and the adjacent flbF promoter and structural gene was determined, and the organization of transcription units within this sequence was investigated. We located the 3' ends of the flaO operon transcript by using a nuclease S1 protection assay, and the 5' end of the flbF transcript was precisely mapped by primer extension analysis. The nucleotide sequence upstream from the 5' end of the flbF transcript contains -10 and -35 elements similar to those found in promoters transcribed by sigma 28 RNA polymerase in other organisms. Mutations that changed nucleotides in the -10 or -35 elements or altered their relative spacing resulted in undetectable levels of flbF transcript, demonstrating that these sequences contain nucleotides essential for promoter function. We identified a 700-codon open reading frame, downstream from the flbF promoter region, that was predicted to be the flbF structural gene. The amino-terminal half of the FlbF amino acid sequence contains eight hydrophobic regions predicted to be membrane-spanning segments, suggesting that the FlbF protein may be an integral membrane protein. The FlbF amino acid sequence is very similar to that of a transcriptional regulatory protein called LcrD that is encoded in the highly conserved low-calcium-response region of virulence plasmid pYVO3 in Yersinia enterocolitica (A.-M. Viitanen, P. Toivanen, and M. Skurnik, J. Bacteriol. 172:3152-3162, 1990).
Full text
PDF

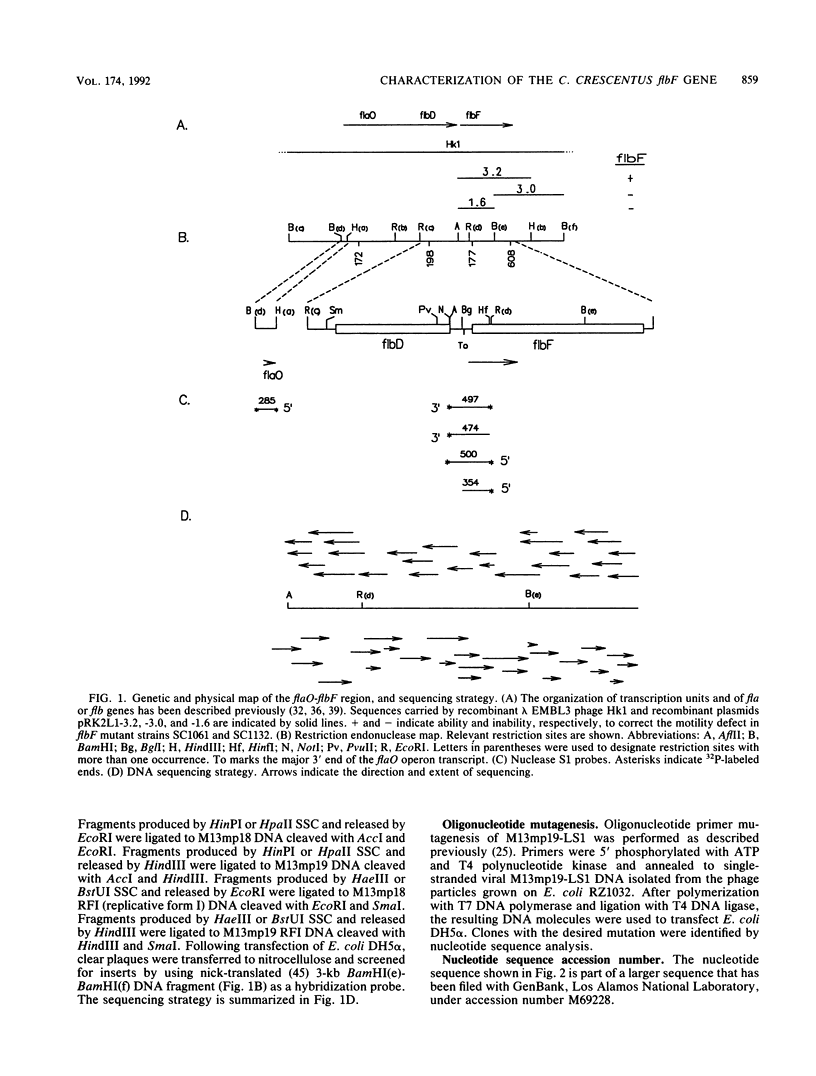

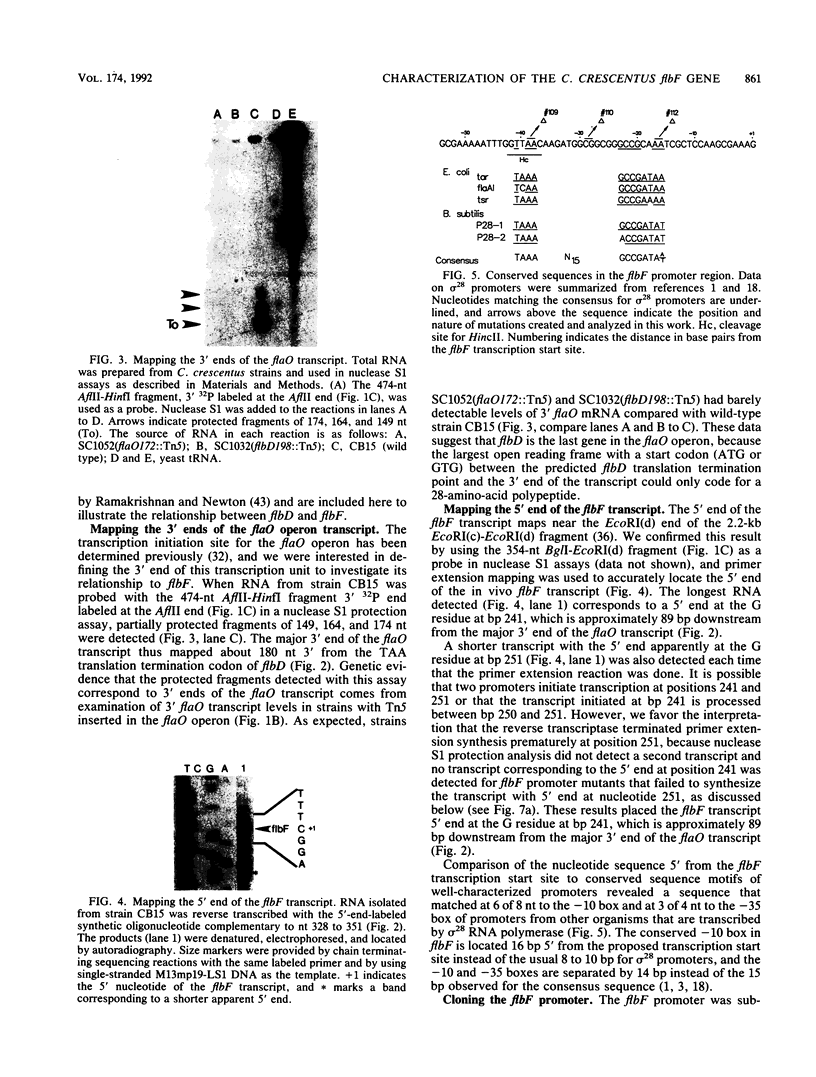
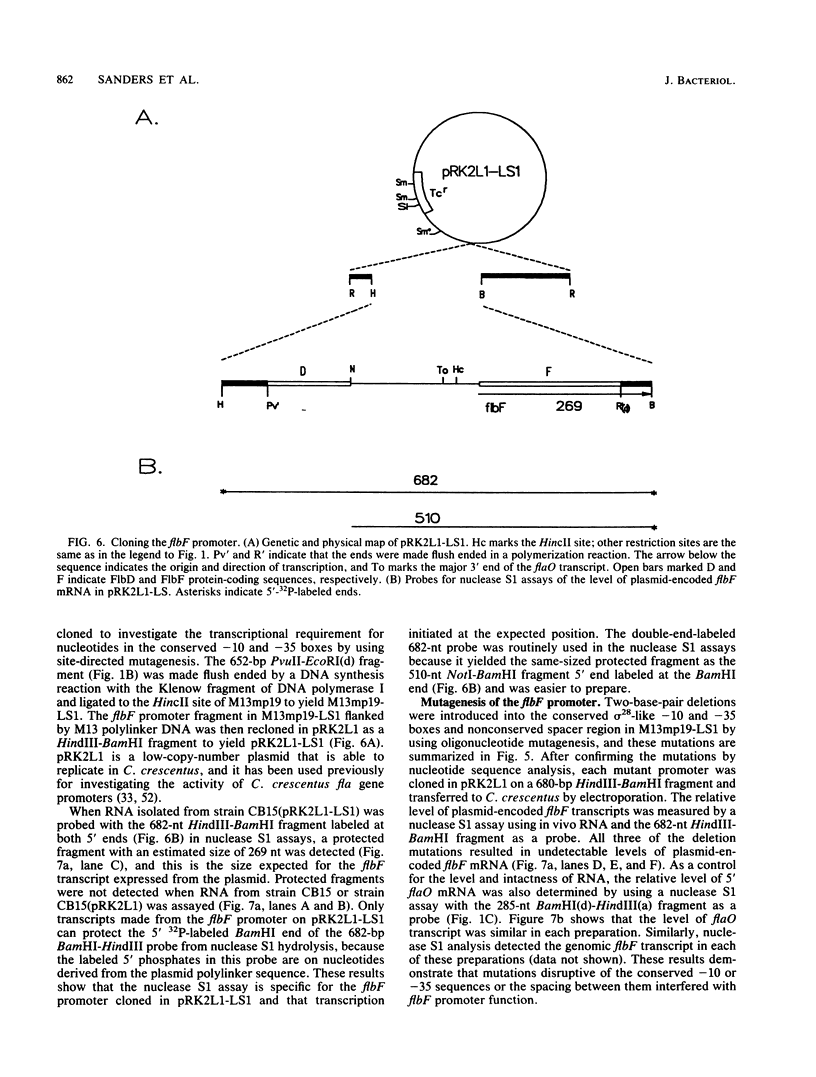




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Arnosti D. N., Chamberlin M. J. Secondary sigma factor controls transcription of flagellar and chemotaxis genes in Escherichia coli. Proc Natl Acad Sci U S A. 1989 Feb;86(3):830–834. doi: 10.1073/pnas.86.3.830. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bartlett D. H., Frantz B. B., Matsumura P. Flagellar transcriptional activators FlbB and FlaI: gene sequences and 5' consensus sequences of operons under FlbB and FlaI control. J Bacteriol. 1988 Apr;170(4):1575–1581. doi: 10.1128/jb.170.4.1575-1581.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berk A. J., Sharp P. A. Sizing and mapping of early adenovirus mRNAs by gel electrophoresis of S1 endonuclease-digested hybrids. Cell. 1977 Nov;12(3):721–732. doi: 10.1016/0092-8674(77)90272-0. [DOI] [PubMed] [Google Scholar]
- Boyer H. W., Roulland-Dussoix D. A complementation analysis of the restriction and modification of DNA in Escherichia coli. J Mol Biol. 1969 May 14;41(3):459–472. doi: 10.1016/0022-2836(69)90288-5. [DOI] [PubMed] [Google Scholar]
- Bryan R., Glaser D., Shapiro L. Genetic regulatory hierarchy in Caulobacter development. Adv Genet. 1990;27:1–31. doi: 10.1016/s0065-2660(08)60022-x. [DOI] [PubMed] [Google Scholar]
- Champer R., Dingwall A., Shapiro L. Cascade regulation of Caulobacter flagellar and chemotaxis genes. J Mol Biol. 1987 Mar 5;194(1):71–80. doi: 10.1016/0022-2836(87)90716-9. [DOI] [PubMed] [Google Scholar]
- Chen L. S., Mullin D., Newton A. Identification, nucleotide sequence, and control of developmentally regulated promoters in the hook operon region of Caulobacter crescentus. Proc Natl Acad Sci U S A. 1986 May;83(9):2860–2864. doi: 10.1073/pnas.83.9.2860. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Doi R. H., Wang L. F. Multiple procaryotic ribonucleic acid polymerase sigma factors. Microbiol Rev. 1986 Sep;50(3):227–243. doi: 10.1128/mr.50.3.227-243.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eisenberg D., Schwarz E., Komaromy M., Wall R. Analysis of membrane and surface protein sequences with the hydrophobic moment plot. J Mol Biol. 1984 Oct 15;179(1):125–142. doi: 10.1016/0022-2836(84)90309-7. [DOI] [PubMed] [Google Scholar]
- Ely B., Ely T. W. Use of pulsed field gel electrophoresis and transposon mutagenesis to estimate the minimal number of genes required for motility in Caulobacter crescentus. Genetics. 1989 Dec;123(4):649–654. doi: 10.1093/genetics/123.4.649. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feingold J., Bellofatto V., Shapiro L., Amemiya K. Organization and nucleotide sequence analysis of an rRNA and tRNA gene cluster from Caulobacter crescentus. J Bacteriol. 1985 Jul;163(1):155–166. doi: 10.1128/jb.163.1.155-166.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Forsberg A., Viitanen A. M., Skurnik M., Wolf-Watz H. The surface-located YopN protein is involved in calcium signal transduction in Yersinia pseudotuberculosis. Mol Microbiol. 1991 Apr;5(4):977–986. doi: 10.1111/j.1365-2958.1991.tb00773.x. [DOI] [PubMed] [Google Scholar]
- Frederikse P. H., Shapiro L. An Escherichia coli chemoreceptor gene is temporally controlled in Caulobacter. Proc Natl Acad Sci U S A. 1989 Jun;86(11):4061–4065. doi: 10.1073/pnas.86.11.4061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gober J. W., Shapiro L. Integration host factor is required for the activation of developmentally regulated genes in Caulobacter. Genes Dev. 1990 Sep;4(9):1494–1504. doi: 10.1101/gad.4.9.1494. [DOI] [PubMed] [Google Scholar]
- HOWARD-FLANDERS P., SIMSON E., THERIOT L. A LOCUS THAT CONTROLS FILAMENT FORMATION AND SENSITIVITY TO RADIATION IN ESCHERICHIA COLI K-12. Genetics. 1964 Feb;49:237–246. doi: 10.1093/genetics/49.2.237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harrison S. C., Aggarwal A. K. DNA recognition by proteins with the helix-turn-helix motif. Annu Rev Biochem. 1990;59:933–969. doi: 10.1146/annurev.bi.59.070190.004441. [DOI] [PubMed] [Google Scholar]
- Hawley D. K., McClure W. R. Compilation and analysis of Escherichia coli promoter DNA sequences. Nucleic Acids Res. 1983 Apr 25;11(8):2237–2255. doi: 10.1093/nar/11.8.2237. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Helmann J. D., Chamberlin M. J. DNA sequence analysis suggests that expression of flagellar and chemotaxis genes in Escherichia coli and Salmonella typhimurium is controlled by an alternative sigma factor. Proc Natl Acad Sci U S A. 1987 Sep;84(18):6422–6424. doi: 10.1073/pnas.84.18.6422. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inouye S., Nakazawa A., Nakazawa T. Expression of the regulatory gene xylS on the TOL plasmid is positively controlled by the xylR gene product. Proc Natl Acad Sci U S A. 1987 Aug;84(15):5182–5186. doi: 10.1073/pnas.84.15.5182. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaplan J. B., Dingwall A., Bryan R., Champer R., Shapiro L. Temporal regulation and overlap organization of two Caulobacter flagellar genes. J Mol Biol. 1989 Jan 5;205(1):71–83. doi: 10.1016/0022-2836(89)90365-3. [DOI] [PubMed] [Google Scholar]
- Komeda Y. Fusions of flagellar operons to lactose genes on a mu lac bacteriophage. J Bacteriol. 1982 Apr;150(1):16–26. doi: 10.1128/jb.150.1.16-26.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Komeda Y. Transcriptional control of flagellar genes in Escherichia coli K-12. J Bacteriol. 1986 Dec;168(3):1315–1318. doi: 10.1128/jb.168.3.1315-1318.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kunkel T. A., Roberts J. D., Zakour R. A. Rapid and efficient site-specific mutagenesis without phenotypic selection. Methods Enzymol. 1987;154:367–382. doi: 10.1016/0076-6879(87)54085-x. [DOI] [PubMed] [Google Scholar]
- Kutsukake K., Ohya Y., Iino T. Transcriptional analysis of the flagellar regulon of Salmonella typhimurium. J Bacteriol. 1990 Feb;172(2):741–747. doi: 10.1128/jb.172.2.741-747.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lederberg E. M., Cohen S. N. Transformation of Salmonella typhimurium by plasmid deoxyribonucleic acid. J Bacteriol. 1974 Sep;119(3):1072–1074. doi: 10.1128/jb.119.3.1072-1074.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Losick R., Pero J. Cascades of Sigma factors. Cell. 1981 Sep;25(3):582–584. doi: 10.1016/0092-8674(81)90164-1. [DOI] [PubMed] [Google Scholar]
- Minnich S. A., Newton A. Promoter mapping and cell cycle regulation of flagellin gene transcription in Caulobacter crescentus. Proc Natl Acad Sci U S A. 1987 Mar;84(5):1142–1146. doi: 10.1073/pnas.84.5.1142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mizusawa S., Nishimura S., Seela F. Improvement of the dideoxy chain termination method of DNA sequencing by use of deoxy-7-deazaguanosine triphosphate in place of dGTP. Nucleic Acids Res. 1986 Feb 11;14(3):1319–1324. doi: 10.1093/nar/14.3.1319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mohana Rao J. K., Argos P. A conformational preference parameter to predict helices in integral membrane proteins. Biochim Biophys Acta. 1986 Jan 30;869(2):197–214. doi: 10.1016/0167-4838(86)90295-5. [DOI] [PubMed] [Google Scholar]
- Mullin D. A., Newton A. Ntr-like promoters and upstream regulatory sequence ftr are required for transcription of a developmentally regulated Caulobacter crescentus flagellar gene. J Bacteriol. 1989 Jun;171(6):3218–3227. doi: 10.1128/jb.171.6.3218-3227.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mullin D., Minnich S., Chen L. S., Newton A. A set of positively regulated flagellar gene promoters in Caulobacter crescentus with sequence homology to the nif gene promoters of Klebsiella pneumoniae. J Mol Biol. 1987 Jun 20;195(4):939–943. doi: 10.1016/0022-2836(87)90497-9. [DOI] [PubMed] [Google Scholar]
- Newton A., Ohta N., Ramakrishnan G., Mullin D., Raymond G. Genetic switching in the flagellar gene hierarchy of Caulobacter requires negative as well as positive regulation of transcription. Proc Natl Acad Sci U S A. 1989 Sep;86(17):6651–6655. doi: 10.1073/pnas.86.17.6651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Newton A., Ohta N. Regulation of the cell division cycle and differentiation in bacteria. Annu Rev Microbiol. 1990;44:689–719. doi: 10.1146/annurev.mi.44.100190.003353. [DOI] [PubMed] [Google Scholar]
- Ninfa A. J., Mullin D. A., Ramakrishnan G., Newton A. Escherichia coli sigma 54 RNA polymerase recognizes Caulobacter crescentus flbG and flaN flagellar gene promoters in vitro. J Bacteriol. 1989 Jan;171(1):383–391. doi: 10.1128/jb.171.1.383-391.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohta N., Chen L. S., Swanson E., Newton A. Transcriptional regulation of a periodically controlled flagellar gene operon in Caulobacter crescentus. J Mol Biol. 1985 Nov 5;186(1):107–115. doi: 10.1016/0022-2836(85)90261-x. [DOI] [PubMed] [Google Scholar]
- Ohta N., Swanson E., Ely B., Newton A. Physical mapping and complementation analysis of transposon Tn5 mutations in Caulobacter crescentus: organization of transcriptional units in the hook gene cluster. J Bacteriol. 1984 Jun;158(3):897–904. doi: 10.1128/jb.158.3.897-904.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- POINDEXTER J. S. BIOLOGICAL PROPERTIES AND CLASSIFICATION OF THE CAULOBACTER GROUP. Bacteriol Rev. 1964 Sep;28:231–295. doi: 10.1128/br.28.3.231-295.1964. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poncz M., Solowiejczyk D., Ballantine M., Schwartz E., Surrey S. "Nonrandom" DNA sequence analysis in bacteriophage M13 by the dideoxy chain-termination method. Proc Natl Acad Sci U S A. 1982 Jul;79(14):4298–4302. doi: 10.1073/pnas.79.14.4298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramakrishnan G., Newton A. FlbD of Caulobacter crescentus is a homologue of the NtrC (NRI) protein and activates sigma 54-dependent flagellar gene promoters. Proc Natl Acad Sci U S A. 1990 Mar;87(6):2369–2373. doi: 10.1073/pnas.87.6.2369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rigby P. W., Dieckmann M., Rhodes C., Berg P. Labeling deoxyribonucleic acid to high specific activity in vitro by nick translation with DNA polymerase I. J Mol Biol. 1977 Jun 15;113(1):237–251. doi: 10.1016/0022-2836(77)90052-3. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schoenlein P. V., Gallman L. S., Winkler M. E., Ely B. Nucleotide sequence of the Caulobacter crescentus flaF and flbT genes and an analysis of codon usage in organisms with G + C-rich genomes. Gene. 1990 Sep 1;93(1):17–25. doi: 10.1016/0378-1119(90)90130-j. [DOI] [PubMed] [Google Scholar]
- Shapiro L. Generation of polarity during Caulobacter cell differentiation. Annu Rev Cell Biol. 1985;1:173–207. doi: 10.1146/annurev.cb.01.110185.001133. [DOI] [PubMed] [Google Scholar]
- Shine J., Dalgarno L. The 3'-terminal sequence of Escherichia coli 16S ribosomal RNA: complementarity to nonsense triplets and ribosome binding sites. Proc Natl Acad Sci U S A. 1974 Apr;71(4):1342–1346. doi: 10.1073/pnas.71.4.1342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stock J. B., Ninfa A. J., Stock A. M. Protein phosphorylation and regulation of adaptive responses in bacteria. Microbiol Rev. 1989 Dec;53(4):450–490. doi: 10.1128/mr.53.4.450-490.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Viitanen A. M., Toivanen P., Skurnik M. The lcrE gene is part of an operon in the lcr region of Yersinia enterocolitica O:3. J Bacteriol. 1990 Jun;172(6):3152–3162. doi: 10.1128/jb.172.6.3152-3162.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wood N. B., Rake A. V., Shapiro L. Structure of Caulobacter deoxyribonucleic acid. J Bacteriol. 1976 Jun;126(3):1305–1315. doi: 10.1128/jb.126.3.1305-1315.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu H., Dingwall A., Shapiro L. Negative transcriptional regulation in the Caulobacter flagellar hierarchy. Proc Natl Acad Sci U S A. 1989 Sep;86(17):6656–6660. doi: 10.1073/pnas.86.17.6656. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]