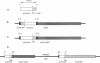Abstract
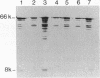
In Methylobacterium extorquens AM1, gene encoding methanol dehydrogenase polypeptides are transcriptionally regulated in response to C1 compounds, including methanol (M. E. Lidstrom and D. I. Stirling, Annu. Rev. Microbiol. 44:27-57, 1990). In order to study this regulation, a transcriptional fusion has been constructed between a beta-galactosidase reporter gene and a 1.55-kb XhoI-SalI fragment of M. extorquens AM1rif DNA encoding the N terminus of the methanol dehydrogenase large subunit (moxF) and 1,289 bp of upstream DNA. The fusion exhibited orientation-specific promoter activity in M. extorquens AM1rif but was expressed constitutively when the transcriptional fusion was located on the plasmid. However, correct regulation was restored when the construction was inserted in the M. extorquens AM1rif chromosome. This DNA fragment was shown to contain both the moxFJGI promoter and the sequences necessary in cis for its transcriptional regulation by methanol. Transcription from this promoter was studied in the M. extorquens AM1rif moxB mutant strains UV4rif and UV25rif, which have a pleiotropic phenotype with regard to the components of methanol oxidation. In these mutants, beta-galactosidase activity from the fusion was reduced to a level equal to that of the vector background when the fusion was present in both plasmid and chromosomal locations. Since both constitutive and methanol-inducible promoter activities were lost in the mutants, moxB appears to be required for transcription of the genes encoding the methanol dehydrogenase polypeptides.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- ARMSTRONG J. M. THE MOLAR EXTINCTION COEFFICIENT OF 2,6-DICHLOROPHENOL INDOPHENOL. Biochim Biophys Acta. 1964 Apr 4;86:194–197. doi: 10.1016/0304-4165(64)90180-1. [DOI] [PubMed] [Google Scholar]
- Anderson D. J., Lidstrom M. E. The moxFG region encodes four polypeptides in the methanol-oxidizing bacterium Methylobacterium sp. strain AM1. J Bacteriol. 1988 May;170(5):2254–2262. doi: 10.1128/jb.170.5.2254-2262.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anderson D. J., Morris C. J., Nunn D. N., Anthony C., Lidstrom M. E. Nucleotide sequence of the Methylobacterium extorquens AM1 moxF and moxJ genes involved in methanol oxidation. Gene. 1990 May 31;90(1):173–176. doi: 10.1016/0378-1119(90)90457-3. [DOI] [PubMed] [Google Scholar]
- Anthony C., Zatman L. J. The microbial oxidation of methanol. 2. The methanol-oxidizing enzyme of Pseudomonas sp. M 27. Biochem J. 1964 Sep;92(3):614–621. doi: 10.1042/bj0920614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Backman K., Ptashne M., Gilbert W. Construction of plasmids carrying the cI gene of bacteriophage lambda. Proc Natl Acad Sci U S A. 1976 Nov;73(11):4174–4178. doi: 10.1073/pnas.73.11.4174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bolivar F., Rodriguez R. L., Greene P. J., Betlach M. C., Heyneker H. L., Boyer H. W., Crosa J. H., Falkow S. Construction and characterization of new cloning vehicles. II. A multipurpose cloning system. Gene. 1977;2(2):95–113. [PubMed] [Google Scholar]
- Ditta G., Schmidhauser T., Yakobson E., Lu P., Liang X. W., Finlay D. R., Guiney D., Helinski D. R. Plasmids related to the broad host range vector, pRK290, useful for gene cloning and for monitoring gene expression. Plasmid. 1985 Mar;13(2):149–153. doi: 10.1016/0147-619x(85)90068-x. [DOI] [PubMed] [Google Scholar]
- Figurski D. H., Helinski D. R. Replication of an origin-containing derivative of plasmid RK2 dependent on a plasmid function provided in trans. Proc Natl Acad Sci U S A. 1979 Apr;76(4):1648–1652. doi: 10.1073/pnas.76.4.1648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fulton G. L., Nunn D. N., Lidstrom M. E. Molecular cloning of a malyl coenzyme A lyase gene from Pseudomonas sp. strain AM1, a facultative methylotroph. J Bacteriol. 1984 Nov;160(2):718–723. doi: 10.1128/jb.160.2.718-723.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harms N., de Vries G. E., Maurer K., Hoogendijk J., Stouthamer A. H. Isolation and nucleotide sequence of the methanol dehydrogenase structural gene from Paracoccus denitrificans. J Bacteriol. 1987 Sep;169(9):3969–3975. doi: 10.1128/jb.169.9.3969-3975.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knecht D. A., Dimond R. L. Visualization of antigenic proteins on Western blots. Anal Biochem. 1984 Jan;136(1):180–184. doi: 10.1016/0003-2697(84)90321-x. [DOI] [PubMed] [Google Scholar]
- La Roche S. D., Leisinger T. Sequence analysis and expression of the bacterial dichloromethane dehalogenase structural gene, a member of the glutathione S-transferase supergene family. J Bacteriol. 1990 Jan;172(1):164–171. doi: 10.1128/jb.172.1.164-171.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lidstrom M. E. Genetics of carbon metabolism in methylotrophic bacteria. FEMS Microbiol Rev. 1990 Dec;7(3-4):431–436. doi: 10.1111/j.1574-6968.1990.tb04949.x. [DOI] [PubMed] [Google Scholar]
- Lidstrom M. E., Stirling D. I. Methylotrophs: genetics and commercial applications. Annu Rev Microbiol. 1990;44:27–58. doi: 10.1146/annurev.mi.44.100190.000331. [DOI] [PubMed] [Google Scholar]
- Machlin S. M., Hanson R. S. Nucleotide sequence and transcriptional start site of the Methylobacterium organophilum XX methanol dehydrogenase structural gene. J Bacteriol. 1988 Oct;170(10):4739–4747. doi: 10.1128/jb.170.10.4739-4747.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McNerney T., O'connor M. L. Regulation of enzymes associated with C-1 metabolism in three facultative methylotrophs. Appl Environ Microbiol. 1980 Aug;40(2):370–375. doi: 10.1128/aem.40.2.370-375.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nunn D. N., Anthony C. The nucleotide sequence and deduced amino acid sequence of the cytochrome cL gene of Methylobacterium extorquens AM1, a novel class of c-type cytochrome. Biochem J. 1988 Dec 1;256(2):673–676. doi: 10.1042/bj2560673. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nunn D. N., Day D., Anthony C. The second subunit of methanol dehydrogenase of Methylobacterium extorquens AM1. Biochem J. 1989 Jun 15;260(3):857–862. doi: 10.1042/bj2600857. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nunn D. N., Lidstrom M. E. Isolation and complementation analysis of 10 methanol oxidation mutant classes and identification of the methanol dehydrogenase structural gene of Methylobacterium sp. strain AM1. J Bacteriol. 1986 May;166(2):581–590. doi: 10.1128/jb.166.2.581-590.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nunn D. N., Lidstrom M. E. Phenotypic characterization of 10 methanol oxidation mutant classes in Methylobacterium sp. strain AM1. J Bacteriol. 1986 May;166(2):591–597. doi: 10.1128/jb.166.2.591-597.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ptashne M. Gene regulation by proteins acting nearby and at a distance. Nature. 1986 Aug 21;322(6081):697–701. doi: 10.1038/322697a0. [DOI] [PubMed] [Google Scholar]
- SAITO H., MIURA K. I. PREPARATION OF TRANSFORMING DEOXYRIBONUCLEIC ACID BY PHENOL TREATMENT. Biochim Biophys Acta. 1963 Aug 20;72:619–629. [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Spanning R. J., Wansell C. W., De Boer T., Hazelaar M. J., Anazawa H., Harms N., Oltmann L. F., Stouthamer A. H. Isolation and characterization of the moxJ, moxG, moxI, and moxR genes of Paracoccus denitrificans: inactivation of moxJ, moxG, and moxR and the resultant effect on methylotrophic growth. J Bacteriol. 1991 Nov;173(21):6948–6961. doi: 10.1128/jb.173.21.6948-6961.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weaver C. A., Lidstrom M. E. Methanol dissimilation in Xanthobacter H4-14: activities, induction and comparison to Pseudomonas AM1 and Paracoccus denitrificans. J Gen Microbiol. 1985 Sep;131(9):2183–2197. doi: 10.1099/00221287-131-9-2183. [DOI] [PubMed] [Google Scholar]