Abstract
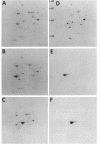
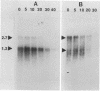
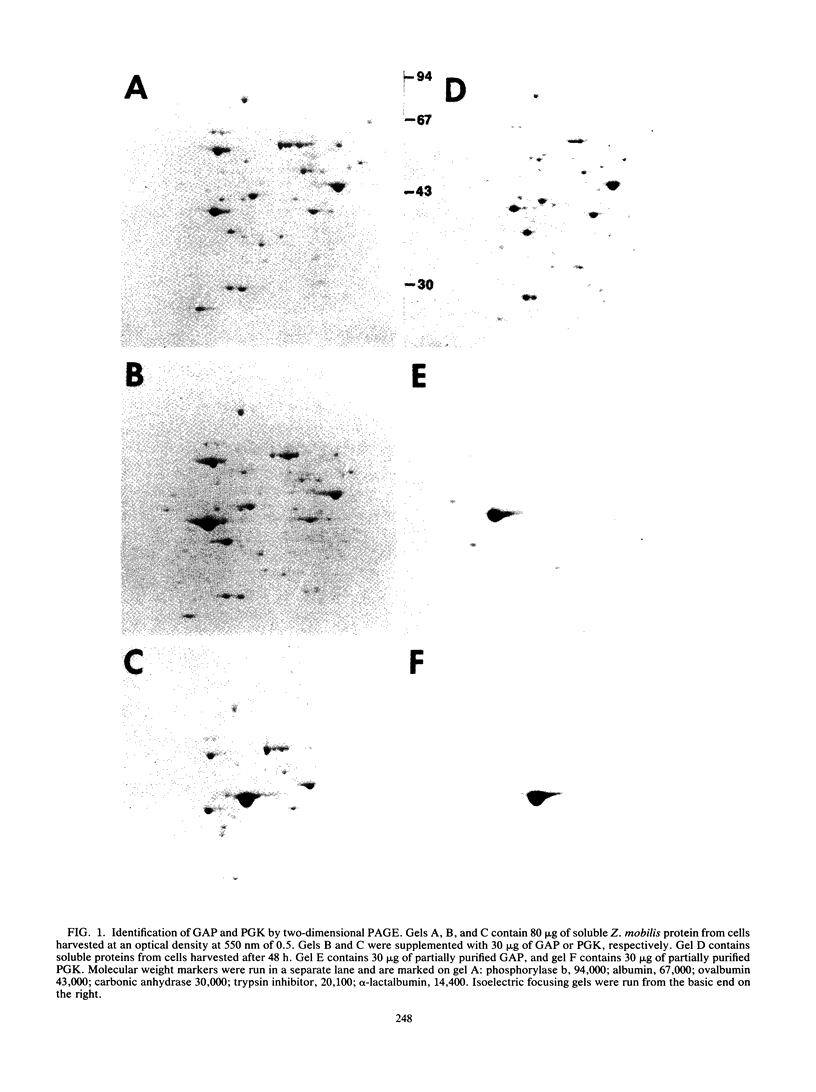
In Zymomonas mobilis, three- to fourfold more glyceraldehyde-3-phosphate dehydrogenase protein than phosphoglycerate kinase is needed for glycolysis because of differences in catalytic efficiency. Consistent with this requirement, higher levels of glyceraldehyde-3-phosphate dehydrogenase were observed with two-dimensional polyacrylamide gel electrophoresis. The genes encoding these enzymes (gap and pgk, respectively) form a bicistronic operon, and some form of regulation is required to provide this differential expression. Two transcripts were observed in Northern RNA analyses with segments of gap as a probe: a more abundant 1.2-kb transcript that contained gap alone and a 2.7-kb transcript that contained both genes. Based on the relative amounts of these transcripts, the coding regions for glyceraldehyde-3-phosphate dehydrogenase were calculated to be fivefold more abundant than those for phosphoglycerate kinase. Assuming equal translational efficiency, this is sufficient to provide the observed differences in expression. Operon fusions with lacZ provided no evidence for intercistronic terminators or attenuation mechanisms. Both gap operon messages were very stable, with half-lives of approximately 16 min (1.2-kb transcript) and 7 min (2.7-kb transcript). Transcript mapping and turnover studies indicated that the shorter gap message was a stable degradation product of the full-length message. Thus differential expression of gap and pgk results primarily from increased translation of the more stable 5' segment of the transcript containing gap. The slow turnover of the messages encoding glyceraldehyde-3-phosphate dehydrogenase and phosphoglycerate kinase is proposed as a major feature contributing to the high level of expression of these essential enzymes.
Full text
PDF








Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Adams C. W., Forrest M. E., Cohen S. N., Beatty J. T. Structural and functional analysis of transcriptional control of the Rhodobacter capsulatus puf operon. J Bacteriol. 1989 Jan;171(1):473–482. doi: 10.1128/jb.171.1.473-482.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alefounder P. R., Perham R. N. Identification, molecular cloning and sequence analysis of a gene cluster encoding the class II fructose 1,6-bisphosphate aldolase, 3-phosphoglycerate kinase and a putative second glyceraldehyde 3-phosphate dehydrogenase of Escherichia coli. Mol Microbiol. 1989 Jun;3(6):723–732. doi: 10.1111/j.1365-2958.1989.tb00221.x. [DOI] [PubMed] [Google Scholar]
- Belasco J. G., Chen C. Y. Mechanism of puf mRNA degradation: the role of an intercistronic stem-loop structure. Gene. 1988 Dec 10;72(1-2):109–117. doi: 10.1016/0378-1119(88)90132-1. [DOI] [PubMed] [Google Scholar]
- Belasco J. G., Higgins C. F. Mechanisms of mRNA decay in bacteria: a perspective. Gene. 1988 Dec 10;72(1-2):15–23. doi: 10.1016/0378-1119(88)90123-0. [DOI] [PubMed] [Google Scholar]
- Bowen D., Littlechild J. A., Fothergill J. E., Watson H. C., Hall L. Nucleotide sequence of the phosphoglycerate kinase gene from the extreme thermophile Thermus thermophilus. Comparison of the deduced amino acid sequence with that of the mesophilic yeast phosphoglycerate kinase. Biochem J. 1988 Sep 1;254(2):509–517. doi: 10.1042/bj2540509. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brendel V., Trifonov E. N. A computer algorithm for testing potential prokaryotic terminators. Nucleic Acids Res. 1984 May 25;12(10):4411–4427. doi: 10.1093/nar/12.10.4411. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen C. Y., Beatty J. T., Cohen S. N., Belasco J. G. An intercistronic stem-loop structure functions as an mRNA decay terminator necessary but insufficient for puf mRNA stability. Cell. 1988 Feb 26;52(4):609–619. doi: 10.1016/0092-8674(88)90473-4. [DOI] [PubMed] [Google Scholar]
- Conway T., Byun M. O., Ingram L. O. Expression Vector for Zymomonas mobilis. Appl Environ Microbiol. 1987 Feb;53(2):235–241. doi: 10.1128/aem.53.2.235-241.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Conway T., Ingram L. O. Phosphoglycerate kinase gene from Zymomonas mobilis: cloning, sequencing, and localization within the gap operon. J Bacteriol. 1988 Apr;170(4):1926–1933. doi: 10.1128/jb.170.4.1926-1933.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Conway T., Osman Y. A., Ingram L. O. Gene expression in Zymomonas mobilis: promoter structure and identification of membrane anchor sequences forming functional lacZ' fusion proteins. J Bacteriol. 1987 Jun;169(6):2327–2335. doi: 10.1128/jb.169.6.2327-2335.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Conway T., Osman Y. A., Konnan J. I., Hoffmann E. M., Ingram L. O. Promoter and nucleotide sequences of the Zymomonas mobilis pyruvate decarboxylase. J Bacteriol. 1987 Mar;169(3):949–954. doi: 10.1128/jb.169.3.949-954.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Conway T., Sewell G. W., Ingram L. O. Glyceraldehyde-3-phosphate dehydrogenase gene from Zymomonas mobilis: cloning, sequencing, and identification of promoter region. J Bacteriol. 1987 Dec;169(12):5653–5662. doi: 10.1128/jb.169.12.5653-5662.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DeHoff B. S., Lee J. K., Donohue T. J., Gumport R. I., Kaplan S. In vivo analysis of puf operon expression in Rhodobacter sphaeroides after deletion of a putative intercistronic transcription terminator. J Bacteriol. 1988 Oct;170(10):4681–4692. doi: 10.1128/jb.170.10.4681-4692.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Devereux J., Haeberli P., Smithies O. A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 1):387–395. doi: 10.1093/nar/12.1part1.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eddy C. K., Mejia J. P., Conway T., Ingram L. O. Differential expression of gap and pgk genes within the gap operon of Zymomonas mobilis. J Bacteriol. 1989 Dec;171(12):6549–6554. doi: 10.1128/jb.171.12.6549-6554.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Freier S. M., Kierzek R., Jaeger J. A., Sugimoto N., Caruthers M. H., Neilson T., Turner D. H. Improved free-energy parameters for predictions of RNA duplex stability. Proc Natl Acad Sci U S A. 1986 Dec;83(24):9373–9377. doi: 10.1073/pnas.83.24.9373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hensel R., Zwickl P., Fabry S., Lang J., Palm P. Sequence comparison of glyceraldehyde-3-phosphate dehydrogenases from the three urkingdoms: evolutionary implication. Can J Microbiol. 1989 Jan;35(1):81–85. doi: 10.1139/m89-012. [DOI] [PubMed] [Google Scholar]
- Holland M. J., Yokoi T., Holland J. P., Myambo K., Innis M. A. The GCR1 gene encodes a positive transcriptional regulator of the enolase and glyceraldehyde-3-phosphate dehydrogenase gene families in Saccharomyces cerevisiae. Mol Cell Biol. 1987 Feb;7(2):813–820. doi: 10.1128/mcb.7.2.813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keshav K. F., Yomano L. P., An H. J., Ingram L. O. Cloning of the Zymomonas mobilis structural gene encoding alcohol dehydrogenase I (adhA): sequence comparison and expression in Escherichia coli. J Bacteriol. 1990 May;172(5):2491–2497. doi: 10.1128/jb.172.5.2491-2497.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kiley P. J., Kaplan S. Molecular genetics of photosynthetic membrane biosynthesis in Rhodobacter sphaeroides. Microbiol Rev. 1988 Mar;52(1):50–69. doi: 10.1128/mr.52.1.50-69.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klug G., Adams C. W., Belasco J., Doerge B., Cohen S. N. Biological consequences of segmental alterations in mRNA stability: effects of deletion of the intercistronic hairpin loop region of the Rhodobacter capsulatus puf operon. EMBO J. 1987 Nov;6(11):3515–3520. doi: 10.1002/j.1460-2075.1987.tb02677.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee J. K., DeHoff B. S., Donohue T. J., Gumport R. I., Kaplan S. Transcriptional analysis of puf operon expression in Rhodobacter sphaeroides 2.4.1 and an intercistronic transcription terminator mutant. J Biol Chem. 1989 Nov 15;264(32):19354–19365. [PubMed] [Google Scholar]
- Mackenzie K. F., Conway T., Aldrich H. C., Ingram L. O. Expression of Zymomonas mobilis adhB (encoding alcohol dehydrogenase II) and adhB-lacZ operon fusions in recombinant Z. mobilis. J Bacteriol. 1989 Sep;171(9):4577–4582. doi: 10.1128/jb.171.9.4577-4582.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mattheakis L. C., Nomura M. Feedback regulation of the spc operon in Escherichia coli: translational coupling and mRNA processing. J Bacteriol. 1988 Oct;170(10):4484–4492. doi: 10.1128/jb.170.10.4484-4492.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Farrell P. H. High resolution two-dimensional electrophoresis of proteins. J Biol Chem. 1975 May 25;250(10):4007–4021. [PMC free article] [PubMed] [Google Scholar]
- Osman Y. A., Conway T., Bonetti S. J., Ingram L. O. Glycolytic flux in Zymomonas mobilis: enzyme and metabolite levels during batch fermentation. J Bacteriol. 1987 Aug;169(8):3726–3736. doi: 10.1128/jb.169.8.3726-3736.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pawluk A., Scopes R. K., Griffiths-Smith K. Isolation and properties of the glycolytic enzymes from Zymomonas mobilis. The five enzymes from glyceraldehyde-3-phosphate dehydrogenase through to pyruvate kinase. Biochem J. 1986 Aug 15;238(1):275–281. doi: 10.1042/bj2380275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pleij C. W. Pseudoknots: a new motif in the RNA game. Trends Biochem Sci. 1990 Apr;15(4):143–147. doi: 10.1016/0968-0004(90)90214-v. [DOI] [PubMed] [Google Scholar]
- Schimmel P. RNA pseudoknots that interact with components of the translation apparatus. Cell. 1989 Jul 14;58(1):9–12. doi: 10.1016/0092-8674(89)90395-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang D., Oyaizu Y., Oyaizu H., Olsen G. J., Woese C. R. Mitochondrial origins. Proc Natl Acad Sci U S A. 1985 Jul;82(13):4443–4447. doi: 10.1073/pnas.82.13.4443. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zuker M., Stiegler P. Optimal computer folding of large RNA sequences using thermodynamics and auxiliary information. Nucleic Acids Res. 1981 Jan 10;9(1):133–148. doi: 10.1093/nar/9.1.133. [DOI] [PMC free article] [PubMed] [Google Scholar]