Abstract
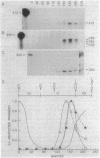
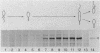
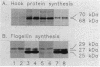
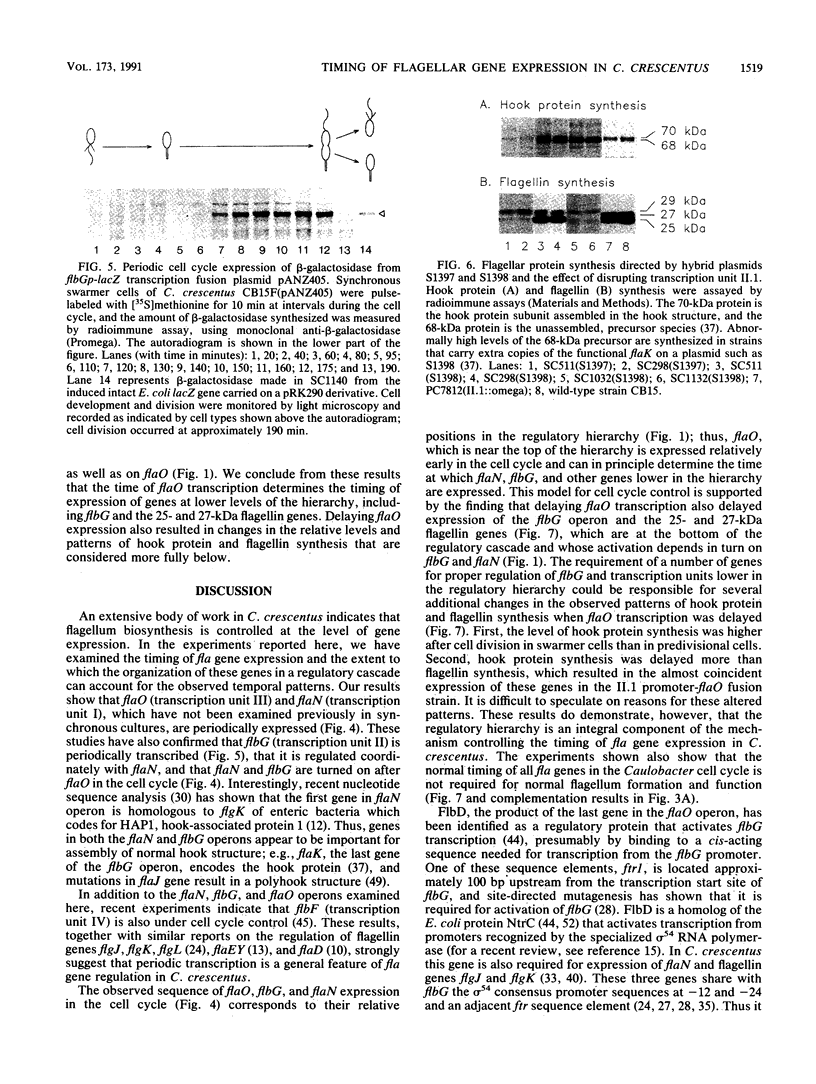
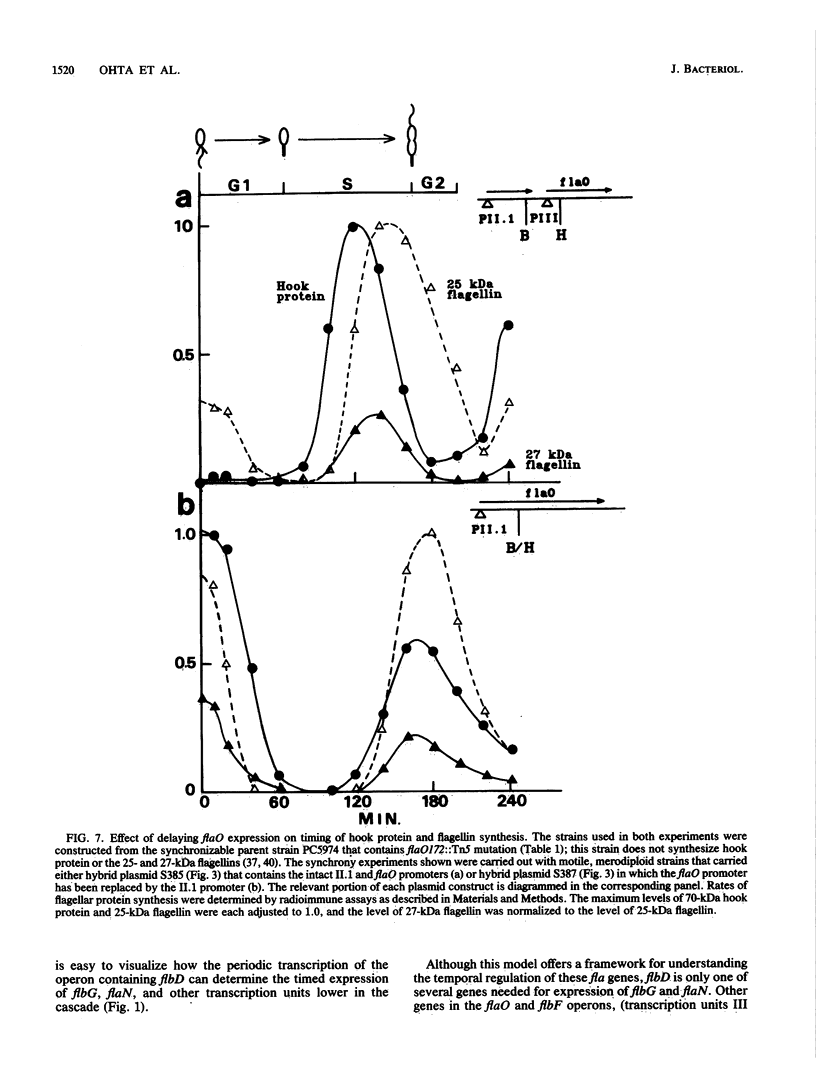
The Caulobacter crescentus flagellar (fla) genes are organized in a regulatory hierarchy in which genes at each level are required for expression of those at the next lower level. To determine the role of this hierarchy in the timing of fla gene expression, we have examined the organization and cell cycle regulation of genes located in the hook gene cluster. As shown here, this cluster is organized into four multicistronic transcription units flaN, flbG, flaO, and flbF that contain fla genes plus a fifth transcription unit II.1 of unknown function. Transcription unit II.1 is regulated independently of the fla gene hierarchy, and it is expressed with a unique pattern of periodicity very late in the cell cycle. The flaN, flbG, and flaO operons are all transcribed periodically, and flaO, which is near the top of the hierarchy and required in trans for the activation of flaN and flbG operons, is expressed earlier in the cell cycle than the other two transcription units. We have shown that delaying flaO transcription by fusing it to the II.1 promoter also delayed the subsequent expression of the flbG operon and the 27- and 25-kDa flagellin genes that are at the bottom of the regulatory hierarchy. Thus, the sequence and timing of fla gene expression in the cell cycle are determined in large measure by the positions of these genes in the regulatory hierarchy. These results also suggest that periodic transcription is a general feature of fla gene expression in C. crescentus.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bryan R., Champer R., Gomes S., Ely B., Shapiro L. Separation of temporal control and trans-acting modulation of flagellin and chemotaxis genes in Caulobacter. Mol Gen Genet. 1987 Feb;206(2):300–306. doi: 10.1007/BF00333588. [DOI] [PubMed] [Google Scholar]
- Bryan R., Purucker M., Gomes S. L., Alexander W., Shapiro L. Analysis of the pleiotropic regulation of flagellar and chemotaxis gene expression in Caulobacter crescentus by using plasmid complementation. Proc Natl Acad Sci U S A. 1984 Mar;81(5):1341–1345. doi: 10.1073/pnas.81.5.1341. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen L. S., Mullin D., Newton A. Identification, nucleotide sequence, and control of developmentally regulated promoters in the hook operon region of Caulobacter crescentus. Proc Natl Acad Sci U S A. 1986 May;83(9):2860–2864. doi: 10.1073/pnas.83.9.2860. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ditta G., Stanfield S., Corbin D., Helinski D. R. Broad host range DNA cloning system for gram-negative bacteria: construction of a gene bank of Rhizobium meliloti. Proc Natl Acad Sci U S A. 1980 Dec;77(12):7347–7351. doi: 10.1073/pnas.77.12.7347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ely B., Ely T. W., Gerardot C. J., Dingwall A. Circularity of the Caulobacter crescentus chromosome determined by pulsed-field gel electrophoresis. J Bacteriol. 1990 Mar;172(3):1262–1266. doi: 10.1128/jb.172.3.1262-1266.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ely B., Ely T. W. Use of pulsed field gel electrophoresis and transposon mutagenesis to estimate the minimal number of genes required for motility in Caulobacter crescentus. Genetics. 1989 Dec;123(4):649–654. doi: 10.1093/genetics/123.4.649. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Evinger M., Agabian N. Envelope-associated nucleoid from Caulobacter crescentus stalked and swarmer cells. J Bacteriol. 1977 Oct;132(1):294–301. doi: 10.1128/jb.132.1.294-301.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hahnenberger K. M., Shapiro L. Identification of a gene cluster involved in flagellar basal body biogenesis in Caulobacter crescentus. J Mol Biol. 1987 Mar 5;194(1):91–103. doi: 10.1016/0022-2836(87)90718-2. [DOI] [PubMed] [Google Scholar]
- Hahnenberger K. M., Shapiro L. Organization and temporal expression of a flagellar basal body gene in Caulobacter crescentus. J Bacteriol. 1988 Sep;170(9):4119–4124. doi: 10.1128/jb.170.9.4119-4124.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Herendeen D. R., Kassavetis G. A., Barry J., Alberts B. M., Geiduschek E. P. Enhancement of bacteriophage T4 late transcription by components of the T4 DNA replication apparatus. Science. 1989 Sep 1;245(4921):952–958. doi: 10.1126/science.2672335. [DOI] [PubMed] [Google Scholar]
- Homma M., DeRosier D. J., Macnab R. M. Flagellar hook and hook-associated proteins of Salmonella typhimurium and their relationship to other axial components of the flagellum. J Mol Biol. 1990 Jun 20;213(4):819–832. doi: 10.1016/S0022-2836(05)80266-9. [DOI] [PubMed] [Google Scholar]
- Kaplan J. B., Dingwall A., Bryan R., Champer R., Shapiro L. Temporal regulation and overlap organization of two Caulobacter flagellar genes. J Mol Biol. 1989 Jan 5;205(1):71–83. doi: 10.1016/0022-2836(89)90365-3. [DOI] [PubMed] [Google Scholar]
- Komeda Y. Fusions of flagellar operons to lactose genes on a mu lac bacteriophage. J Bacteriol. 1982 Apr;150(1):16–26. doi: 10.1128/jb.150.1.16-26.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kustu S., Santero E., Keener J., Popham D., Weiss D. Expression of sigma 54 (ntrA)-dependent genes is probably united by a common mechanism. Microbiol Rev. 1989 Sep;53(3):367–376. doi: 10.1128/mr.53.3.367-376.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lagenaur C., Agabian N. Caulobacter flagellar organelle: synthesis, compartmentation, and assembly. J Bacteriol. 1978 Sep;135(3):1062–1069. doi: 10.1128/jb.135.3.1062-1069.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Loewy Z. G., Bryan R. A., Reuter S. H., Shapiro L. Control of synthesis and positioning of a Caulobacter crescentus flagellar protein. Genes Dev. 1987 Aug;1(6):626–635. doi: 10.1101/gad.1.6.626. [DOI] [PubMed] [Google Scholar]
- Lott T., Ohta N., Newton A. Order of gene replication in Caulobacter crescentus; use of in vivo labeled genomic DNA as a probe. Mol Gen Genet. 1987 Dec;210(3):543–550. doi: 10.1007/BF00327210. [DOI] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- Minnich S. A., Newton A. Promoter mapping and cell cycle regulation of flagellin gene transcription in Caulobacter crescentus. Proc Natl Acad Sci U S A. 1987 Mar;84(5):1142–1146. doi: 10.1073/pnas.84.5.1142. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Minnich S. A., Ohta N., Taylor N., Newton A. Role of the 25-, 27-, and 29-kilodalton flagellins in Caulobacter crescentus cell motility: method for construction of deletion and Tn5 insertion mutants by gene replacement. J Bacteriol. 1988 Sep;170(9):3953–3960. doi: 10.1128/jb.170.9.3953-3960.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mullin D. A., Newton A. Ntr-like promoters and upstream regulatory sequence ftr are required for transcription of a developmentally regulated Caulobacter crescentus flagellar gene. J Bacteriol. 1989 Jun;171(6):3218–3227. doi: 10.1128/jb.171.6.3218-3227.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mullin D., Minnich S., Chen L. S., Newton A. A set of positively regulated flagellar gene promoters in Caulobacter crescentus with sequence homology to the nif gene promoters of Klebsiella pneumoniae. J Mol Biol. 1987 Jun 20;195(4):939–943. doi: 10.1016/0022-2836(87)90497-9. [DOI] [PubMed] [Google Scholar]
- Newton A., Ohta N., Ramakrishnan G., Mullin D., Raymond G. Genetic switching in the flagellar gene hierarchy of Caulobacter requires negative as well as positive regulation of transcription. Proc Natl Acad Sci U S A. 1989 Sep;86(17):6651–6655. doi: 10.1073/pnas.86.17.6651. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Newton A., Ohta N. Regulation of the cell division cycle and differentiation in bacteria. Annu Rev Microbiol. 1990;44:689–719. doi: 10.1146/annurev.mi.44.100190.003353. [DOI] [PubMed] [Google Scholar]
- Ninfa A. J., Mullin D. A., Ramakrishnan G., Newton A. Escherichia coli sigma 54 RNA polymerase recognizes Caulobacter crescentus flbG and flaN flagellar gene promoters in vitro. J Bacteriol. 1989 Jan;171(1):383–391. doi: 10.1128/jb.171.1.383-391.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohta N., Chen L. S., Newton A. Isolation and expression of cloned hook protein gene from Caulobacter crescentus. Proc Natl Acad Sci U S A. 1982 Aug;79(16):4863–4867. doi: 10.1073/pnas.79.16.4863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohta N., Chen L. S., Swanson E., Newton A. Transcriptional regulation of a periodically controlled flagellar gene operon in Caulobacter crescentus. J Mol Biol. 1985 Nov 5;186(1):107–115. doi: 10.1016/0022-2836(85)90261-x. [DOI] [PubMed] [Google Scholar]
- Ohta N., Masurekar M., Newton A. Cloning and cell cycle-dependent expression of DNA replication gene dnaC from Caulobacter crescentus. J Bacteriol. 1990 Dec;172(12):7027–7034. doi: 10.1128/jb.172.12.7027-7034.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohta N., Mullin D. A., Tarleton J., Ely B., Newton A. Identification, distribution, and sequence analysis of new insertion elements in Caulobacter crescentus. J Bacteriol. 1990 Jan;172(1):236–242. doi: 10.1128/jb.172.1.236-242.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohta N., Swanson E., Ely B., Newton A. Physical mapping and complementation analysis of transposon Tn5 mutations in Caulobacter crescentus: organization of transcriptional units in the hook gene cluster. J Bacteriol. 1984 Jun;158(3):897–904. doi: 10.1128/jb.158.3.897-904.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Osley M. A., Sheffery M., Newton A. Regulation of flagellin synthesis in the cell cycle of caulobacter: dependence on DNA replication. Cell. 1977 Oct;12(2):393–400. doi: 10.1016/0092-8674(77)90115-5. [DOI] [PubMed] [Google Scholar]
- Prentki P., Krisch H. M. In vitro insertional mutagenesis with a selectable DNA fragment. Gene. 1984 Sep;29(3):303–313. doi: 10.1016/0378-1119(84)90059-3. [DOI] [PubMed] [Google Scholar]
- Purucker M., Bryan R., Amemiya K., Ely B., Shapiro L. Isolation of a Caulobacter gene cluster specifying flagellum production by using nonmotile Tn5 insertion mutants. Proc Natl Acad Sci U S A. 1982 Nov;79(22):6797–6801. doi: 10.1073/pnas.79.22.6797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ramakrishnan G., Newton A. FlbD of Caulobacter crescentus is a homologue of the NtrC (NRI) protein and activates sigma 54-dependent flagellar gene promoters. Proc Natl Acad Sci U S A. 1990 Mar;87(6):2369–2373. doi: 10.1073/pnas.87.6.2369. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roberts D., Hoopes B. C., McClure W. R., Kleckner N. IS10 transposition is regulated by DNA adenine methylation. Cell. 1985 Nov;43(1):117–130. doi: 10.1016/0092-8674(85)90017-0. [DOI] [PubMed] [Google Scholar]
- Shapiro L. Generation of polarity during Caulobacter cell differentiation. Annu Rev Cell Biol. 1985;1:173–207. doi: 10.1146/annurev.cb.01.110185.001133. [DOI] [PubMed] [Google Scholar]
- Sheffery M., Newton A. Purification and characterization of a polyhook protein from Caulobacter crescentus. J Bacteriol. 1979 May;138(2):575–583. doi: 10.1128/jb.138.2.575-583.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sheffery M., Newton A. Regulation of periodic protein synthesis in the cell cycle: control of initiation and termination of flagellar gene expression. Cell. 1981 Apr;24(1):49–57. doi: 10.1016/0092-8674(81)90500-6. [DOI] [PubMed] [Google Scholar]
- Xu H., Dingwall A., Shapiro L. Negative transcriptional regulation in the Caulobacter flagellar hierarchy. Proc Natl Acad Sci U S A. 1989 Sep;86(17):6656–6660. doi: 10.1073/pnas.86.17.6656. [DOI] [PMC free article] [PubMed] [Google Scholar]