Abstract
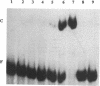
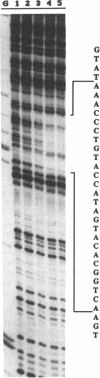
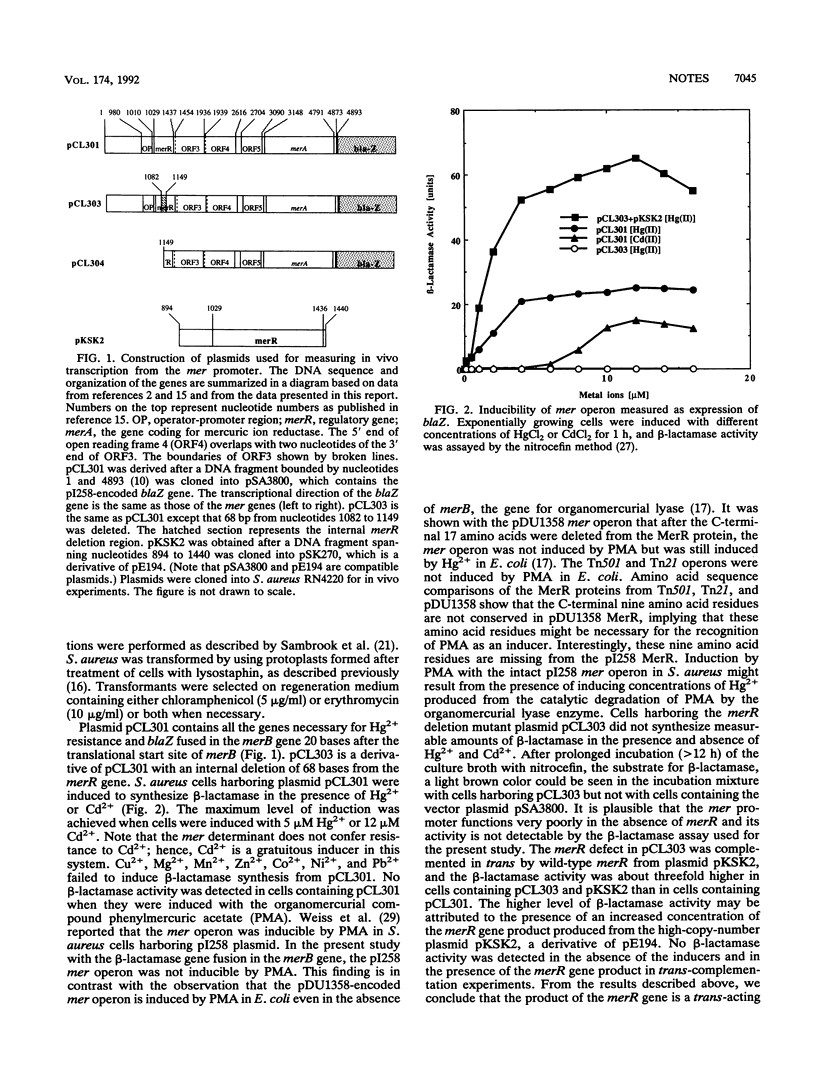
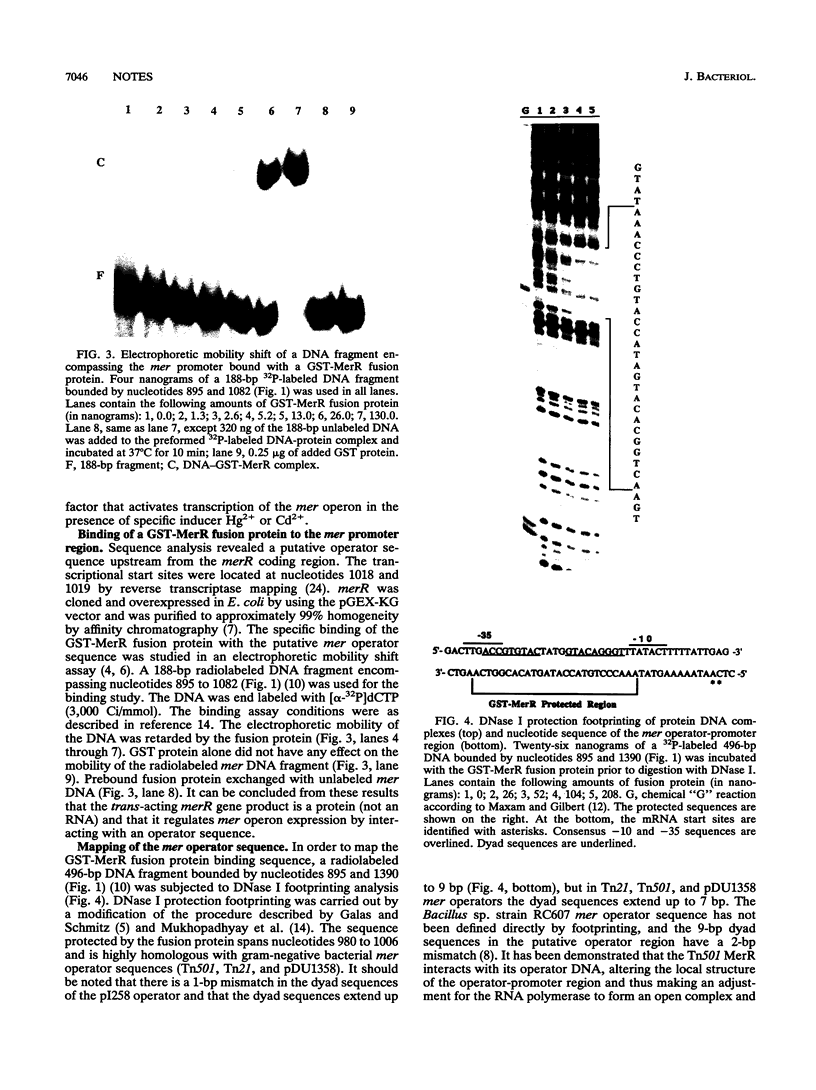
Experiments involving fusion between the Staphylococcus aureus plasmid pI258-encoded mer operon and the reporter gene beta-lactamase, mutational analysis, and trans-complementation studies have shown that the merR gene of pI258, which shows DNA sequence similarity with known merR genes from other bacteria, regulates the expression of the mer operon in vivo. The merR gene product is a trans-acting protein that activates mer operon transcription in the presence of the inducers Hg2+ and Cd2+. A glutathione-S-transferase-MerR fusion protein specifically bound and protected a 27-nucleotide operator sequence from DNase I digestion. This operator sequence is highly homologous with mer operator sequences of other known systems.
Full text
PDF

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ansari A. Z., Chael M. L., O'Halloran T. V. Allosteric underwinding of DNA is a critical step in positive control of transcription by Hg-MerR. Nature. 1992 Jan 2;355(6355):87–89. doi: 10.1038/355087a0. [DOI] [PubMed] [Google Scholar]
- Babich K., Engle M., Skinner J. S., Laddaga R. A. Deletion mutant analysis of the Staphylococcus aureus plasmid pI258 mercury-resistance determinant. Can J Microbiol. 1991 Aug;37(8):624–631. doi: 10.1139/m91-106. [DOI] [PubMed] [Google Scholar]
- Foster T. J. The genetics and biochemistry of mercury resistance. Crit Rev Microbiol. 1987;15(2):117–140. doi: 10.3109/10408418709104455. [DOI] [PubMed] [Google Scholar]
- Fried M., Crothers D. M. Equilibria and kinetics of lac repressor-operator interactions by polyacrylamide gel electrophoresis. Nucleic Acids Res. 1981 Dec 11;9(23):6505–6525. doi: 10.1093/nar/9.23.6505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Galas D. J., Schmitz A. DNAse footprinting: a simple method for the detection of protein-DNA binding specificity. Nucleic Acids Res. 1978 Sep;5(9):3157–3170. doi: 10.1093/nar/5.9.3157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garner M. M., Revzin A. A gel electrophoresis method for quantifying the binding of proteins to specific DNA regions: application to components of the Escherichia coli lactose operon regulatory system. Nucleic Acids Res. 1981 Jul 10;9(13):3047–3060. doi: 10.1093/nar/9.13.3047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guan K. L., Dixon J. E. Eukaryotic proteins expressed in Escherichia coli: an improved thrombin cleavage and purification procedure of fusion proteins with glutathione S-transferase. Anal Biochem. 1991 Feb 1;192(2):262–267. doi: 10.1016/0003-2697(91)90534-z. [DOI] [PubMed] [Google Scholar]
- Helmann J. D., Wang Y., Mahler I., Walsh C. T. Homologous metalloregulatory proteins from both gram-positive and gram-negative bacteria control transcription of mercury resistance operons. J Bacteriol. 1989 Jan;171(1):222–229. doi: 10.1128/jb.171.1.222-229.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kreiswirth B. N., Löfdahl S., Betley M. J., O'Reilly M., Schlievert P. M., Bergdoll M. S., Novick R. P. The toxic shock syndrome exotoxin structural gene is not detectably transmitted by a prophage. Nature. 1983 Oct 20;305(5936):709–712. doi: 10.1038/305709a0. [DOI] [PubMed] [Google Scholar]
- Laddaga R. A., Chu L., Misra T. K., Silver S. Nucleotide sequence and expression of the mercurial-resistance operon from Staphylococcus aureus plasmid pI258. Proc Natl Acad Sci U S A. 1987 Aug;84(15):5106–5110. doi: 10.1073/pnas.84.15.5106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lund P. A., Brown N. L. Regulation of transcription in Escherichia coli from the mer and merR promoters in the transposon Tn501. J Mol Biol. 1989 Jan 20;205(2):343–353. doi: 10.1016/0022-2836(89)90345-8. [DOI] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- Misra T. K. Bacterial resistances to inorganic mercury salts and organomercurials. Plasmid. 1992 Jan;27(1):4–16. doi: 10.1016/0147-619x(92)90002-r. [DOI] [PubMed] [Google Scholar]
- Mukhopadhyay D., Yu H. R., Nucifora G., Misra T. K. Purification and functional characterization of MerD. A coregulator of the mercury resistance operon in gram-negative bacteria. J Biol Chem. 1991 Oct 5;266(28):18538–18542. [PubMed] [Google Scholar]
- Novick R. P., Iordanescu S., Projan S. J., Kornblum J., Edelman I. pT181 plasmid replication is regulated by a countertranscript-driven transcriptional attenuator. Cell. 1989 Oct 20;59(2):395–404. doi: 10.1016/0092-8674(89)90300-0. [DOI] [PubMed] [Google Scholar]
- Novick R., Sanchez-Rivas C., Gruss A., Edelman I. Involvement of the cell envelope in plasmid maintenance: plasmid curing during the regeneration of protoplasts. Plasmid. 1980 May;3(3):348–358. doi: 10.1016/0147-619x(80)90048-7. [DOI] [PubMed] [Google Scholar]
- Nucifora G., Chu L., Silver S., Misra T. K. Mercury operon regulation by the merR gene of the organomercurial resistance system of plasmid pDU1358. J Bacteriol. 1989 Aug;171(8):4241–4247. doi: 10.1128/jb.171.8.4241-4247.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Halloran T. V., Frantz B., Shin M. K., Ralston D. M., Wright J. G. The MerR heavy metal receptor mediates positive activation in a topologically novel transcription complex. Cell. 1989 Jan 13;56(1):119–129. doi: 10.1016/0092-8674(89)90990-2. [DOI] [PubMed] [Google Scholar]
- Parkhill J., Brown N. L. Site-specific insertion and deletion mutants in the mer promoter-operator region of Tn501; the nineteen base-pair spacer is essential for normal induction of the promoter by MerR. Nucleic Acids Res. 1990 Sep 11;18(17):5157–5162. doi: 10.1093/nar/18.17.5157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ralston D. M., O'Halloran T. V. Metalloregulatory proteins and molecular mechanisms of heavy metal signal transduction. Adv Inorg Biochem. 1990;8:1–31. [PubMed] [Google Scholar]
- Silver S., Misra T. K. Plasmid-mediated heavy metal resistances. Annu Rev Microbiol. 1988;42:717–743. doi: 10.1146/annurev.mi.42.100188.003441. [DOI] [PubMed] [Google Scholar]
- Silver S., Walderhaug M. Gene regulation of plasmid- and chromosome-determined inorganic ion transport in bacteria. Microbiol Rev. 1992 Mar;56(1):195–228. doi: 10.1128/mr.56.1.195-228.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Skinner J. S., Ribot E., Laddaga R. A. Transcriptional analysis of the Staphylococcus aureus plasmid pI258 mercury resistance determinant. J Bacteriol. 1991 Aug;173(16):5234–5238. doi: 10.1128/jb.173.16.5234-5238.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Summers A. O. Untwist and shout: a heavy metal-responsive transcriptional regulator. J Bacteriol. 1992 May;174(10):3097–3101. doi: 10.1128/jb.174.10.3097-3101.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walsh C. T., Distefano M. D., Moore M. J., Shewchuk L. M., Verdine G. L. Molecular basis of bacterial resistance to organomercurial and inorganic mercuric salts. FASEB J. 1988 Feb;2(2):124–130. doi: 10.1096/fasebj.2.2.3277886. [DOI] [PubMed] [Google Scholar]
- Wang P. Z., Projan S. J., Leason K. R., Novick R. P. Translational fusion with a secretory enzyme as an indicator. J Bacteriol. 1987 Jul;169(7):3082–3087. doi: 10.1128/jb.169.7.3082-3087.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Y., Moore M., Levinson H. S., Silver S., Walsh C., Mahler I. Nucleotide sequence of a chromosomal mercury resistance determinant from a Bacillus sp. with broad-spectrum mercury resistance. J Bacteriol. 1989 Jan;171(1):83–92. doi: 10.1128/jb.171.1.83-92.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss A. A., Murphy S. D., Silver S. Mercury and organomercurial resistances determined by plasmids in Staphylococcus aureus. J Bacteriol. 1977 Oct;132(1):197–208. doi: 10.1128/jb.132.1.197-208.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]