Abstract
Lactobacillus rhamnosus GG (ATCC 53103) is one of the clinically best-studied probiotic organisms. Moreover, L. rhamnosus GG displays very good in vitro adherence to epithelial cells and mucus. Here, we report that L. rhamnosus GG is able to form biofilms on abiotic surfaces, in contrast to other strains of the Lactobacillus casei group tested under the same conditions. Microtiter plate biofilm assays indicated that in vitro biofilm formation by L. rhamnosus GG is strongly modulated by culture medium factors and conditions related to the gastrointestinal environment, including low pH; high osmolarity; and the presence of bile, mucins, and nondigestible polysaccharides. Additionally, phenotypic analysis of mutants affected in exopolysaccharides (wzb), lipoteichoic acid (dltD), and central metabolism (luxS) showed their relative importance in biofilm formation by L. rhamnosus GG.
One of the clinically best-studied probiotic organisms is Lactobacillus rhamnosus GG (ATCC 53103), which was isolated from a healthy human gut microbiota (12, 15). Various health effects have been attributed to L. rhamnosus GG, such as the prevention and treatment of acute diarrhea in children, the prevention of antibiotic-associated diarrhea, and the prevention and treatment of allergy, as well as occasionally beneficial effects for other disorders (12). However, the mode of action of L. rhamnosus GG related to these health-promoting effects is mainly unknown. Adherence and colonization capacity are considered as contributing factors for immune modulation, pathogen exclusion, and enhanced contact with the mucosa by probiotic bacteria (47, 48). In this way, probiotics would fortify the resident microbiota that forms an integral part of the mucosal barrier and colonization resistance against pathogens.
In comparative studies, L. rhamnosus GG performs well in in vitro adherence experiments with epithelial cells (48) and mucus (47). In vivo, L. rhamnosus GG is able to adhere to the human intestinal mucosa and to persist for more than 1 week after oral intake by healthy adults (1). Moreover, oral administration of L. rhamnosus GG to pregnant women has been reported to result in colonization of their infants for up to 24 months of age (39). Additionally, L. rhamnosus GG is also able to colonize the mouth and can be cultured from saliva for 2 weeks after ingestion (30). This colonization capacity was suggested to be related to the reported beneficial effect of L. rhamnosus GG on the clinical development of dental caries in children (33). Although L. rhamnosus GG was originally isolated from humans, it is also able to colonize the digestive tract of germfree C3H mice (19). L. rhamnosus GG was found to be associated with the mucosa of both the stomach and the intestine of these mice, and this association increased from the proximal to the distal intestine (19).
Many adherent bacteria occur in natural environments as surface-attached biofilms, where they are contained within a self-produced extracellular matrix that protects them against hostile environmental conditions (6, 26). Biofilms also play a role in the intimate relationship between the human body and its resident microbes, for example, in the gut (26). In this study, we investigated the impact of a number of environmental and genetic factors on L. rhamnosus GG biofilm formation. Furthermore, we compared the biofilm formation capacity of L. rhamnosus GG with those of related Lactobacillus strains.
MATERIALS AND METHODS
Bacterial strains and culture conditions.
Lactobacillus strains (Table 1) were routinely grown at 37°C in de Man-Rogosa-Sharpe (MRS) medium (Difco) (11) under static (i.e., microaerobic) conditions. Additionally, Lactobacilli AOAC medium (Difco) (15 g/liter peptonized milk, 5 g/liter yeast extract, 10 g/liter glucose, 5 g/liter tomato juice, 2 g/liter monopotassium phosphate, and 1 g/liter Tween 80), and Trypticase soy broth (TSB) medium (BD Biosciences) (17 g/liter pancreatic digest of casein, 3 g/liter enzymatic digest of soybean meal, 5 g/liter sodium chloride, 2.5 g/liter dipotassium phosphate, and 2.5 g/liter dextrose) were used in this study. Modified TSB (mTSB) medium consisted of 15 g/liter TSB (BD Biosciences) enriched with 20 g/liter Bacto proteose peptone no. 3 (BD Biosciences). Anaerobic conditions were achieved by the use of anaerobic jars (BBL Gas-Pack anaerobic systems; VWR International, Haasrode, Belgium). Other compounds tested in this study were mucin from porcine stomach type III (Sigma), lactoferrin (bovine) (Sigma), bile (bovine, minimum 50% bile acids, mixture of free and conjugated bile acids) (Sigma), and inulin (Beneo HP; degree of polymerization, 12 to 60 [Orafti]) or its derivatives oligofructose (OF) (Beneo P95; degree of polymerization, 3 to 7 [Orafti]) and Synergy1 (Beneo Synergy 1, an OF-inulin mixture [Orafti]). The pH of the media was adjusted with 1.0 M HCl or 1.0 M NaOH before autoclaving. If required, erythromycin was added to the cultures in a final concentration of 5 μg/ml.
TABLE 1.
Bacterial strains used in this study
| Strain or plasmid | Relevant characteristics | Source or reference |
|---|---|---|
| Strains | ||
| E. coli DH5α | F− φ80dlacZΔM15 Δ(lacZYA-argF)U169 deoR recA1 endA1 hsdR17(rk− mk−) supE44 λ−thi-1 girA96 relA1 | Gibco-BRL |
| L. rhamnosus GG | ||
| ATCC 53103 | Wild type, human isolate | ATCC |
| CMPG5412 | luxS-knockout mutant of L. rhamnosus GG; Tetr | 27 |
| CMPG5540 | dltD-knockout mutant of L. rhamnosus GG; Tetr | 35 |
| CMPG5344 | wzb-knockdown mutant of L. rhamnosus GG by expression of antisense RNA after introduction of plasmid pCMPG5344; Eryr | This study |
| Other Lactobacillus strains | ||
| L. rhamnosus | ||
| ATCC 595 | Wild-type strain | ATCC |
| ATCC 7469 | Wild-type strain, type strain | ATCC |
| ATCC 10863 | Wild-type strain | ATCC |
| L. casei | ||
| ATCC 334 | Wild-type strain, neotype strain | ATCC |
| ATCC 393 | Wild-type strain, also classified as L. zeae | ATCC |
| Immunitas | Isolated from Actimel | Danone |
| Shirota | Isolated from Yakult | Yakult |
| Plasmids | ||
| pLAB1301 | E. coli-Lactobacillus shuttle vector; Apr Eryr | 23 |
| pCMPG5334 | pLAB1301 derivative for expression of antisense wzb RNA of L. rhamnosus GG driven by ldhL promoter; Apr Eryr | This study |
In vitro biofilm assay.
A method for assaying biofilm formation of L. rhamnosus GG was based on reference 10 with minor modifications. Briefly, the device used for biofilm formation is a platform carrying 96 polystyrene pegs (Nunc no. 445497) that fits as a microtiter plate lid with a peg hanging into each microtiter plate well (Nunc no. 269789). For biofilm formation, the device was placed in its original sterile tray filled with 200 μl medium. Ca. 3 × 107 CFU were added and incubated without shaking for 72 h at 37°C. To quantify biofilm formation, the pegs were briefly washed in phosphate-buffered saline (PBS). The remaining attached bacteria were stained for 30 min with 200 μl 0.1% (wt/vol) crystal violet in an isopropanol-methanol-PBS solution (1:1:18 [vol/vol]). Excess stain was rinsed off by placing the pegs into a 96-well plate filled with 200 μl water per well. After the pegs were air dried (30 min), the dye bound to the adherent cells was extracted with 200 μl ethanol-acetone (80:20) or 30% glacial acetic acid for thick biofilms. The optical density (OD) of 135 μl of each well was measured at 570 nm using a VERSAmax (Molecular Devices). Each strain and/or condition was tested in at least three independent experiments, each with eight biological replicates. Data were normalized to the indicated positive control, which was taken as 100% to compare different experiments. The results are presented as means ± standard deviations. Additionally, sterile medium was always included (negative control) to ensure that the influence on biofilm formation by mucus, inulin, and bile was not attributed to a nonspecific binding effect to crystal violet.
Monitoring bacterial growth in suspension.
Lactobacillus cultures were grown overnight in MRS medium, washed with PBS, brought to equal cell densities (5 × 108 CFU/ml), and diluted 15,000-fold in 300 μl fresh medium. Three replicates of each 300-μl culture were inoculated into the wells of sterile Honeycomb plates. These plates were incubated at 37°C with continuous shaking, and the OD at 600 nm (OD600) was measured every 30 min in a Bioscreen C microbiology workstation (Labsystems Oy, Zellik, Belgium). For each time point, the average OD was calculated from three independent measurements.
EPS isolation and quantification.
A protocol was followed to differentiate bound exopolysaccharide (EPS-b) from EPS released into the culture medium (EPS-r). EPS-r was precipitated from culture supernatant with ethanol, while EPS-b was first extracted from the bacterial cells with 0.05 M EDTA prior to ethanol precipitation as previously described (45). To compare EPS production in different culture media, EPS amounts were expressed as the amount of equivalents of glucose produced per 109 CFU. As a negative control, EPS content of sterile growth medium was assessed. Each experiment was repeated at least three times.
DNA manipulations.
Routine molecular biology techniques were performed according to standard procedures (38). Restriction and modifying enzymes (from New England Biolabs) were used as recommended by the manufacturer. Plasmid DNA was prepared from Escherichia coli cells by QIAGEN miniprep kits. Chromosomal DNA and plasmid DNA were isolated from L. rhamnosus GG as previously described (9).
Cloning of the L. rhamnosus GG wzb gene.
Primers Pro-0005 (5′-AGTTGATTAATACCGCGACAACAA-3′) and Pro-0058 (5′-GAAGGGGAACTGAATGATGATTGATG-3′) were designed to identify the wzb gene of L. rhamnosus GG based on the published wzb sequence of the closely related strain L. rhamnosus ATCC 9595 (NCBI accession no. AY659976) (34). The PCR product was cloned into pCRII-TOPO vector (Invitrogen), resulting in plasmid pCMPG5921, and sequenced. After BLASTx analysis, a 762-bp DNA fragment was identified that putatively encodes the phosphoprotein phosphatase Wzb of L. rhamnosus GG.
Construction of wzb antisense RNA expression plasmid pCMPG5344.
To achieve overexpression of antisense RNA, the wzb gene was cloned in a high-copy vector in reverse orientation controlled by a constitutive promoter. Therefore, the promoter region of the ldhL gene of L. rhamnosus GG was amplified from chromosomal DNA with primers Pro-127 (5′-CTGAGCTCCTTGTCACAGGATTCACAAGTCTTGC-3′) and Pro-128 (5′-GTCATATGGATATCATCCTTTCTTATGTGCATGC-3′). The 0.2-kb amplicon was cloned into a pCRII-TOPO vector (Invitrogen), resulting in plasmid pCMPG5901. Subsequently, a fragment containing wzb was cut with EcoRI from pCMPG5921 and blunt ligated in the reverse direction in pCMPG5901, which was digested with NdeI (blunt), resulting in plasmid pCMPG5922. A ca. 1-kb fragment containing the ldhL promoter and wzb gene in the antisense direction was subsequently cut from pCMPG5922 with EcoRI and ligated in the EcoRI site of the high-copy vector pLAB1301 (23), resulting in plasmid pCMPG5344. This vector was transferred to L. rhamnosus GG as previously described (9). The presence of antisense RNA was confirmed by Northern blot analysis with specific probes.
Nucleotide sequence accession number.
The sequence of the putative wzb gene has been submitted to the NCBI database (accession no. EF690379).
RESULTS
In vitro biofilm formation by L. rhamnosus GG.
A microtiter plate format assay was used to assay biofilm formation. Three different growth media were tested: the standard medium for lactobacilli (i.e., nutrient-rich MRS medium) (11), TSB with slight modifications (mTSB) (see Materials and Methods), and Lactobacilli AOAC medium (Difco). No biofilm formation could be observed within 24 to 96 h in the MRS medium (Fig. 1). Confluent biofilm growth could be observed after 72 h of growth in mTSB medium (Fig. 1). Additionally, L. rhamnosus GG was able to form biofilms in AOAC medium (Difco), as we previously reported (27). Biofilm formation is less pronounced in AOAC medium than in mTSB medium (Fig. 1). Biofilms were formed under both microaerobic (Fig. 1) and anaerobic (data not shown) conditions. Biofilm formation of L. rhamnosus GG could also be observed on negatively charged glass surfaces (data not shown).
FIG. 1.
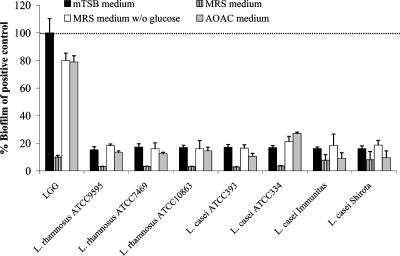
Comparison of capacities of biofilm formation by different lactobacilli. Capacities of biofilm formation by eight Lactobacillus strains were compared under different culture conditions: i.e., mTSB medium, MRS medium (with and without [w/o] glucose), and AOAC medium. The results are expressed compared to biofilm formation of L. rhamnosus GG in mTSB medium (positive control), which was taken as 100% (dotted line). The error bars represent standard deviations of eight biological repeats. The data shown are representative of at least three independent experiments.
Comparison of differences in biofilm formation by different Lactobacillus strains.
To detect possible Lactobacillus strain differences in biofilm formation, as was done previously for adhesion properties (47, 48), we chose seven closely related strains of the Lactobacillus casei group for a comparative study. Strains were selected that showed comparable rates of planktonic growth in the different media used in the biofilm assay (data not shown). The ability to form biofilm in vitro varied extensively among L. rhamnosus and related L. casei strains. Under all conditions tested, L. rhamnosus GG showed by far the best biofilm formation properties on polystyrene (Fig. 1).
Influence of growth medium components.
Since L. rhamnosus GG does not form biofilms in the standard MRS medium, the influence of some key components was investigated. Therefore, compounds were systematically omitted from MRS medium and were used to supplement mTSB medium. When glucose, the main carbon source for lactic acid bacteria (24), was omitted from MRS medium, biofilm formation could be observed (Fig. 2A). This biofilm-repressing effect of glucose was confirmed by addition of glucose to mTSB medium, which resulted in reduction of biofilm formation (Fig. 2A). In contrast to L. rhamnosus GG, no biofilm formation in glucose-depleted MRS medium could be observed for the other L. rhamnosus and L. casei strains tested (Fig. 1).
FIG. 2.
Influence of MRS medium factors on biofilm formation by L. rhamnosus GG. (A) Different components were omitted (−) from MRS medium to investigate their effect on biofilm formation. Additionally, the influence of these components was investigated after addition (+) to mTSB medium in the same concentration as that present in MRS medium. The results are expressed compared to biofilm formation of L. rhamnosus GG in mTSB medium (positive control), which was taken as 100% (dotted line). (B) The effect of the same components on 72-h growth in suspension (OD600) was investigated after omission from MRS medium (−) and addition to mTSB medium (+).
Divalent cations are known to affect bacterial adhesion by a direct effect on electrostatic interactions. MnSO4 (0.05 g/liter) exerted an inhibitory effect on biofilm formation, while no major effect of MgSO4 (0.1 g/liter) on biofilm formation could be observed (Fig. 2A). This difference is most likely due to a different growth-stimulating effect. As was previously described for lactobacilli (22) and confirmed in this study for L. rhamnosus GG, Mn2+ strongly stimulated growth in suspension, while no major effect was seen for Mg2+ (Fig. 2B). The surfactant Tween 80 present in MRS medium also inhibited biofilm formation (Fig. 2A). However, the effect of Tween 80 appears to be medium specific since its addition to mTSB medium did not reduce biofilm formation of L. rhamnosus GG.
Influence of gastrointestinal environment-related conditions.
The influence of some factors was tested in both mTSB and AOAC medium since the growth yield of L. rhamnosus GG is quite different in these media (i.e., 0.5 and 1.5 as the final OD600 reached, respectively).
(i) pH and osmolarity.
Important stressful conditions that are encountered by intestinal bacteria include changing pH and osmolarity. To test their impact on biofilm formation, conditions were chosen that did not considerably influence growth in suspension (data not shown). Biofilm formation by L. rhamnosus GG was inhibited at an initial pH of 4.0 in all media tested, in contrast to neutral pH in the control media (Fig. 3). Increasing osmolarity and ionic strength to 0.3 M NaCl only slightly diminished biofilm formation by L. rhamnosus GG (Fig. 3).
FIG. 3.
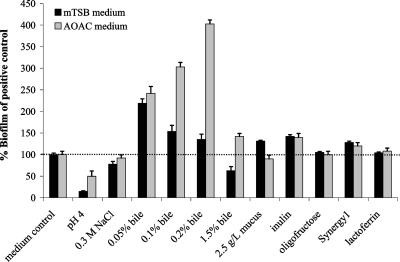
Influence of gastrointestinal tract-mimicking conditions on biofilm formation by L. rhamnosus GG. The effect of addition of mucus (2.5 g/liter), inulin-type prebiotics (20 g/liter), bile (0.05 to 2.0%), and lactoferrin (100 μg/ml) to mTSB (black) and AOAC medium (gray) was investigated. Additionally, the influence of changing the pH and osmolarity to 0.3 M NaCl of the biofilm medium was assessed. Biofilm formations in the supplemented mTSB and AOAC media were compared to those of L. rhamnosus GG in unsupplemented mTSB and AOAC media (positive controls, taken as 100% [dotted line]), respectively.
(ii) Bile.
Bile acids are surface-active molecules present in the human intestine in physiological concentrations ranging from 0.1 to 2.0% (20). They possess potent antimicrobial activity, but many enteric and probiotic bacteria such as L. rhamnosus GG have developed mechanisms to resist bile (12). Bile acids have been previously shown to increase adhesion and biofilm formation of enteric pathogens (20, 37). We also observed a two- to fourfold increase in biofilm formation in AOAC medium after addition of 0.05 to 0.2% bile acids to the medium. The effect was less pronounced when 1.5% bile acids was added (Fig. 3), probably because the tolerance of bile by gram-positive bacteria rapidly decreases at concentrations over 0.3% (2). Addition of 0.05% bile to mTSB medium also increased biofilm formation around twofold, but increasing the concentration of bile gradually decreased biofilm formation (Fig. 3). Therefore, in mTSB medium, the activity of the surface-active bile acids seems to be partly masked. Soy proteins, present in TSB, have been shown to bind bile acids and aggregate them (2).
(iii) Mucins.
The mucus layer covering the epithelial cells is considered an important site for bacterial adhesion and colonization (40, 43). Mucus has previously been reported to have positive and negative effects on biofilm formation, depending on the bacterial species tested. Mucins stimulated biofilm formation by commensal E. coli (4), while they inhibited biofilm formation of the stomach colonizer Helicobacter pylori (8). When we added mucins to mTSB medium in a concentration of 2.5 g/liter, estimated to be present in the transverse colon (31), we observed an increase of biofilm formation by L. rhamnosus GG of more than 20% (Fig. 3). However, the effect of mucus on L. rhamnosus GG seems to be medium dependent. In AOAC medium, mucins did not stimulate biofilm formation (Fig. 3).
(iv) Nondigestible carbohydrates.
Undigested complex carbohydrates especially have a role in the lower gastrointestinal environment. Here, we investigated the influence of inulin-type prebiotics that cannot be fermented by L. rhamnosus GG (25). Addition of inulin or its derivative Synergy1 to mTSB or AOAC medium at 20 g/liter increased biofilm formation up to 1.5-fold (Fig. 3). This biofilm-enhancing effect was mainly observed for the long-chain polysaccharides (Fig. 3).
(v) Antimicrobial peptides.
The presence of different antimicrobial peptides close to the gut epithelium probably plays an important role in the absence of commensal bacteria directly adherent to the gut epithelium in healthy subjects (44). The iron-chelating peptide lactoferrin, expressed by glandular epithelial cells and neutrophils, was recently reported to block biofilm development on mucosal surfaces by the opportunistic pathogen Pseudomonas aeruginosa (concentration tested, 20 μg/ml) (42). However, no effect of lactoferrin on biofilm development of the probiotic bacterium L. rhamnosus GG could be observed at concentrations of 20 and 100 μg/ml (Fig. 3).
Role of EPS.
EPS play a crucial role in biofilm development (6). We chose to analyze the effect of knocking down the wzb homologue, since Wzb is a phosphotyrosine protein phosphatase that has been shown to regulate EPS biosynthesis and polymer size in combination with a protein tyrosine kinase in different bacteria such as E. coli (50) and Streptococcus pneumoniae (32). In Burkholderia cepacia, biofilm formation is reduced when the wzb homologue bceD is inactivated (14). We first isolated and identified the L. rhamnosus wzb homologue and investigated its role in L. rhamnosus GG biofilm formation by antisense RNA technology as previously described (5). Overexpression of antisense RNA of wzb by pCMPG5344 showed medium-dependent effects on biofilm formation. Silencing of wzb expression resulted in the largest reduction in biofilm formation in AOAC medium, while in other media only minor effects were observed (Fig. 4A). To investigate a possible link between biofilm formation and EPS production, EPS fractions of L. rhamnosus GG grown in different media were isolated (Fig. 5). EPS production was shown to be medium dependent. AOAC medium induced the highest EPS production by L. rhamnosus GG (Fig. 5). This suggests, together with the results for pCMPG5344 (wzb antisense), that biofilm formation seems to be especially EPS dependent in AOAC medium.
FIG. 4.
Influence of genetic factors on biofilm formation by L. rhamnosus GG. (A) The influence of EPS was investigated after overexpression of antisense wzb RNA in strain CMPG5344 of L. rhamnosus GG in different growth media: mTSB, AOAC, and MRS medium without glucose (MRS − glc). Biofilm formation was then compared to that of L. rhamnosus GG transformed with the empty cloning vector pLAB1301 (positive control). Erythromycin was added for stable maintenance of the plasmids. (B) The influence of d-alanylation of LTA on biofilm formation was investigated by analysis of the phenotype of a dltD mutant, CMPG5540, under the same conditions. Biofilm formation was then compared to that of wild-type L. rhamnosus GG grown under the same conditions (positive control). (C) The influence of central metabolism was investigated by analysis of the phenotype of the luxS mutant CMPG5412 under different conditions. Biofilm formation of the mutants was compared to that of wild-type L. rhamnosus GG (positive control) under the same conditions, which was taken as 100% (dotted line).
FIG. 5.
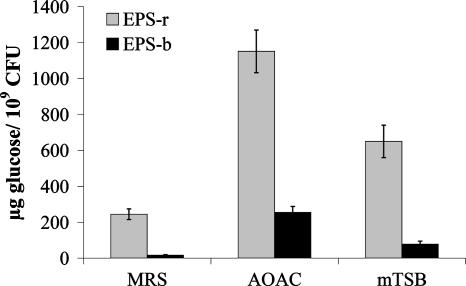
Comparison of levels of EPS production by L. rhamnosus GG in different media. Two EPS fractions were differentiated: EPS-b and EPS-r. EPS fractions were isolated from stationary-phase cultures of L. rhamnosus GG grown in MRS medium (OD600, ∼2.0), AOAC medium (OD600, ∼1.5), and mTSB medium (OD600, ∼0.5). Since the final OD differs considerably in the different media, results are expressed as μg of glucose equivalents produced per 109 CFU.
Role of LTA.
It has previously been suggested that lipoteichoic acids (LTA) are involved in biofilm formation of Lactobacillus strains on the mouse gastric epithelium (41, 51) and in adhesion of L. johnsonii La1 to human enterocyte-like Caco-2 cells (16). Teichoic acids constitute up to 50% of the cell wall dry weight of gram-positive bacteria, and d-alanyl ester substitutions of LTA are directly related to surface charge. We have recently shown that insertional inactivation of the gene encoding one of the enzymes important for LTA synthesis (i.e., dltD) resulted in complete absence of d-alanyl esters in the LTA and in modification of different cell surface properties of L. rhamnosus GG (35). Here, the biofilm formation capacity of the dltD mutant CMPG5540 was investigated under different culture conditions (Fig. 4B). Surprisingly, in contrast to similar mutations in Staphylococcus aureus (18) and Enterococcus faecalis (13), the L. rhamnosus GG dltD mutant showed more biofilm formation to polystyrene, especially in mTSB medium and MRS medium without glucose.
Role of LuxS and central metabolism.
The activity of LuxS is widely investigated in pathogenic bacteria in relation to biofilm formation due to its role in the biosynthesis of the interspecies signaling molecule AI-2 (49). In the recycling of S-adenosylmethionine (SAM), the LuxS enzyme catalyzes the conversion of S-ribosylhomocysteine, yielding AI-2 and homocysteine. However, since the LuxS enzyme forms an integral part of the activated methyl cycle and amino acid metabolism of methionine and cysteine, the analysis of phenotypes of luxS mutants is multifaceted (52). Previously, we have shown that the L. rhamnosus GG luxS mutant CMPG5412 is affected in biofilm formation in AOAC medium. However, this defect could not be complemented by adding the signaling molecule AI-2, whereas it could be nutritionally complemented with cysteine (27). In this study, we compared the biofilm formation capacities of CMPG5412 in the different media used in this study (Fig. 4C). The biofilm defect of the luxS mutant was medium dependent. In a nitrogen-rich environment such as mTSB medium, CMPG5412 was not impaired in biofilm formation, in contrast to the situation in AOAC medium (Fig. 4C). This is in agreement with our previously published results where we showed that the biofilm defect of CMPG5412 has an important metabolic nature and is not merely due to disrupted AI-2-mediated communication (27).
DISCUSSION
The colonization capacity of probiotic strains is generally studied in vitro by using mucus or intestinal epithelial cell lines like Caco-2 and HT-29 cells (47, 48). These models are representative for the initial attachment of ingested probiotics to the intestinal wall but are not suitable for studying further more dynamic steps in colonization: i.e., microcolony formation and structural development to biofilm-like communities. Evaluation of biofilm formation in different media is a complementary approach to better understand the mechanisms by which bacteria adapt to environmental stresses and colonize different niches. Here, we report that the widely investigated probiotic strain L. rhamnosus GG is able to form biofilms in vitro, in contrast to related lactobacilli tested. In addition, we studied several factors influencing biofilm formation. To our knowledge, the present study constitutes the most extensive analysis of in vitro biofilm formation by lactobacilli published thus far.
We can conclude from our comparative analysis that the intrinsic biofilm formation capacity of L. rhamnosus GG is strongly dependent on environmental factors and the culture medium used. In general, there appeared to be an inverse relationship between extent of biofilm formation and growth in suspension (Fig. 2). Nutrient limitation and especially low availability of fermentable carbon sources, such as in mTSB medium with its low C/N ratio, stimulated biofilm growth of L. rhamnosus GG. However, growth limitation does not seem to be sufficient to induce biofilm formation in Lactobacillus strains, as shown by a comparative study with other Lactobacillus strains (Fig. 1).
The underlying genetic factors determining the good biofilm formation capacity of L. rhamnosus GG still need to be characterized, since it is probably mediated by various genetic pathways. In this study, we investigated the role of EPS (wzb), LTA (dltD), and central metabolism (luxS). In most bacteria, EPS production is important for biofilm formation (6). L. rhamnosus GG is not known as a high EPS producer in MRS medium in comparison with other lactobacilli (36), although we have demonstrated in this study that EPS production of L. rhamnosus GG is highly medium dependent (Fig. 5). The importance of EPS in biofilm formation by L. rhamnosus GG seems to be dependent on the culture conditions. Indeed, AOAC medium induced the highest EPS production (Fig. 5), while the best biofilm formation was observed in mTSB medium (Fig. 1). Also, a wzb-knockdown mutant of L. rhamnosus GG was shown to be particularly impaired in biofilm formation in AOAC medium. Therefore, EPS production is clearly not the only factor determining biofilm formation of L. rhamnosus GG. For instance, besides EPS production quantity, the polymer size and specific chemical structure and composition of the EPS molecules could also affect biofilm formation. Moreover, in addition to EPS, other surface components are most likely involved in biofilm formation, as exemplified by the dltD mutant. Although dltD inactivation resulted in increased negative surface charge (35), we observed more biofilm formation on polystyrene, depending on the culture medium. This is probably due to indirect effects of altered LTA on biofilm development. For instance, absence of d-alanyl esters in teichoic acids has been shown to alter folding of exoproteins in Bacillus subtilis (21) and to cause loss of large surface protein adhesins in Streptococcus gordonii (7). Altered surface proteins might lead to altered physicochemical properties of the cell surface. Additionally, differences in central metabolism could also partly account for the variation in biofilm formation capacity between lactobacillus strains, as exemplified by contrasting biofilm phenotypes in Lactobacillus reuteri (46) and L. rhamnosus GG (27) luxS mutants.
Biofilms of lactobacilli can be found in many natural environments. Since the gastrointestinal tract is an important target for probiotics, some factors related to this niche were investigated in this study. Our data show that conditions such as low pH, high osmolarity, and the presence of bile and mucins highly modulate biofilm formation of L. rhamnosus GG, but the effect of each factor depends on the microenvironment (i.e., the culture medium). Some factors such as mucins probably facilitate binding of L. rhamnosus GG to substrates. L. rhamnosus GG was even previously shown to induce mucin expression (29), which could be a way for surface conditioning of the microhabitat to increase its colonization capacity. Additionally, L. rhamnosus GG has excellent in vitro mucus-adhering properties (47), which are mediated in many lactobacilli by specific mucus-binding proteins (3). The fact that the biofilm-promoting effect of mucus is less pronounced in the EPS-promoting AOAC medium might be due to shielding of these mucus-binding proteins by EPS. Additionally, undigested food particles may affect the structural arrangement of the microbiota in the intestine (43). It seems plausible that the observed increased biofilm formation of L. rhamnosus GG by inulin is due to stimulation of aggregation. These complex polysaccharides might be incorporated in the extracellular matrix, thereby enhancing biofilm formation. Other factors such as pH and bile acids may greatly affect the cell surface of L. rhamnosus GG itself and thereby influence biofilm development. Bile stimulated biofilm formation of L. rhamnosus GG in concentrations not affecting growth. As shown for Vibrio cholerae (20), bile might therefore be a signal for L. rhamnosus GG to form biofilms in the small intestine. It will be of interest to investigate in future studies differential gene expression of L. rhamnosus GG in the presence of bile. In some bacteria, bile induced the expression of specific adhesins such as EPS (20) and fimbria-like appendages (37). Additionally, biofilm formation of L. rhamnosus GG was shown to be more sensitive to low pH than growth in suspension, as was also reported for Streptococcus gordonii (28). However, biofilm formation is a complex process. The crystal violet method used in this study assesses the final outcome of initial bacterial attachment and biofilm maturation. It might well be that low pH stimulates initial adhesion, as was shown for adhesion to epithelial cells (17), but that the development to mature biofilms is inhibited by low pH. Interestingly, some lactobacilli are able to colonize in biofilm-like communities the acidic stomach of rodents (46). This clearly demonstrates that further studies, including in vivo imaging such as fluorescence in situ hybridization, are needed to investigate the biofilm formation capacities of exogenously applied and endogenous lactobacilli in different compartments (e.g., stomach, small intestine, and colon) and niches (e.g., mucous layer, mucosa, and food residues) and even at other sites outside the gastrointestinal tract.
Acknowledgments
S.L. and S.C.J.D.K. are research assistants of the Fund for Scientific Research (FWO-Vlaanderen).
We thank E. Steenackers and S. Tejero for valuable technical assistance. We gratefully acknowledge D. Bosscher from ORAFTI (Belgium) for kindly providing the inulin-type prebiotics used in this study.
Footnotes
Published ahead of print on 7 September 2007.
REFERENCES
- 1.Alander, M., R. Satokari, R. Korpela, M. Saxelin, T. Vilpponen-Salmela, T. Mattila-Sandholm, and A. von Wright. 1999. Persistence of colonization of human colonic mucosa by a probiotic strain, Lactobacillus rhamnosus GG, after oral consumption. Appl. Environ. Microbiol. 65:351-354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Begley, M., C. G. Gahan, and C. Hill. 2005. The interaction between bacteria and bile. FEMS Microbiol. Rev. 29:625-651. [DOI] [PubMed] [Google Scholar]
- 3.Boekhorst, J., Q. Helmer, M. Kleerebezem, and R. J. Siezen. 2006. Comparative analysis of proteins with a mucus-binding domain found exclusively in lactic acid bacteria. Microbiology 152:273-280. [DOI] [PubMed] [Google Scholar]
- 4.Bollinger, R. R., M. L. Everett, D. Palestrant, S. D. Love, S. S. Lin, and W. Parker. 2003. Human secretory immunoglobulin A may contribute to biofilm formation in the gut. Immunology 109:580-587. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Bouazzaoui, K., and G. LaPointe. 2006. Use of antisense RNA to modulate glycosyltransferase gene expression and exopolysaccharide molecular mass in Lactobacillus rhamnosus. J. Microbiol. Methods 65:216-225. [DOI] [PubMed] [Google Scholar]
- 6.Branda, S. S., S. Vik, L. Friedman, and R. Kolter. 2005. Biofilms: the matrix revisited. Trends Microbiol. 13:20-26. [DOI] [PubMed] [Google Scholar]
- 7.Clemans, D. L., P. E. Kolenbrander, D. V. Debabov, Q. Zhang, R. D. Lunsford, H. Sakone, C. J. Whittaker, M. P. Heaton, and F. C. Neuhaus. 1999. Insertional inactivation of genes responsible for the d-alanylation of lipoteichoic acid in Streptococcus gordonii DL1 (Challis) affects intrageneric coaggregations. Infect. Immun. 67:2464-2474. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Cole, S. P., J. Harwood, R. Lee, R. She, and D. G. Guiney. 2004. Characterization of monospecies biofilm formation by Helicobacter pylori. J. Bacteriol. 186:3124-3132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.De Keersmaecker, S. C. J., K. Braeken, T. L. A. Verhoeven, M. Perea Vélez, S. Lebeer, J. Vanderleyden, and P. Hols. 2006. Flow cytometric testing of green fluorescent protein-tagged Lactobacillus rhamnosus GG for response to defensins. Appl. Environ. Microbiol. 72:4923-4930. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.De Keersmaecker, S. C., C. Varszegi, N. van Boxel, L. W. Habel, K. Metzger, R. Daniels, K. Marchal, D. De Vos, and J. Vanderleyden. 2005. Chemical synthesis of (S)-4,5-dihydroxy-2,3-pentanedione, a bacterial signal molecule precursor, and validation of its activity in Salmonella typhimurium. J. Biol. Chem. 280:19563-19568. [DOI] [PubMed] [Google Scholar]
- 11.de Man, J. C., M. Rogosa, and M. E. Sharpe. 1960. A medium for the cultivation of lactobacilli. J. Appl. Bacteriol. 23:130-135. [Google Scholar]
- 12.Doron, S., D. R. Snydman, and S. L. Gorbach. 2005. Lactobacillus GG: bacteriology and clinical applications. Gastroenterol. Clin. N. Am. 34:483-498. [DOI] [PubMed] [Google Scholar]
- 13.Fabretti, F., C. Theilacker, L. Baldassarri, Z. Kaczynski, A. Kropec, O. Holst, and J. Huebner. 2006. Alanine esters of enterococcal lipoteichoic acid play a role in biofilm formation and resistance to antimicrobial peptides. Infect. Immun. 74:4164-4171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Ferreira, A. S., J. H. Leitão, S. A. Sousa, A. M. Cosme, I. Sá-Correia, and L. M. Moreira. 2007. Functional analysis of Burkholderia cepacia genes bceD and bceF, encoding a phosphotyrosine phosphatase and a tyrosine autokinase, respectively: role in exopolysaccharide biosynthesis and biofilm formation. Appl. Environ. Microbiol. 73:524-534. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gorbach, S. L. 1996. The discovery of Lactobacillus GG. Nutr. Today 31:2S-4S. [Google Scholar]
- 16.Granato, D., F. Perotti, I. Masserey, M. Rouvet, M. Golliard, A. Servin, and D. Brassart. 1999. Cell surface-associated lipoteichoic acid acts as an adhesion factor for attachment of Lactobacillus johnsonii La1 to human enterocyte-like Caco-2 cells. Appl. Environ. Microbiol. 65:1071-1077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Greene, J. D., and T. R. Klaenhammer. 1994. Factors involved in adherence of lactobacilli to human Caco-2 cells. Appl. Environ. Microbiol. 60:4487-4494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Gross, M., S. E. Cramton, F. Götz, and A. Peschel. 2001. Key role of teichoic acid net charge in Staphylococcus aureus colonization of artificial surfaces. Infect. Immun. 69:3423-3426. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Hudault, S., V. Lievin, M.-F. Bernet-Camard, and A. L. Servin. 1997. Antagonistic activity exerted in vitro and in vivo by Lactobacillus casei (strain GG) against Salmonella typhimurium C5 infection. Appl. Environ. Microbiol. 63:513-518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Hung, D. T., J. Zhu, D. Sturtevant, and J. J. Mekalanos. 2006. Bile acids stimulate biofilm formation in Vibrio cholerae. Mol. Microbiol. 59:193-201. [DOI] [PubMed] [Google Scholar]
- 21.Hyyrylainen, H. L., M. Vitikainen, J. Thwaite, H. Wu, M. Sarvas, C. R. Harwood, V. P. Kontinen, and K. Stephenson. 2000. D-Alanine substitution of teichoic acids as a modulator of protein folding and stability at the cytoplasmic membrane/cell wall interface of Bacillus subtilis. J. Biol. Chem. 275:26696-26703. [DOI] [PubMed] [Google Scholar]
- 22.Imbert, M., and R. Blondeau. 1998. On the iron requirement of lactobacilli grown in chemically defined medium. Curr. Microbiol. 37:64-66. [DOI] [PubMed] [Google Scholar]
- 23.Josson, K., T. Scheirlinck, F. Michiels, C. Platteeuw, P. Stanssens, H. Joos, P. Dhaese, M. Zabeau, and J. Mahillon. 1989. Characterization of a gram-positive broad-host-range plasmid isolated from Lactobacillus hilgardii. Plasmid 21:9-20. [DOI] [PubMed] [Google Scholar]
- 24.Kandler, O., and N. Weiss. 1986. Genus Lactobacillus, p. 1063-1065. In P. H. A. Sneath, N. S. Mair, M. S. Sharpe, and J. G. Holt (ed.), Bergey's manual of systematic bacteriology. Williams & Wilkins, Baltimore, MD.
- 25.Kaplan, H., and R. W. Hutkins. 2000. Fermentation of fructooligosaccharides by lactic acid bacteria and bifidobacteria. Appl. Environ. Microbiol. 66:2682-2684. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Kolter, R., and E. P. Greenberg. 2006. Microbial sciences: the superficial life of microbes. Nature 441:300-302. [DOI] [PubMed] [Google Scholar]
- 27.Lebeer, S., S. C. J. De Keersmaecker, T. L. A. Verhoeven, A. A. Fadda, K. Marchal, and J. Vanderleyden. 2007. Functional analysis of luxS in the probiotic strain Lactobacillus rhamnosus GG reveals a central metabolic role important for growth and biofilm formation. J. Bacteriol. 189:860-871. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Loo, C. Y., D. A. Corliss, and N. Ganeshkumar. 2000. Streptococcus gordonii biofilm formation: identification of genes that code for biofilm phenotypes. J. Bacteriol. 182:1374-1382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Mattar, A. F., D. H. Teitelbaum, R. A. Drongowski, F. Yongyi, C. M. Harmon, and A. G. Coran. 2002. Probiotics up-regulate MUC-2 mucin gene expression in a Caco-2 cell-culture model. Pediatr. Surg. Int. 18:586-590. [DOI] [PubMed] [Google Scholar]
- 30.Meurman, J. H., H. Antila, and S. Salminen. 1994. Recovery of Lactobacillus strain GG (ATCC53103) from saliva of healthy volunteers after consumption of yoghurt prepared with the bacterium. Microb. Ecol. Health Dis. 7:295-298. [Google Scholar]
- 31.Molly, K., M. Vande Woestyne, and W. Verstraete. 1993. Development of a 5-step multi-chamber reactor as a simulation of the human intestinal microbial ecosystem. Appl. Microbiol. Biotechnol. 39:254-258. [DOI] [PubMed] [Google Scholar]
- 32.Morona, J. K., R. Morona, D. C. Miller, and J. C. Paton. 2002. Streptococcus pneumoniae capsule biosynthesis protein CpsB is a novel manganese-dependent phosphotyrosine-protein phosphatase. J. Bacteriol. 184:577-583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Nase, L., K. Hatakka, E. Savilahti, M. Saxelin, A. Pönkä, T. Poussa, R. Korpela, and J. H. Meurman. 2001. Effect of long-term consumption of a probiotic bacterium, Lactobacillus rhamnosus GG, in milk on dental caries and caries risk in children. Caries Res. 35:412-420. [DOI] [PubMed] [Google Scholar]
- 34.Peant, B., G. LaPointe, C. Gilbert, D. Atlan, P. Ward, and D. Roy. 2005. Comparative analysis of the exopolysaccharide biosynthesis gene clusters from four strains of Lactobacillus rhamnosus. Microbiology 151:1839-1851. [DOI] [PubMed] [Google Scholar]
- 35.Perea Vélez, M., T. L. A. Verhoeven, C. Draing, S. Von Aulock, M. Pfitzenmaier, A. Geyer, I. Lambrichts, C. Grangette, B. Pot, J. Vanderleyden, and S. C. J. De Keersmaecker. 2007. Functional analysis of d-alanylation of lipoteichoic acid in the probiotic strain Lactobacillus rhamnosus GG. Appl. Environ. Microbiol. 73:3595-3604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Provencher, C., G. LaPointe, S. Sirois, M.-R. Van Calsteren, and D. Roy. 2003. Consensus-degenerate hybrid oligonucleotide primers for amplification of priming glycosyltransferase genes of the exopolysaccharide locus in strains of the Lactobacillus casei group. Appl. Environ. Microbiol. 69:3299-3307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Pumbwe, L., C. A. Skilback, V. Nakano, M. J. Avila-Campos, R. M. Piazza, and H. M. Wexler. 2007. Bile salts enhance bacterial co-aggregation, bacterial-intestinal epithelial cell adhesion, biofilm formation and antimicrobial resistance of Bacteroides fragilis. Microb. Pathog. 43:78-87. [DOI] [PubMed] [Google Scholar]
- 38.Sambrook, J., E. F. Fritsch, and T. Maniatis. 1989. Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY.
- 39.Schultz, M., C. Gottl, R. J. Young, P. Iwen, and J. A. Vanderhoof. 2004. Administration of oral probiotic bacteria to pregnant women causes temporary infantile colonization. J. Pediatr. Gastroenterol. Nutr. 38:293-297. [DOI] [PubMed] [Google Scholar]
- 40.Servin, A. L., and M. H. Coconnier. 2003. Adhesion of probiotic strains to the intestinal mucosa and interaction with pathogens. Best Pract. Res. Clin. Gastroenterol. 17:741-754. [DOI] [PubMed] [Google Scholar]
- 41.Sherman, L. A., and D. C. Savage. 1986. Lipoteichoic acids in Lactobacillus strains that colonize the mouse gastric epithelium. Appl. Environ. Microbiol. 52:302-304. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Singh, P. K., M. R. Parsek, E. P. Greenberg, and M. J. Welsh. 2002. A component of innate immunity prevents bacterial biofilm development. Nature 417:552-555. [DOI] [PubMed] [Google Scholar]
- 43.Sonnenburg, J. L., L. T. Angenent, and J. I. Gordon. 2004. Getting a grip on things: how do communities of bacterial symbionts become established in our intestine? Nat. Immunol. 5:569-573. [DOI] [PubMed] [Google Scholar]
- 44.Swidsinski, A., J. Weber, V. Loening-Baucke, L. P. Hale, and H. Lochs. 2005. Spatial organization and composition of the mucosal flora in patients with inflammatory bowel disease. J. Clin. Microbiol. 43:3380-3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Tallon, R., P. Bressollier, and M. C. Urdaci. 2003. Isolation and characterization of two exopolysaccharides produced by Lactobacillus plantarum EP56. Res. Microbiol. 154:705-712. [DOI] [PubMed] [Google Scholar]
- 46.Tannock, G. W., S. Ghazally, J. Walter, D. Loach, H. Brooks, G. Cook, M. Surette, C. Simmers, P. Bremer, F. Dal Bello, and C. Hertel. 2005. Ecological behavior of Lactobacillus reuteri 100-23 is affected by mutation of the luxS gene. Appl. Environ. Microbiol. 71:8419-8425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Tuomola, E. M., A. C. Ouwehand, and S. J. Salminen. 1999. The effect of probiotic bacteria on the adhesion of pathogens to human intestinal mucus. FEMS Immunol. Med. Microbiol. 26:137-142. [DOI] [PubMed] [Google Scholar]
- 48.Tuomola, E. M., and S. J. Salminen. 1998. Adhesion of some probiotic and dairy Lactobacillus strains to Caco-2 cell cultures. Int. J. Food Microbiol. 41:45-51. [DOI] [PubMed] [Google Scholar]
- 49.Vendeville, A., K. Winzer, K. Heurlier, C. M. Tang, and K. R. Hardie. 2005. Making ‘sense’ of metabolism: autoinducer-2, LuxS and pathogenic bacteria. Nat. Rev. Microbiol. 3:383-396. [DOI] [PubMed] [Google Scholar]
- 50.Vincent, C., P. Doublet, C. Grangeasse, E. Vaganay, A. J. Cozzone, and B. Duclos. 1999. Cells of Escherichia coli contain a protein-tyrosine kinase, Wzc, and a phosphotyrosine-protein phosphatase, Wzb. J. Bacteriol. 181:3472-3477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Walter, J., D. M. Loach, M. Alqumber, C. Rockel, C. Hermann, M. Pfitzenmaier, and G. W. Tannock. 2007. d-Alanyl ester depletion of teichoic acids in Lactobacillus reuteri 100-23 results in impaired colonization of the mouse gastrointestinal tract. Environ. Microbiol. 9:1750-1760. [DOI] [PubMed] [Google Scholar]
- 52.Winzer, K., K. R. Hardie, and P. Williams. 2003. LuxS and autoinducer-2: their contribution to quorum sensing and metabolism in bacteria. Adv. Appl. Microbiol. 53:291-396. [DOI] [PubMed] [Google Scholar]