Abstract
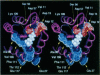
CheY is the response regulator protein that interacts with the flagellar switch apparatus to modulate flagellar rotation during chemotactic signaling. CheY can be phosphorylated and dephosphorylated in vitro, and evidence indicates that CheY-P is the activated form that induces clockwise flagellar rotation, resulting in a tumble in the cell's swimming pattern. The flagellar switch apparatus is a complex macromolecular structure composed of at least three gene products, FliG, FliM, and FliN. Genetic analysis of Escherichia coli has identified fliG and fliM as genes in which mutations occur that allele specifically suppress cheY mutations, indicating interactions among these gene products. We have generated a class of cheY mutations selected for dominant suppression of fliG mutations. Interestingly, these cheY mutations dominantly suppressed both fliG and fliM mutations; this is consistent with the idea that the CheY protein interacts with both switch gene products during signaling. Biochemical characterization of wild-type and suppressor CheY proteins did not reveal altered phosphorylation properties or evidence for phosphorylation-dependent CheY multimerization. These data indicate that suppressor CheY proteins are specifically altered in the ability to transduce chemotactic signals to the switch at some point subsequent to phosphorylation. Physical mapping of suppressor amino acid substitutions on the crystal structure of CheY revealed a high degree of spatial clustering, suggesting that this region of CheY is a signaling surface that transduces chemotactic signals to the switch.
Full text
PDF



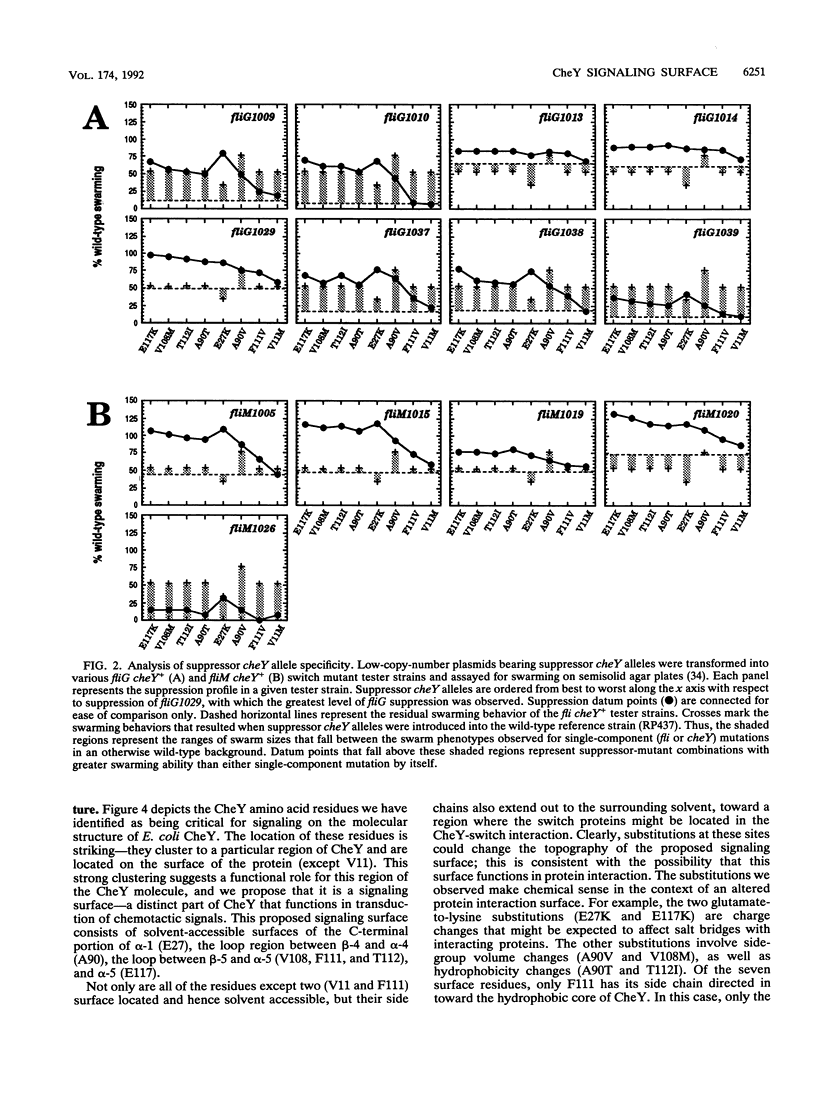




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Adams A. E., Botstein D. Dominant suppressors of yeast actin mutations that are reciprocally suppressed. Genetics. 1989 Apr;121(4):675–683. doi: 10.1093/genetics/121.4.675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Adams A. E., Botstein D., Drubin D. G. A yeast actin-binding protein is encoded by SAC6, a gene found by suppression of an actin mutation. Science. 1989 Jan 13;243(4888):231–233. doi: 10.1126/science.2643162. [DOI] [PubMed] [Google Scholar]
- Albright L. M., Huala E., Ausubel F. M. Prokaryotic signal transduction mediated by sensor and regulator protein pairs. Annu Rev Genet. 1989;23:311–336. doi: 10.1146/annurev.ge.23.120189.001523. [DOI] [PubMed] [Google Scholar]
- Amaya E., Musci T. J., Kirschner M. W. Expression of a dominant negative mutant of the FGF receptor disrupts mesoderm formation in Xenopus embryos. Cell. 1991 Jul 26;66(2):257–270. doi: 10.1016/0092-8674(91)90616-7. [DOI] [PubMed] [Google Scholar]
- Block S. M., Segall J. E., Berg H. C. Adaptation kinetics in bacterial chemotaxis. J Bacteriol. 1983 Apr;154(1):312–323. doi: 10.1128/jb.154.1.312-323.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Botstein D., Maurer R. Genetic approaches to the analysis of microbial development. Annu Rev Genet. 1982;16:61–83. doi: 10.1146/annurev.ge.16.120182.000425. [DOI] [PubMed] [Google Scholar]
- Bourret R. B., Borkovich K. A., Simon M. I. Signal transduction pathways involving protein phosphorylation in prokaryotes. Annu Rev Biochem. 1991;60:401–441. doi: 10.1146/annurev.bi.60.070191.002153. [DOI] [PubMed] [Google Scholar]
- Bourret R. B., Hess J. F., Simon M. I. Conserved aspartate residues and phosphorylation in signal transduction by the chemotaxis protein CheY. Proc Natl Acad Sci U S A. 1990 Jan;87(1):41–45. doi: 10.1073/pnas.87.1.41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bowie J. U., Lüthy R., Eisenberg D. A method to identify protein sequences that fold into a known three-dimensional structure. Science. 1991 Jul 12;253(5016):164–170. doi: 10.1126/science.1853201. [DOI] [PubMed] [Google Scholar]
- Clegg D. O., Koshland D. E., Jr The role of a signaling protein in bacterial sensing: behavioral effects of increased gene expression. Proc Natl Acad Sci U S A. 1984 Aug;81(16):5056–5060. doi: 10.1073/pnas.81.16.5056. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Das A., Merril C., Adhya S. Interaction of RNA polymerase and rho in transcription termination: coupled ATPase. Proc Natl Acad Sci U S A. 1978 Oct;75(10):4828–4832. doi: 10.1073/pnas.75.10.4828. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guarente L. P., Beckwith J. Mutant RNA polymerase of Escherichia coli terminates transcription in strains making defective rho factor. Proc Natl Acad Sci U S A. 1978 Jan;75(1):294–297. doi: 10.1073/pnas.75.1.294. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hartman P. E., Roth J. R. Mechanisms of suppression. Adv Genet. 1973;17:1–105. doi: 10.1016/s0065-2660(08)60170-4. [DOI] [PubMed] [Google Scholar]
- Herskowitz I. Functional inactivation of genes by dominant negative mutations. Nature. 1987 Sep 17;329(6136):219–222. doi: 10.1038/329219a0. [DOI] [PubMed] [Google Scholar]
- Hess J. F., Oosawa K., Kaplan N., Simon M. I. Phosphorylation of three proteins in the signaling pathway of bacterial chemotaxis. Cell. 1988 Apr 8;53(1):79–87. doi: 10.1016/0092-8674(88)90489-8. [DOI] [PubMed] [Google Scholar]
- Hess J. F., Oosawa K., Matsumura P., Simon M. I. Protein phosphorylation is involved in bacterial chemotaxis. Proc Natl Acad Sci U S A. 1987 Nov;84(21):7609–7613. doi: 10.1073/pnas.84.21.7609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huffaker T. C., Hoyt M. A., Botstein D. Genetic analysis of the yeast cytoskeleton. Annu Rev Genet. 1987;21:259–284. doi: 10.1146/annurev.ge.21.120187.001355. [DOI] [PubMed] [Google Scholar]
- Jarvik J., Botstein D. Conditional-lethal mutations that suppress genetic defects in morphogenesis by altering structural proteins. Proc Natl Acad Sci U S A. 1975 Jul;72(7):2738–2742. doi: 10.1073/pnas.72.7.2738. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koop A. H., Hartley M. E., Bourgeois S. A low-copy-number vector utilizing beta-galactosidase for the analysis of gene control elements. Gene. 1987;52(2-3):245–256. doi: 10.1016/0378-1119(87)90051-5. [DOI] [PubMed] [Google Scholar]
- Kossmann M., Wolff C., Manson M. D. Maltose chemoreceptor of Escherichia coli: interaction of maltose-binding protein and the tar signal transducer. J Bacteriol. 1988 Oct;170(10):4516–4521. doi: 10.1128/jb.170.10.4516-4521.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuo S. C., Koshland D. E., Jr Multiple kinetic states for the flagellar motor switch. J Bacteriol. 1989 Nov;171(11):6279–6287. doi: 10.1128/jb.171.11.6279-6287.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu J. D., Parkinson J. S. Genetic evidence for interaction between the CheW and Tsr proteins during chemoreceptor signaling by Escherichia coli. J Bacteriol. 1991 Aug;173(16):4941–4951. doi: 10.1128/jb.173.16.4941-4951.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lukat G. S., Lee B. H., Mottonen J. M., Stock A. M., Stock J. B. Roles of the highly conserved aspartate and lysine residues in the response regulator of bacterial chemotaxis. J Biol Chem. 1991 May 5;266(13):8348–8354. [PubMed] [Google Scholar]
- Magariyama Y., Yamaguchi S., Aizawa S. Genetic and behavioral analysis of flagellar switch mutants of Salmonella typhimurium. J Bacteriol. 1990 Aug;172(8):4359–4369. doi: 10.1128/jb.172.8.4359-4369.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin C., Okamura S., Young R. Genetic exploration of interactive domains in RNA polymerase II subunits. Mol Cell Biol. 1990 May;10(5):1908–1914. doi: 10.1128/mcb.10.5.1908. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsumura P., Rydel J. J., Linzmeier R., Vacante D. Overexpression and sequence of the Escherichia coli cheY gene and biochemical activities of the CheY protein. J Bacteriol. 1984 Oct;160(1):36–41. doi: 10.1128/jb.160.1.36-41.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McNally D. F., Matsumura P. Bacterial chemotaxis signaling complexes: formation of a CheA/CheW complex enhances autophosphorylation and affinity for CheY. Proc Natl Acad Sci U S A. 1991 Jul 15;88(14):6269–6273. doi: 10.1073/pnas.88.14.6269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morris N. R., Lai M. H., Oakley C. E. Identification of a gene for alpha-tubulin in Aspergillus nidulans. Cell. 1979 Feb;16(2):437–442. doi: 10.1016/0092-8674(79)90019-9. [DOI] [PubMed] [Google Scholar]
- Mortin M. A. Use of second-site suppressor mutations in Drosophila to identify components of the transcriptional machinery. Proc Natl Acad Sci U S A. 1990 Jun;87(12):4864–4868. doi: 10.1073/pnas.87.12.4864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakashima K., Kanamaru K., Aiba H., Mizuno T. Signal transduction and osmoregulation in Escherichia coli. A novel type of mutation in the phosphorylation domain of the activator protein, OmpR, results in a defect in its phosphorylation-dependent DNA binding. J Biol Chem. 1991 Jun 15;266(17):10775–10780. [PubMed] [Google Scholar]
- Novick P., Osmond B. C., Botstein D. Suppressors of yeast actin mutations. Genetics. 1989 Apr;121(4):659–674. doi: 10.1093/genetics/121.4.659. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oosawa K., Hess J. F., Simon M. I. Mutants defective in bacterial chemotaxis show modified protein phosphorylation. Cell. 1988 Apr 8;53(1):89–96. doi: 10.1016/0092-8674(88)90490-4. [DOI] [PubMed] [Google Scholar]
- Parkinson J. S. Complementation analysis and deletion mapping of Escherichia coli mutants defective in chemotaxis. J Bacteriol. 1978 Jul;135(1):45–53. doi: 10.1128/jb.135.1.45-53.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parkinson J. S., Houts S. E. Isolation and behavior of Escherichia coli deletion mutants lacking chemotaxis functions. J Bacteriol. 1982 Jul;151(1):106–113. doi: 10.1128/jb.151.1.106-113.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parkinson J. S., Parker S. R. Interaction of the cheC and cheZ gene products is required for chemotactic behavior in Escherichia coli. Proc Natl Acad Sci U S A. 1979 May;76(5):2390–2394. doi: 10.1073/pnas.76.5.2390. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parkinson J. S., Parker S. R., Talbert P. B., Houts S. E. Interactions between chemotaxis genes and flagellar genes in Escherichia coli. J Bacteriol. 1983 Jul;155(1):265–274. doi: 10.1128/jb.155.1.265-274.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parkinson J. S. cheA, cheB, and cheC genes of Escherichia coli and their role in chemotaxis. J Bacteriol. 1976 May;126(2):758–770. doi: 10.1128/jb.126.2.758-770.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ravid S., Matsumura P., Eisenbach M. Restoration of flagellar clockwise rotation in bacterial envelopes by insertion of the chemotaxis protein CheY. Proc Natl Acad Sci U S A. 1986 Oct;83(19):7157–7161. doi: 10.1073/pnas.83.19.7157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanders D. A., Gillece-Castro B. L., Stock A. M., Burlingame A. L., Koshland D. E., Jr Identification of the site of phosphorylation of the chemotaxis response regulator protein, CheY. J Biol Chem. 1989 Dec 25;264(36):21770–21778. [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Slauch J. M., Russo F. D., Silhavy T. J. Suppressor mutations in rpoA suggest that OmpR controls transcription by direct interaction with the alpha subunit of RNA polymerase. J Bacteriol. 1991 Dec;173(23):7501–7510. doi: 10.1128/jb.173.23.7501-7510.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sockett H., Yamaguchi S., Kihara M., Irikura V. M., Macnab R. M. Molecular analysis of the flagellar switch protein FliM of Salmonella typhimurium. J Bacteriol. 1992 Feb;174(3):793–806. doi: 10.1128/jb.174.3.793-806.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spurlino J. C., Lu G. Y., Quiocho F. A. The 2.3-A resolution structure of the maltose- or maltodextrin-binding protein, a primary receptor of bacterial active transport and chemotaxis. J Biol Chem. 1991 Mar 15;266(8):5202–5219. doi: 10.2210/pdb1mbp/pdb. [DOI] [PubMed] [Google Scholar]
- Stock A. M., Mottonen J. M., Stock J. B., Schutt C. E. Three-dimensional structure of CheY, the response regulator of bacterial chemotaxis. Nature. 1989 Feb 23;337(6209):745–749. doi: 10.1038/337745a0. [DOI] [PubMed] [Google Scholar]
- Stock J. B., Stock A. M., Mottonen J. M. Signal transduction in bacteria. Nature. 1990 Mar 29;344(6265):395–400. doi: 10.1038/344395a0. [DOI] [PubMed] [Google Scholar]
- Treptow N. A., Shuman H. A. Allele-specific malE mutations that restore interactions between maltose-binding protein and the inner-membrane components of the maltose transport system. J Mol Biol. 1988 Aug 20;202(4):809–822. doi: 10.1016/0022-2836(88)90560-8. [DOI] [PubMed] [Google Scholar]
- Volz K., Matsumura P. Crystal structure of Escherichia coli CheY refined at 1.7-A resolution. J Biol Chem. 1991 Aug 15;266(23):15511–15519. doi: 10.2210/pdb3chy/pdb. [DOI] [PubMed] [Google Scholar]
- Wolfe A. J., Berg H. C. Migration of bacteria in semisolid agar. Proc Natl Acad Sci U S A. 1989 Sep;86(18):6973–6977. doi: 10.1073/pnas.86.18.6973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wolfe A. J., Conley M. P., Kramer T. J., Berg H. C. Reconstitution of signaling in bacterial chemotaxis. J Bacteriol. 1987 May;169(5):1878–1885. doi: 10.1128/jb.169.5.1878-1885.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wylie D., Stock A., Wong C. Y., Stock J. Sensory transduction in bacterial chemotaxis involves phosphotransfer between Che proteins. Biochem Biophys Res Commun. 1988 Mar 15;151(2):891–896. doi: 10.1016/s0006-291x(88)80365-6. [DOI] [PubMed] [Google Scholar]
- Yamaguchi S., Aizawa S., Kihara M., Isomura M., Jones C. J., Macnab R. M. Genetic evidence for a switching and energy-transducing complex in the flagellar motor of Salmonella typhimurium. J Bacteriol. 1986 Dec;168(3):1172–1179. doi: 10.1128/jb.168.3.1172-1179.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamaguchi S., Fujita H., Ishihara A., Aizawa S., Macnab R. M. Subdivision of flagellar genes of Salmonella typhimurium into regions responsible for assembly, rotation, and switching. J Bacteriol. 1986 Apr;166(1):187–193. doi: 10.1128/jb.166.1.187-193.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]