Abstract
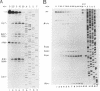
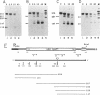
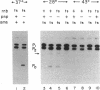
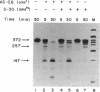
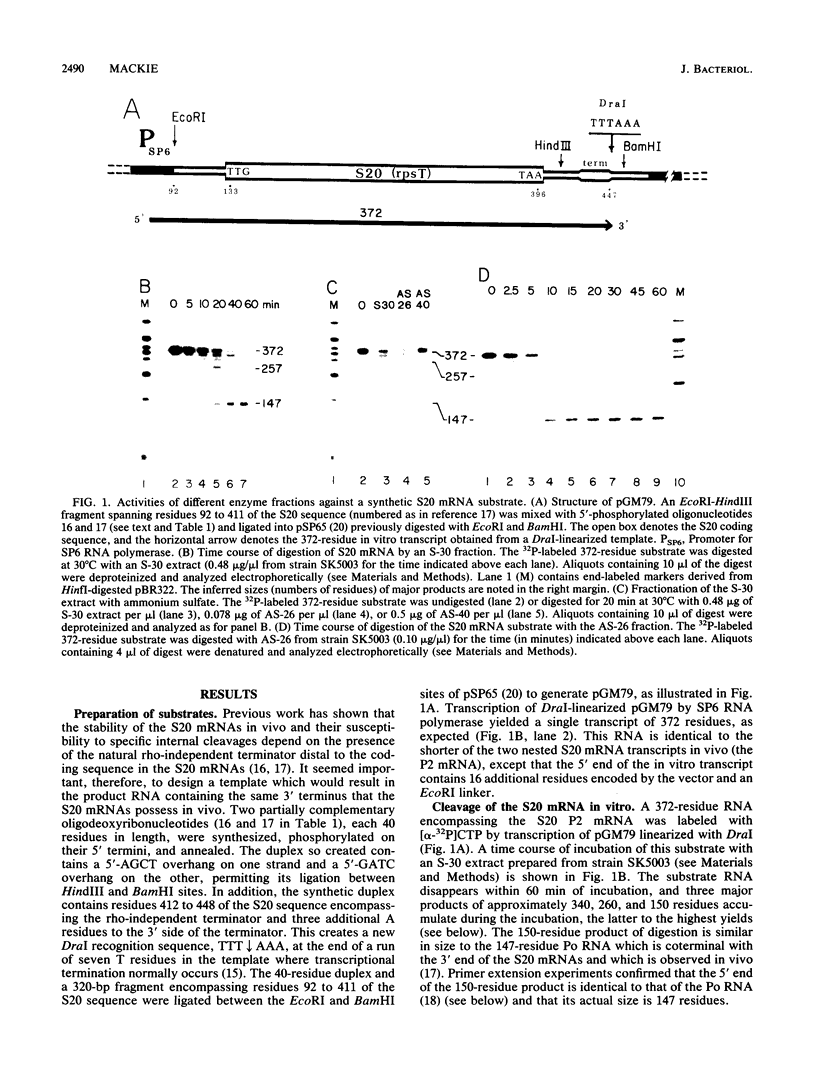
Endonucleolytic cleavage is believed to initiate the degradation of most bacterial mRNAs, but with several exceptions, the enzymes responsible have yet to be identified. Crude (S-30) or partially fractionated extracts of Escherichia coli strains with reduced exonuclease activities catalyze the cleavage of a 372-residue RNA substrate containing the sequences coding for ribosomal protein S20 to yield a number of discrete products. The major product of 147 residues is obtained in 60 to 70% yield, is coterminal with the 3' end of the substrate, and is identical to an mRNA fragment previously characterized in vivo (G. A. Mackie, J. Bacteriol. 171:4112-4120, 1989). A number of other products of 150 to 340 residues are also formed, and the cleavage sites, typically N decreases AU sequences, have been identified in the S20 mRNA substrate by Northern (RNA) blotting and primer extension. All cleavages required a native rather than a denatured RNA substrate. The rate of cutting of the S20 mRNA substrate at the site yielding the prominent 147-residue product appears to be independent of cleavages at other sites. In addition, the activity of the putative endonuclease(s) depends strongly, both in vivo and in vitro, on the product of the ams gene, which is known to influence mRNA lifetimes in vivo. Taken together, the data show that the fractionated extract described here reproduces steps in the degradation of some mRNAs which occur in living cells.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Apirion D. Degradation of RNA in Escherichia coli. A hypothesis. Mol Gen Genet. 1973 May 28;122(4):313–322. doi: 10.1007/BF00269431. [DOI] [PubMed] [Google Scholar]
- Arraiano C. M., Yancey S. D., Kushner S. R. Stabilization of discrete mRNA breakdown products in ams pnp rnb multiple mutants of Escherichia coli K-12. J Bacteriol. 1988 Oct;170(10):4625–4633. doi: 10.1128/jb.170.10.4625-4633.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barry G., Squires C., Squires C. L. Attenuation and processing of RNA from the rplJL--rpoBC transcription unit of Escherichia coli. Proc Natl Acad Sci U S A. 1980 Jun;77(6):3331–3335. doi: 10.1073/pnas.77.6.3331. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belasco J. G., Higgins C. F. Mechanisms of mRNA decay in bacteria: a perspective. Gene. 1988 Dec 10;72(1-2):15–23. doi: 10.1016/0378-1119(88)90123-0. [DOI] [PubMed] [Google Scholar]
- Cannistraro V. J., Kennell D. Purification and characterization of ribonuclease M and mRNA degradation in Escherichia coli. Eur J Biochem. 1989 May 1;181(2):363–370. doi: 10.1111/j.1432-1033.1989.tb14733.x. [DOI] [PubMed] [Google Scholar]
- Cannistraro V. J., Subbarao M. N., Kennell D. Specific endonucleolytic cleavage sites for decay of Escherichia coli mRNA. J Mol Biol. 1986 Nov 20;192(2):257–274. doi: 10.1016/0022-2836(86)90363-3. [DOI] [PubMed] [Google Scholar]
- Claverie-Martin F., Diaz-Torres M. R., Yancey S. D., Kushner S. R. Cloning of the altered mRNA stability (ams) gene of Escherichia coli K-12. J Bacteriol. 1989 Oct;171(10):5479–5486. doi: 10.1128/jb.171.10.5479-5486.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deutscher M. P. E. coli RNases: making sense of alphabet soup. Cell. 1985 Apr;40(4):731–732. doi: 10.1016/0092-8674(85)90330-7. [DOI] [PubMed] [Google Scholar]
- Donovan W. P., Kushner S. R. Polynucleotide phosphorylase and ribonuclease II are required for cell viability and mRNA turnover in Escherichia coli K-12. Proc Natl Acad Sci U S A. 1986 Jan;83(1):120–124. doi: 10.1073/pnas.83.1.120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dunn J. J., Studier F. W. T7 early RNAs are generated by site-specific cleavages. Proc Natl Acad Sci U S A. 1973 May;70(5):1559–1563. doi: 10.1073/pnas.70.5.1559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- King T. C., Sirdeskmukh R., Schlessinger D. Nucleolytic processing of ribonucleic acid transcripts in procaryotes. Microbiol Rev. 1986 Dec;50(4):428–451. doi: 10.1128/mr.50.4.428-451.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lundberg U., von Gabain A., Melefors O. Cleavages in the 5' region of the ompA and bla mRNA control stability: studies with an E. coli mutant altering mRNA stability and a novel endoribonuclease. EMBO J. 1990 Sep;9(9):2731–2741. doi: 10.1002/j.1460-2075.1990.tb07460.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mackie G. A. Nucleotide sequence of the gene for ribosomal protein S20 and its flanking regions. J Biol Chem. 1981 Aug 10;256(15):8177–8182. [PubMed] [Google Scholar]
- Mackie G. A. Posttranscriptional regulation of ribosomal protein S20 and stability of the S20 mRNA species. J Bacteriol. 1987 Jun;169(6):2697–2701. doi: 10.1128/jb.169.6.2697-2701.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mackie G. A. Stabilization of the 3' one-third of Escherichia coli ribosomal protein S20 mRNA in mutants lacking polynucleotide phosphorylase. J Bacteriol. 1989 Aug;171(8):4112–4120. doi: 10.1128/jb.171.8.4112-4120.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mackie G. A. Structure of the DNA distal to the gene for ribosomal protein S20 in Escherichia coli K12: presence of a strong terminator and an IS1 element. Nucleic Acids Res. 1986 Sep 11;14(17):6965–6981. doi: 10.1093/nar/14.17.6965. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Melton D. A., Krieg P. A., Rebagliati M. R., Maniatis T., Zinn K., Green M. R. Efficient in vitro synthesis of biologically active RNA and RNA hybridization probes from plasmids containing a bacteriophage SP6 promoter. Nucleic Acids Res. 1984 Sep 25;12(18):7035–7056. doi: 10.1093/nar/12.18.7035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Misra T. K., Apirion D. Characterization of an endoribonuclease, RNase N, from Escherichia coli. J Biol Chem. 1978 Aug 25;253(16):5594–5599. [PubMed] [Google Scholar]
- Misra T. K., Apirion D. RNase E, an RNA processing enzyme from Escherichia coli. J Biol Chem. 1979 Nov 10;254(21):11154–11159. [PubMed] [Google Scholar]
- Mudd E. A., Carpousis A. J., Krisch H. M. Escherichia coli RNase E has a role in the decay of bacteriophage T4 mRNA. Genes Dev. 1990 May;4(5):873–881. doi: 10.1101/gad.4.5.873. [DOI] [PubMed] [Google Scholar]
- Mudd E. A., Krisch H. M., Higgins C. F. RNase E, an endoribonuclease, has a general role in the chemical decay of Escherichia coli mRNA: evidence that rne and ams are the same genetic locus. Mol Microbiol. 1990 Dec;4(12):2127–2135. doi: 10.1111/j.1365-2958.1990.tb00574.x. [DOI] [PubMed] [Google Scholar]
- Mudd E. A., Prentki P., Belin D., Krisch H. M. Processing of unstable bacteriophage T4 gene 32 mRNAs into a stable species requires Escherichia coli ribonuclease E. EMBO J. 1988 Nov;7(11):3601–3607. doi: 10.1002/j.1460-2075.1988.tb03238.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nilsson G., Lundberg U., von Gabain A. In vivo and in vitro identity of site specific cleavages in the 5' non-coding region of ompA and bla mRNA in Escherichia coli. EMBO J. 1988 Jul;7(7):2269–2275. doi: 10.1002/j.1460-2075.1988.tb03067.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ono M., Kuwano M. A conditional lethal mutation in an Escherichia coli strain with a longer chemical lifetime of messenger RNA. J Mol Biol. 1979 Apr 15;129(3):343–357. doi: 10.1016/0022-2836(79)90500-x. [DOI] [PubMed] [Google Scholar]
- Parsons G. D., Donly B. C., Mackie G. A. Mutations in the leader sequence and initiation codon of the gene for ribosomal protein S20 (rpsT) affect both translational efficiency and autoregulation. J Bacteriol. 1988 Jun;170(6):2485–2492. doi: 10.1128/jb.170.6.2485-2492.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Portier C., Dondon L., Grunberg-Manago M., Régnier P. The first step in the functional inactivation of the Escherichia coli polynucleotide phosphorylase messenger is a ribonuclease III processing at the 5' end. EMBO J. 1987 Jul;6(7):2165–2170. doi: 10.1002/j.1460-2075.1987.tb02484.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spahr P. F., Gesteland R. F. Specific cleavage of bacteriophage R17 RNA by an endonuclease isolated from E. coli MRE-600. Proc Natl Acad Sci U S A. 1968 Mar;59(3):876–883. doi: 10.1073/pnas.59.3.876. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhu L. Q., Gangopadhyay T., Padmanabha K. P., Deutscher M. P. Escherichia coli rna gene encoding RNase I: cloning, overexpression, subcellular distribution of the enzyme, and use of an rna deletion to identify additional RNases. J Bacteriol. 1990 Jun;172(6):3146–3151. doi: 10.1128/jb.172.6.3146-3151.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zoller M. J., Smith M. Oligonucleotide-directed mutagenesis: a simple method using two oligonucleotide primers and a single-stranded DNA template. DNA. 1984 Dec;3(6):479–488. doi: 10.1089/dna.1.1984.3.479. [DOI] [PubMed] [Google Scholar]