Abstract
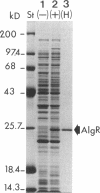
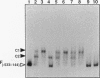
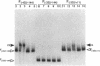
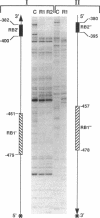
Strong transcriptional activation of algD, a key event in the overproduction of alginate and establishment of mucoidy in Pseudomonas aeruginosa, depends on the functional algR gene. The predicted gene product of algR shows homologies to response regulators from bacterial signal transduction systems. The algR gene was overexpressed in Escherichia coli, its product (AlgR) was purified by utilizing its apparent affinity for heparin, and its sequence was verified by partial amino acid sequence analysis. AlgR was found to interact directly with the algD promoter. Deletion mapping analysis, in conjunction with mobility shift DNA-binding assays, indicated the presence of three regions within the algD promoter capable of specifically binding AlgR. A relatively weak interaction was observed with the algD promoter fragment containing the region immediately upstream of the algD mRNA start site (-144 to +11). However, when fragments spanning regions located very far upstream from the algD mRNA initiation site (-533 and -332) were used, strong specific binding was observed. These regions were separated by a DNA segment not binding AlgR and spanning positions -332 to -144. DNase I footprinting analysis further established the presence of discrete AlgR binding sites overlapping with FUS, the far-upstream sites required for full induction of algD transcription and its environmental modulation. There were two distinct binding sites: RB1, spanning nucleotides -479 to -457, and RB2, spanning nucleotides -400 to -380. Both of these sequences shared a highly conserved core region, ACCGTTCGTC. These results established a direct interaction of AlgR with the algD promoter and revealed an arrangement of binding sites highly unusual for response regulators of the AlgR type.
Full text
PDF

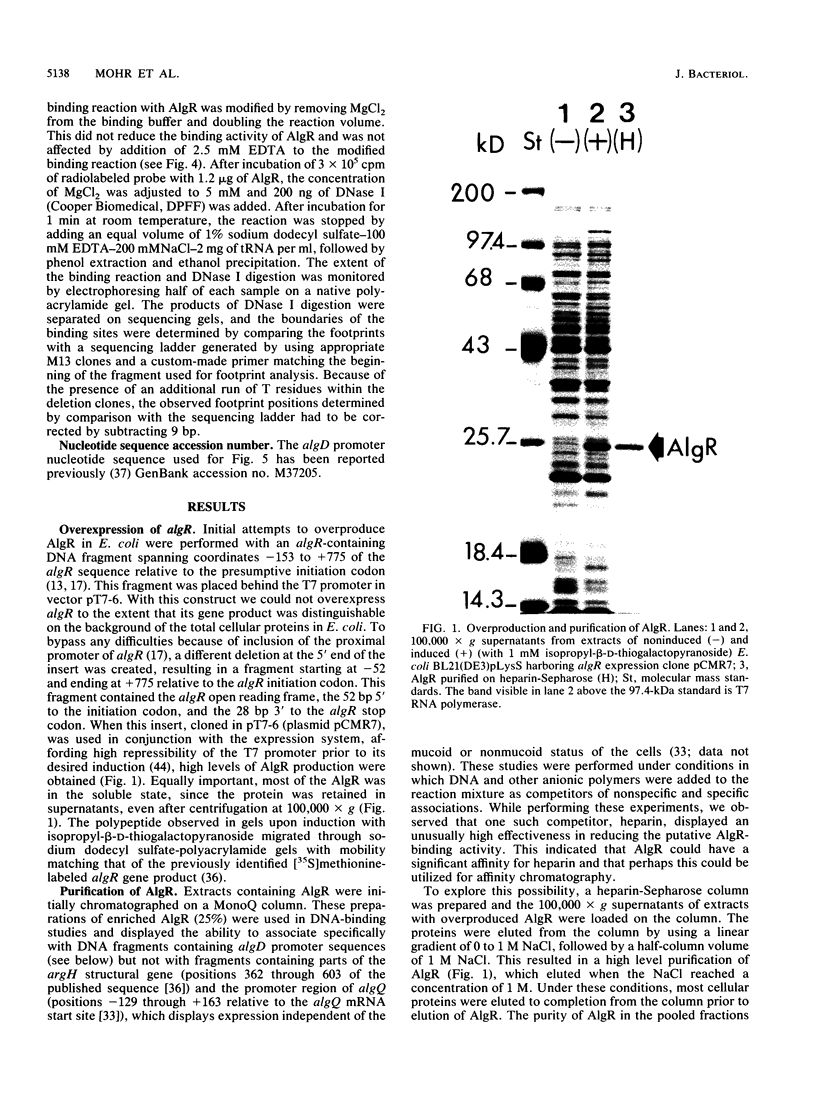
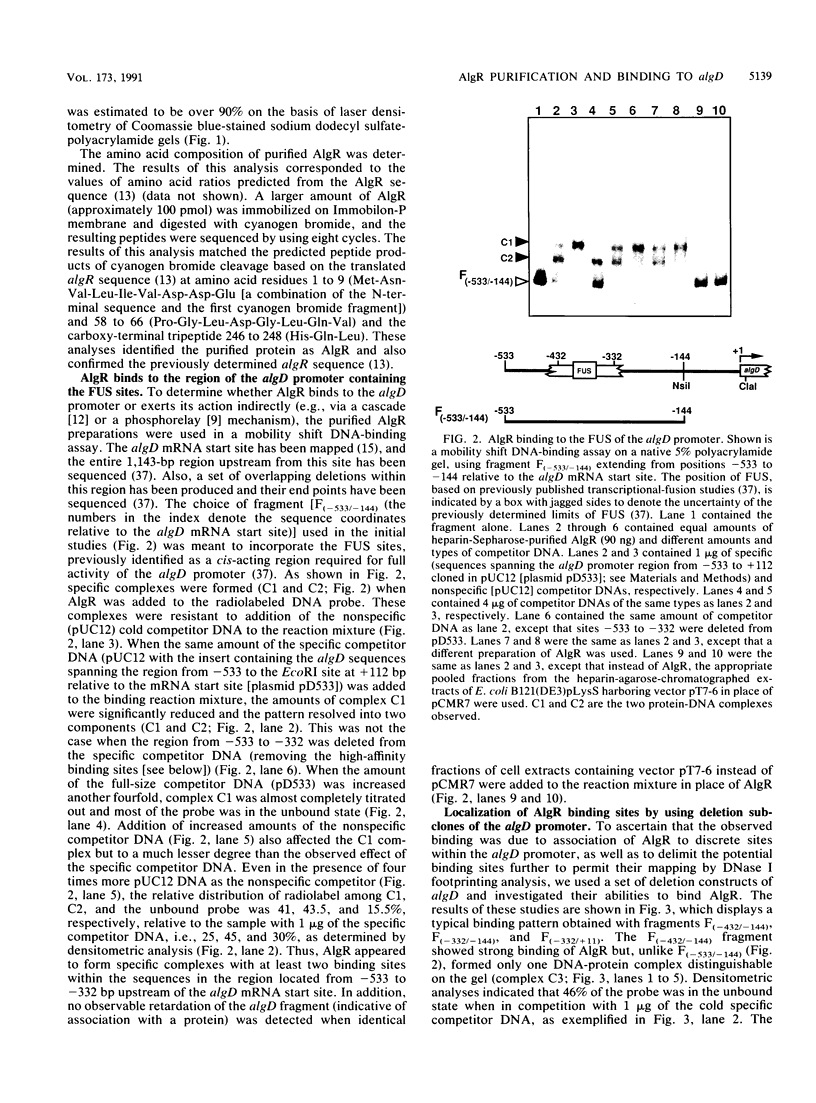
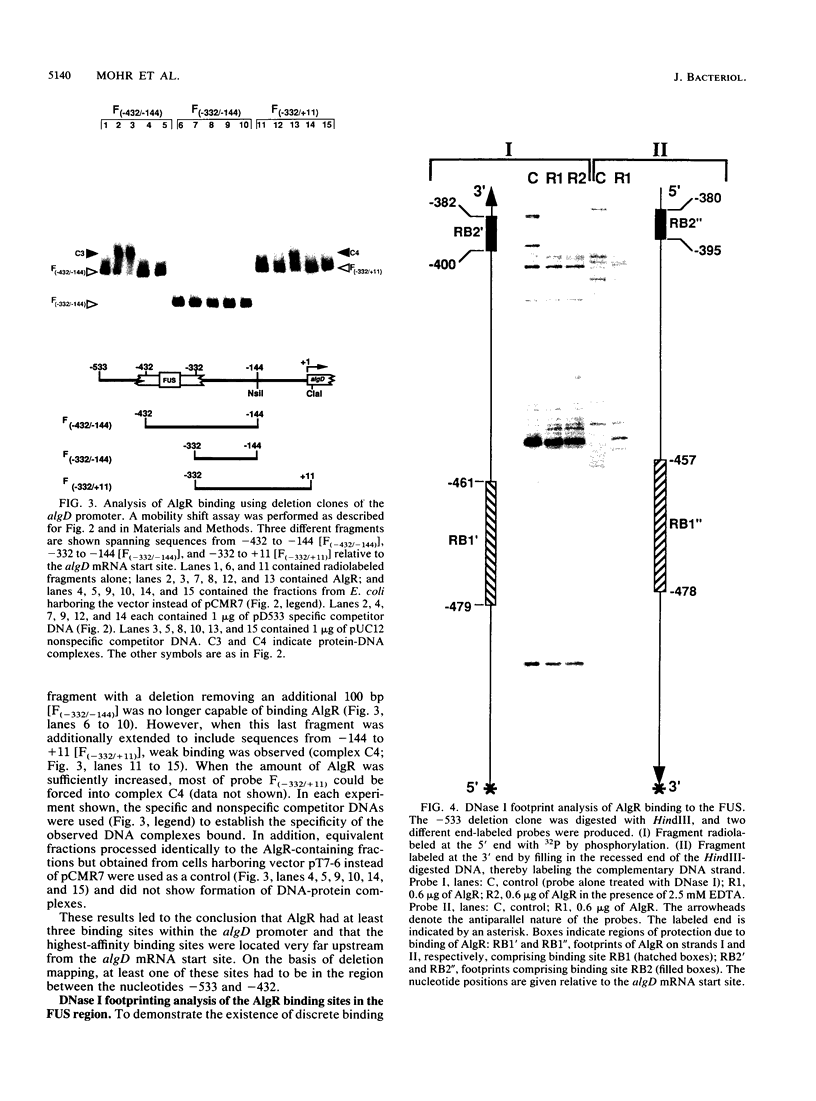



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Allan J., Mitchell T., Harborne N., Bohm L., Crane-Robinson C. Roles of H1 domains in determining higher order chromatin structure and H1 location. J Mol Biol. 1986 Feb 20;187(4):591–601. doi: 10.1016/0022-2836(86)90337-2. [DOI] [PubMed] [Google Scholar]
- Amemura M., Makino K., Shinagawa H., Nakata A. Cross talk to the phosphate regulon of Escherichia coli by PhoM protein: PhoM is a histidine protein kinase and catalyzes phosphorylation of PhoB and PhoM-open reading frame 2. J Bacteriol. 1990 Nov;172(11):6300–6307. doi: 10.1128/jb.172.11.6300-6307.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Anastassiou E. D., Mintzas A. C., Kounavis C., Dimitracopoulos G. Alginate production by clinical nonmucoid Pseudomonas aeruginosa strains. J Clin Microbiol. 1987 Apr;25(4):656–659. doi: 10.1128/jcm.25.4.656-659.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bayer A. S., Eftekhar F., Tu J., Nast C. C., Speert D. P. Oxygen-dependent up-regulation of mucoid exopolysaccharide (alginate) production in Pseudomonas aeruginosa. Infect Immun. 1990 May;58(5):1344–1349. doi: 10.1128/iai.58.5.1344-1349.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berry A., DeVault J. D., Chakrabarty A. M. High osmolarity is a signal for enhanced algD transcription in mucoid and nonmucoid Pseudomonas aeruginosa strains. J Bacteriol. 1989 May;171(5):2312–2317. doi: 10.1128/jb.171.5.2312-2317.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bradford M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976 May 7;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Burbulys D., Trach K. A., Hoch J. A. Initiation of sporulation in B. subtilis is controlled by a multicomponent phosphorelay. Cell. 1991 Feb 8;64(3):545–552. doi: 10.1016/0092-8674(91)90238-t. [DOI] [PubMed] [Google Scholar]
- Dale R. M., McClure B. A., Houchins J. P. A rapid single-stranded cloning strategy for producing a sequential series of overlapping clones for use in DNA sequencing: application to sequencing the corn mitochondrial 18 S rDNA. Plasmid. 1985 Jan;13(1):31–40. doi: 10.1016/0147-619x(85)90053-8. [DOI] [PubMed] [Google Scholar]
- Darzins A., Wang S. K., Vanags R. I., Chakrabarty A. M. Clustering of mutations affecting alginic acid biosynthesis in mucoid Pseudomonas aeruginosa. J Bacteriol. 1985 Nov;164(2):516–524. doi: 10.1128/jb.164.2.516-524.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- David M., Daveran M. L., Batut J., Dedieu A., Domergue O., Ghai J., Hertig C., Boistard P., Kahn D. Cascade regulation of nif gene expression in Rhizobium meliloti. Cell. 1988 Aug 26;54(5):671–683. doi: 10.1016/s0092-8674(88)80012-6. [DOI] [PubMed] [Google Scholar]
- Deretic V., Dikshit R., Konyecsni W. M., Chakrabarty A. M., Misra T. K. The algR gene, which regulates mucoidy in Pseudomonas aeruginosa, belongs to a class of environmentally responsive genes. J Bacteriol. 1989 Mar;171(3):1278–1283. doi: 10.1128/jb.171.3.1278-1283.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deretic V., Gill J. F., Chakrabarty A. M. Gene algD coding for GDPmannose dehydrogenase is transcriptionally activated in mucoid Pseudomonas aeruginosa. J Bacteriol. 1987 Jan;169(1):351–358. doi: 10.1128/jb.169.1.351-358.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deretic V., Gill J. F., Chakrabarty A. M. Pseudomonas aeruginosa infection in cystic fibrosis: nucleotide sequence and transcriptional regulation of the algD gene. Nucleic Acids Res. 1987 Jun 11;15(11):4567–4581. doi: 10.1093/nar/15.11.4567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deretic V., Govan J. R., Konyecsni W. M., Martin D. W. Mucoid Pseudomonas aeruginosa in cystic fibrosis: mutations in the muc loci affect transcription of the algR and algD genes in response to environmental stimuli. Mol Microbiol. 1990 Feb;4(2):189–196. doi: 10.1111/j.1365-2958.1990.tb00586.x. [DOI] [PubMed] [Google Scholar]
- Deretic V., Konyecsni W. M. A procaryotic regulatory factor with a histone H1-like carboxy-terminal domain: clonal variation of repeats within algP, a gene involved in regulation of mucoidy in Pseudomonas aeruginosa. J Bacteriol. 1990 Oct;172(10):5544–5554. doi: 10.1128/jb.172.10.5544-5554.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deretic V., Konyecsni W. M. Control of mucoidy in Pseudomonas aeruginosa: transcriptional regulation of algR and identification of the second regulatory gene, algQ. J Bacteriol. 1989 Jul;171(7):3680–3688. doi: 10.1128/jb.171.7.3680-3688.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deretic V., Mohr C. D., Martin D. W. Mucoid Pseudomonas aeruginosa in cystic fibrosis: signal transduction and histone-like elements in the regulation of bacterial virulence. Mol Microbiol. 1991 Jul;5(7):1577–1583. doi: 10.1111/j.1365-2958.1991.tb01903.x. [DOI] [PubMed] [Google Scholar]
- Doggett R. G., Harrison G. M., Carter R. E. Mucoid Pseudomonas aeruginosa in patients with chronic illnesses. Lancet. 1971 Jan 30;1(7692):236–237. doi: 10.1016/s0140-6736(71)90973-1. [DOI] [PubMed] [Google Scholar]
- Flynn J. L., Ohman D. E. Use of a gene replacement cosmid vector for cloning alginate conversion genes from mucoid and nonmucoid Pseudomonas aeruginosa strains: algS controls expression of algT. J Bacteriol. 1988 Jul;170(7):3228–3236. doi: 10.1128/jb.170.7.3228-3236.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fyfe J. A., Govan J. R. Alginate synthesis in mucoid Pseudomonas aeruginosa: a chromosomal locus involved in control. J Gen Microbiol. 1980 Aug;119(2):443–450. doi: 10.1099/00221287-119-2-443. [DOI] [PubMed] [Google Scholar]
- Hawkins A. R., Wootton J. C. A single DNA-binding protein from Pseudomonas aeruginosa homologous to proteins NS1 and NS2 (HU proteins) of Escherichia coli and other bacteria. FEBS Lett. 1981 Aug 3;130(2):275–278. doi: 10.1016/0014-5793(81)81138-6. [DOI] [PubMed] [Google Scholar]
- Hoover T. R., Santero E., Porter S., Kustu S. The integration host factor stimulates interaction of RNA polymerase with NIFA, the transcriptional activator for nitrogen fixation operons. Cell. 1990 Oct 5;63(1):11–22. doi: 10.1016/0092-8674(90)90284-l. [DOI] [PubMed] [Google Scholar]
- Huang Y., Morel P., Powell B., Kado C. I. VirA, a coregulator of Ti-specified virulence genes, is phosphorylated in vitro. J Bacteriol. 1990 Feb;172(2):1142–1144. doi: 10.1128/jb.172.2.1142-1144.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Høiby N., Döring G., Schiøtz P. O. Pathogenic mechanisms of chronic Pseudomonas aeruginosa infections in cystic fibrosis patients. Antibiot Chemother (1971) 1987;39:60–76. [PubMed] [Google Scholar]
- Igo M. M., Ninfa A. J., Stock J. B., Silhavy T. J. Phosphorylation and dephosphorylation of a bacterial transcriptional activator by a transmembrane receptor. Genes Dev. 1989 Nov;3(11):1725–1734. doi: 10.1101/gad.3.11.1725. [DOI] [PubMed] [Google Scholar]
- Jin S., Roitsch T., Ankenbauer R. G., Gordon M. P., Nester E. W. The VirA protein of Agrobacterium tumefaciens is autophosphorylated and is essential for vir gene regulation. J Bacteriol. 1990 Feb;172(2):525–530. doi: 10.1128/jb.172.2.525-530.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kato J., Misra T. K., Chakrabarty A. M. AlgR3, a protein resembling eukaryotic histone H1, regulates alginate synthesis in Pseudomonas aeruginosa. Proc Natl Acad Sci U S A. 1990 Apr;87(8):2887–2891. doi: 10.1073/pnas.87.8.2887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Konyecsni W. M., Deretic V. Broad-host-range plasmid and M13 bacteriophage-derived vectors for promoter analysis in Escherichia coli and Pseudomonas aeruginosa. Gene. 1988 Dec 30;74(2):375–386. doi: 10.1016/0378-1119(88)90171-0. [DOI] [PubMed] [Google Scholar]
- Konyecsni W. M., Deretic V. DNA sequence and expression analysis of algP and algQ, components of the multigene system transcriptionally regulating mucoidy in Pseudomonas aeruginosa: algP contains multiple direct repeats. J Bacteriol. 1990 May;172(5):2511–2520. doi: 10.1128/jb.172.5.2511-2520.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- MacGeorge J., Korolik V., Morgan A. F., Asche V., Holloway B. W. Transfer of a chromosomal locus responsible for mucoid colony morphology in Pseudomonas aeruginosa isolated from cystic fibrosis patients to P. aeruginosa PAO. J Med Microbiol. 1986 Jun;21(4):331–336. doi: 10.1099/00222615-21-4-331. [DOI] [PubMed] [Google Scholar]
- Matsudaira P. Sequence from picomole quantities of proteins electroblotted onto polyvinylidene difluoride membranes. J Biol Chem. 1987 Jul 25;262(21):10035–10038. [PubMed] [Google Scholar]
- Mohr C. D., Deretic V. Gene-scrambling mutagenesis: generation and analysis of insertional mutations in the alginate regulatory region of Pseudomonas aeruginosa. J Bacteriol. 1990 Nov;172(11):6252–6260. doi: 10.1128/jb.172.11.6252-6260.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mohr C. D., Martin D. W., Konyecsni W. M., Govan J. R., Lory S., Deretic V. Role of the far-upstream sites of the algD promoter and the algR and rpoN genes in environmental modulation of mucoidy in Pseudomonas aeruginosa. J Bacteriol. 1990 Nov;172(11):6576–6580. doi: 10.1128/jb.172.11.6576-6580.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ninfa A. J., Reitzer L. J., Magasanik B. Initiation of transcription at the bacterial glnAp2 promoter by purified E. coli components is facilitated by enhancers. Cell. 1987 Sep 25;50(7):1039–1046. doi: 10.1016/0092-8674(87)90170-x. [DOI] [PubMed] [Google Scholar]
- Norioka S., Ramakrishnan G., Ikenaka K., Inouye M. Interaction of a transcriptional activator, OmpR, with reciprocally osmoregulated genes, ompF and ompC, of Escherichia coli. J Biol Chem. 1986 Dec 25;261(36):17113–17119. [PubMed] [Google Scholar]
- Pier G. B., Matthews W. J., Jr, Eardley D. D. Immunochemical characterization of the mucoid exopolysaccharide of Pseudomonas aeruginosa. J Infect Dis. 1983 Mar;147(3):494–503. doi: 10.1093/infdis/147.3.494. [DOI] [PubMed] [Google Scholar]
- Reitzer L. J., Magasanik B. Transcription of glnA in E. coli is stimulated by activator bound to sites far from the promoter. Cell. 1986 Jun 20;45(6):785–792. doi: 10.1016/0092-8674(86)90553-2. [DOI] [PubMed] [Google Scholar]
- Ronson C. W., Nixon B. T., Ausubel F. M. Conserved domains in bacterial regulatory proteins that respond to environmental stimuli. Cell. 1987 Jun 5;49(5):579–581. doi: 10.1016/0092-8674(87)90530-7. [DOI] [PubMed] [Google Scholar]
- Stock J. B., Ninfa A. J., Stock A. M. Protein phosphorylation and regulation of adaptive responses in bacteria. Microbiol Rev. 1989 Dec;53(4):450–490. doi: 10.1128/mr.53.4.450-490.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Studier F. W., Moffatt B. A. Use of bacteriophage T7 RNA polymerase to direct selective high-level expression of cloned genes. J Mol Biol. 1986 May 5;189(1):113–130. doi: 10.1016/0022-2836(86)90385-2. [DOI] [PubMed] [Google Scholar]
- Su W., Porter S., Kustu S., Echols H. DNA-looping and enhancer activity: association between DNA-bound NtrC activator and RNA polymerase at the bacterial glnA promoter. Proc Natl Acad Sci U S A. 1990 Jul;87(14):5504–5508. doi: 10.1073/pnas.87.14.5504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Terry J. M., Piña S. E., Mattingly S. J. Environmental conditions which influence mucoid conversion Pseudomonas aeruginosa PAO1. Infect Immun. 1991 Feb;59(2):471–477. doi: 10.1128/iai.59.2.471-477.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vieira J., Messing J. The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene. 1982 Oct;19(3):259–268. doi: 10.1016/0378-1119(82)90015-4. [DOI] [PubMed] [Google Scholar]
- Wozniak D. J., Ohman D. E. Pseudomonas aeruginosa AlgB, a two-component response regulator of the NtrC family, is required for algD transcription. J Bacteriol. 1991 Feb;173(4):1406–1413. doi: 10.1128/jb.173.4.1406-1413.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]