Abstract
Fluorescent-dye-conjugated oligonucleotides were used to classify 14 Fibrobacter strains by fluorescence microscopy. On the basis of partial 16S rRNA sequences of six Fibrobacter strains, four hybridization probes were designed to discriminate between the species Fibrobacter succinogenes and Fibrobacter intestinalis and to identify F. succinogenes subsp. succinogenes. After in situ hybridization to whole cells of the six sequenced strains, epifluorescence microscopy confirmed probe specificity. The four probes were then used to make presumptive species and subspecies assignments of eight additional Fibrobacter strains not previously characterized by comparative sequencing. These assignments were confirmed by comparative sequencing of the 16S rRNA target regions from the additional organisms. Single-mismatch discrimination between certain probe and nontarget sequences was demonstrated, and fluorescent intensity was shown to be enhanced by hybridization to multiple probes of the same specificity. The direct detection of F. intestinalis in mouse cecum samples demonstrated the application of this technique to the characterization of complex natural samples.
Full text
PDF






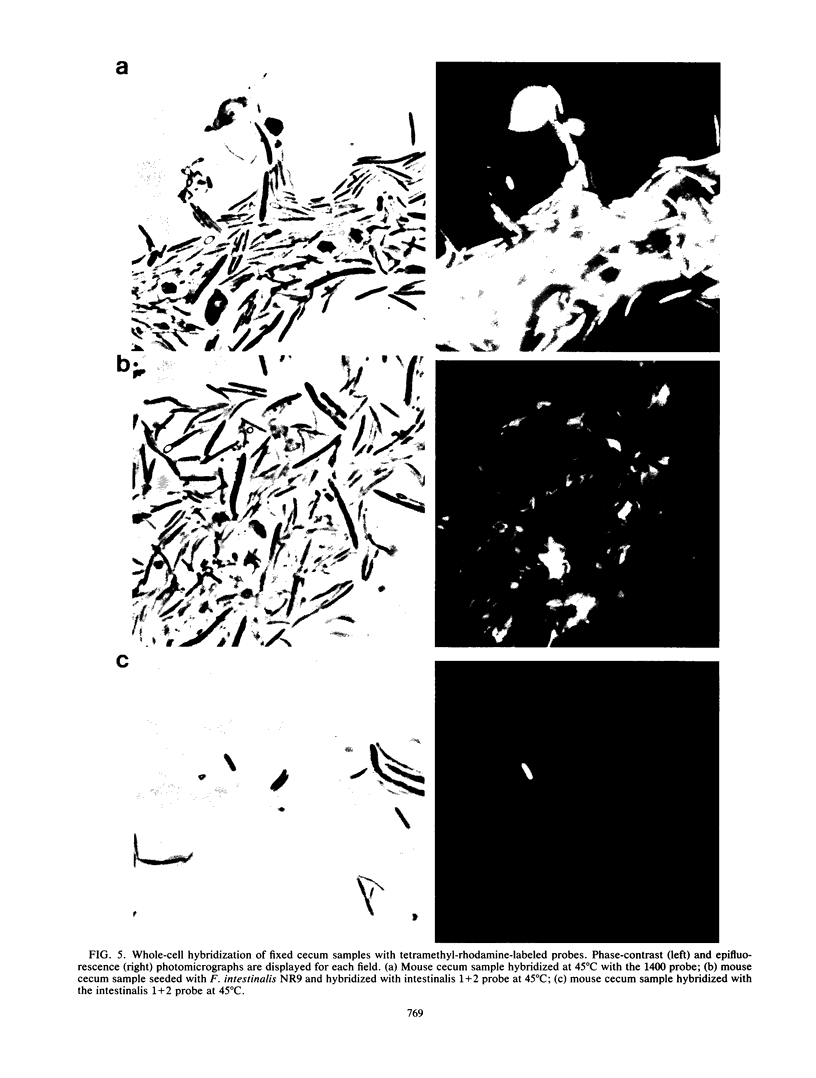

Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Brosius J., Palmer M. L., Kennedy P. J., Noller H. F. Complete nucleotide sequence of a 16S ribosomal RNA gene from Escherichia coli. Proc Natl Acad Sci U S A. 1978 Oct;75(10):4801–4805. doi: 10.1073/pnas.75.10.4801. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carbon P., Ebel J. P., Ehresmann C. The sequence of the ribosomal 16S RNA from Proteus vulgaris. Sequence comparison with E. coli 16S RNA and its use in secondary model building. Nucleic Acids Res. 1981 May 25;9(10):2325–2333. doi: 10.1093/nar/9.10.2325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DeLong E. F., Wickham G. S., Pace N. R. Phylogenetic stains: ribosomal RNA-based probes for the identification of single cells. Science. 1989 Mar 10;243(4896):1360–1363. doi: 10.1126/science.2466341. [DOI] [PubMed] [Google Scholar]
- Giovannoni S. J., DeLong E. F., Olsen G. J., Pace N. R. Phylogenetic group-specific oligodeoxynucleotide probes for identification of single microbial cells. J Bacteriol. 1988 Feb;170(2):720–726. doi: 10.1128/jb.170.2.720-726.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Green C. J., Stewart G. C., Hollis M. A., Vold B. S., Bott K. F. Nucleotide sequence of the Bacillus subtilis ribosomal RNA operon, rrnB. Gene. 1985;37(1-3):261–266. doi: 10.1016/0378-1119(85)90281-1. [DOI] [PubMed] [Google Scholar]
- Jablonski E., Moomaw E. W., Tullis R. H., Ruth J. L. Preparation of oligodeoxynucleotide-alkaline phosphatase conjugates and their use as hybridization probes. Nucleic Acids Res. 1986 Aug 11;14(15):6115–6128. doi: 10.1093/nar/14.15.6115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lane D. J., Pace B., Olsen G. J., Stahl D. A., Sogin M. L., Pace N. R. Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc Natl Acad Sci U S A. 1985 Oct;82(20):6955–6959. doi: 10.1073/pnas.82.20.6955. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Macy J. M., Farrand J. R., Montgomery L. Cellulolytic and non-cellulolytic bacteria in rat gastrointestinal tracts. Appl Environ Microbiol. 1982 Dec;44(6):1428–1434. doi: 10.1128/aem.44.6.1428-1434.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Montgomery L., Macy J. M. Characterization of rat cecum cellulolytic bacteria. Appl Environ Microbiol. 1982 Dec;44(6):1435–1443. doi: 10.1128/aem.44.6.1435-1443.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olsen G. J., Lane D. J., Giovannoni S. J., Pace N. R., Stahl D. A. Microbial ecology and evolution: a ribosomal RNA approach. Annu Rev Microbiol. 1986;40:337–365. doi: 10.1146/annurev.mi.40.100186.002005. [DOI] [PubMed] [Google Scholar]
- Pace B., Matthews E. A., Johnson K. D., Cantor C. R., Pace N. R. Conserved 5S rRNA complement to tRNA is not required for protein synthesis. Proc Natl Acad Sci U S A. 1982 Jan;79(1):36–40. doi: 10.1073/pnas.79.1.36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schildkraut C. Dependence of the melting temperature of DNA on salt concentration. Biopolymers. 1965;3(2):195–208. doi: 10.1002/bip.360030207. [DOI] [PubMed] [Google Scholar]
- Stahl D. A., Flesher B., Mansfield H. R., Montgomery L. Use of phylogenetically based hybridization probes for studies of ruminal microbial ecology. Appl Environ Microbiol. 1988 May;54(5):1079–1084. doi: 10.1128/aem.54.5.1079-1084.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stahl D. A., Urbance J. W. The division between fast- and slow-growing species corresponds to natural relationships among the mycobacteria. J Bacteriol. 1990 Jan;172(1):116–124. doi: 10.1128/jb.172.1.116-124.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stewart C. S., Paniagua C., Dinsdale D., Cheng K. J., Garrow S. H. Selective isolation and characteristics of Bacteriodes succinogenes from the rumen of a cow. Appl Environ Microbiol. 1981 Feb;41(2):504–510. doi: 10.1128/aem.41.2.504-510.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Varel V. H., Fryda S. J., Robinson I. M. Cellulolytic bacteria from pig large intestine. Appl Environ Microbiol. 1984 Jan;47(1):219–221. doi: 10.1128/aem.47.1.219-221.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weisburg W. G., Oyaizu Y., Oyaizu H., Woese C. R. Natural relationship between bacteroides and flavobacteria. J Bacteriol. 1985 Oct;164(1):230–236. doi: 10.1128/jb.164.1.230-236.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weller R., Ward D. M. Selective Recovery of 16S rRNA Sequences from Natural Microbial Communities in the Form of cDNA. Appl Environ Microbiol. 1989 Jul;55(7):1818–1822. doi: 10.1128/aem.55.7.1818-1822.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woese C. R. Bacterial evolution. Microbiol Rev. 1987 Jun;51(2):221–271. doi: 10.1128/mr.51.2.221-271.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woese C. R., Stackebrandt E., Macke T. J., Fox G. E. A phylogenetic definition of the major eubacterial taxa. Syst Appl Microbiol. 1985;6:143–151. doi: 10.1016/s0723-2020(85)80047-3. [DOI] [PubMed] [Google Scholar]
- Yang D., Oyaizu Y., Oyaizu H., Olsen G. J., Woese C. R. Mitochondrial origins. Proc Natl Acad Sci U S A. 1985 Jul;82(13):4443–4447. doi: 10.1073/pnas.82.13.4443. [DOI] [PMC free article] [PubMed] [Google Scholar]






