Abstract
Induction of nitric-oxide synthase-2 (iNOS) by cytokines and bacterial products is associated with protein binding at the proximal promoter and in an upstream enhancer region of the Nos2 gene. To clarify how ethanol suppresses rat iNOS activity, we constructed several deletion mutants of the Nos2 promoter fused to the luciferase gene and transfected the constructs into C6 glial cells. Acute ethanol exposure of stably transfected cells for 24 h inhibits induced activity of Nos2 promoter constructs containing deletions in the 5′ flanking region, including a 94 bp promoter that lacks any known NF-κB site but which carries a C/EBPβ and overlapping γ-IRE, GAS and Oct motifs. Ethanol failed to inhibit the endogenous activity of a smaller, 78 bp promoter that lacks the C/EBPβ and overlapping, γ-IRE and GAS motifs and showed no inducible activity. As another approach, in vivo DNA footprinting was used and identified protein protections at five regions of the proximal Nos2 promoter in induced cells. Exposure to acute ethanol diminished protein occupation in the five promoter regions including the γ-IRE/NF-κB and the overlapping γIRE/GAS/Oct sites. Site-directed mutagenesis in the octamer domain of the γIRE/GAS/Oct motifs was studied in a 1002 bp promoter to examine its role in ethanol inhibition of cytokine and lipopolysaccharide induced activity. The data indicate ethanol failed to inhibit promoter activity when the Oct motif is missing. Electrophoretic mobility shift assays performed using a 22-mer probe containing the overlapping γ-IRE/GAS/Oct sites showed three complexes with one of the complexes being competed by an octamer-1 antibody. These observations demonstrate the role of protein-DNA binding at the core promoter, and the likely involvement of the octamer motif, in ethanol modulation of cytokine and lipopolysaccharide induced iNOS expression.
Keywords: ethanol, Nos2, octamer, astrocyte, promoter
1. Introduction
Transactivation of the Nos2 gene, expression of inducible nitric-oxide synthase (iNOS) and the resulting production of nitric oxide (NO) is an integral inflammatory response in the CNS (Murphy and Grzybicki, 1996). Nitric oxide generated from iNOS has been implicated in many CNS pathologies including brain infections, and neurological disorders such as acquired immunodeficiency syndrome and multiple sclerosis (Adamson, et al., 1996, Bagasra, et al., 1995). In these pathological and other inflammatory conditions, iNOS is expressed mainly by the two glial cell types involved in neuroimmune regulation, activated astrocytes and microglia (Merrill, et al., 1997, Mucke and Eddleston, 1993, Perry, et al., 2003). Induction of iNOS in isolated glial cells treated with proinflammatory cytokines, microbial and viral products, and abnormal protein aggregates is well documented (Adamson, et al., 1996, Brosnan, et al., 1997, Meda, et al., 1999, Militante, et al., 1997, Murphy, et al., 1993, Syapin, et al., 2001). The mechanisms by which these agents regulate expression of the Nos2 gene have been the focus of intense studies to develop strategies to inhibit NO synthesis and thereby suppress inflammatory reactions and subsequent brain damage.
There are multiple regulatory elements found in the rat Nos2 promoter (Eberhardt, et al., 1996, Gavrilyuk, et al., 2001, Zhang, et al., 1998). However, the molecular details regulating expression of the gene in glia is incompletely understood and the level of involvement of each of the regulatory elements found in the promoter is presently unclear. We have previously shown the presence of positive and negative cis-acting regions in the proximal promoter of the rat Nos2 gene (Sanchez, et al., 2003). In that study we demonstrated that unlike the murine Nos2 promoter where a downstream NF-κB site plays a key role for lipopolysaccharide (LPS) responsiveness (Xie, et al., 1994), the proximal NF-κB site in the rat Nos2 promoter is not required for transcriptional activation in C6 glial cells (Sanchez, et al., 2003).
Extensive information is available on Nos2 transcription, particularly from murine macrophage (Lowenstein, et al., 1993, Xie, et al., 1993). However, the majority of these studies have been carried out in vitro using the electrophoretic mobility shift assay (EMSA) or reporter genes. Despite the usefulness of these studies, these techniques do not take into account actual protein-DNA interaction in intact cells. For these reasons, we investigated rat Nos2 transcription using in vivo DNA footprinting with ligation-mediated PCR (Garrity and Wold, 1992, Mueller and Wold, 1989) in addition to EMSA and reporter gene studies. In vivo DNA footprinting provides the strongest evidence yet available for a functional consequence to in vivo occupation of gene promoter sequences by proteins when used in combination with reporter gene and EMSA data.
In the mouse macrophage cell line RAW 264.7, studies found that a putative, non-consensus single-mismatch octamer (Oct) motif, found 15 bp downstream of the proximal NF-κB site of the mouse Nos2 promoter, is required for iNOS activation by LPS (Xie, 1997) and LPS + interferon (IFN) γ (Kim, et al., 1999). The rat Nos2 promoter also contains this non-consensus octamer motif (ATGCAAAA) found 24 bp upstream from the TATA box. This octamer motif has been shown to be critical for interleukin (IL)-1β and IFNγ mediated promoter activity in rat primary β-cells and in RINm5Fcells (Darville, et al., 2004).
Long-term chronic and acute, binge-type alcohol abuse, cause changes in CNS gene expression that are partly responsible for the development of pathophysiological changes such as alcoholic tolerance, dependence, and brain damage (Fan, et al., 2004, Liu, et al., 2006). Previous studies indicate that ethanol functions transcriptionally to inhibit expression of the Nos2 gene in C6 glia (Syapin, et al., 2001), although the mechanism by which ethanol inhibits Nos2 transcription remains unclear. In general, ethanol causes inhibition of cytokine-induced iNOS expression in a time- and concentration-dependent manner independent of cell or tissue type (Davis and Syapin, 2005). In the case of cultured glial cells, ethanol is effective at concentrations ranging from 25 mM to 200 mM (Lee, et al., 2004, Militante, et al., 1997, Syapin, et al., 2001, Wang and Sun, 2001, Wang, et al., 1998). The biological relevance of this concentration range is supported by results of both animal and human studies. In animal models, blood ethanol concentrations of 50 mM are routinely exceeded (Nixon and Crews, 2002). Similarly, blood ethanol levels in human subjects have been reported to be 50 mM or higher (Urso, et al., 1981, Wells and Barnhill, 1996), and in at least one case in excess of 300 mM (Johnson, et al., 1982). For our current study, we exposed cultured cells to a relatively high ethanol concentration to ensure robust ethanol inhibition of iNOS expression that would increase the likelihood of observing relevant transcriptional effects.
The aim of the present study was to further define the key regulatory elements for Nos2 induction in a glial phenotype and to characterize the functional role of the octamer motif contained within the overlapping γ-IRE, GAS and Oct binding sites of the proximal promoter of the rat Nos2 gene. Data presented herein revealed novel inducible interactions with the rat Nos2 core promoter regions. We have identified a region in the Nos2 promoter from −86 to −65 bp, where cytokine and LPS induced protein-DNA interactions are altered in cells exposed to ethanol. These observations lend significant insight into the mechanism by which ethanol alters Nos2 expression in C6 glia.
2. Results
2.1 The piNOS-luc-78 promoter has no stimulated transcriptional activity
We have previously characterized the reporter gene activity of serial 5′ flanking region deletions in the rat promoter to identify cis-acting elements relevant to Nos2 induction and regulation (Sanchez, et al., 2003). Reporter gene data indicate five regions in the core promoter relevant to Nos2 induction (Figure 1). Stimulated activity observed from the 94 bp construct which lacks the putative proximal NF-κB binding site at −107 to −97 bp indicate that this proximal NF-κB is not an absolute requirement for promoter activity. As previously demonstrated, the piNOS-luc-94 promoter showed highly significant inducible activity for each set of stimuli tested (P<0.0001, paired t-test, n=6). Therefore, binding sites present in the 94 bp promoter appear to be sufficient for rat glial Nos2 activation. Previous EMSA and in vivo footprinting data also showed protein binding to the −86 to −64 bp region of the Nos2 promoter (Sanchez, et al., 2003), further supporting the importance of this region for inducible promoter activity.
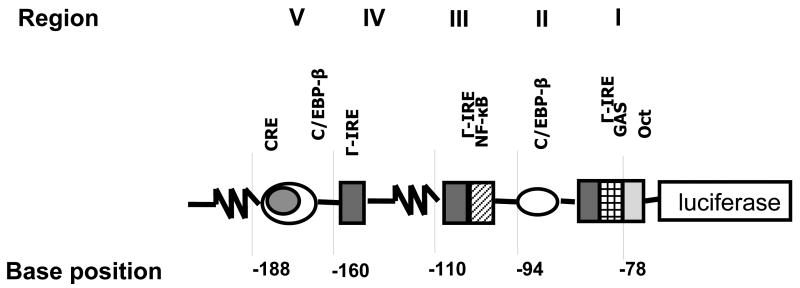
Figure 1.
Schematic showing the rat core promoter-luciferase expression vector with the identified five regions containing the relevant transcription factor binding sites (modified from Sanchez et al., 2003).
A further 5′-deletion of 16 bases from the piNOS-luc-94 promoter to the shorter piNOS-luc-78 promoter (−78 to +133 bp) abolished the ability of the plasmid to respond to appropiriate induction stimuli, i.e., exposure to CM, LPS+IFNγ, or LPS+PMA. With all 3 set of stimuli the piNOS-luc-78 promoter showed only basal, and not stimulated activity (see control data in Figure 3).
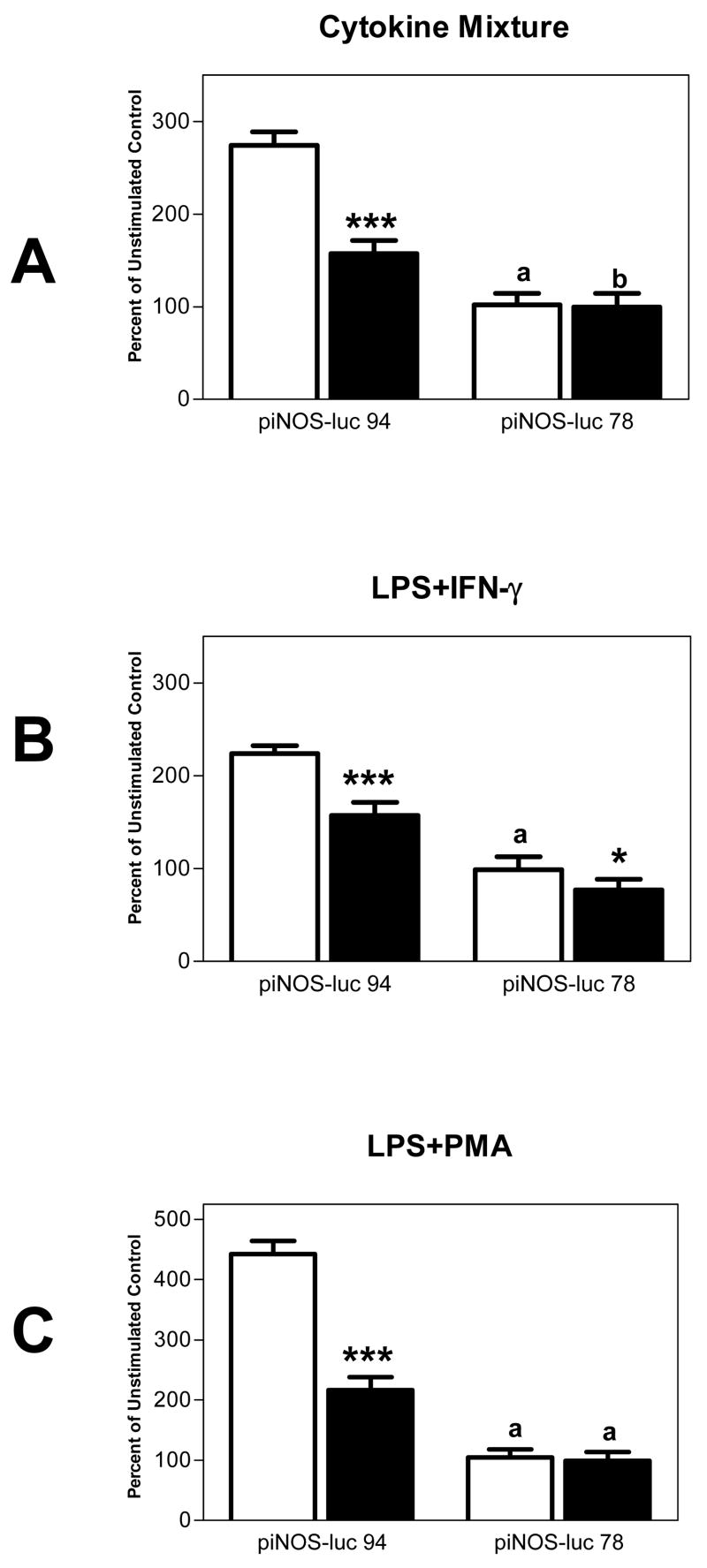
Figure 3.
Deletion of 16 bp from piNOS-luc-94 (−94 to −78 bp) abolished inducible promoter activity and ethanol suppression. Values are mean RLU/μg protein + SEM expressed as percent of the unstimulated no ethanol control. White bars are cells stimulated in the absence of ethanol and black bars represent cells stimulated in the presence of 150 mM ethanol for 24 h. One-way ANOVA indicated no significant differences (P=0.348) between unstimulated and stimulated groups for cells expressing the piNOS-luc-78 construct.
*, P<0.05 (paired t-test) versus corresponding no ethanol condition; n=10
***, P<0.0001 (paired t-test) versus corresponding no ethanol condition except for LPS+IFNγ where P=0.0006; n=6
a, P<0.0001; b. P<0.05 (Bonferroni post-hoc tests) versus corresponding piNOS-luc-94 condition
2.2 Acute ethanol inhibits piNOS-luc-94 but not piNOS-luc-78 promoter activity
Figure 2 shows acute ethanol exposure (150 mM) for 24 h inhibits LPS+IFNγ-induced promoter activity of all the constructs studied, including the piNOS-luc-94 promoter construct. The same results are observed using CM and LPS+PMA to induce activity (data not shown). When the effect of ethanol on piNOS-luc-78 promoter activity was examined, there was neither a stimulation nor inhibition of activity (Figure 3), except for a slight, but statistically significant reduction in basal activity of cells exposed to LPS+IFNγ (Figure 3B). These data confirm that ethanol sensitive cis-acting motif(s) mediating alcohol inhibition of transcriptionally induced Nos2 activity reside between nucleotides −94 to −1 bp of the rat Nos2 promoter. Furthermore, the results suggests that the promoter region from bases −87 to −66, a region known to be involved in Nos2 gene activation as shown by in vivo DNA footprinting (Sanchez, et al, 2003), may contain the site where ethanol modulation occurs.
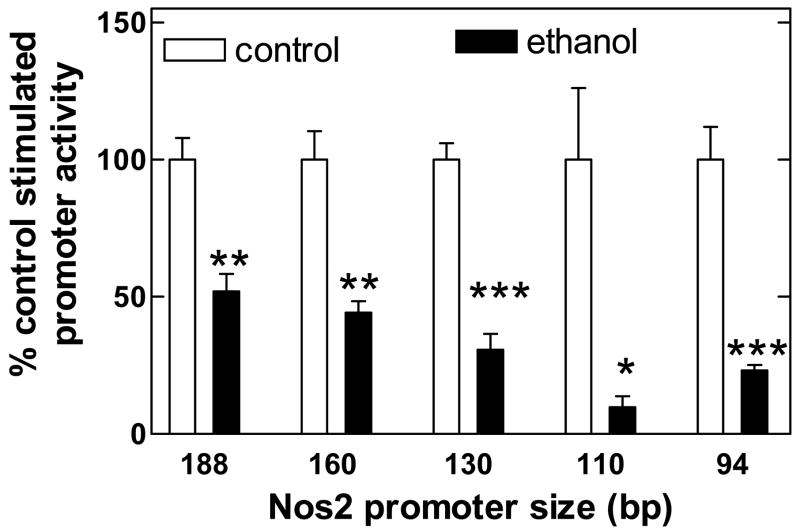
Figure 2.
Ethanol (150 mM for 24 h) inhibits LPS+IFNγ-stimulated promoter activity including the piNOS-luc-94. Values are mean + SEM for pooled data samples from 5, 4, 5, 2 and 4 experiments (left to right) on at least 2 separate stable transfected C6 cell lines normalized to their own controls. *, P=0.008 versus control (paired t-test); **, P=0.0001 versus control (paired t-test); ***, P<0.0001 versus control (paired t-test).
2.3 Oct motif mediates alcohol inhibition of iNOS promoter induction
The piNOS-luc-94 promoter carries a C/EBPβ site and overlapping sites for γ-IRE/GAS/Oct. The smaller piNOS-luc-78 promoter does not have the C/EBPβ, γ-IRE or GAS sites, but retains the minimal Oct motif. Ethanol inhibition of the stimulated activity of the 94 bp promoter construct implies that the C/EBPβ and/or overlapping γ-IRE, GAS and Oct motifs comprise an ethanol sensitive region (ESR), and are crucial for ethanol-mediated modulation of rat glial Nos2 expression. To further clarify the cis-acting motif(s) involved in the ethanol inhibition, we investigated the role of the Oct site in the context of a longer, inducible promoter. C6 cells were stably transfected with a construct of the 1002 bp of the wild-type rat Nos2 promoter or with piNOS-1002Oct/M2 that has a double mutation in the 3′ end dinucleotide AA residues (A to C at position –71 and A to G at position –70) of the Oct motif (Darville, et al., 2004) contained in the implicated ESR. The wild type promoter (piNOS-1002luc) showed significant inducible activity in cells stimulated with CM or LPS+PMA, that was sensitive to inhibition by ethanol exposure (Fig. 4A and C). Interestingly, piNOS-1002luc reporter gene activity induced by LPS+IFNγ was not inhibited by ethanol exposure (Fig. 4B). In contrast, mutation of the dinucleotide AA residues in the Oct motif (piNOS-1002Oct/M2) resulted in a plasmid that was more responsive to CM induced promoter activation, showed equivalent activation by LPS+IFNγ as the wild-type promoter, but was drastically less responsive to LPS+PMA induced activation. Although to a lesser degree relative to cells expressing the wild-type piNOS-1002luc reporter gene, treatment of cells expressing the piNOS-1002Oct/M2 plasmid with LPS+PMA resulted in a small (22%), but statistically significant inducible activity (P=0.0172, paired t-test, n=7). Importantly, the site-directed dinucleotide mutation of the Oct motif, designated as Oct/M2 herein, eliminated ethanol inhibition of CM and LPS+PMA induced promoter activity, without alteration of the C/EBPβ, γIRE or GAS sites (Fig. 4A and C). These results suggest a critical role for factors binding to the Oct site during alcohol mediated inhibition of Nos2 promoter activity in glial cells.
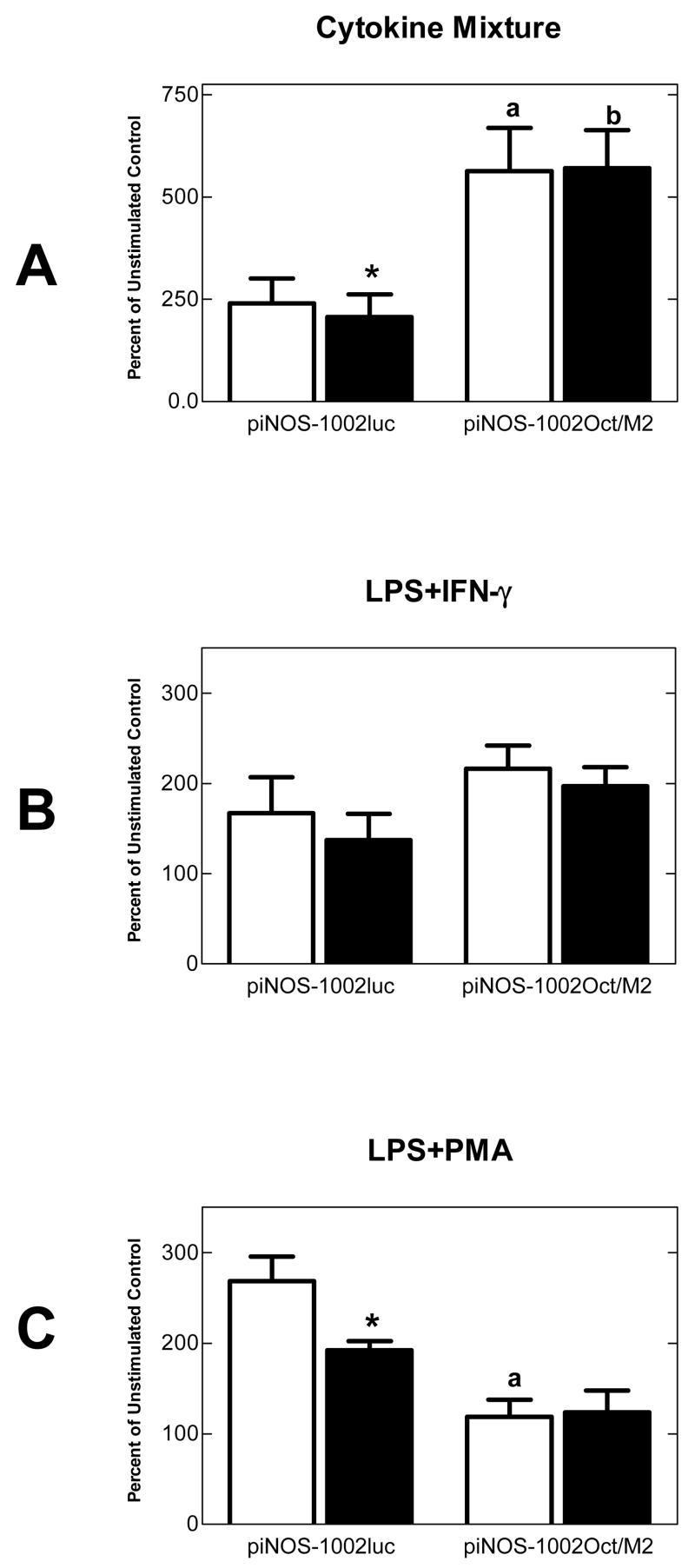
Figure 4.
Effect of octamer site mutation on ethanol suppression of induced Nos2 promoter activity. The Oct mutation drastically reduced LPS+PMA induced promoter activity while enhancing cytokine mediated promoter activity. The Oct mutation also abolished ethanol suppression of induced Nos2 promoter activity. Repeated measures ANOVA indicated significant stimulated activity, with or without 150 mM ethanol, for cells expressing the piNOS-1002luc construct when exposed to cytokine mix (P=0.0276, n=6) or LPS+PMA (P=0.0082, n=4). Cells expressing the piNOS-1002Oct/M2 construct showed significant stimulated activity when exposed to cytokine mix (P=0.0002, n=6), LPS+IFNγ (P=0.0001, n=6), or LPS+PMA (P=0.0288, n=5).
*, P<0.05 (paired t-test) versus corresponding no ethanol condition
a, P<0.001; b. P<0.01 (Bonferroni post-hoc tests) versus corresponding piNOS-1002luc condition
2.4 EMSA data support DNA-protein interaction in the core promoter region
We previously reported EMSA data showing active in vitro protein binding in four of the five identified regions in the core promoter (Sanchez, et al., 2003). DNA-protein interaction was detectable for the probe designed for the γ-IRE/NF-κB (−115 to −94), CRE/CEBPβ site (−159 to −130), and for the γ-IRE site in CM induced cells. No protein-DNA interaction was observed for the proximal C/EBPβ site (−96 to −83) probe. Essentially the same pattern was observed for LPS+IFNγ induced cells (data not shown).
DNA-protein interactions were also observed using a probe for theγ-IRE/GAS/Oct (−86 to −65) ESR sequence, denoted ESR-1. Three complexes, two major and one minor were obtained (Figure 5). Although the data in Figure 5 appears to show an increase in protein binding to the ESR-1 probe from extracts prepared from stimulated control cells, and a decrease in extracts from ethanol treated cells, this was not observed consistently (see Figure 7). Instead, any changes, if present, in protein binding to the ESR-1 probe due to cell stimulation or ethanol treatment were inconsistent and subtle across at least 6 independent experiments (data not shown). To further investigate binding to the ESR-1 sequence, several 14-bp oligonucleotide probes were prepared, each carrying point mutations for one of 5 putative binding sites, and studied by EMSA. Point mutations within the −80 to −67 bp region practically abolished protein binding, with the γ-IRE mutation being least disruptive to protein-DNA binding (data not shown). However, because the 14-bp mutant probes lacked sequences found in the wild-type ESR-1 probe, a new set of 22-bp probes was generated for EMSA (Table 1). This set of ESR-1 probes also contains point mutations within each of the putative binding motifs and was used for binding competition studies against the wild-type ESR-1 probe in EMSA experiments. Two different mutations in the Oct motif were studied. One, designated OCT/M1, was a single A to T change at position –72. The second, designated Oct/M2, was the double mutation in the 3′ end dinucleotide AA residues (A to C at position –71 and A to G at position –70) that corresponded to the mutation present in the piNOS-1002Oct/M2 reporter gene studied previously (see Figure 4). The rationale for these studies was that if a mutation in the ESR-1 reduces protein:DNA complex formation, the ESR-1 oligonucleotide containing that mutation is expected to be a weak competitor for binding in the EMSA. On the other hand, if a mutation in the ESR-1 still supports complex formation, an ESR-1 oligonucleotide with that mutation should strongly compete for protein binding in the EMSA. Figure 6A shows that protein binding to the ESR-1 region was strongly disrupted by competitor probes containing mutations in the IRS, γ-IRE, or position –72 of the Oct motif (the OCT/M1 probe), indicating these mutations do not alter protein binding to ESR-1. In contrast, the Oct/M2 dinucleotide ESR-1 mutant was a poor binding competitor, indicating the Oct/M2 mutation disrupts protein binding to the ESR-1 sequence. These results were reproducible, as shown in Figure 6B using control nuclear extracts from an independent experiment. Figure 6B also shows that protein binding to the ESR-1 region was strongly competed by DNA containing the consensus sequence for the octamer-1 motif, but not by DNA containing the AP2 consensus sequence.
Figure 5.
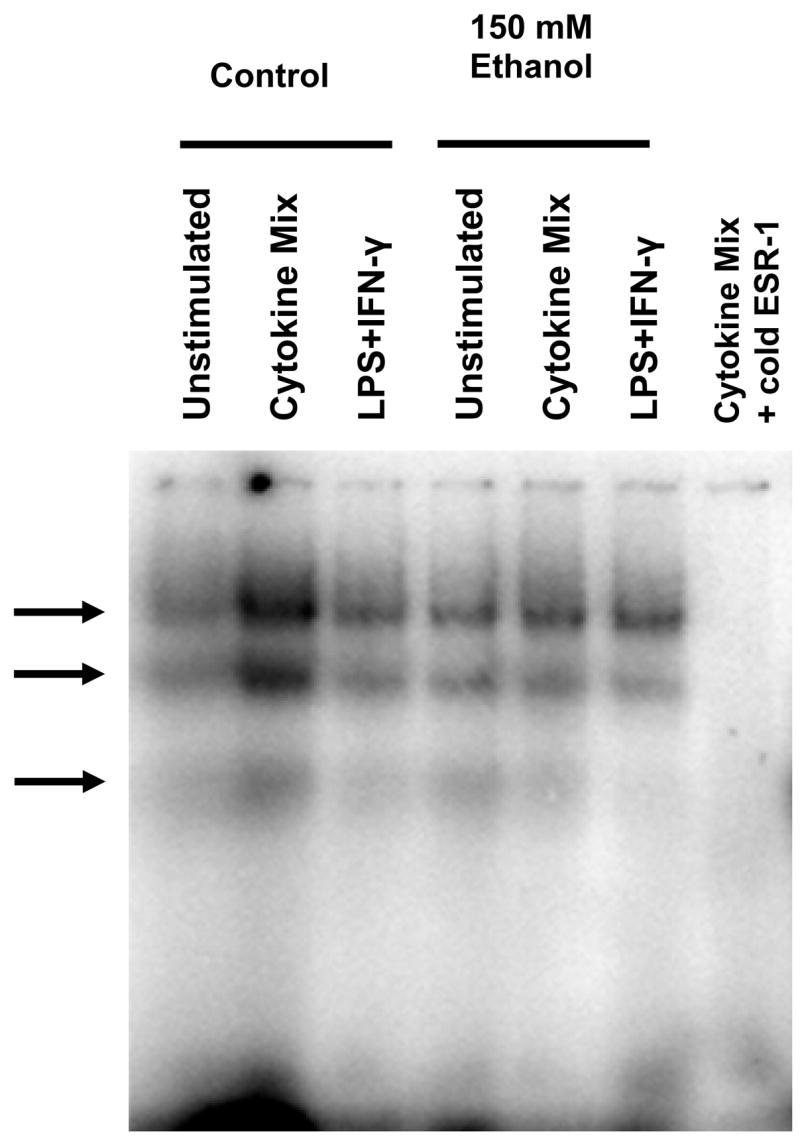
Protein-DNA interactions using ESR-1 as probe (−86 to −65 bp). Arrows denote location of specific complexes repeatedly observed with C6 cell nuclear extracts. Essentially the same two major and one minor complex are formed when cells are stimulated with cytokine mixture or LPS+IFNγ.
Figure 7.
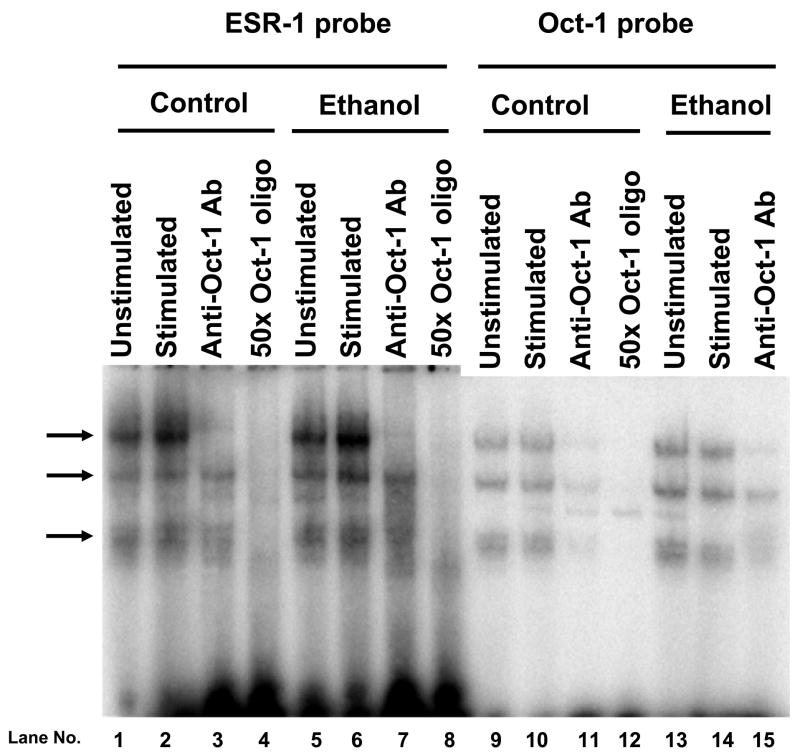
The overlapping wild-type γ-IRE/GAS/Oct sites in the proximal rat Nos2 promoter bind α-Oct-1 and Oct-1 related protein(s) to form 3 complexes. EMSA was performed using the 22-bp γ-IRE/GAS/Oct oligonucleotide probe incubated with nuclear extract from unstimulated C6 cells and cells stimulated with cytokine mixture for 3 h. Arrows denote location of specific complexes repeatedly observed with C6 cell nuclear extracts. 50x Oct-1 oligo denotes competition with a 50-molar excess of an octamer-1 consensus oligonucleotide (see Table 1). Figure is representative of results obtained on 3 or 4 independent nuclear extracts.
Table 1.
Oligonucleotide sequences of electrophoretic mobility shift assay probes. The DNA binding site motif is shown in bold font, while mutated nucleotides are italicized and bolded.
| Region | Probe Name | Length (bp) | Position | Sequence |
|---|---|---|---|---|
| V | CRE-C/EBPβ | 22 | −183 to −162 | 5′ T ACA CCA CAG AG T GAC TA ATA |
| IV | IRE | 30 | −159 to −130 | 5′ GCA TAC AGA CTA GGA GTG CC ATC GCG AAT |
| III | Proximal NF-kB | 22 | −115 to −94 | 5′ A CCC TA C TGG GGA CTC CC CTT |
| II | Proximal C/EBPβ | 14 | −96 to −83 | 5′ CTT TGG GAA CAG TG |
| I | γ-IRE/GAS/Oct wild-type probe (ESR-1) | 22 | −86 to −65 | 5′ AGT GAC TTT ATG CAA AAC AGC T |
| I | γ-IRE mutant probe | 22 | −86 to −65 | 5′-AGT GAG GTT ATG CAA AAC AGC T |
| I | GAS mutant probe | −86 to −65 | 5′-AGT GAC TGG ATG CAA AAC AGC T | |
| I | IRS mutant probe | 22 | −86 to −65 | 5′-AGT GAC TTT ATG CAA AAG TGC T |
| I | OCT/M1 mutant probe | 22 | −86 to −65 | 5′-AGT GAC TTT ATG CTA AAC AGC T |
| I | Oct/M2 mutant probe | 22 | −86 to −65 | 5′-AGT GAC TTT ATG CAC GAC AGC T |
| OCT1 consensus probe | 22 | 5′-TGT CGA ATG CAA ATC ACT AGA A | ||
| AP2 consensus probe | 5′-GAT CGA ACT GAC CGC CCG CGG CCC GT |
Figure 6.
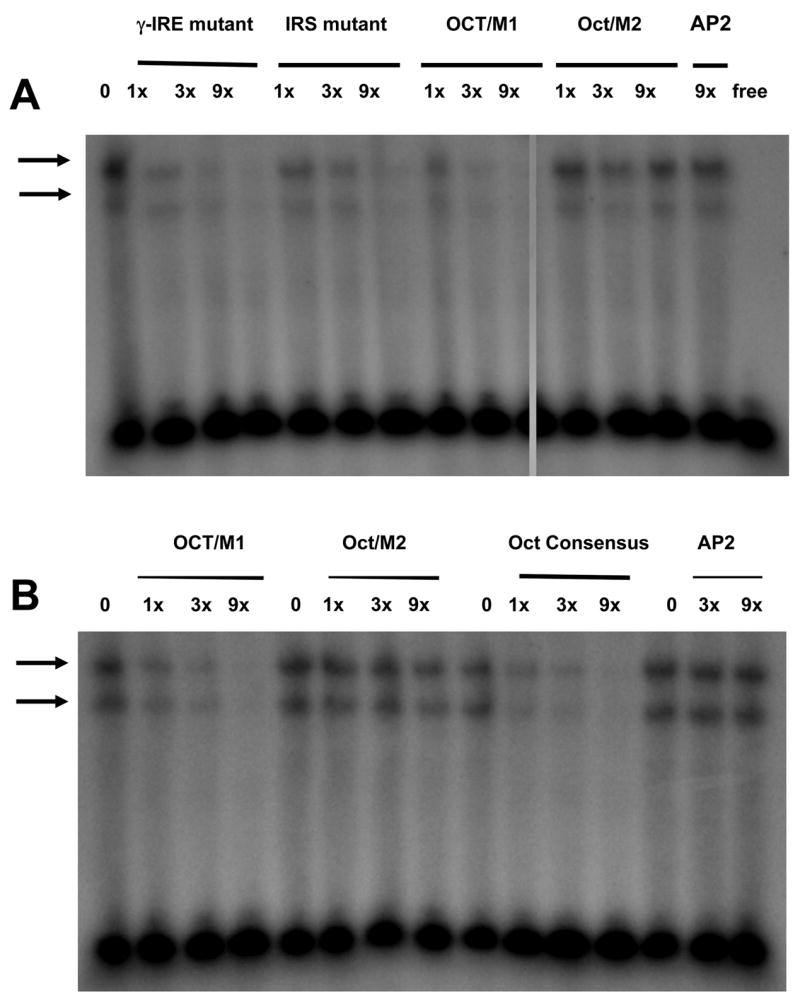
Binding competition of the wild-type ESR-1 probe with indicated molar excess (1x–9x) of unlabeled probes, each containing point mutations within an overlapping binding site (see Table 1). Arrows denote location of specific complexes repeatedly observed with C6 cell nuclear extracts. The fastest migrating complex is absent because the nuclear extracts contained phosphatase inhibitors. Figure is representative of results obtained on 2 or 3 independent nuclear extracts, including some without phosphatase inhibitors. ‘free’ denotes lane contains radioactive γ-IRE/GAS/Oct probe only.
In addition to the three ESR-1 complexes being competed by a 50x molar excess of oligonucleotide containing a consensus octamer motif (Figure 7, lanes 4 and 8), the consensus Oct probe also formed 3 complexes (Figure 7, lane10). These complexe were also competed by 50x molar excess of cold probe (Figure 7, lane 12), indicating binding of octamer related factors in the three complexes. These results are consistent with previous results for the binding of nuclear proteins from C6 cells to an Oct-1 consensus sequence (Schreiber et al., 1990). However, only the protein-DNA interaction in the slowest migrating ESR-1 band (complex 1) was inhibited by an Oct-1 antibody (Figure 7, lanes 3 and 7), similar to results from rat astrocytes (Schreiber et al., 1990). These results show that the octamer site binds an Oct-1 related protein. As mentioned previously, no detectable effect of exposing cells to ethanol were observed in these in vitro DNA-protein interactions (Figure 7, lanes 5–8 and 13–15). These results are representative of at least 3 independent experiments.
The direct effect of 200 mM ethanol on the in vitro binding of control nuclear extracts to the ESR-1 probe was also tested and found to have no effect on the number or pattern of binding complexes formed in the EMSA (data not shown).
2.5 Acute ethanol blocks protein occupation in the promoter in vivo
Following the identification of DNA binding motifs relevant to Nos2 activation by reporter gene assay and protein-DNA interaction using EMSA, we carried out in vivo footprinting with ligation-mediated PCR to confirm and/or define in vivo protein-DNA binding events in the core promoter region of the rat Nos2 gene of C6 glial cells. This procedure provides a qualitative analysis of chromosomal gene promoter occupancy by DNA binding proteins under in vivo conditions (for reviews see: Warshawsky and Miller, 1999; Cartwright and Kelly, 1991). We limit the discussion of results to those guanine residues consistently protected or hypermethylated in three separate samples and in independent experiments.
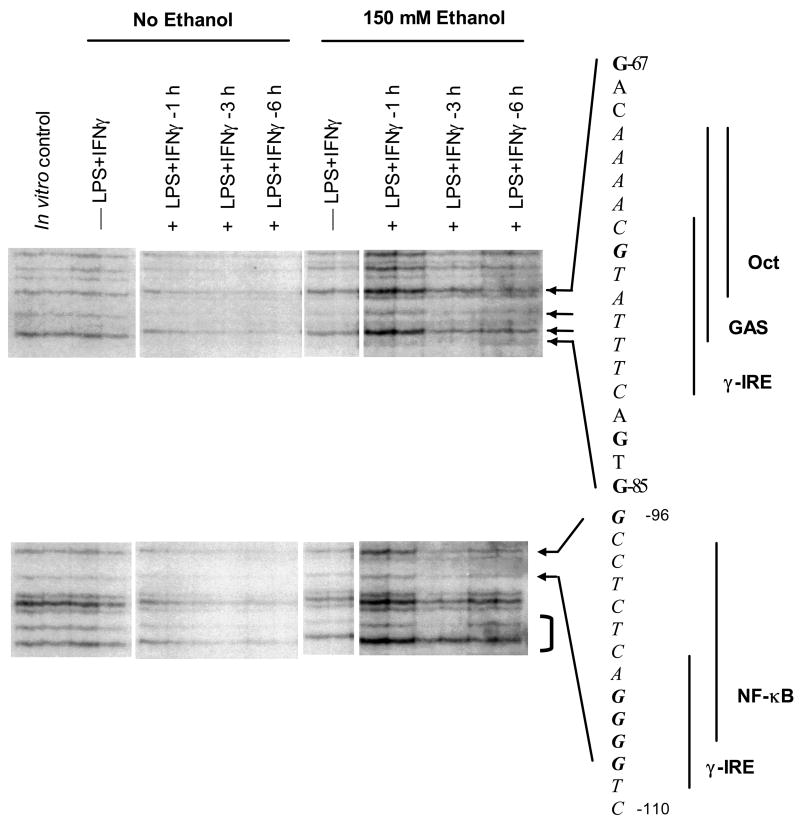
Acute exposure to ethanol allowed greater dimethyl sulfate (DMS) reactivity in LPS+IFNγ stimulated cells as indicated by darker bands in ethanol treated cells (Figure 8 No ethanol vs. 150 mM ethanol lanes), consistent with less protein occupation of these sites. The guanine residues in the overlapping γ-IRE/GAS/Oct site (ESR-1) and in the γ-IRE/NF-κB sites in the coding strand both appear to be strongly ethanol sensitive. The faint bands in ethanol treated cells corresponding to guanine residues in both the proximal C/EBPβ (region II) and the upstream CRE-C/EBPβ (region V) sites, and darker bands corresponding to guanines in the γIRE sites (bases –144, –146, –147 and bases –152 and –159) suggest binding to these sites is also inhibited by exposure to acute ethanol. Increased DMS sensitivity mediated by ethanol appears to be maximal at 3 h after stimulation and less pronounced after 6 h of stimulation. Footprinting of the non-coding strand identified similar DNA-protein interactions in the same sites upon stimulation and similar effects of acute exposure to ethanol (data not shown). DMS-sensitivity of the same sites was observed in CM induced cells (Sanchez, et al., 2003), indicating a general role of these sites in Nos2 gene induction.
Figure 8.
In vivo DNA footprinting by ligation-mediated PCR. Results show effect of 150 mM ethanol on sensitivity to DMS methylation of guanine residues in the rat Nos2 core promoter sequence in C6 wild type cells after indicated time of stimulation with LPS+IFNγ. Arrows denote bands that disappear over time following stimulation, indicating in vivo protein protection of guanine residues in the corresponding site on the 5′ flanking region of the endogenous rat core promoter region. Bracket denotes protection of guanine residues immediately upstream of the NF-κB site. Samples were run as side-by-side duplicate lanes on the electrophoresis separating gel.
3. Discussion
The identification of relevant DNA regulatory sequences involved in transcriptional control is a prerequisite to identify proteins mediating cell-specific gene expression. In vivo footprinting reveals protein-DNA interactions as they actually occur in intact cells and is considered a benchmark approach to investigating mechanism of gene regulation in situ. We have used in vivo DNA footprinting to complement in vitro studies of the rat Nos2 regulatory regions to identify regulatory sequences crucial for ethanol-mediated suppression of Nos2 expression. In the present study, we provide evidence for the involvement of several factors binding within the rat Nos2 core promoter region. Previous studies in the mouse have implicated NF-κB (Lowenstein, et al., 1993, Sherman, et al., 1993) and IRF-1 (Kamijo, et al., 1994, Martin, et al., 1994) in the transcriptional response of the Nos2 gene to LPS+IFNγ. In the rat, NF-κB, C/EBPβ and ATF-2, and CREB (Bhat, et al., 2002, Gavrilyuk, et al., 2001) have been implicated in transcriptional activation of Nos2 by LPS+IFNγ in glial cells. Our data suggest an important role for the ESR-1 region comprised of the overlapping γ-IRE/GAS/Oct triad, and particularly the Oct motif, both in Nos2 activation and in ethanol mediated suppression of its expression. The data show protein protection of a guanine residue (–75 bp) in the ESR-1 region during LPS+IFNγ stimulation. The same guanine residue showed increased DMS sensitivity in corresponding conditions under acute ethanol, indicating ethanol is blocking the protein-DNA interaction in the region. Increased DMS reactivity of guanines found in the γ-IRE sites (bases –144, –146, –147 and bases –152 and –159) suggests binding to these sites are also inhibited by exposure to acute ethanol.
Although the importance of NF-κB in the Nos2 promoter has been implicated from previous studies (Bhat, et al., 2002, Gavrilyuk, et al., 2001), our reporter gene data suggest an indirect role for NF-κB during Nos2 induction in rat C6 cells, based on activity of the 94 bp promoter construct which lacks any known NF-κB site. However, in vivo footprinting showed protein occupation in the overlapping γ-IRE/NF-κB region and previous EMSA data showed weak binding during stimulation using a combination of cytokines (Sanchez, et al., 2003). Hence, it is possible that the necessary protein-DNA conformations occurring in vivo could not be fully replicated by the in vitro EMSA conditions.
The present study shows for the first time that an octamer binding site contained within an overlapping γ-IRE/GAS/Oct site in the rat proximal Nos2 promoter plays an important role for gene induction activity in the rat C6 glial cell line. The octamer site under consideration carries the dinucleotide LPS-response element (LREAA) reported previously in the mouse. The LREAA together with NF-κB is reported to be required for LPS-induced Nos2 promoter activity in the mouse RAW 264.7 cell line (Xie, 1997). Transfection experiments carried out in rat C6 cells using a promoter construct with an intact Oct site, but with deletion of nucleotides immediately upstream of this Oct site (piNOS-luc-78), completely abolished induced promoter activity. This indicates that besides the octamer binding site itself, other unidentified sites located in the −94 to −79 bp region of the Nos2 promoter, (e.g., a putative C/EBPβ site) contribute to induced promoter activity. The importance of flanking regions around octamer binding sites has been previously noted (Herr and Cleary, 1995, Walker, et al., 1994). The data confirm the presence of in vitro nuclear protein binding activity, and in vivo protein binding for the Nos2 gene DNA region at −87 to −66 bp from the transcriptional start site demonstrated previously (Sanchez, et al., 2003). In this aspect, the rat promoter is responding in an equivalent manner as the mouse Nos2 promoter.
Site-directed mutation of the octamer motif in the context of a longer, intact promoter (piNOS-1002Oct) has a complicated effect on ethanol induced inhibition of Nos2 promoter activity. Interestingly, the plasmid carrying the −1002 to +132 bp portion of the wild-type Nos2 promoter was not as strongly activated by cytokines as some other deletion mutants (Sanchez et al, 2003), and appeared to be less sensitive to ethanol, depending on the specific inducers used, although this has not been studied further. While these complexities could hinder interpretation of some data, the results indicate that in those cases where ethanol significantly inhibitied inducible activity of the wild-type promoter fragment, that inhibition was no longer present when the promoter fragment contained a double mutation in the proximal Oct1 site, herein designated as Oct/M2, thereby supporting a role for the proximal Oc1 site in mediating the ethanol inhibition. On the other hand, EMSA experiments showed that mutations in other putative transcription factor binding sites within the overlapping γ-IRE/GAS/Oct motif of the ESR-1 sequence, including those designed to distrupt protein binding to the IRS, γ-IRE, and GAS sites, did not change the ability of the mutant ESR-1 to complete for binding against the wilde-type sequence. However, the Oct/M2 mutation of the dinucleotide AA in the octamer motif alone abolished the ability of the mutant probe to compete for protein binding to the ESR-1 probe. This further underscores the critical role of the octamer motif for actual protein-DNA binding.
The EMSA binding competition experiment using an Oct-1 antibody indicated that the protein binding the DNA in complex 1 is the ubiquitously expressed transcription factor Oct-1 (Kambe, et al., 1993). It is interesting to note that while mutation of the octamer site greatly reduces LPS+PMA induced promoter activity, the same mutation appears to enhance cytokine induced promoter activity (Figure 4, Panel A versus Panel C). This result is in contrast to that observed in rat RINm5F cells wherein mutation of the same octamer site reduced both IL-1β and IL-1β+IFNγ induced promoter activity and caused a 5-fold decrease of the IL-1β induced promoter activity in rat primary β cells (Darville, et al., 2004).
That mutation of the proximal Oct motif can enhance transcriptional activation of the Nos2 promoter in response to certain stimuli and limit transcriptional activation to other stimuli, suggests that this site may serve a dual role in Nos2 promoter activity. Dual transcriptional functions for Oct-1 (as an activator or repressor) have been reported in the regulation of the human GnRH receptor gene (Cheng, et al., 2002, Cheng, et al., 2001), although the effects are mediated through different promoter regions. A dual function for Oct-1 has been explained by the fact that Oct-1 phosphorylation alters its DNA binding activity (Belsham and Mellon, 2000) and that the protein can adopt site-specific conformations, depending on the exact binding motif and flanking sequences involved (Walker, et al., 1994). Furthermore, other members in the POU family, originally called POU for the Pit-1 factor in the pitituary, the octamer-binding proteins Oct-1 and Oct-2, and the Unc-86 gene of Caenorhabditis elegans (see Andersen and Rosenfeld, 2001 for a review) can interact with Oct-1 (Bendall, et al., 1993, Verrijzer, et al., 1992) and other POU family members can bind the octamer sequence (Gay, et al., 1998). Therefore, the precise DNA and protein-protein contacts made by Oct-1 at different promoter sites appear capable of imparting unique regulatory properties to this ubiquitous transcription factor.
In summary, the regulation of the Nos2 gene is clearly complex and brought about by the action of multiple cis-acting sites in the promoter and their corresponding binding factors. We have identified an octamer regulatory sequence as a key cis-acting element having a dual function within the rat Nos2 promoter in cells with a glial phenotype. Furthermore, the octamer sequence resides within an ethanol sensitive region (ESR-1) of the proximal Nos2 promoter that appears necessary and sufficient for ethanol to inhibit the Nos2 promoter activity of C6 glial cells. The POU domain transcription factor Oct-1, through binding to this octamer sequence, plays an essential role in the transcriptional induction of the Nos2 gene. Our observations confirm and extend previous results that ethanol reduces Nos2 gene expression (Militante, et al., 1997, Syapin, et al., 2001). Furthermore, in vivo DNA footprinting of the Nos2 promoter region revealed several regulatory regions relevant to ethanol-mediated suppression of iNOS expression.
4. Experimental Procedures
4.1 Cell Culture
Procedures for cell culture were done as described previously (Sanchez, et al., 2003, Syapin, et al., 2001). All cell cultures were maintained at 37°C inside humidified incubators with 5% CO2:95% air.
4.2 Stable Transfection and Reporter Genes
Rat C6 cells were transfected with rat Nos2 promoter constructs prepared in the pGL3 vector (Promega, Madison, WI, USA). The co-precipitation of calcium phosphate and DNA method was used to prepare stable transfectants following standard protocols for adherent cells incorporating minor modifications for C6 cells (Feinstein, et al., 1996). The C6 cells were co-transfected with pBK-CMV plasmid to allow selection by growth in 0.6 or 0.3 mg/mL geneticin. Stable transfection was chosen to enable the use of the same cells for unstimulated controls, allowing analysis of actual stimulated reporter gene activity.
The various piNOS-luc-promoter constructs were prepared according to protocols reported previously (Sanchez, et al., 2003). The smaller piNOS-luc-78 promoter was generated from the piNOS-luc-94 promoter construct using the Erase-A-Base system (Promega) following the general procedures recommended by the manufacturer. Briefly, the piNOS-luc-94 promoter construct was linearized and an ExoIII resistant 3′ overhang was generated by digestion with KpnI. An ExoIII susceptible 5′ overhang was then generated by digestion with MluI. The DNA was then precipitated by ethanol and subjected to a controlled ExoIII digestion/deletion reaction. The resulting ExoIII digests were treated with S1 nuclease to remove single strands of DNA. Klenow DNA polymerase was added after the S1 nuclease was neutralized and heat inactivated. The deletion-containing vectors were then ligated and used directly for transformation of competent cells. Several subclones were screened and sequenced to select for the appropriate piNOS-luc-78 promoter construct.
The piNOS-1002luc (wild type) and piNOS-1002Oct (herein designated piNOS-1002Oct/M2, which contains a double mutation in the 3′ end dinucleotide AA residues (A to C at position –71 and A to G at position –70) in the octamer site that is identical to the mutation present in the Oct/M2 mutant ESR-1 oligonuceotide probe used in the EMSA) promoter constructs were kindly provided by Dr. Martine Darville from the Université Libre de Bruxelles, Belgium.
4.2 Nos2 Induction
C6 cells were activated in serum-free media for 24 h using the following conditions: LPS+IFNγ; 10 μg/mL LPS (E. coli serotype 055:B5; Sigma, St. Louis, MO, USA) +100 U/mL rat IFNγ: LPS+PMA; 500 ng/mL LPS with 400 ng/mL PMA: and CM (2 ng/mL human IL-1β, 100 U/mL rat IFNγ, and human TNFα) These 3 set of stimuli are routinely used by our laboratory because they activate different, but synergistic, signaling pathways leading to Nos2 gene induction and iNOS protein expression with different time courses (Sanchez et al., 2003), and sensitivity to ethanol exposure (Syapin et al., 2001). The CM used to induce Nos2 expression for cells transfected with the various piNOS-luc promoter deletion constructs contain 60 ng/mL TNFα. CM for the cells transfected with piNOS-1002luc and piNOS-1002Oct promoter constructs contained a lower TNFα concentration (5 ng/mL), following our observation that this lowered concentration still induced a maximum response.
Luciferase Activity Assay
Transfected cells were grown in 12-well plates and stimulated for Nos2 induction as detailed above (4.2). The cells were then extracted with 250 μl per well of 1x Passive Lysis Buffer (Promega, Madison, WI, USA) according to the manufacturer’s instructions. Luciferase activity in the cell lysate was determined using a TD-2020 luminometer (Turner Designs, Sunnyvale, CA, USA). Expression of the Nos2 promoter is reported as relative light units (RLU)/μg protein of the stimulated cells expressed as percent RLU/μg protein of the unstimulated control for the same cells. Protein values were determined by the bicinchoninic acid method (Smith, et al., 1985).
4.3 Electrophoretic Mobility Shift Assay
The assay was run according to the protocol described in the Gel Shift Assay Systems kit (Promega, Madison, WI), with minor modifications as described previously (Sanchez, et al., 2003). Briefly, an aliquot of nuclear extract containing 8 μg protein was incubated for 20 min at room temperature in binding buffer [10 mM Tris-HCl, pH 7.5, 50 mM NaCl, 0.5 mM EDTA, 0.5 mM DTT, 4% glycerol, and 0.05 mg/ml poly(dI-dC)· poly(dI-dC)] in a final volume of 10 μl. A 1μl aliquot of a 32P end-labeled double-stranded oligonucleotide probe (labeled with [32P] γ-ATP using T4 polynucleotide kinase per the manufacturer’s instructions) was added and the mixture incubated for 20 min at room temperature. The incubation mixture was then separated by electrophoresis under non-denaturing conditions on a native 4% acrylamide gel (1 h at 250 V; in 0.5x TBE buffer). A polyclonal anti Oct-1 antibody (was included in the incubation mixture The gel was processed and the image obtained using a Molecular Dynamics model 445 SI phosphorimager and ImageQuaNT version 5.2 software. Table 1 shows the probe sequences used for the gel shift analysis and their relative positions in the core Nos2 promoter (Figure 1).
4.4 In vivo DNA Footprinting
C6 cells for experimental use were grown to near confluency as described previously (Sanchez, et al., 2003) and then stimulated as discussed above (see 4.2) in the absence or presence of 150 mM ethanol. After the specified length of incubation, cells were washed with pre-warmed (37°C) PBS. DNA extraction was done following the protocols of Hornstra and Yang (1993).
DNA methlylation in vivo and in vitro was carried out following the protocols by Goldring et al. (1996). Briefly, in vivo methylation was carried out at 20°C for 1 min by adding 0.5% DMS (Sigma-Aldrich, Saint Louis, MO, USA) in PBS to intact cells and stopped by adding 4 ml of 2% β-mercaptoethanol in ice-cold PBS. The cells were then washed with the same solution and DNA was isolated. In vitro methylation of naked genomic DNA in 200 μl TE buffer was carried out using 0.5% DMS for 1 min at 20°C. The reaction was stopped by addition of 50 μl 1.5 M NaCl and 3.4% β-mercaptoethanol. DNA was ethanol-precipitated, washed with 75% ethanol and resuspended in 200 μl nuclease-free water. Cleavage reaction was carried out by adding 10% piperidine and incubation at 90°C for 30 min. DNA was then cooled to room temperature, precipitated using ethanol and sodium acetate, washed with 75% ethanol and resuspended in TE buffer.
Ligation-mediated PCR was carried out following the method of Garrity and Wold (1992) with the protocols optimized for the RapidCycler (Idaho Technology, ID) as described (Sanchez, et al., 2003). The primers used to analyze both strands of the promoter region, the annealing temperature and positions relative to the transcriptional start site, and the conditions for electrophoretic separation of the DNA fragments were those described previously (Sanchez, et al., 2003).
4.5 Statistical Analyses
Prism version 4.0 software (GraphPad Inc., San Diego, CA) was used for graphic presentation and statistical analysis. Data are expressed as percent control and presented as mean activity + S.E.M. of at least three independent experiments. Analyses performed included paired t-test for comparisons within a construct, and one-way ANOVA followed by the Bonferroni multiple comparison test for comparisons between promoter constructs. A probability (P) of < 0.05 was accepted as demonstrating statistically significant differences between groups. Specific analysis is described for each experiment in the respective figure legend.
Acknowledgments
The α-Oct-1 antibody was a kind gift from Dr. Winship Herr, Cold Spring Harbor Laboratory, New York, USA. This work was supported by a School of Medicine Research Seed Grant and NIH grant AA11953 to PJS, and NRSA AA13221 to RLD.
Abbreviations
- LPS
lipopolysaccharide
- EMSA
electrophoretic mobility shift assay
- iNOS
nitric-oxide synthase-2
- Oct
octamer
- IFN
interferon
- IL
interleukin
- CM
cytokine mixture
- PMA
phorbol 12-myristate 13-acetate
- ESR
ethanol sensitive region
- DMS
dimethyl sulfate
- TNF
tumor necrosis factor
- RLU
relative light units
- PBS
phosphate buffered saline
- TE
Tris-EDTA
- TBE
Tris-Borate-EDTA
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Adamson DC, Wildemann B, Sasaki M, Glass JD, McArthur JC, Christov VI, Dawson TM, Dawson VL. Immunologic NO synthase: elevation in severe AIDS dementia and induction by HIV-1 gp41. Science. 1996;274:1917–1921. doi: 10.1126/science.274.5294.1917. [DOI] [PubMed] [Google Scholar]
- Bagasra O, Michaels FH, Zheng YM, Bobroski LE, Spitsin SV, Fu ZF, Tawadros R, Koprowski H. Activation of the inducible form of nitric oxide synthase in the brains of patients with multiple sclerosis. Proc Natl Acad Sci USA. 1995;92:12041–12045. doi: 10.1073/pnas.92.26.12041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belsham DD, Mellon PL. Transcription factors Oct-1 and C/EBPβ (CCAAT/enhancer-binding protein-beta) are involved in the glutamate/nitric oxide/cyclic-guanosine 5′-monophosphate-mediated repression of mediated repression of gonadotropin-releasing hormone gene expression. Mol Endocrinol. 2000;14:212–228. doi: 10.1210/mend.14.2.0418. [DOI] [PubMed] [Google Scholar]
- Bendall AJ, Sturm RA, Danoy PA, Molloy PL. Broad binding-site specificity and affinity properties of octamer 1 and brain octamer-binding proteins. Eur J Biochem. 1993;217:799–811. doi: 10.1111/j.1432-1033.1993.tb18308.x. [DOI] [PubMed] [Google Scholar]
- Bhat NR, Feinstein DL, Shen Q, Bhat AN. p38 MAPK-mediated transcriptional activation of inducible nitric-oxide synthase in glial cells. Roles of nuclear factors, nuclear factor κB, cAMP response element-binding protein, CCAAT/enhancer-binding protein-β, and activating transcription factor-2. J Biol Chem. 2002;277:29584–29592. doi: 10.1074/jbc.M204994200. [DOI] [PubMed] [Google Scholar]
- Brosnan CF, Lee SC, Liu J. Regulation of inducible nitric oxide synthase expression in human glia: implications for inflammatory central nervous system diseases. Biochem Soc Trans. 1997;25:679–683. doi: 10.1042/bst0250679. [DOI] [PubMed] [Google Scholar]
- Cartwright IL, Kelly SE. Probing the nature of chromosomal DNA-protein contacts by in vivo footprinting. Biotrechniques. 1991;11:188–203. [PubMed] [Google Scholar]
- Cheng CK, Yeung CM, Hoo RL, Chow BK, Leung PC. Oct-1 is involved in the transcriptional repression of the gonadotropin-releasing hormone receptor gene. Endocrinol. 2002;143:4693–4701. doi: 10.1210/en.2002-220576. [DOI] [PubMed] [Google Scholar]
- Cheng KW, Chow BK, Leung PC. Functional mapping of a placenta-specific upstream promoter for human gonadotropin-releasing hormone receptor gene. Endocrinol. 2001;142:1506–1516. doi: 10.1210/endo.142.4.8104. [DOI] [PubMed] [Google Scholar]
- Darville MI, Terryn S, Eizirik DL. An Octamer Motif Is Required for Activation of the Inducible Nitric Oxide Synthase Promoter in Pancreatic β-Cells. Endocrinol. 2004;145:1130–1136. doi: 10.1210/en.2003-1200. [DOI] [PubMed] [Google Scholar]
- Davis RL, Syapin PJ. Interactions of alcohol and nitric-oxide synthase in the brain. Brain Res Brain Res Rev. 2005;49:494–504. doi: 10.1016/j.brainresrev.2005.01.008. [DOI] [PubMed] [Google Scholar]
- Eberhardt W, Kunz D, Hummel R, Pfeilschifter J. Molecular cloning of the rat inducible nitric oxide synthase gene promoter. Biochem Biophys Res Commun. 1996;223:752–756. doi: 10.1006/bbrc.1996.0968. [DOI] [PubMed] [Google Scholar]
- Fan L, Bellinger F, Ge YL, Wilce P. Genetic study of alcoholism and novel gene expression in the alcoholic brain. Addict Biol. 2004;9:11–18. doi: 10.1080/13556210410001674040. [DOI] [PubMed] [Google Scholar]
- Feinstein DL, Galea E, Aquino DA, Li GC, Xu H, Reis DJ. Heat shock protein 70 suppresses astroglial-inducible nitric-oxide synthase expression by decreasing NFκB activation. J Biol Chem. 1996;271:17724–17732. doi: 10.1074/jbc.271.30.17724. [DOI] [PubMed] [Google Scholar]
- Garrity PA, Wold BJ. Effects of different DNA polymerases in ligation-mediated PCR: enhanced genomic sequencing and in vivo footprinting. Proc Natl Acad Sci USA. 1992;89:1021–1025. doi: 10.1073/pnas.89.3.1021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gavrilyuk V, Horvath P, Weinberg G, Feinstein DL. A 27-bp region of the inducible nitric oxide synthase promoter regulates expression in glial cells. J Neurochem. 2001;78:129–140. doi: 10.1046/j.1471-4159.2001.00375.x. [DOI] [PubMed] [Google Scholar]
- Gay RD, Dawson SJ, Murphy WJ, Russell SW, Latchman DS. Activation of the iNOS gene promoter by Brn-3 POU family transcription factors is dependent upon the octamer motif in the promoter. Biochim Biophys Acta. 1998;1443:315–322. doi: 10.1016/s0167-4781(98)00234-6. [DOI] [PubMed] [Google Scholar]
- Goldring CE, Reveneau S, Algarte M, Jeannin JF. In vivo footprinting of the mouse inducible nitric oxide synthase gene: inducible protein occupation of numerous sites including Oct and NF-IL6. Nucleic Acids Res. 1996;24:1682–1687. doi: 10.1093/nar/24.9.1682. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Herr W, Cleary MA. The POU domain: versatility in transcriptional regulation by a flexible two-in-one DNA-binding domain. Genes Dev. 1995;9:1679–1693. doi: 10.1101/gad.9.14.1679. [DOI] [PubMed] [Google Scholar]
- Hornstra IK, Yang TP. In vivo footprinting and genomic sequencing by ligation-mediated PCR. Anal Biochem. 1993;213:179–193. doi: 10.1006/abio.1993.1407. [DOI] [PubMed] [Google Scholar]
- Johnson RA, Noll EC, Rodney WM. Survival after a serum ethanol concentration of 1 1/2% Lancet. 1982;2:1394. doi: 10.1016/s0140-6736(82)91285-5. [DOI] [PubMed] [Google Scholar]
- Kambe F, Tsukahara S, Kato T, Seo H. The POU-domain protein Oct-1 is widely expressed in adult rat organs. Biochim Biophys Acta. 1993;1171:307–310. doi: 10.1016/0167-4781(93)90071-k. [DOI] [PubMed] [Google Scholar]
- Kamijo R, Harada H, Matsuyama T, Bosland M, Gerecitano J, Shapiro D, Le J, Koh SI, Kimura T, Green SJ, Mak TW, Taniguchi T, Vilcek J. Requirement for transcription factor IRF-1 in NO synthase induction in macrophages. Science. 1994;263:1612–1615. doi: 10.1126/science.7510419. [DOI] [PubMed] [Google Scholar]
- Kim YM, Ko CB, Park YP, Kim YJ, Paik SG. Octamer motif is required for the NF-κB-mediated induction of the inducible nitric oxide synthase gene expression in RAW 264.7 macrophages. Mol Cells. 1999;9:99–109. [PubMed] [Google Scholar]
- Lee H, Jeong J, Son E, Mosa A, Cho GJ, Choi WS, Ha JH, Kim IK, Lee MG, Kim CY, Suk K. Ethanol selectively modulates inflammatory activation signaling of brain microglia. J Neuroimmunol. 2004;156:88–95. doi: 10.1016/j.jneuroim.2004.07.008. [DOI] [PubMed] [Google Scholar]
- Liu J, Lewohl JM, Harris RA, Iyer VR, Dodd PR, Randall PK, Mayfield RD. Patterns of gene expression in the frontal cortex discriminate alcoholic from nonalcoholic individuals. Neuropsychopharmacol. 2006;31:1574–1582. doi: 10.1038/sj.npp.1300947. [DOI] [PubMed] [Google Scholar]
- Lowenstein CJ, Alley EW, Raval P, Snowman AM, Snyder SH, Russell SW, Murphy WJ. Macrophage nitric oxide synthase gene: two upstream regions mediate induction by interferon gamma and lipopolysaccharide. Proc Natl Acad Sci USA. 1993;90:9730–9734. doi: 10.1073/pnas.90.20.9730. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Martin E, Nathan C, Xie QW. Role of interferon regulatory factor 1 in induction of nitric oxide synthase. J Exp Med. 1994;180:977–984. doi: 10.1084/jem.180.3.977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meda L, Baron P, Prat E, Scarpini E, Scarlato G, Cassatella MA, Rossi F. Proinflammatory profile of cytokine production by human monocytes and murine microglia stimulated with beta-amyloid[25–35] J Neuroimmunol. 1999;93:45–52. doi: 10.1016/s0165-5728(98)00188-x. [DOI] [PubMed] [Google Scholar]
- Merrill JE, Murphy SP, Mitrovic B, Mackenzie-Graham A, Dopp JC, Ding MZ, Griscavage J, Ignarro LJ, Lowenstein CJ. Inducible nitric oxide synthase and nitric oxide production by oligodendrocytes. J Neurosci Res. 1997;48:372–384. [PubMed] [Google Scholar]
- Militante JD, Feinstein DL, Syapin PJ. Suppression by ethanol of inducible nitric oxide synthase expression in C6 glioma cells. J Pharmacol Exp Ther. 1997;281:558–565. [PubMed] [Google Scholar]
- Mucke L, Eddleston M. Astrocytes in infectious and immune-mediated diseases of the central nervous system. FASEB J. 1993;7:1226–1232. doi: 10.1096/fasebj.7.13.8405808. [DOI] [PubMed] [Google Scholar]
- Mueller PR, Wold B. In vivo footprinting of a muscle specific enhancer by ligation mediated PCR. Science. 1989;246:780–786. doi: 10.1126/science.2814500. [DOI] [PubMed] [Google Scholar]
- Murphy S, Grzybicki D. Glial NO: Normal and pathological roles. Neuroscientist. 1996;2:90–99. [Google Scholar]
- Murphy S, Simmons ML, Agullo L, Garcia A, Feinstein DL, Galea E, Reis DJ, Minc-Golomb D, Schwartz JP. Synthesis of nitric oxide in CNS glial cells. Trends Neurosci. 1993;16:323–328. doi: 10.1016/0166-2236(93)90109-y. [DOI] [PubMed] [Google Scholar]
- Nixon K, Crews FT. Binge ethanol exposure decreases neurogenesis in adult rat hippocampus. J Neurochem. 2002;83:1087–1093. doi: 10.1046/j.1471-4159.2002.01214.x. [DOI] [PubMed] [Google Scholar]
- Perry VH, Newman TA, Cunningham C. The impact of systemic infection on the progression of neurodegenerative disease. Nat Rev Neurosci. 2003;4:103–112. doi: 10.1038/nrn1032. [DOI] [PubMed] [Google Scholar]
- Sanchez AC, Davis RL, Syapin PJ. Identification of cis-regulatory regions necessary for robust NOS2 promoter activity in glial cells: Indirect role for NF-κB. J Neurochem. 2003;86:1379–1390. doi: 10.1046/j.1471-4159.2003.01943.x. [DOI] [PubMed] [Google Scholar]
- Schreiber E, Harshman K, Kemler I, Malipiero U, Schaffner W, Fontana A. Astrocytes and glioblastoma cells express novel octamer-DNA binding proteins distinct from the ubiquitous Oct-1 and B cell type Oct-2 proteins. Nuc Acids Res. 1990;18:5495–5503. doi: 10.1093/nar/18.18.5495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sherman MP, Aeberhard EE, Wong VZ, Griscavage JM, Ignarro LJ. Pyrrolidine dithiocarbamate inhibits induction of nitric oxide synthase activity in rat alveolar macrophages. Biochem Biophys Res Commun. 1993;191:1301–1308. doi: 10.1006/bbrc.1993.1359. [DOI] [PubMed] [Google Scholar]
- Smith PK, Krohn RI, Hermanson GT, Mallia AK, Gartner FH, Provenzano MD, Fujimoto EK, Goeke NM, Olson BJ, Klenk DC. Measurement of protein using bicinchoninic acid. Anal Biochem. 1985;150:76–85. doi: 10.1016/0003-2697(85)90442-7. [DOI] [PubMed] [Google Scholar]
- Syapin PJ, Militante JD, Garrett DK, Ren L. Cytokine-induced iNOS expression in C6 glial cells: transcriptional inhibition by ethanol. J Pharmacol Exp Ther. 2001;298:744–752. [PubMed] [Google Scholar]
- Urso T, Gavaler JS, Van Thiel DH. Blood ethanol levels in sober alcohol users seen in an emergency room. Life Sci. 1981;28:1053–1056. doi: 10.1016/0024-3205(81)90752-9. [DOI] [PubMed] [Google Scholar]
- Verrijzer CP, van Oosterhout JA, van der Vliet PC. The Oct-1 POU domain mediates interactions between Oct-1 and other POU proteins. Mol Cell Biol. 1992;12:542–551. doi: 10.1128/mcb.12.2.542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walker S, Hayes S, O’Hare P. Site-specific conformational alteration of the Oct-1 POU domain-DNA complex as the basis for differential recognition by Vmw65 (VP16) Cell. 1994;79:841–852. doi: 10.1016/0092-8674(94)90073-6. [DOI] [PubMed] [Google Scholar]
- Wang JH, Sun GY. Ethanol inhibits cytokine-induced iNOS and sPLA2 in immortalized astrocytes: evidence for posttranscriptional site of ethanol action. J Biomed Sci. 2001;8:126–133. doi: 10.1007/BF02255981. [DOI] [PubMed] [Google Scholar]
- Wang JY, Shum AY, Hwang CP. Ethanol modulates induction of nitric oxide synthase in glial cells by endotoxin. Life Sci. 1998;63:1571–1583. doi: 10.1016/s0024-3205(98)00424-x. [DOI] [PubMed] [Google Scholar]
- Warshawsky D, Miller L. Mapping protein-DNA interactions using in vivo footprinting. Methods Mol Biol. 1999;127:199–212. doi: 10.1385/1-59259-678-9:199. [DOI] [PubMed] [Google Scholar]
- Wells DJ, Jr, Barnhill MT., Jr Unusually high ethanol levels in two emergency medicine patients. J Anal Toxicol. 1996;20:272. doi: 10.1093/jat/20.4.272. [DOI] [PubMed] [Google Scholar]
- Xie Q. A novel lipopolysaccharide-response element contributes to induction of nitric oxide synthase. J Biol Chem. 1997;272:14867–14872. doi: 10.1074/jbc.272.23.14867. [DOI] [PubMed] [Google Scholar]
- Xie QW, Kashiwabara Y, Nathan C. Role of transcription factor NF-κB/Rel in induction of nitric oxide synthase. J Biol Chem. 1994;269:4705–4708. [PubMed] [Google Scholar]
- Xie QW, Whisnant R, Nathan C. Promoter of the mouse gene encoding calcium-independent nitric oxide synthase confers inducibility by interferon γ and bacterial lipopolysaccharide. J Exp Med. 1993;177:1779–1784. doi: 10.1084/jem.177.6.1779. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhang H, Chen X, Teng X, Snead C, Catravas JD. Molecular cloning and analysis of the rat inducible nitric oxide synthase gene promoter in aortic smooth muscle cells. Biochem Pharmacol. 1998;55:1873–1880. doi: 10.1016/s0006-2952(98)00078-1. [DOI] [PubMed] [Google Scholar]