Abstract
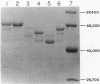
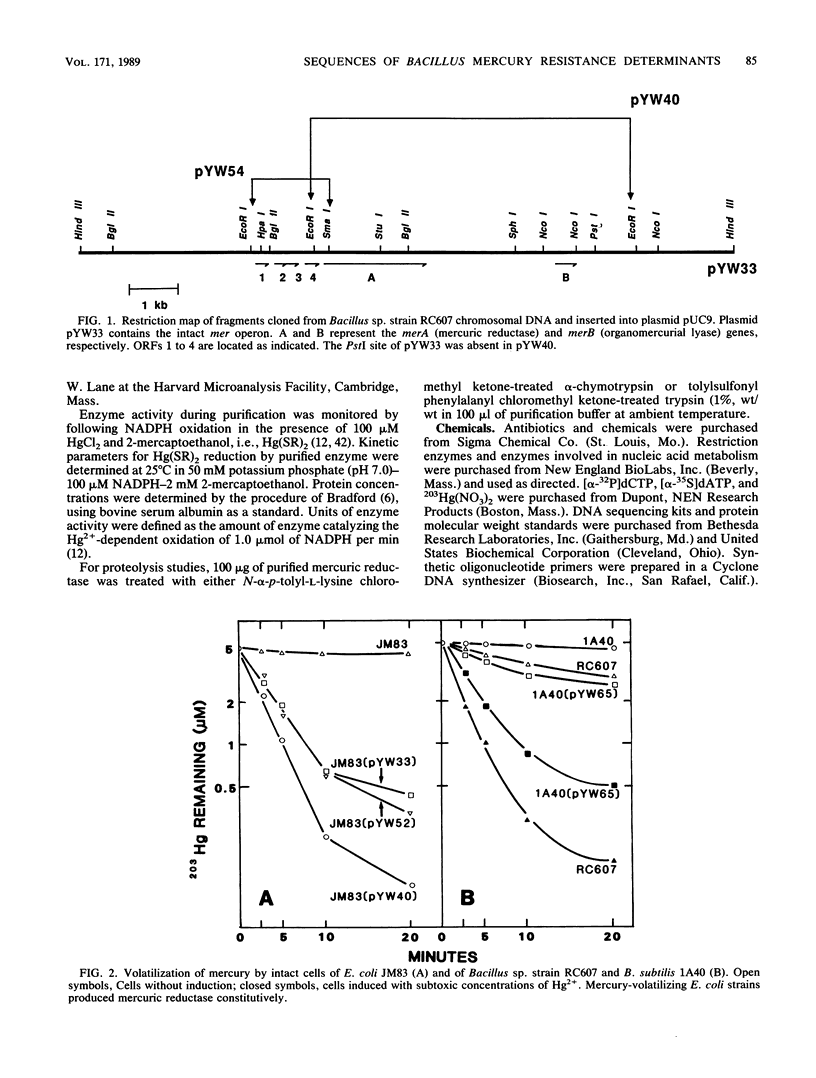
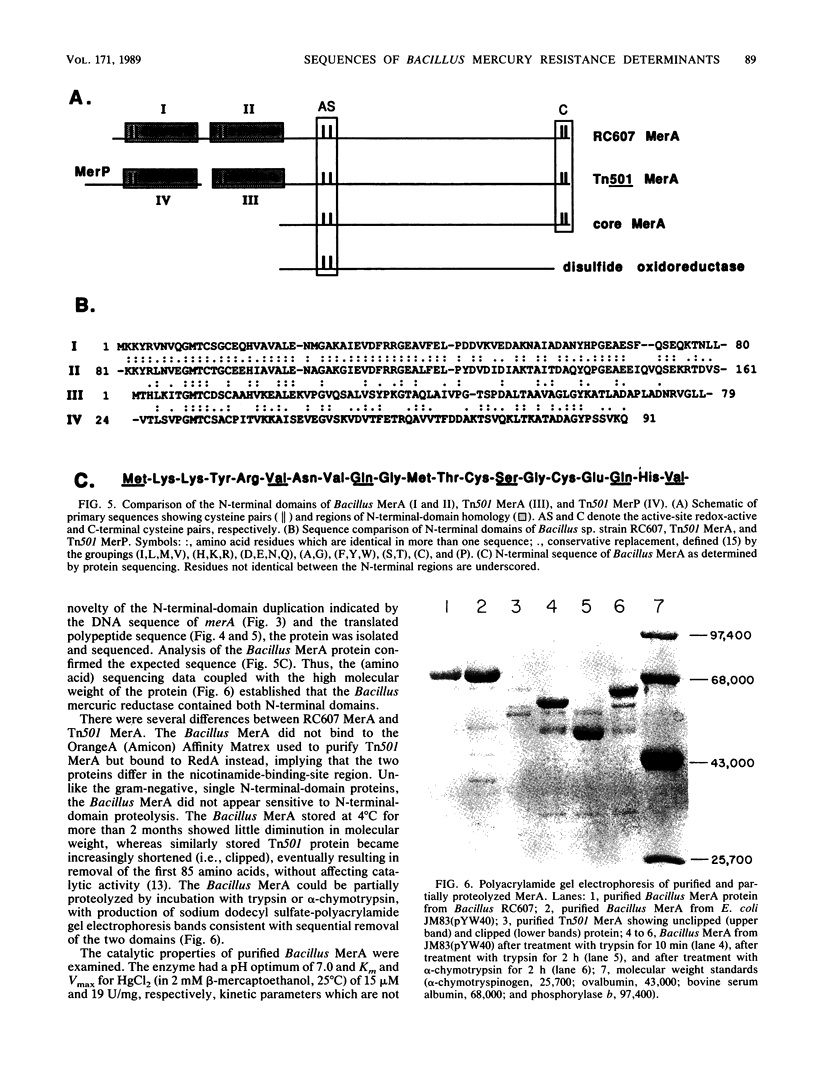
A 13.5-kilobase HindIII fragment, bearing an intact mercury resistance (mer) operon, was isolated from chromosomal DNA of broad-spectrum mercury-resistant Bacillus sp. strain RC607 by using as a probe a clone containing the mercury reductase (merA) gene. The new clone, pYW33, expressed broad-spectrum mercury resistance both in Escherichia coli and in Bacillus subtilis, but only in B. subtilis was the mercuric reductase activity inducible. Sequencing of a 1.8-kilobase mercury hypersensitivity-producing fragment revealed four open reading frames (ORFs). ORF1 may code for a regulatory protein (MerR). ORF2 and ORF4 were associated with cellular transport function and the hypersensitivity phenotype. DNA fragments encompassing the merA and the merB genes were sequenced. The predicted Bacillus sp. strain RC607 MerA (mercuric reductase) and MerB (organomercurial lyase) were similar to those predicted from Staphylococcus aureus plasmid pI258 (67 and 73% amino acid identities, respectively); however, only 40% of the amino acid residues of RC607 MerA were identical to those of the mercuric reductase from gram-negative bacteria. A 69-kilodalton polypeptide was isolated and identified as the merA gene product by examination of its amino-terminal sequence.
Full text
PDF







Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Barkay T., Fouts D. L., Olson B. H. Preparation of a DNA gene probe for detection of mercury resistance genes in gram-negative bacterial communities. Appl Environ Microbiol. 1985 Mar;49(3):686–692. doi: 10.1128/aem.49.3.686-692.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barrineau P., Gilbert P., Jackson W. J., Jones C. S., Summers A. O., Wisdom S. The DNA sequence of the mercury resistance operon of the IncFII plasmid NR1. J Mol Appl Genet. 1984;2(6):601–619. [PubMed] [Google Scholar]
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bott K. F., Wilson G. A. Development of competence in the Bacillus subtilis transformation system. J Bacteriol. 1967 Sep;94(3):562–570. doi: 10.1128/jb.94.3.562-570.1967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bradford M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976 May 7;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Brown N. L., Misra T. K., Winnie J. N., Schmidt A., Seiff M., Silver S. The nucleotide sequence of the mercuric resistance operons of plasmid R100 and transposon Tn501: further evidence for mer genes which enhance the activity of the mercuric ion detoxification system. Mol Gen Genet. 1986 Jan;202(1):143–151. doi: 10.1007/BF00330531. [DOI] [PubMed] [Google Scholar]
- Chen E. Y., Seeburg P. H. Supercoil sequencing: a fast and simple method for sequencing plasmid DNA. DNA. 1985 Apr;4(2):165–170. doi: 10.1089/dna.1985.4.165. [DOI] [PubMed] [Google Scholar]
- Feng D. F., Doolittle R. F. Progressive sequence alignment as a prerequisite to correct phylogenetic trees. J Mol Evol. 1987;25(4):351–360. doi: 10.1007/BF02603120. [DOI] [PubMed] [Google Scholar]
- Foster T. J. The genetics and biochemistry of mercury resistance. Crit Rev Microbiol. 1987;15(2):117–140. doi: 10.3109/10408418709104455. [DOI] [PubMed] [Google Scholar]
- Fox B. S., Walsh C. T. Mercuric reductase: homology to glutathione reductase and lipoamide dehydrogenase. Iodoacetamide alkylation and sequence of the active site peptide. Biochemistry. 1983 Aug 16;22(17):4082–4088. doi: 10.1021/bi00286a014. [DOI] [PubMed] [Google Scholar]
- Fox B., Walsh C. T. Mercuric reductase. Purification and characterization of a transposon-encoded flavoprotein containing an oxidation-reduction-active disulfide. J Biol Chem. 1982 Mar 10;257(5):2498–2503. [PubMed] [Google Scholar]
- Griffin H. G., Foster T. J., Silver S., Misra T. K. Cloning and DNA sequence of the mercuric- and organomercurial-resistance determinants of plasmid pDU1358. Proc Natl Acad Sci U S A. 1987 May;84(10):3112–3116. doi: 10.1073/pnas.84.10.3112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Helmann J. D., Chamberlin M. J. Structure and function of bacterial sigma factors. Annu Rev Biochem. 1988;57:839–872. doi: 10.1146/annurev.bi.57.070188.004203. [DOI] [PubMed] [Google Scholar]
- Laddaga R. A., Chu L., Misra T. K., Silver S. Nucleotide sequence and expression of the mercurial-resistance operon from Staphylococcus aureus plasmid pI258. Proc Natl Acad Sci U S A. 1987 Aug;84(15):5106–5110. doi: 10.1073/pnas.84.15.5106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lund P. A., Brown N. L. Role of the merT and merP gene products of transposon Tn501 in the induction and expression of resistance to mercuric ions. Gene. 1987;52(2-3):207–214. doi: 10.1016/0378-1119(87)90047-3. [DOI] [PubMed] [Google Scholar]
- Mahler I., Levinson H. S., Wang Y., Halvorson H. O. Cadmium- and mercury-resistant Bacillus strains from a salt marsh and from Boston Harbor. Appl Environ Microbiol. 1986 Dec;52(6):1293–1298. doi: 10.1128/aem.52.6.1293-1298.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsudaira P. Sequence from picomole quantities of proteins electroblotted onto polyvinylidene difluoride membranes. J Biol Chem. 1987 Jul 25;262(21):10035–10038. [PubMed] [Google Scholar]
- Misra T. K., Brown N. L., Fritzinger D. C., Pridmore R. D., Barnes W. M., Haberstroh L., Silver S. Mercuric ion-resistance operons of plasmid R100 and transposon Tn501: the beginning of the operon including the regulatory region and the first two structural genes. Proc Natl Acad Sci U S A. 1984 Oct;81(19):5975–5979. doi: 10.1073/pnas.81.19.5975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Misra T. K., Brown N. L., Haberstroh L., Schmidt A., Goddette D., Silver S. Mercuric reductase structural genes from plasmid R100 and transposon Tn501: functional domains of the enzyme. Gene. 1985;34(2-3):253–262. doi: 10.1016/0378-1119(85)90134-9. [DOI] [PubMed] [Google Scholar]
- Nakahara H., Silver S., Miki T., Rownd R. H. Hypersensitivity to Hg2+ and hyperbinding activity associated with cloned fragments of the mercurial resistance operon of plasmid NR1. J Bacteriol. 1979 Oct;140(1):161–166. doi: 10.1128/jb.140.1.161-166.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Halloran T., Walsh C. Metalloregulatory DNA-binding protein encoded by the merR gene: isolation and characterization. Science. 1987 Jan 9;235(4785):211–214. doi: 10.1126/science.3798107. [DOI] [PubMed] [Google Scholar]
- Ogawa H. I., Tolle C. L., Summers A. O. Physical and genetic map of the organomercury resistance (Omr) and inorganic mercury resistance (Hgr) loci of the IncM plasmid R831b. Gene. 1984 Dec;32(3):311–320. doi: 10.1016/0378-1119(84)90006-4. [DOI] [PubMed] [Google Scholar]
- Perlman D., Halvorson H. O. A putative signal peptidase recognition site and sequence in eukaryotic and prokaryotic signal peptides. J Mol Biol. 1983 Jun 25;167(2):391–409. doi: 10.1016/s0022-2836(83)80341-6. [DOI] [PubMed] [Google Scholar]
- Rigby P. W., Dieckmann M., Rhodes C., Berg P. Labeling deoxyribonucleic acid to high specific activity in vitro by nick translation with DNA polymerase I. J Mol Biol. 1977 Jun 15;113(1):237–251. doi: 10.1016/0022-2836(77)90052-3. [DOI] [PubMed] [Google Scholar]
- Rinderle S. J., Booth J. E., Williams J. W. Mercuric reductase from R-plasmid NR1: characterization and mechanistic study. Biochemistry. 1983 Feb 15;22(4):869–876. doi: 10.1021/bi00273a025. [DOI] [PubMed] [Google Scholar]
- Rosenberg M., Court D. Regulatory sequences involved in the promotion and termination of RNA transcription. Annu Rev Genet. 1979;13:319–353. doi: 10.1146/annurev.ge.13.120179.001535. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schottel J. L. The mercuric and organomercurial detoxifying enzymes from a plasmid-bearing strain of Escherichia coli. J Biol Chem. 1978 Jun 25;253(12):4341–4349. [PubMed] [Google Scholar]
- Schultz P. G., Au K. G., Walsh C. T. Directed mutagenesis of the redox-active disulfide in the flavoenzyme mercuric ion reductase. Biochemistry. 1985 Nov 19;24(24):6840–6848. doi: 10.1021/bi00345a016. [DOI] [PubMed] [Google Scholar]
- Silver S., Misra T. K. Plasmid-mediated heavy metal resistances. Annu Rev Microbiol. 1988;42:717–743. doi: 10.1146/annurev.mi.42.100188.003441. [DOI] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Sullivan M. A., Yasbin R. E., Young F. E. New shuttle vectors for Bacillus subtilis and Escherichia coli which allow rapid detection of inserted fragments. Gene. 1984 Jul-Aug;29(1-2):21–26. doi: 10.1016/0378-1119(84)90161-6. [DOI] [PubMed] [Google Scholar]
- Summers A. O. Organization, expression, and evolution of genes for mercury resistance. Annu Rev Microbiol. 1986;40:607–634. doi: 10.1146/annurev.mi.40.100186.003135. [DOI] [PubMed] [Google Scholar]
- Tabor S., Richardson C. C. DNA sequence analysis with a modified bacteriophage T7 DNA polymerase. Proc Natl Acad Sci U S A. 1987 Jul;84(14):4767–4771. doi: 10.1073/pnas.84.14.4767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tanaka T., Kuroda M., Sakaguchi K. Isolation and characterization of four plasmids from Bacillus subtilis. J Bacteriol. 1977 Mar;129(3):1487–1494. doi: 10.1128/jb.129.3.1487-1494.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vieira J., Messing J. The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene. 1982 Oct;19(3):259–268. doi: 10.1016/0378-1119(82)90015-4. [DOI] [PubMed] [Google Scholar]
- Wang Y., Mahler I., Levinson H. S., Halvorson H. O. Cloning and expression in Escherichia coli of chromosomal mercury resistance genes from a Bacillus sp. J Bacteriol. 1987 Oct;169(10):4848–4851. doi: 10.1128/jb.169.10.4848-4851.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss A. A., Murphy S. D., Silver S. Mercury and organomercurial resistances determined by plasmids in Staphylococcus aureus. J Bacteriol. 1977 Oct;132(1):197–208. doi: 10.1128/jb.132.1.197-208.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]
- von Heijne G. Signal sequences. The limits of variation. J Mol Biol. 1985 Jul 5;184(1):99–105. doi: 10.1016/0022-2836(85)90046-4. [DOI] [PubMed] [Google Scholar]