Abstract
The disassembly of promoter nucleosomes appears to be a general property of highly transcribed eukaryotic genes. We have previously shown that the disassembly of chromatin from the promoters of the Saccharomyces cerevisiae PHO5 and PHO8 genes, mediated by the histone chaperone anti-silencing function 1 (Asf1), is essential for transcriptional activation upon phosphate depletion. This mechanism of transcriptional regulation is shared with the ADY2 and ADH2 genes upon glucose removal. Promoter chromatin disassembly by Asf1 is required for recruitment of TBP and RNA polymerase II, but not the Pho4 and Pho2 activators. Furthermore, accumulation of SWI/SNF and SAGA at the PHO5 promoter requires promoter chromatin disassembly. By contrast, the requirement for SWI/SNF and SAGA to facilitate Pho4 activator recruitment to the nucleosome-buried binding site in the PHO5 promoter occurs prior to chromatin disassembly and is distinct from the stable recruitment of SWI/SNF and SAGA that occurs after chromatin disassembly.
The packaging of the eukaryotic genome into chromatin is essential for normal growth, development, and differentiation. The basic repeating unit of chromatin, the nucleosome, comprises 147 bp of DNA wrapped around a core histone octamer containing two molecules each of histones H2A, H2B, H3, and H4 (55). A growing body of data indicates that chromatin is a highly dynamic structure that tightly regulates all nuclear processes that use DNA as a substrate, including transcription, DNA replication, repair, and recombination.
Chromatin structure plays a fundamental role in the establishment and maintenance of transcriptional programs. This occurs at many different levels, including ATP-dependent chromatin remodeling, histone modifications, and chromatin disassembly during gene expression (32, 59). Nucleosome disassembly is arguably the most profound manner in which the cell can modify the chromatin structure. Transcriptional activation is accompanied by the disassembly of nucleosomes from promoter regions during transcriptional induction (8, 11, 42, 46, 62). This is likely to be a general characteristic of eukaryotic gene activation, because global analyses of histone occupancy in Saccharomyces cerevisiae, Drosophila, and humans demonstrate that transcribing promoters tend to be depleted of nucleosomes compared to inactive promoters (6, 7, 28, 33, 40).
Chromatin disassembly from promoter regions was first observed at the S. cerevisiae PHO5 gene (2, 8, 9, 26, 42). The PHO5 gene encodes an acid phosphatase that is required for phosphate production and is tightly regulated in response to the intracellular phosphate concentration (43, 52). There are two regulatory elements in the PHO5 promoter, designated upstream activating sequences p1 and p2 (UASp1 and UASp2), which correspond to the major binding sites for the activators Pho2 and Pho4. Under high-phosphate conditions, Pho4 is highly phosphorylated by the Pho80-Pho85 complex, which causes Pho4 to be exported to the cytoplasm (22, 39). When phosphate is low, a signaling cascade activates Pho81 to inhibit the kinase activity of Pho80-Pho85 (38, 44). Upon inhibition of the Pho80-Pho85 kinase, Pho4 becomes localized to the nucleus, where it interacts with another activator, Pho2, and together they bind the PHO5 promoter (3, 39). The PHO5 promoter has two chromatin conformations. Under repressing (high-phosphate) conditions, it is assembled into four-positioned nucleosomes (nuc −4 to −1) that contain UASp2 and the TATA box (49). Upon activation (in response to phosphate depletion), these four-positioned nucleosomes are disassembled (9). The histone acetyltransferase (HAT) complex SAGA and the ATP-dependent chromatin remodeler SWI/SNF contribute to activation of PHO5 (4, 5) in a manner that is dependent on the phosphate concentration. The low level of nuclear Pho4 that occurs under low-phosphate conditions absolutely requires SWI/SNF and SAGA for its ability to bind to the PHO5 promoter, while the high level of nuclear Pho4 that occurs under phosphate-free conditions bypasses the need for SWI/SNF and SAGA to promote Pho4 binding to the PHO5 promoter (12).
Chromatin disassembly is mediated by histone binding proteins termed “histone chaperones.” Histone removal from the yeast PHO5 promoter is mediated by the histone H3/H4 chaperone anti-silencing function 1 (Asf1) in response to phosphate depletion (2). We previously showed that Asf1-mediated promoter chromatin disassembly is essential for activation of the yeast PHO5 and PHO8 genes following phosphate depletion (2), but the molecular reason for this was unclear. In the current work, we examined how promoter chromatin influences the recruitment of the activators, coactivators, and general transcription machinery to the PHO5 promoter. We found that while promoter chromatin disassembly is not a requirement for Pho4 and Pho2 activator recruitment, it is essential for the recruitment of the general transcription machinery. Furthermore, this work uncovered a stepwise process of events: nucleosome remodeling and histone modification enable activator binding, followed by nucleosome destabilization and histone removal by Asf1. This in turn is followed by stable recruitment of SWI/SNF and SAGA to the nucleosome-depleted promoter and recruitment of the general transcription machinery.
MATERIALS AND METHODS
Yeast strains and media.
Yeast cultures and genetic manipulations were performed following standard methods (1). The genotypes of all strains used in this study are described in Table 1. Yeast strains were grown to a density of approximately 1 × 107 cells/ml in yeast extract-peptone-dextrose (YPD) for all analyses. Phosphate-depleted medium (−Pi) was prepared as follows. For 200 ml of −Pi solution, 13.2 g Bacto Peptone and 6.8 g yeast extract were dissolved in 200 ml distilled H2O (dH2O). Once dissolved, 3 g MgSO4·7H2O was added, and the pH was adjusted to 8.0 with concentrated NH4OH. This solution was placed on ice for ∼30 min to precipitate the phosphate. After 30 min, the suspension was centrifuged at 10,000 × g at 4°C for 10 min, and the supernatant was carefully poured into new bottles. The supernatant was centrifuged again at 10,000 × g at 4°C for 10 min and then filtered through Whatman's paper and adjusted to pH 7 with 2 M HCl. For 1 liter of medium, 76 ml of −Pi solution, 6 g (NH4)2SO4, 0.5 g MgSO4·7H2O, 0.326 g CaCl2·2H2O, 20 mg adenine, and 20 mg uracil were combined, the volume was adjusted to 1 liter with dH2O, and the mixture was dispensed into bottles and autoclaved. After the autoclaving, the appropriate glucose was added. As a standard, we determined the phosphate content of our phosphate-depleted medium using the Malachite Green Phosphate Assay kits from Bioassay Systems and consistently achieved depletion to close to 0.1 mM. For repression of PHO5, phosphate was added to a 13.4 mM final concentration.
TABLE 1.
Genotypes of yeast strains used in these studies
| Name | Background | Genotype | Source or reference |
|---|---|---|---|
| JKT0010 | W303 | MATahis3-11 leu2-3,112 lys2 trp1-1 ura3-1 bar1::LEU2 | 2 |
| JKT0018 | W303 | MATahis3-11 leu2-3,112 lys2 trp1-1 ura3-1 bar1::LEU2 asf1::his5+ | 2 |
| LKY069 | W303 | MATaade2 can1-100 his3-11.15 leu2-13,112 trp1-1 ura3-1 3HA-TBP URA3 | 27 |
| MAY0063 | W303 | MATaade2 can1-100 his3-11,15 leu2-13,112 trp1-1 ura3-1 3HA-TBP asf1::KAN | This study |
| MAY0068 | W303 | MATahis3-11 leu2-3,112 lys2 trp1-1 ura3-1 bar1::LEU2 PHO4-HA-TRP | This study |
| MAY0074 | W303 | MATahis3-11 leu2-3,112 lys2 trp1-1 ura3-1 bar1::LEU2 asf1::his5+ PHO4-HA-TRP | This study |
| Z1466 | W303 | MATaade2-1 trp1-1 can1-100 leu2-3,112 his3-11,15 ura3 GAL+psi+ GCN5-MYC9-TRP1 | 53 |
| JCY0011 | W303 | MATaade2-1 trp1-1 can1-100 leu2-3,112 his3-11,15 ura3 GAL+psi+ GCN5-MYC9-TRP1 asf1::KAN | This study |
| BY4741 | S288C | MATahis3D1 leu2Δ met15Δ ura3Δ | ResGena |
| BY4741pho4Δ | S288C | MATahis3D1 leu2Δ met15Δ ura3Δ pho4::KanMX4 | ResGen |
| MAY1000 | S288C | MATahis3D1 leu2Δ met15Δ ura3Δ pho4::KanMX4 (pPho4-VP16) | This study |
| MAY1001 | S288C | MATahis3D1 leu2Δ met15Δ ura3Δ pho4::KanMX4 asf1::his5+ (pPho4-VP16) | This study |
| BY4741spt3Δ | S288C | MATahis3D1 leu2Δ met15Δ ura3Δ spt3::kanMX4 | ResGen |
| BY4741gcn5Δ | S288C | MATahis3D1 leu2Δ met15Δ ura3Δ gcn5::kanMX4 | ResGen |
| BY4741spt7Δ | S288C | MATahis3D1 leu2Δ met15Δ ura3Δ spt7::kanMX4 | ResGen |
| BY4741snf2Δ | S288C | MATahis3D1 leu2Δ met15Δ ura3Δ snf2::kanMX4 | ResGen |
| BY4741asf1Δ | S288C | MATahis3D1 leu2Δ met15Δ ura3Δ asf1::kanMX4 | ResGen |
| JLY098 | S288C | MATahis3D1 leu2Δ met15Δ ura3Δ PHO4-HA-HIS-TRP | This study |
| SKW015 | S288C | MATahis3D1 leu2Δ met15Δ ura3Δ snf2::kanMX4 PHO4-HA-HIS-TRP | This study |
| SKW025 | S288C | MATahis3D1 leu2Δ met15Δ ura3Δ gcn5::kanMX4 PHO4-HA-HIS-TRP | This study |
ResGen, Research Genetics genome-wide deletion strains.
Endogenous PHO4 and RPB3 were tagged on the C terminus with hemagglutinin (HA) using PCR-based gene modification (29). For PHO5 experiments, strains were grown up to a density of approximately 1 × 107 cells/ml in phosphate-depleted YPD containing phosphate to a 0.0134 M final concentration from a stock of 1.34 M KH2PO4 (i.e., PHO5-repressing conditions). To induce PHO5, the cells were then shifted to phosphate-depleted medium as previously described (17). For ADH2 and ADY2 analyses, strains were grown in YPD-2% glucose (final concentration) for repressing conditions and then shifted to 0.05% glucose medium for activating conditions for 4 hours. The VP16-Pho4 plasmid was previously described (20).
Chromatin immunoprecipitation (ChIP) analysis.
Cells (1.25 × 108) were treated with 1% formaldehyde (final concentration) for 20 min at room temperature. Cross-linking was quenched by the addition of glycine to a final concentration of 125 mM. The cells were sedimented and washed twice in ice-cold TBS (150 mM NaCl, 20 mM Tris-HCl, pH 7.6). The cells were resuspended in 400 μl lysis buffer (0.1% deoxycholate [DOC], 1 mM EDTA, 50 mM HEPES, pH 7.5, 140 mM NaCl, 1% Triton X-100), an equal volume of 0.5-mm glass beads was added, and the cells were vortexed for 10 min at 4°C. The lysate was removed from the beads and placed into a new tube. The beads were rinsed with an additional 400 μl of lysis buffer, which was collected as well. Chromatin was sheared with a Branson 350 Sonifier to an average size of 500 bp. Following the shearing, samples were centrifuged for 15 min at 4°C at 13,000 rpm. Then, the lysates (∼750 μl) were transferred to a new Eppendorf tube; 50 μl of each lysate was moved to a new tube for input, and 750 μl was used for each immunoprecipitation (IP). IPs were performed using the following antibodies: (i) for histone H3, histone H3 antibody (Abcam no. ab-1791) at 2.5 μl per IP (this antibody recognizes the C terminus and therefore is not affected by the modification state of the histones, which are found predominantly in the N terminus); (ii) for HA-tagged proteins, HA antibody (Covance no. MMS-101R) at 4 μl per IP; (iii) for Pho2, anti-Pho2 antibody at 1 μl per IP; (iv) for Snf2, anti-Snf2 antibody at 2 μl per IP (kindly provided by Joseph Reese, Penn State University); and (v) for MYC-tagged proteins, MYC antibody (Covance no. MMS-164P) at 4 μl per IP. The lysates were incubated overnight at 4°C with rocking on a nutator. After overnight incubation, a 50/50 (wt/vol) slurry of protein A-Sepharose was added and incubated for 1 h at 4°C with rocking. The samples were centrifuged at 7,500 rpm and washed with 1,000 μl of each of the following: (i) lysis buffer (0.1% DOC, 1 mM EDTA, 50 mM HEPES, pH 7.5, 140 mM NaCl, 1% Triton X-100), (ii) lysis 500 buffer (0.1% DOC, 1 mM EDTA, 50 mM HEPES, pH 7.5, 500 mM NaCl, 1% Triton X-100), (iii) LiCl detergent (0.5% DOC, 1 mM EDTA, 250 mM LiCl, 0.5% NP-50, 10 mM Tris, pH 7.9), and (iv) TBS (150 mM NaCl, 20 mM Tris, pH 7.5); 100 μl of 1% sodium dodecyl sulfate (SDS)-1× Tris-EDTA (TE) was added after the final wash and incubated at 65°C for 10 min. Samples were centrifuged at 13,000 rpm for 15 seconds, and the eluates were removed from the beads with a gel-loading tip and transferred to a new Eppendorf tube; 150 μl of 0.67% SDS-1× TE was added to the beads, vortexed, and centrifuged at 13,000 rpm for 15 seconds. The remaining eluate was removed from the beads and added to the Eppendorf tube containing the previous eluate. To reverse cross-linking, samples were incubated for at least 8 h at 65°C. Following reversal of the cross-linking, DNA was deproteinized by the addition of 250 μl of proteinase K solution (2 μg proteinase K, 0.4 μg glycogen, 1× TE) and incubated at 37°C for 2 h, followed by phenol-chloroform extraction (equal volumes of phenol-chloroform and 55 μl of 4 M LiCl) and ethanol precipitation. DNA was resuspended in 50 μl TE.
The data in Fig. 1 to 4, with the exception of Fig. 2E, 3E, and 4D, were generated by standard PCR analysis and agarose gel quantification as follows. For PCR analysis, the dilution of each sample was determined empirically; generally, IP samples were diluted 1:5 in double-distilled H2O (ddH2O), and input samples were diluted 1:20 in ddH2O. PCR cycles included 95°C for 5 min, followed by 24 cycles of 95°C for 30 seconds, 45°C for 30 seconds, and 72°C for 1 min. The linear range of templates for multiplex PCR was determined empirically, and PCR-amplified products were quantitatively measured using Labworks (UVP Inc.). The PCR products were resolved on a 3% agarose gel (1.5% agarose/1.5% low-melt Nusieve agarose) and visualized with ethidium bromide. The amount of immunoprecipitated DNA was determined by quantitative PCR with primer pairs directed against the indicated regions. As a control, primer sets were used for a region proximal to the right telomere of chromosome 6 (TELVIR). Relative occupancy was determined by the ratio of IP experimental product relative to the IP control product divided by the ratio of input experimental product relative to the input control product. The values shown are averages of three independent experiments, and error bars indicate the 95% confidence interval. Primers used for all ChIP PCR experiments are available upon request.
FIG. 1.
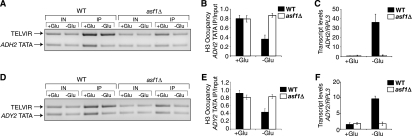
Asf1 mediates nucleosome disassembly at the ADH2 and ADY2 promoters. (A) ChIP analysis of histone H3 levels over the ADH2 TATA region. Histone H3 levels were measured by ChIP analysis in wild-type (WT) (JKT0010) and asf1Δ (JKT0018) strains growing in 2% glucose-containing media (+Glu) and 0.05% glucose-containing media (−Glu). As a control, primer sets were also used for the TELVIR region. (B) Quantitation of histone H3 ChIP analyses at the ADH2 TATA. (C) Quantitation of ADH2 transcript levels. Shown is quantitative RT-PCR analysis of ADH2 transcript levels in wild-type and asf1Δ strains grown as described for panel A. (D) ChIP analysis of histone levels over the ADY2 TATA region as described for panel A. (E) Quantitation of histone H3 ChIP as described for panel B, with primers directed against the TATA box region of ADY2. (F) Quantitation of ADY2 transcript levels as described for panel C. All histograms shown present the mean and standard deviation of three or more independent experiments.
FIG. 4.
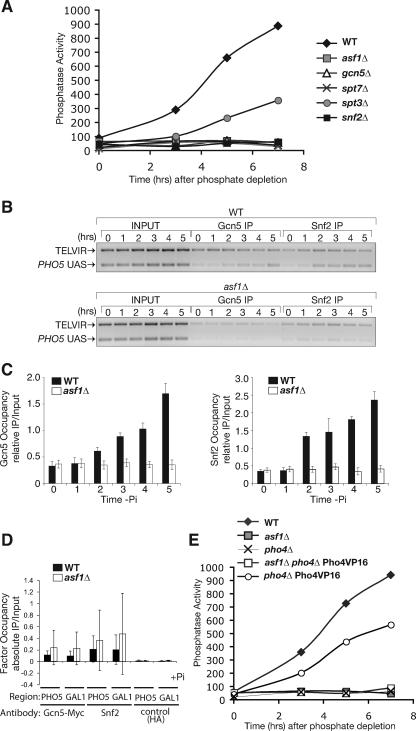
Asf1-mediated nucleosome disassembly is required for stable Gcn5 and Snf2 recruitment. (A) SWI/SNF and SAGA are essential for PHO5 activation following phosphate depletion. Activation of the PHO5 gene was measured by acid phosphatase activity determinations in strain BY4741 (wild type [WT]) and the isogenic asf1Δ, gcn5Δ, spt7Δ, spt3Δ, and snf2Δ versions of BY4741 at the indicated times after transfer to phosphate-depleted media. (B) ChIP analysis of Gcn5/MYC and Snf2 levels at the PHO5 promoter. Gcn5 and Snf2 levels were measured by ChIP analysis in WT (Z1466) and asf1Δ (JCY0011) strains growing in phosphate-containing media and then shifted to a phosphate-depleted medium for the indicated times. (C) Quantitation of ChIP analysis of Gcn5/MYC and Snf2 levels over the PHO5 UAS region, normalized to input and the control region as described in the legend to Fig. 2. (D) ChIP analyses using the same strains as for panels B and C at the PHO5 UAS and the GAL1 UAS under repressing conditions (+Pi, +Glu) but including an unrelated antibody (HA) as a control. The large error bars reflect the extremely low signal that is essentially measuring background and is at the limit of quantitative detection for the real-time PCR analysis. (E) Asf1-mediated chromatin disassembly is required for activation by Pho4-VP16. Acid phosphatase levels were measured in strains BY4741 (WT), BY4741asf1Δ (asf1Δ), BY4741pho4Δ (pho4Δ), MAY1000 (pho4Δ Pho4VP16), and MAY10001 (asf1Δ pho4Δ Pho4VP16) at the indicated times after transfer to phosphate-depleted media to provide a measurement of PHO5 induction. All histograms shown present the mean and standard deviation of three independent experiments.
FIG. 2.
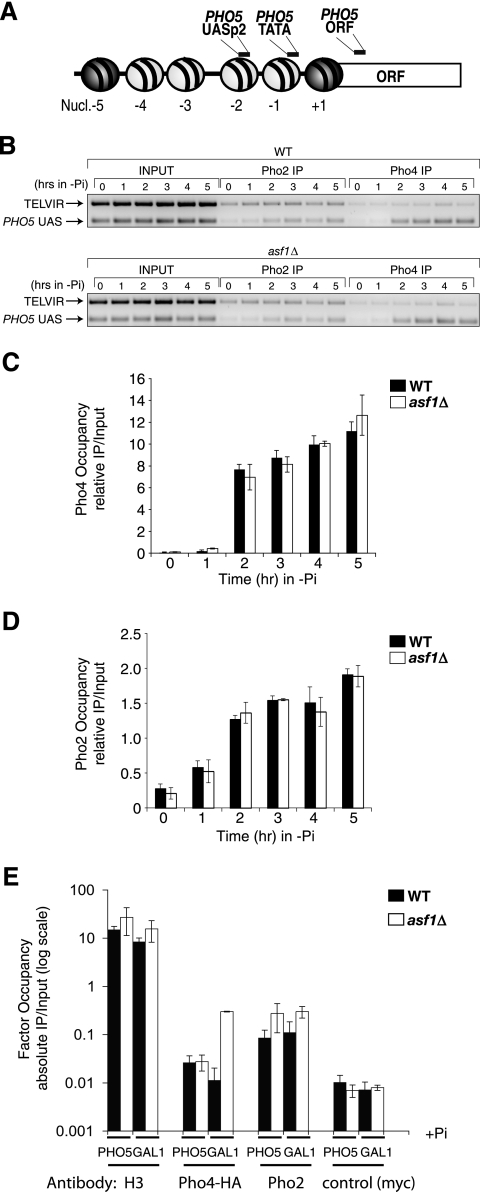
Pho2 and Pho4 recruitment is not impaired in asf1 mutants. (A) Schematic of the PHO5 promoter region. The light gray circles represent nucleosomes that are disassembled upon transcriptional activation. Indicated are the locations of regions that are PCR amplified in the ChIP analyses. ORF, open reading frame. (B) ChIP analyses were performed in wild-type (WT) (MAY0068) and asf1Δ (MAY0074) strains growing in phosphate-containing media and then shifted to a phosphate-depleted medium for the indicated times. (C) Quantitation of ChIP analysis of Pho4 levels over the PHO5 UAS region, using Pho4:HA, normalized to the input and relative to the TELVIR control region. (D) Quantitation of ChIP analysis of Pho2 levels over the PHO5 UAS region, normalized as described above. (E) ChIP analyses using the same strains as panels C and D at the PHO5 UAS and the GAL1 UAS under repressing conditions (+Pi, +Glu), but including an unrelated antibody (myc) as a control. Note that the y axis is a log scale to allow comparison of the ChIP antibodies to the control antibody. All histograms shown present the mean and standard deviation of three independent experiments.
FIG. 3.
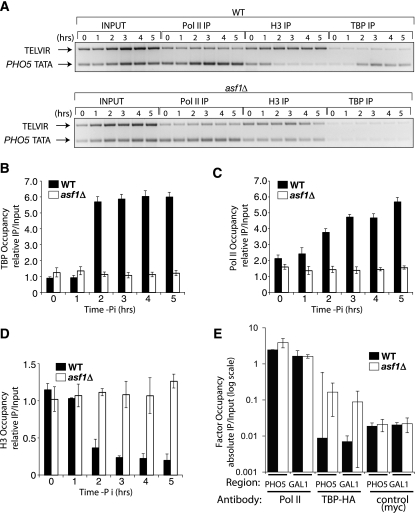
Asf1-mediated nucleosome disassembly is required for TBP and Pol II recruitment to the PHO5 promoter. (A) ChIP analysis of Pol II, histone H3, and TBP levels over the PHO5 promoter, measured by ChIP analysis in wild-type (WT) (LKY0069) and asf1Δ (MAY0063) strains growing in phosphate-containing media and then shifted to phosphate-depleted media for the indicated times. As a control, primer sets were used for a region in TELVIR. (B to D) Quantitation of ChIP analysis of TBP recruitment (B), Pol II recruitment (C), and histone H3 disassembly (D) over the PHO5 TATA region, normalized to input and control regions as described in the legend to Fig. 2. (E) ChIP analyses using the same strains as in panels B to D at the PHO5 TATA and GAL1 TATA under repressing conditions (+Pi, +Glu) but including an unrelated antibody (myc) as a control. Note that the y axis is a log scale to allow comparison of the ChIP antibodies to the control antibody. All histograms shown present the mean and standard deviation of three independent experiments.
The data in Fig. 2E, 3E, 4D, and 5 were generated by real-time PCR analysis as follows. Analysis was performed using a Roche Applied Sciences Light Cycler 480. The linear range of PCR templates was determined by performing a twofold serial dilution to find where the threshold cycle changed by one cycle. Using a dilution lying in the middle of this linear range, which was typically 1:10, each sample was analyzed in triplicate using a total volume of 10-μl reaction mixtures in a 384-well format. The thermal profile was always as follows: denaturation at 95°C for 10 min, amplification with 50 cycles of 95°C for 15 seconds and then 60°C for 1 min, followed by cooling at 40°C for 10 min. All probes (30 nucleotides in length or less) used either a 6-carboxyfluorescein or VIC 5′ label and a nonfluorescent quencher on the 3′ end. Each IP sample was normalized to its respective input sample (to account for the number of cells), as well as a control region that was known to be unaffected by phosphate levels (the GAL1 promoter or the TELVIR region, to account for nonspecific background signal). Primers and TaqMan probe sequences will be provided upon request.
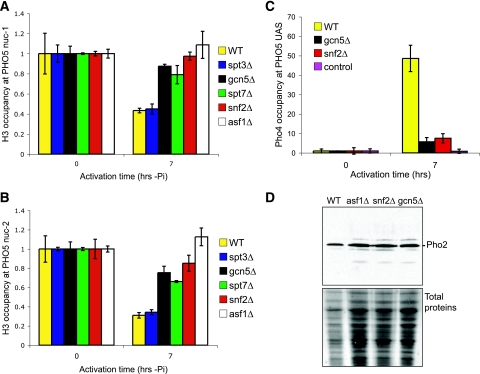
FIG. 5.
SWI/SNF and SAGA are required for chromatin disassembly to allow Pho4 recruitment prior to their stable recruitment. (A) Chromatin disassembly from nucleosome −1 of the PHO5 promoter in SAGA and SWI/SNF mutants. Histone occupancy at nuc −1 was determined by H3 ChIP analysis in strain BY4741 (wild type [WT]) and the isogenic asf1Δ, gcn5Δ, spt7Δ, spt3Δ, and snf2Δ versions of BY4741 at the indicated times after transfer to phosphate-depleted media. The data were normalized as described in the legend to Fig. 2. (B) Chromatin disassembly from nucleosome −2 of the PHO5 promoter in SAGA and SWI/SNF mutants, performed as described for panel A. (C) SWI/SNF and SAGA contribute to Pho4 recruitment to the PHO5 promoter. Pho4 occupancy at the PHO5 UAS was determined by ChIP analysis in strain JLY098 (WT) and isogenic SKW0015 (gcn5Δ) and SKW0025 (snf2Δ) strains at the indicated times after transfer to phosphate-depleted media. An unrelated antibody (myc) was used as a control. The data were normalized as described in the legend to Fig. 2. (D) Pho2 levels are not affected by deletion of SNF2 or GCN5. Western blot analysis in strain BY4741 (WT) and the isogenic gcn5Δ and snf2Δ versions of BY4741. Amido black was included as a loading control. All histograms shown present the mean and standard deviation of three independent experiments.
All histograms shown present the mean and standard deviation of three independent experiments.
Acid phosphatase activity assays.
Cells were assayed for phosphatase activity as described previously (35). Briefly, 5 ml of 1 × 107 cells/ml were spun down and washed twice with cold 0.1 M sodium acetate, pH 3.6. The cells were resuspended in 500 μl of 0.1 M sodium acetate, pH 3.6. To determine the number of cells used for each reaction, 20 to 100 μl of cell lysate was diluted 1:10 in ddH2O, and the optical density at 600 nm (OD600) was read. For experimental samples, 20 μl of cell lysate (−Pi) or 100 μl of cell lysate (with phosphate [+Pi]) was added to 480 μl or 400 μl of 0.1 M sodium acetate, pH 3.6. For a control, 500 μl of 0.1 M sodium acetate, pH 3.6, was used. Samples were incubated at 30°C for 10 min with an appropriate volume (500 μl for each reaction) of freshly made nitrophenyl phosphate (0.0742 g/10 ml 0.1 M sodium acetate, pH 3.6). Reactions were stopped by adding 250 μl 1 M Na2CO3. Samples were centrifuged at 13,000 rpm for 2 min, and 1 ml of supernatant was removed to a cuvette. The OD was read at 420 nm and calculated as phosphatase activity as follows: (OD420 × 1,000)/(OD600 × volume cell lysate used[μl] × minutes incubated).
Reverse transcriptase PCR (RT-PCR) analysis.
To extract the RNA, 10 ml of cells (2 × 107 cells/ml) was harvested by centrifugation and quickly frozen in liquid N2. The cells were thawed briefly on ice and moved to a microcentrifuge tube. Acid phenol (700 μl; 1:1 phenol, pH 5.2) and 550 μl of 50 mM sodium acetate, pH 5.2-10 mM EDTA-1% (final concentration) SDS was added to the samples and vortexed for 15 seconds. The samples were incubated at 65°C for 10 min with vortexing every minute for 10 seconds. Following incubation, the samples were placed on ice for 15 min. The samples were centrifuged for 3 min at 14,000 rpm, and 600 μl of the aqueous layer was transferred to a new tube containing 600 μl of phenol-chloroform-isoamyl alcohol (25:24:1) and incubated on a nutator for 5 min at room temperature. Samples were centrifuged for 3 min at 14,000 rpm, and 500 μl of the aqueous layer was transferred to a new tube containing 500 μl of chloroform-isoamyl alcohol and incubated on a nutator for 5 min at room temperature. Samples were then centrifuged for 3 min at 14,000 rpm, and 400 μl of the aqueous layer was transferred to a new tube containing 40 μl of 3 M sodium acetate. One milliliter of ethanol was added, and the sample was inverted to mix it and incubated at −20°C for 1 h, followed by centrifugation at 16,000 rpm for 20 min at 4°C. The supernatant was removed with a filter tip and discarded, followed by air drying the pellets for 10 min. The samples were resuspended in 100 μl of ddH2O with 1:1,000 diethyl pyrocarbonate.
First-strand synthesis was performed with gene-specific 3′ primers and Stratagene RTase according to Stratagene's protocol. The subsequent PCR used 2 μl of the cDNA reaction mixture and included a deoxynucleoside triphosphate mixture containing [α32p]dATP. The linear amplification range was empirically determined for each control primer set. RPL3 was used as a control for an RNA transcript whose copy number does not change as a consequence of changing phosphate levels. PCR products were resolved on a 3% agarose gel. The dried gel was exposed to a phosphor screen, and the image was quantitated using ImageQuant. The average with 95% confidence interval for transcript levels, normalized to levels of the RPL3 control, from four independent RT-PCR analyses is shown for each experiment. Primers for RT-PCRs are available upon request.
RESULTS
Asf1-mediated nucleosome disassembly is required for activation of the ADH2 and ADY2 genes.
We have previously shown that Asf1-mediated disassembly of promoter nucleosomes is essential for transcriptional activation of the yeast PHO5 and PHO8 genes during phosphate depletion (2). In order to determine whether Asf1-mediated chromatin disassembly was specific to the promoters of phosphate-regulated genes, we examined the glucose-regulated ADH2 and ADY2 genes. The nucleosome presiding over the TATA box region in the ADH2 promoter is remodeled during activation in low-glucose media (56). Using ChIP analysis with an antibody to the C terminus of histone H3 that is not influenced by histone modifications (2), we observed that nucleosome occupancy in the ADH2 TATA box region is reduced approximately twofold during transcriptional activation (Fig. 1A and B). However, in yeast with ASF1 deleted (asf1Δ), histone levels in the ADH2 TATA box region did not decrease under activating conditions (minus Glu) (Fig. 1A and B). Furthermore, when we examined ADH2 transcript levels in low-glucose medium, it was apparent that the asf1 mutant did not activate ADH2 transcription (Fig. 1C). Similar to the ADH2 promoter, the ADY2 promoter has a nucleosome positioned over the TATA box that is remodeled upon activation in low-glucose medium (14, 56). We found that activation of the ADY2 gene by glucose depletion was accompanied by an approximately twofold reduction in nucleosome occupancy over the TATA box region in the wild type, but not in the asf1Δ strain (Fig. 1D and E). In addition, we did not observe an increase in ADY2 transcription in asf1 mutants compared to the wild type upon the switch to activating conditions (Fig. 1F). These data demonstrate that Asf1 is also required for histone disassembly from the ADH2 and ADY2 promoters and that this Asf1-mediated chromatin disassembly is required for transcriptional activation of these genes.
Promoter chromatin disassembly is not required for recruitment of the Pho4 and Pho2 activators to the PHO5 promoter.
In order to establish why promoter chromatin disassembly is essential for transcriptional activation, we examined factor recruitment in wild-type and asf1Δ cells at the well-characterized PHO5 promoter (Fig. 2A). Asf1-mediated chromatin disassembly is required for activation of the PHO5 promoter upon phosphate depletion (2). We previously demonstrated by in vivo dimethyl sulfate footprinting that the Pho4 activator could gain access to its UASp1 and UASp2 binding sites in the PHO5 promoter without chromatin disassembly in an asf1Δ strain (2). However, it was possible that the amount of Pho4 recruitment was reduced in an asf1 mutant, as in vivo dimethyl sulfate footprinting is not quantitative. Therefore, to obtain a quantitative measure of Pho4 recruitment to the PHO5 promoter, we performed ChIP analysis of Pho4 recruitment to the PHO5 promoter using an HA epitope-tagged Pho4 strain (Pho4:HA) (Fig. 2). The resolution of our ChIP analysis is not sufficient to distinguish between Pho4 binding to UASp2 and to the UASp1 site that is located in the adjacent linker region. However, the available evidence in vivo and in vitro suggests that Pho4 binding to the low-affinity UASp1 is approximately nine times weaker than to the high-affinity UASp2 site (37, 57). As such, the signal that we see is largely due to Pho4 recruitment to the UASp2 site. The signal obtained for Pho4 occupancy of the repressed PHO5 UAS was not significantly different from the very low signal obtained with a control antibody (Fig. 2E), indicating that negligible amounts of Pho4 were bound to the repressed PHO5 promoter. Upon depletion of phosphate from the media, we found recruitment of Pho4 to the PHO5 promoter in the wild-type and asf1Δ strain (Fig. 2B and C). Furthermore, the kinetics and extents of Pho4 recruitment to the PHO5 promoter were indistinguishable in wild-type and asf1Δ yeast (Fig. 2B and C and data not shown). These results clearly demonstrate that the Pho4 activator efficiently binds to the PHO5 promoter prior to Asf1-dependent chromatin disassembly.
Recruitment of the Pho2 activator is also required for activation of the PHO5 gene (3). Therefore, we investigated by ChIP analysis whether Asf1-mediated promoter chromatin disassembly is required for recruitment of Pho2 to the PHO5 promoter, utilizing an antibody to Pho2 (kindly provided by David Stillman, University of Utah). We found that the amounts of Pho2 recruitment were equivalent in wild-type and asf1Δ yeast (Fig. 2B and D). We noted that the signal obtained for Pho2 occupancy of the repressed PHO5 UAS was higher than that obtained with a control antibody (Fig. 2E). However, this signal was similar to the signal for Pho2 occupancy of the GAL1 promoter, which lacks Pho2 binding sites, and therefore represents nonspecific background signal. This background is about eightfold lower than the signal from the induced PHO5 promoter (Fig. 2D). These data show that Asf1-dependent promoter chromatin disassembly is not required for recruitment of Pho2 or Pho4 to the PHO5 promoter. Therefore, the failure to activate transcription in the absence of promoter chromatin disassembly (i.e., in the asf1 mutant) is due to the blockage of a step in transcription activation after activator recruitment.
Asf1-mediated promoter nucleosome disassembly is required for recruitment of the general transcription machinery.
To address why transcription is blocked by promoter chromatin even though the activators occupy the promoter, we examined the recruitment of the general transcription machinery. Specifically, we performed ChIP analysis using antibodies against the HA-tagged TATA-binding protein (TBP) and the carboxyl-terminal domain of RNA polymerase II (Pol II) at increasing times after phosphate depletion in wild-type and asf1 mutant strains. This analysis showed clear recruitment of TBP and Pol II to the PHO5 promoter following phosphate depletion in wild-type cells (Fig. 3A, B, and C). Moreover the timing of the appearance of TBP and Pol II at the PHO5 promoter correlated well with the timing of the loss of histones from the PHO5 promoter (Fig. 3A, B, C, and D). By contrast, no detectable recruitment of TBP or Pol II occurred in the asf1 mutant after phosphate depletion (Fig. 3A, B, and C). Figure 3E shows a ChIP analysis of Pol II and TBP under repressing conditions, including an unrelated antibody as a negative control. Due to the high signal from the Pol II antibody at both the repressed PHO5 and GAL1 promoters (Fig. 3E), we confirmed by Rpb3-HA ChIP analysis that Pol II does not localize to the PHO5 promoter in asf1 mutants (data not shown). This failure to recruit the general transcription machinery agrees well with the failure to activate transcription in the asf1 mutants (2). From these data, we conclude that promoter chromatin disassembly is required for the binding of the general transcription machinery.
Asf1-mediated chromatin disassembly is required for the stable recruitment of the chromatin modifiers SAGA and SWI/SNF.
To further examine the effect of promoter chromatin on transcriptional activation, we measured the recruitment of the coactivators SWI/SNF and SAGA to the PHO5 promoter. SWI/SNF and SAGA are essential for activation of the PHO5 promoter under phosphate-depleted conditions, as measured by the ability to detect Pho5 phosphatase activity (12) (Fig. 4A). Specifically, deletion of the gene encoding the Snf2 ATPase of the SWI/SNF complex, or the Gcn5 HAT or the Spt7 structural component of the SAGA complex (48), entirely blocked induction of PHO5 (Fig. 4A). By contrast, deletion of the gene encoding the Spt3 component of SAGA, which is known to recruit TBP to the PHO5 promoter (4), only reduced PHO5 induction. Next, we examined whether SWI/SNF and SAGA recruitment to the PHO5 promoter was blocked by promoter chromatin (in asf1 mutants following phosphate depletion). This was done by ChIP analysis using antibodies against the Myc-tagged Gcn5 subunit of SAGA and against Snf2. Although SWI/SNF and SAGA were clearly recruited to the PHO5 promoter following phosphate depletion in wild-type yeast, no recruitment was observed in the asf1Δ yeast (Fig. 4B and C). The amount of signal corresponding to SWI/SNF and SAGA occupying the repressed PHO5 promoter was not significantly different from the signal from a control antiserum (Fig. 4D). Taken together, these results indicate that Asf1-mediated nucleosome disassembly is required for the recruitment of the chromatin modifiers SAGA and SWI/SNF to the PHO5 promoter.
SAGA and SWI/SNF are known to be recruited to promoters via interactions with the activation domains of activators (18). Accordingly, SWI/SNF has been shown to bind directly to the activation domain of Pho4 (36). To investigate the mechanism whereby the Pho4/Pho2 activators that are bound to the PHO5 promoter fail to recruit coactivators in asf1 mutants, we asked whether this mechanism was specific to the Pho4 activation domain. The herpes simplex virus type 1 activation domain of VP16, when fused to the Pho4 activator, also drives promoter chromatin remodeling/chromatin disassembly and transcriptional activation of PHO5 (50). Therefore, we tested whether the ability of the VP16 activation domain to induce PHO5 transcription was also blocked by promoter chromatin. We found that, like the Pho4 and Gal4 activation domains (25), the VP16 activation domain also failed to induce PHO5 transcription in the presence of promoter chromatin (in asf1 mutants) (Fig. 4E). Thus, the mechanism whereby promoter chromatin blocks transcriptional activation is not specific to the Pho4 activation domain but is likely shared with at least the VP16 and Gal4 activation domains.
SAGA and SWI/SNF contribute to chromatin disassembly from the PHO5 promoter as an indirect consequence of their roles in Pho4 activator recruitment.
To further examine the requirement for SWI/SNF and SAGA in PHO5 activation following phosphate depletion (Fig. 4A), we examined their influence on promoter chromatin disassembly. We observed a significant reduction in disassembly of the −1 and −2 nucleosomes of the PHO5 promoter upon inactivation of SWI/SNF and SAGA by ChIP analysis with the H3 C-terminal antibody (Fig. 5A and B). The fact that we saw a greater degree of disassembly of nucleosome −2 than of −1 nucleosomes in the wild-type strain at 7 h, as had been seen previously in single-molecule analyses (21), indicates that we were getting at least some resolution of these two adjacent nuclesomes in our ChIP analysis. In addition, the kinetics of disassembly of nuc −1 versus nuc −2 were not differentially affected by mutation of SWI/SNF and SAGA (data not shown), indicating that SWI/SNF and SAGA contribute equally to disassembly of the two nucleosomes. Inactivation of the Spt3 subunit of SAGA that recruits TBP (4, 5) did not lead to a significant defect in promoter chromatin disassembly (Fig. 5A and B) during transcriptional activation, indicating that the role of SAGA in chromatin disassembly is likely to reflect a requirement for its role as a HAT rather than its role in TBP recruitment.
Chromatin disassembly at the PHO5 promoter is a consequence of Pho4 binding. Therefore, in order to distinguish between a direct and an indirect role for SWI/SNF and SAGA in promoter chromatin disassembly, we examined recruitment of the Pho4 activator in mutants of SWI/SNF and SAGA. A previous report had shown a requirement for SWI/SNF and SAGA for Pho4 binding under intermediate (0.05 to 0.25 mM) phosphate conditions (12). Accordingly, by ChIP analysis, we found that in contrast to the efficient recruitment of Pho4 that occurs in wild-type cells, deletion of GCN5 or SNF2 greatly reduced Pho4 binding to the PHO5 promoter upon phosphate depletion (Fig. 5C). We interpret this as failure to recruit Pho4 to the nucleosome-occluded UASp2 site for the reasons described above for Fig. 2. It is likely that the small increase in Pho4 binding over time in the snf2 and gcn5 mutants is due to Pho4 binding to the exposed UASp1 site. Importantly, the contributions of SWI/SNF and SAGA to Pho4 recruitment are not indirect consequences of reduced Pho2 or Pho4 protein levels upon deletion of GCN5 or SNF2 because the levels of these protein are unaffected by loss of Gcn5 or Snf2 (Fig. 5D) (12). Therefore, we conclude that the defect in promoter chromatin disassembly in phosphate-depleted media in the SWI/SNF and SAGA mutants is an indirect consequence of defective Pho4 recruitment to the PHO5 promoter upon inactivation of SWI/SNF and SAGA. Taken together, these data indicate that SWI/SNF and SAGA promote Pho4 activator binding to DNA in the absence of their detectable recruitment to the PHO5 promoter, because detectable recruitment of SWI/SNF and SAGA does not occur until after both Pho4 binding and chromatin disassembly have occurred (Fig. 6).
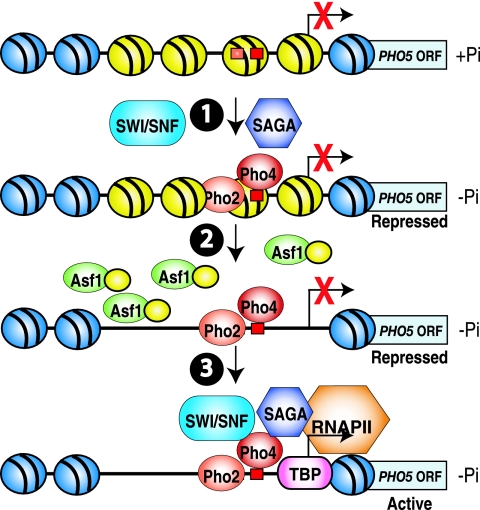
FIG. 6.
Stepwise model for transcriptional activation by chromatin disassembly. At the top is shown a schematic depicting the repressed PHO5 promoter. The binding sites for Pho4 and Pho2 are shown by red boxes. (Step 1) Upon phosphate depletion (activating conditions), the activators bind the DNA with the assistance of SWI/SNF and SAGA in wild-type strains, but in the absence of stable recruitment of SWI/SNF and SAGA to the promoter. (Step 2) Following activator recruitment, the promoter nucleosomes are disassembled by Asf1. (Step 3) Chromatin disassembly allows the recruitment of the general transcriptional machinery and stable recruitment of SAGA and SWI/SNF. However, upon phosphate depletion in asf1 mutants, nucleosome disassembly does not occur and SWI/SNF and/or SAGA can promote Pho4 binding to the promoter (step 1) but are unable to interact with the activator Pho4 to enable their recruitment to the PHO5 promoter (step 3).
DISCUSSION
This work establishes the generality of promoter-chromatin disassembly as a potent mechanism of transcriptional activation used by multiple activators and at multiple promoters. Using yeast PHO5 as our model system, we have discerned that Asf1-dependent promoter chromatin disassembly enables the recruitment of the general transcription machinery and subsequent transcriptional initiation. In addition, this work has separated the functional requirement of the SWI/SNF and SAGA coactivators from their detectable recruitment to chromatin.
Promoter chromatin disassembly as a global mechanism for transcriptional activation.
In addition to chromatin disassembly from the PHO5 promoter being essential for transcriptional activation (2), we now know that nucleosome disassembly from the PHO8, ADY2, and ADH2 promoters is also essential for transcriptional activation (2) (Fig. 1). Thus, promoter nucleosome disassembly is not specific to genes activated by Pho4, as the ADH2 and ADY2 promoters are activated by the Adr1 and Cat8 activators (58). Indeed, Asf1-mediated promoter chromatin disassembly and transcriptional activation are also driven by the VP16 (Fig. 4E) and Gal4 (25) activation domains. Accordingly, at least five different activators have the ability to trigger the promoter chromatin disassembly that is mediated via Asf1. It is intriguing to speculate that the many genes that are transcriptionally downregulated upon inactivation of Asf1 (61) may also be regulated by Asf1-mediated promoter chromatin disassembly. Consistent with previous results (45), our ChIP analyses of Asf1 localization found Asf1 to be located everywhere that we examined, including active and inactive promoters, Asf1-dependent and Asf1-independent promoters, open reading frames, and intergenic regions (data not shown). This is consistent with Asf1 being an opportunist, waiting on the chromatin for activators to initiate the cascade of chromatin changes that ultimately lead to loosening of histone-DNA contacts, which are required to enable Asf1 to bind to histones H3/H4 and remove them from the DNA (13). This function for Asf1 in chromatin disassembly-mediated gene activation is likely to be conserved in higher eukaryotes. For example, Asf1 is localized to the active interband regions of Drosophila polytene chromosomes (34, 54). In addition, Asf1 is associated with the transcription-specific histone variant H3.3 (51). Moreover, the genome-wide analyses of histone occupancy in yeast, Drosophila, and humans clearly demonstrates that highly transcribed genes are depleted of nucleosomes over the core promoter (6, 7, 28, 33, 40), suggesting that all eukaryotes may widely use promoter chromatin disassembly to activate transcription.
Transcriptional regulation through chromatin-regulated accessibility of the general transcription machinery.
We have found that the critical reason for chromatin disassembly from the promoter is to create a nucleosome-depleted core promoter-binding site for the general transcription machinery (Fig. 3). This is consistent with the previous studies that showed that TBP is unable to bind a nucleosome-occluded TATA box (15, 19, 60). The relatively low sequence specificity of the general transcription machinery is not sufficient to drive accurate transcriptional initiation from a genome-size DNA template, so that a naked genome would undoubtedly lead to transcription from cryptic initiation sites at inappropriate times and from inappropriate sequences, even in the absence of activators. Clearly, a key reason for packaging the genome into chromatin is to “hide” potential cryptic binding sites from the general transcription machinery. This is apparent from the cryptic transcriptional initiation that results from within open reading frames when chromatin is disrupted (10, 23, 31). Thus, it appears that the role of activators in generating a nucleosome-depleted DNA region by triggering nucleosome disassembly is critical to reveal a binding site for the general transcription machinery. This role of the activator inducing a nucleosome-free region indirectly dictates the specificity of transcriptional initiation.
Separable dual roles for coactivators at promoters.
Our results indicate that the roles of SWI/SNF and SAGA in promoting activator binding are distinct from the recruitment of remodelers by activators. This was shown by trapping a transcription intermediate in our asf1 mutant that still required SWI/SNF and SAGA to promote Pho4 binding to the nucleosome-buried PHO5 UASp2 site yet failed to stably recruit SWI/SNF or SAGA. In this way, we can order the events occurring during transcriptional activation as follows (Fig. 6): (i) chromatin remodeling and histone acetylation facilitate activator binding to nucleosome-buried DNA binding sites in the absence of detectable recruitment of SWI/SNF or SAGA (conversely, SWI/SNF and SAGA are not required for activator binding when the binding sites are located between nucleosomes, as is the case for PHO8) (16), (ii) chromatin disassembly from the promoter, (iii) stable recruitment of SWI/SNF and SAGA to the DNA-bound activator. The question that remains is whether SWI/SNF and SAGA contribute to PHO5 transcription following their recruitment to the nucleosome-depleted promoter by the DNA-bound activators. We know that the Spt3-dependent recruitment of TBP by SAGA contributes to transcription of some promoters, such as PHO5 (4), but it is unclear what function, if any, the chromatin-remodeling or HAT activities of SWI/SNF and SAGA (respectively) have on the nucleosome-depleted PHO5 promoter following their recruitment by activators. It is possible that SWI/SNF and SAGA recruitment to the nucleosome-depleted PHO5 promoter may promote chromatin remodeling within the adjacent open reading frame, as has been observed previously at HIS3 (24). Notably, our study demonstrates the danger of using coactivator recruitment (as measured by ChIP) as the sole measure of their function, as SWI/SNF and SAGA clearly function to enable activator recruitment in the absence of their detectable recruitment (in the asf1 mutants). It will be interesting to determine in the future whether the chromatin remodeling that results in activator recruitment to DNA requires interaction with the activators, for example, using mutants of SWI/SNF that remodel yet fail to interact with activators (41).
Our data indicate that Pho4 is unable to stably recruit the SWI/SNF and SAGA complexes to the PHO5 promoter in asf1 mutants, despite the localization of Pho4 to the PHO5 promoter. This result indicates that the promoter-bound nucleosomes are either directly or indirectly inhibitory to recruitment of SWI/SNF and SAGA. There are several possible scenarios that could explain this observation, but one possibility is that in the absence of nucleosome disassembly, the histones sterically hinder the ability of SWI/SNF and SAGA to bind to the activation domain of Pho4. It will be interesting to determine in the future if promoter chromatin is generally inhibitory to the stable recruitment of other activator-binding proteins and activators.
The current data indicate that SWI/SNF is not directly required as a nonredundant ATP-dependent remodeling activity during chromatin disassembly from the PHO5 promoter. Consistent with the idea that SWI/SNF is not involved in promoter chromatin disassembly, it has recently been shown that SWI/SNF and Asf1 work in independent pathways toward PHO5 activation (25). Modeling of our recently solved structure of Asf1 bound to histones H3/H4 (13) onto the nucleosome structure (30) revealed that histone H3/H4 removal from the DNA by Asf1 would occur only after H2A/H2B had been removed and the DNA had been at least partially unwound from the nucleosome, because they are otherwise physically in the way of the Asf1-histone interaction (13). Undoubtedly, therefore, the action of an ATP-dependent chromatin remodeler(s) and H2A/H2B chaperones is required prior to H3/H4 removal by Asf1 during chromatin disassembly. It will be interesting to test whether other ATP-dependent remodeling activities, such as the INO80 complex that is required for remodeling of the PHO5 promoter (47), assist Asf1 and an as-yet-unidentified H2A/H2B chaperone during promoter chromatin assembly.
Acknowledgments
We are grateful to Jason Feser for critical reading of the manuscript and to Joshua Carson for technical support. We thank David Bentley, Joseph Reese, and David Stillman for the antibodies to CTD, Snf2, and Pho2, respectively. We thank Colin Goding for the Pho4-VP16 plasmid.
This work was supported by a grant from the NIH to J.K.T. (GM64475). J.K.T. is a Scholar of the Leukemia and Lymphoma Society.
Footnotes
Published ahead of print on 9 July 2007.
REFERENCES
- 1.Adams, A. G., D. E. Kaiser, and C. A. T. Stearns. 1997. Methods in yeast genetics. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
- 2.Adkins, M. W., S. R. Howar, and J. K. Tyler. 2004. Chromatin disassembly mediated by the histone chaperone Asf1 is essential for transcriptional activation of the yeast PHO5 and PHO8 genes. Mol. Cell 14:657-666. [DOI] [PubMed] [Google Scholar]
- 3.Barbaric, S., M. Munsterkotter, J. Svaren, and W. Horz. 1996. The homeodomain protein Pho2 and the basic-helix-loop-helix protein Pho4 bind DNA cooperatively at the yeast PHO5 promoter. Nucleic Acids Res. 24:4479-4486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Barbaric, S., H. Reinke, and W. Horz. 2003. Multiple mechanistically distinct functions of SAGA at the PHO5 promoter. Mol. Cell. Biol. 23:3468-3476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Barbaric, S., J. Walker, A. Schmid, J. Q. Svejstrup, and W. Horz. 2001. Increasing the rate of chromatin remodeling and gene activation—a novel role for the histone acetyltransferase Gcn5. EMBO J. 20:4944-4951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Barrera, L. O., and B. Ren. 2006. The transcriptional regulatory code of eukaryotic cells—insights from genome-wide analysis of chromatin organization and transcription factor binding. Curr. Opin. Cell Biol. 18:291-298. [DOI] [PubMed] [Google Scholar]
- 7.Bernstein, B. E., C. L. Liu, E. L. Humphrey, E. O. Perlstein, and S. L. Schreiber. 2004. Global nucleosome occupancy in yeast. Genome Biol. 5:R62. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Boeger, H., J. Griesenbeck, J. S. Strattan, and R. D. Kornberg. 2003. Nucleosomes unfold completely at a transcriptionally active promoter. Mol. Cell 11:1587-1598. [DOI] [PubMed] [Google Scholar]
- 9.Boeger, H., J. Griesenbeck, J. S. Strattan, and R. D. Kornberg. 2004. Removal of promoter nucleosomes by disassembly rather than sliding in vivo. Mol. Cell 14:667-673. [DOI] [PubMed] [Google Scholar]
- 10.Carrozza, M. J., B. Li, L. Florens, T. Suganuma, S. K. Swanson, K. K. Lee, W. J. Shia, S. Anderson, J. Yates, M. P. Washburn, and J. L. Workman. 2005. Histone H3 methylation by Set2 directs deacetylation of coding regions by Rpd3S to suppress spurious intragenic transcription. Cell 123:581-592. [DOI] [PubMed] [Google Scholar]
- 11.Chen, X., J. Wang, D. Woltring, S. Gerondakis, and M. F. Shannon. 2005. Histone dynamics on the interleukin-2 gene in response to T-cell activation. Mol. Cell. Biol. 25:3209-3219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Dhasarathy, A., and M. P. Kladde. 2005. Promoter occupancy is a major determinant of chromatin remodeling enzyme requirements. Mol. Cell. Biol. 25:2698-2707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.English, C. M., M. W. Adkins, J. J. Carson, M. E. A. Churchill, and J. K. Tyler. 2006. Structural basis for the histone chaperone activity of Asf1. Cell 127:495-508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Gavin, I. M., and R. T. Simpson. 1997. Interplay of yeast global transcriptional regulators Ssn6p-Tup1p and Swi-Snf and their effect on chromatin structure. EMBO J. 16:6263-6271. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Godde, J. S., and A. P. Wolffe. 1995. Disruption of reconstituted nucleosomes. The effect of particle concentration, MgCl2 and KCl concentration, the histone tails, and temperature. J. Biol. Chem. 270:27399-27402. [DOI] [PubMed] [Google Scholar]
- 16.Gregory, P. D., A. Schmid, M. Zavari, M. Munsterkotter, and W. Horz. 1999. Chromatin remodelling at the PHO8 promoter requires SWI-SNF and SAGA at a step subsequent to activator binding. EMBO J. 18:6407-6414. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Han, M., U. J. Kim, P. Kayne, and M. Grunstein. 1988. Depletion of histone H4 and nucleosomes activates the PHO5 gene in Saccharomyces cerevisiae. EMBO J. 7:2221-2228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Hassan, A. H., K. E. Neely, M. Vignali, J. C. Reese, and J. L. Workman. 2001. Promoter targeting of chromatin-modifying complexes. Front. Biosci. 6:D1054-D1064. [DOI] [PubMed] [Google Scholar]
- 19.Imbalzano, A. N., H. Kwon, M. R. Green, and R. E. Kingston. 1994. Facilitated binding of TATA-binding protein to nucleosomal DNA. Nature 370:481-485. [DOI] [PubMed] [Google Scholar]
- 20.Jayaraman, P. S., K. Hirst, and C. R. Goding. 1994. The activation domain of a basic helix-loop-helix protein is masked by repressor interaction with domains distinct from that required for transcription regulation. EMBO J. 13:2192-2199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Jessen, W. J., S. A. Hoose, J. A. Kilgore, and M. P. Kladde. 2006. Active PHO5 chromatin encompasses variable numbers of nucleosomes at individual promoters. Nat. Struct. Mol. Biol. 13:256-263. [DOI] [PubMed] [Google Scholar]
- 22.Kaffman, A., I. Herskowitz, R. Tjian, and E. K. O'Shea. 1994. Phosphorylation of the transcription factor PHO4 by a cyclin-CDK complex, PHO80-PHO85. Science 263:1153-1156. [DOI] [PubMed] [Google Scholar]
- 23.Kaplan, C. D., L. Laprade, and F. Winston. 2003. Transcription elongation factors repress transcription initiation from cryptic sites. Science 301:1096-1099. [DOI] [PubMed] [Google Scholar]
- 24.Kim, Y., N. McLaughlin, K. Lindstrom, T. Tsukiyama, and D. J. Clark. 2006. Activation of Saccharomyces cerevisiae HIS3 results in Gcn4p-dependent, SWI/SNF-dependent mobilization of nucleosomes over the entire gene. Mol. Cell. Biol. 26:8607-8622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Korber, P., S. Barbaric, T. Luckenbach, A. Schmid, U. J. Schermer, D. Blaschke, and W. Horz. 2006. The histone chaperone Asf1 increases the rate of histone eviction at the yeast PHO5 and PHO8 promoters. J. Biol. Chem. 3:5539-5545. [DOI] [PubMed] [Google Scholar]
- 26.Kristjuhan, A., and J. Q. Svejstrup. 2004. Evidence for distinct mechanisms facilitating transcript elongation through chromatin in vivo. EMBO J. 23:4243-4252. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Kuras, L., and K. Struhl. 1999. Binding of TBP to promoters in vivo is stimulated by activators and requires Pol II holoenzyme. Nature 399:609-613. [DOI] [PubMed] [Google Scholar]
- 28.Lee, C. K., Y. Shibata, B. Rao, B. D. Strahl, and J. D. Lieb. 2004. Evidence for nucleosome depletion at active regulatory regions genome-wide. Nat. Genet. 36:900-905. [DOI] [PubMed] [Google Scholar]
- 29.Longtine, M. S., A. McKenzie III, D. J. Demarini, N. G. Shah, A. Wach, A. Brachat, P. Philippsen, and J. R. Pringle. 1998. Additional modules for versatile and economical PCR-based gene deletion and modification in Saccharomyces cerevisiae. Yeast 14:953-961. [DOI] [PubMed] [Google Scholar]
- 30.Luger, K., A. W. Mader, R. K. Richmond, D. F. Sargent, and T. J. Richmond. 1997. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389:251-260. [DOI] [PubMed] [Google Scholar]
- 31.Mason, P. B., and K. Struhl. 2003. The FACT complex travels with elongating RNA polymerase II and is important for the fidelity of transcriptional initiation in vivo. Mol. Cell. Biol. 23:8323-8333. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Mellor, J. 2006. Dynamic nucleosomes and gene transcription. Trends Genet. 22:320-329. [DOI] [PubMed] [Google Scholar]
- 33.Mito, Y., J. G. Henikoff, and S. Henikoff. 2005. Genome-scale profiling of histone H3.3 replacement patterns. Nat. Genet. 37:1090-1097. [DOI] [PubMed] [Google Scholar]
- 34.Moshkin, Y. M., J. A. Armstrong, R. K. Maeda, J. W. Tamkun, P. Verrijzer, J. A. Kennison, and F. Karch. 2002. Histone chaperone ASF1 cooperates with the Brahma chromatin-remodelling machinery. Genes Dev. 16:2621-2626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Neef, D. W., and M. P. Kladde. 2003. Polyphosphate loss promotes SNF/SWI- and Gcn5-dependent mitotic induction of PHO5. Mol. Cell. Biol. 23:3788-3797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Neely, K. E., A. H. Hassan, C. E. Brown, L. Howe, and J. L. Workman. 2002. Transcription activator interactions with multiple SWI/SNF subunits. Mol. Cell. Biol. 22:1615-1625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Nourani, A., R. T. Utley, S. Allard, and J. Cote. 2004. Recruitment of the NuA4 complex poises the PHO5 promoter for chromatin remodeling and activation. EMBO J. 23:2597-2607. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Ogawa, N., K. Noguchi, H. Sawai, Y. Yamashita, C. Yompakdee, and Y. Oshima. 1995. Functional domains of Pho81p, an inhibitor of Pho85p protein kinase, in the transduction pathway of Pi signals in Saccharomyces cerevisiae. Mol. Cell. Biol. 15:997-1004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.O'Neill, E. M., A. Kaffman, E. R. Jolly, and E. K. O'Shea. 1996. Regulation of PHO4 nuclear localization by the PHO80-PHO85 cyclin-CDK complex. Science 271:209-212. [DOI] [PubMed] [Google Scholar]
- 40.Pokholok, D. K., C. T. Harbison, S. Levine, M. Cole, N. M. Hannett, T. I. Lee, G. W. Bell, K. Walker, P. A. Rolfe, E. Herbolsheimer, J. Zeitlinger, F. Lewitter, D. K. Gifford, and R. A. Young. 2005. Genome-wide map of nucleosome acetylation and methylation in yeast. Cell 122:517-527. [DOI] [PubMed] [Google Scholar]
- 41.Prochasson, P., K. E. Neely, A. H. Hassan, B. Li, and J. L. Workman. 2003. Targeting activity is required for SWI/SNF function in vivo and is accomplished through two partially redundant activator-interaction domains. Mol. Cell 12:983-990. [DOI] [PubMed] [Google Scholar]
- 42.Reinke, H., and W. Horz. 2003. Histones are first hyperacetylated and then lose contact with the activated PHO5 promoter. Mol. Cell 11:1599-1607. [DOI] [PubMed] [Google Scholar]
- 43.Rogers, D. T., J. M. Lemire, and K. A. Bostian. 1982. Acid phosphatase polypeptides in Saccharomyces cerevisiae are encoded by a differentially regulated multigene family. Proc. Natl. Acad. Sci. USA 79:2157-2161. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Schneider, K. R., R. L. Smith, and E. K. O'Shea. 1994. Phosphate-regulated inactivation of the kinase PHO80-PHO85 by the CDK inhibitor PHO81. Science 266:122-126. [DOI] [PubMed] [Google Scholar]
- 45.Schwabish, M. A., and K. Struhl. 2006. Asf1 mediates histone eviction and deposition during elongation by RNA polymerase II. Mol. Cell 22:415-422. [DOI] [PubMed] [Google Scholar]
- 46.Schwabish, M. A., and K. Struhl. 2004. Evidence for eviction and rapid deposition of histones upon transcriptional elongation by RNA polymerase II. Mol. Cell. Biol. 24:10111-10117. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Steger, D. J., E. S. Haswell, A. L. Miller, S. R. Wente, and E. K. O'Shea. 2003. Regulation of chromatin remodeling by inositol polyphosphates. Science 299:114-116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Sterner, D. E., P. A. Grant, S. M. Roberts, L. J. Duggan, R. Belotserkovskaya, L. A. Pacella, F. Winston, J. L. Workman, and S. L. Berger. 1999. Functional organization of the yeast SAGA complex: distinct components involved in structural integrity, nucleosome acetylation, and TATA-binding protein interaction. Mol. Cell. Biol. 19:86-98. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Svaren, J., and W. Horz. 1995. Interplay between nucleosomes and transcription factors at the yeast PHO5 promoter. Semin. Cell Biol. 6:177-183. [DOI] [PubMed] [Google Scholar]
- 50.Svaren, J., J. Schmitz, and W. Horz. 1994. The transactivation domain of Pho4 is required for nucleosome disruption at the PHO5 promoter. EMBO J. 13:4856-4862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Tagami, H., D. Ray-Gallet, G. Almouzni, and Y. Nakatani. 2004. Histone H3.1 and H3.3 complexes mediate nucleosome assembly pathways dependent or independent of DNA synthesis. Cell 116:51-61. [DOI] [PubMed] [Google Scholar]
- 52.Tait-Kamradt, A. G., K. J. Turner, R. A. Kramer, Q. D. Elliott, S. J. Bostian, G. P. Thill, D. T. Rogers, and K. A. Bostian. 1986. Reciprocal regulation of the tandemly duplicated PHO5/PHO3 gene cluster within the acid phosphatase multigene family of Saccharomyces cerevisiae. Mol. Cell. Biol. 6:1855-1865. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Tamburini, B. A., and J. K. Tyler. 2005. Localized histone acetylation and deacetylation triggered by the homologous recombination pathway of double-strand DNA repair. Mol. Cell. Biol. 25:4903-4913. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Tyler, J. K., K. A. Collins, J. Prasad-Sinha, E. Amiott, M. Bulger, P. J. Harte, R. Kobayashi, and J. T. Kadonaga. 2001. Interaction between the Drosophila CAF-1 and ASF1 chromatin assembly factors. Mol. Biol. Cell 21:6574-6584. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Van Holde, K. E. 1989. Chromatin. Springer-Verlag, New York, NY.
- 56.Verdone, L., G. Camilloni, E. Di Mauro, and M. Caserta. 1996. Chromatin remodeling during Saccharomyces cerevisiae ADH2 gene activation. Mol. Cell. Biol. 16:1978-1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Vogel, K., W. Horz, and A. Hinnen. 1989. The two positively acting regulatory proteins PHO2 and PHO4 physically interact with PHO5 upstream activation regions. Mol. Cell. Biol. 9:2050-2057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Walther, K., and H. J. Schuller. 2001. Adr1 and Cat8 synergistically activate the glucose-regulated alcohol dehydrogenase gene ADH2 of the yeast Saccharomyces cerevisiae. Microbiology 147:2037-2044. [DOI] [PubMed] [Google Scholar]
- 59.Workman, J. L. 2006. Nucleosome displacement in transcription. Genes Dev. 20:2009-2017. [DOI] [PubMed] [Google Scholar]
- 60.Workman, J. L., and R. G. Roeder. 1987. Binding of transcription factor TFIID to the major late promoter during in vitro nucleosome assembly potentiates subsequent initiation by RNA polymerase II. Cell 51:613-622. [DOI] [PubMed] [Google Scholar]
- 61.Zabaronick, S. R., and J. K. Tyler. 2005. The histone chaperone anti-silencing function 1 is a global regulator of transcription independent of passage through S phase. Mol. Cell. Biol. 25:652-660. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Zhao, J., J. Herrera-Diaz, and D. S. Gross. 2005. Domain-wide displacement of histones by activated heat shock factor occurs independently of Swi/Snf and is not correlated with RNA polymerase II density. Mol. Cell. Biol. 25:8985-8999. [DOI] [PMC free article] [PubMed] [Google Scholar]