Abstract
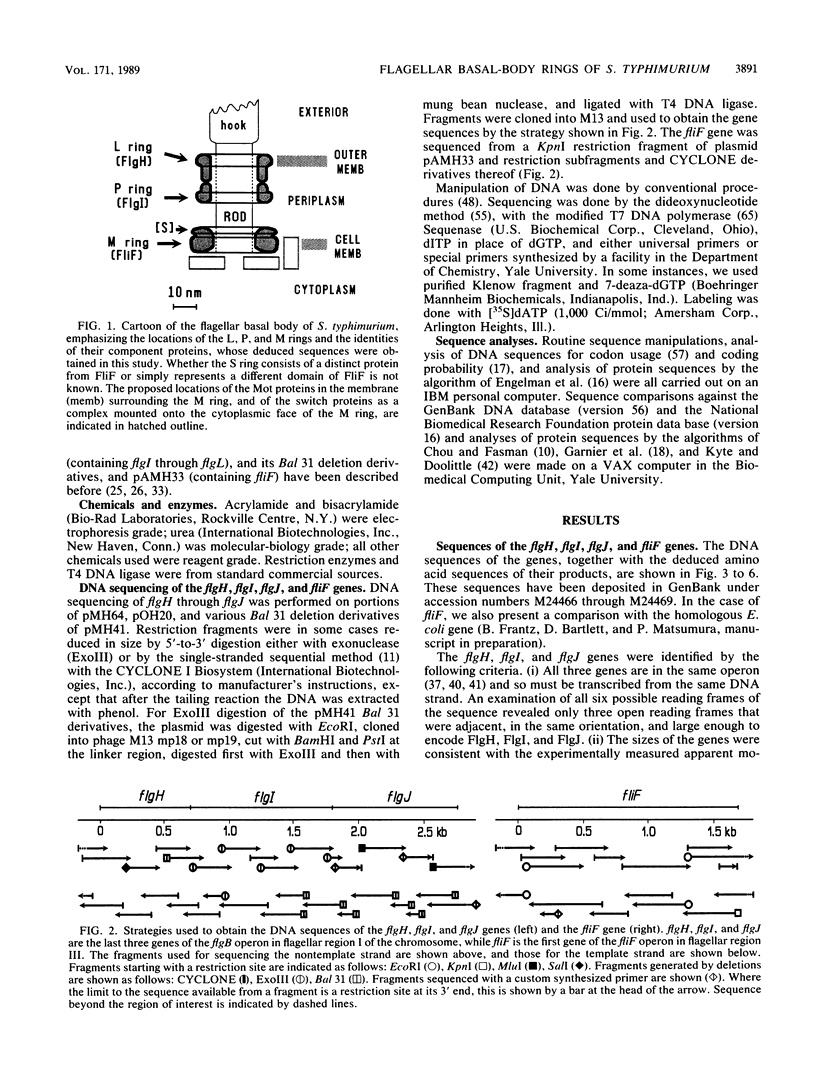
The flgH, flgI, and fliF genes of Salmonella typhimurium encode the major proteins for the L, P, and M rings of the flagellar basal body. We have determined the sequences of these genes and the flgJ gene and examined the deduced amino acid sequences of their products. FlgH and FlgI, which are exported across the cell membrane to their destinations in the outer membrane and periplasmic space, respectively, both had typical N-terminal cleaved signal-peptide sequences. FlgH is predicted to have a considerable amount of beta-sheet structure, as has been noted for other outer membrane proteins. FlgI is predicted to have an even greater amount of beta-structure. FliF, as is usual for a cytoplasmic membrane protein of a procaryote, lacked a signal peptide; it is predicted to have considerable alpha-helical structure, including an N-terminal sequence that is likely to be membrane-spanning. However, it had overall a quite hydrophilic sequence with a high charge density, especially towards its C terminus. The flgJ gene, immediately adjacent to flgI and the last gene of the flgB operon, encodes a flagellar protein of unknown function whose deduced sequence was hydrophilic and may correspond to a cytoplasmic protein. Several aspects of the DNA sequence of these genes and their surrounds suggest complex regulation of the flagellar gene system. A notable example occurs within the flgB operon, where between the end of flgG (encoding the distal rod protein of the basal body) and the start of flgH (encoding the L-ring protein) there was an unusually long noncoding region containing a potential stem-loop sequence, which could attenuate termination of transcription or stabilize part of the transcript against degradation. Another example is the interface between the flgB and flgK operons, where transcription termination of the former may occur within the coding region of the latter.
Full text
PDF









Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aizawa S. I., Dean G. E., Jones C. J., Macnab R. M., Yamaguchi S. Purification and characterization of the flagellar hook-basal body complex of Salmonella typhimurium. J Bacteriol. 1985 Mar;161(3):836–849. doi: 10.1128/jb.161.3.836-849.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Apirion D., Watson N. Ribonuclease III is involved in motility of Escherichia coli. J Bacteriol. 1978 Mar;133(3):1543–1545. doi: 10.1128/jb.133.3.1543-1545.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bartlett D. H., Frantz B. B., Matsumura P. Flagellar transcriptional activators FlbB and FlaI: gene sequences and 5' consensus sequences of operons under FlbB and FlaI control. J Bacteriol. 1988 Apr;170(4):1575–1581. doi: 10.1128/jb.170.4.1575-1581.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bartlett D. H., Matsumura P. Identification of Escherichia coli region III flagellar gene products and description of two new flagellar genes. J Bacteriol. 1984 Nov;160(2):577–585. doi: 10.1128/jb.160.2.577-585.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Belasco J. G., Beatty J. T., Adams C. W., von Gabain A., Cohen S. N. Differential expression of photosynthesis genes in R. capsulata results from segmental differences in stability within the polycistronic rxcA transcript. Cell. 1985 Jan;40(1):171–181. doi: 10.1016/0092-8674(85)90320-4. [DOI] [PubMed] [Google Scholar]
- Braun V. Covalent lipoprotein from the outer membrane of Escherichia coli. Biochim Biophys Acta. 1975 Oct 31;415(3):335–377. doi: 10.1016/0304-4157(75)90013-1. [DOI] [PubMed] [Google Scholar]
- Burton Z. F., Gross C. A., Watanabe K. K., Burgess R. R. The operon that encodes the sigma subunit of RNA polymerase also encodes ribosomal protein S21 and DNA primase in E. coli K12. Cell. 1983 Feb;32(2):335–349. doi: 10.1016/0092-8674(83)90453-1. [DOI] [PubMed] [Google Scholar]
- Champer R., Dingwall A., Shapiro L. Cascade regulation of Caulobacter flagellar and chemotaxis genes. J Mol Biol. 1987 Mar 5;194(1):71–80. doi: 10.1016/0022-2836(87)90716-9. [DOI] [PubMed] [Google Scholar]
- Chen R., Schmidmayr W., Krämer C., Chen-Schmeisser U., Henning U. Primary structure of major outer membrane protein II (ompA protein) of Escherichia coli K-12. Proc Natl Acad Sci U S A. 1980 Aug;77(8):4592–4596. doi: 10.1073/pnas.77.8.4592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chou P. Y., Fasman G. D. Prediction of the secondary structure of proteins from their amino acid sequence. Adv Enzymol Relat Areas Mol Biol. 1978;47:45–148. doi: 10.1002/9780470122921.ch2. [DOI] [PubMed] [Google Scholar]
- Dale R. M., McClure B. A., Houchins J. P. A rapid single-stranded cloning strategy for producing a sequential series of overlapping clones for use in DNA sequencing: application to sequencing the corn mitochondrial 18 S rDNA. Plasmid. 1985 Jan;13(1):31–40. doi: 10.1016/0147-619x(85)90053-8. [DOI] [PubMed] [Google Scholar]
- DePamphilis M. L., Adler J. Attachment of flagellar basal bodies to the cell envelope: specific attachment to the outer, lipopolysaccharide membrane and the cyoplasmic membrane. J Bacteriol. 1971 Jan;105(1):396–407. doi: 10.1128/jb.105.1.396-407.1971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DePamphilis M. L., Adler J. Fine structure and isolation of the hook-basal body complex of flagella from Escherichia coli and Bacillus subtilis. J Bacteriol. 1971 Jan;105(1):384–395. doi: 10.1128/jb.105.1.384-395.1971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dean G. E., Macnab R. M., Stader J., Matsumura P., Burks C. Gene sequence and predicted amino acid sequence of the motA protein, a membrane-associated protein required for flagellar rotation in Escherichia coli. J Bacteriol. 1984 Sep;159(3):991–999. doi: 10.1128/jb.159.3.991-999.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Emerson S. U., Tokuyasu K., Simon M. I. Bacterial flagella: polarity of elongation. Science. 1970 Jul 10;169(3941):190–192. doi: 10.1126/science.169.3941.190. [DOI] [PubMed] [Google Scholar]
- Fickett J. W. Recognition of protein coding regions in DNA sequences. Nucleic Acids Res. 1982 Sep 11;10(17):5303–5318. doi: 10.1093/nar/10.17.5303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Garnier J., Osguthorpe D. J., Robson B. Analysis of the accuracy and implications of simple methods for predicting the secondary structure of globular proteins. J Mol Biol. 1978 Mar 25;120(1):97–120. doi: 10.1016/0022-2836(78)90297-8. [DOI] [PubMed] [Google Scholar]
- Gren E. J. Recognition of messenger RNA during translational initiation in Escherichia coli. Biochimie. 1984 Jan;66(1):1–29. doi: 10.1016/0300-9084(84)90188-3. [DOI] [PubMed] [Google Scholar]
- Hahnenberger K. M., Shapiro L. Organization and temporal expression of a flagellar basal body gene in Caulobacter crescentus. J Bacteriol. 1988 Sep;170(9):4119–4124. doi: 10.1128/jb.170.9.4119-4124.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Helmann J. D., Chamberlin M. J. DNA sequence analysis suggests that expression of flagellar and chemotaxis genes in Escherichia coli and Salmonella typhimurium is controlled by an alternative sigma factor. Proc Natl Acad Sci U S A. 1987 Sep;84(18):6422–6424. doi: 10.1073/pnas.84.18.6422. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Homma M., Aizawa S., Dean G. E., Macnab R. M. Identification of the M-ring protein of the flagellar motor of Salmonella typhimurium. Proc Natl Acad Sci U S A. 1987 Nov;84(21):7483–7487. doi: 10.1073/pnas.84.21.7483. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Homma M., Komeda Y., Iino T., Macnab R. M. The flaFIX gene product of Salmonella typhimurium is a flagellar basal body component with a signal peptide for export. J Bacteriol. 1987 Apr;169(4):1493–1498. doi: 10.1128/jb.169.4.1493-1498.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Homma M., Kutsukake K., Iino T. Structural genes for flagellar hook-associated proteins in Salmonella typhimurium. J Bacteriol. 1985 Aug;163(2):464–471. doi: 10.1128/jb.163.2.464-471.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Homma M., Ohnishi K., Iino T., Macnab R. M. Identification of flagellar hook and basal body gene products (FlaFV, FlaFVI, FlaFVII and FlaFVIII) in Salmonella typhimurium. J Bacteriol. 1987 Aug;169(8):3617–3624. doi: 10.1128/jb.169.8.3617-3624.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iino T., Komeda Y., Kutsukake K., Macnab R. M., Matsumura P., Parkinson J. S., Simon M. I., Yamaguchi S. New unified nomenclature for the flagellar genes of Escherichia coli and Salmonella typhimurium. Microbiol Rev. 1988 Dec;52(4):533–535. doi: 10.1128/mr.52.4.533-535.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iino T. Polarity of flagellar growth in salmonella. J Gen Microbiol. 1969 May;56(2):227–239. doi: 10.1099/00221287-56-2-227. [DOI] [PubMed] [Google Scholar]
- Ikemura T. Correlation between the abundance of Escherichia coli transfer RNAs and the occurrence of the respective codons in its protein genes: a proposal for a synonymous codon choice that is optimal for the E. coli translational system. J Mol Biol. 1981 Sep 25;151(3):389–409. doi: 10.1016/0022-2836(81)90003-6. [DOI] [PubMed] [Google Scholar]
- Jones C. J., Homma M., Macnab R. M. Identification of proteins of the outer (L and P) rings of the flagellar basal body of Escherichia coli. J Bacteriol. 1987 Apr;169(4):1489–1492. doi: 10.1128/jb.169.4.1489-1492.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kagawa H., Aizawa S. I., Yamaguchi S., Ishizu J. I. Isolation and characterization of bacterial flagellar hook proteins from salmonellae and Escherichia coli. J Bacteriol. 1979 Apr;138(1):235–240. doi: 10.1128/jb.138.1.235-240.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Khan S., Dapice M., Reese T. S. Effects of mot gene expression on the structure of the flagellar motor. J Mol Biol. 1988 Aug 5;202(3):575–584. doi: 10.1016/0022-2836(88)90287-2. [DOI] [PubMed] [Google Scholar]
- Kihara M., Homma M., Kutsukake K., Macnab R. M. Flagellar switch of Salmonella typhimurium: gene sequences and deduced protein sequences. J Bacteriol. 1989 Jun;171(6):3247–3257. doi: 10.1128/jb.171.6.3247-3257.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kleffel B., Garavito R. M., Baumeister W., Rosenbusch J. P. Secondary structure of a channel-forming protein: porin from E. coli outer membranes. EMBO J. 1985 Jun;4(6):1589–1592. doi: 10.1002/j.1460-2075.1985.tb03821.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Komeda Y. Fusions of flagellar operons to lactose genes on a mu lac bacteriophage. J Bacteriol. 1982 Apr;150(1):16–26. doi: 10.1128/jb.150.1.16-26.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Komeda Y., Kutsukake K., Iino T. Definition of additional flagellar genes in Escherichia coli K12. Genetics. 1980 Feb;94(2):277–290. doi: 10.1093/genetics/94.2.277. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Komeda Y. Transcriptional control of flagellar genes in Escherichia coli K-12. J Bacteriol. 1986 Dec;168(3):1315–1318. doi: 10.1128/jb.168.3.1315-1318.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Konigsberg W., Godson G. N. Evidence for use of rare codons in the dnaG gene and other regulatory genes of Escherichia coli. Proc Natl Acad Sci U S A. 1983 Feb;80(3):687–691. doi: 10.1073/pnas.80.3.687. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuo S. C., Koshland D. E., Jr Sequence of the flaA (cheC) locus of Escherichia coli and discovery of a new gene. J Bacteriol. 1986 Jun;166(3):1007–1012. doi: 10.1128/jb.166.3.1007-1012.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kutsukake K., Iino T., Komeda Y., Yamaguchi S. Functional homology of fla genes between Salmonella typhimurium and Escherichia coli. Mol Gen Genet. 1980 Apr;178(1):59–67. doi: 10.1007/BF00267213. [DOI] [PubMed] [Google Scholar]
- Kutsukake K., Ohya Y., Yamaguchi S., Iino T. Operon structure of flagellar genes in Salmonella typhimurium. Mol Gen Genet. 1988 Sep;214(1):11–15. doi: 10.1007/BF00340172. [DOI] [PubMed] [Google Scholar]
- Kyte J., Doolittle R. F. A simple method for displaying the hydropathic character of a protein. J Mol Biol. 1982 May 5;157(1):105–132. doi: 10.1016/0022-2836(82)90515-0. [DOI] [PubMed] [Google Scholar]
- Lupski J. R., Smiley B. L., Godson G. N. Regulation of the rpsU-dnaG-rpoD macromolecular synthesis operon and the initiation of DNA replication in Escherichia coli K-12. Mol Gen Genet. 1983;189(1):48–57. doi: 10.1007/BF00326054. [DOI] [PubMed] [Google Scholar]
- Macnab R. M. The end of the line in bacterial sensing: the flagellar motor. Cold Spring Harb Symp Quant Biol. 1988;53(Pt 1):67–75. doi: 10.1101/sqb.1988.053.01.011. [DOI] [PubMed] [Google Scholar]
- Malakooti J., Komeda Y., Matsumura P. DNA sequence analysis, gene product identification, and localization of flagellar motor components of Escherichia coli. J Bacteriol. 1989 May;171(5):2728–2734. doi: 10.1128/jb.171.5.2728-2734.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mott J. E., Galloway J. L., Platt T. Maturation of Escherichia coli tryptophan operon mRNA: evidence for 3' exonucleolytic processing after rho-dependent termination. EMBO J. 1985 Jul;4(7):1887–1891. doi: 10.1002/j.1460-2075.1985.tb03865.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohnishi K., Homma M., Kutsukake K., Iino T. Formation of flagella lacking outer rings by flaM, flaU, and flaY mutants of Escherichia coli. J Bacteriol. 1987 Apr;169(4):1485–1488. doi: 10.1128/jb.169.4.1485-1488.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Okino H., Isomura M., Yamaguchi S., Magariyama Y., Kudo S., Aizawa S. I. Release of flagellar filament-hook-rod complex by a Salmonella typhimurium mutant defective in the M ring of the basal body. J Bacteriol. 1989 Apr;171(4):2075–2082. doi: 10.1128/jb.171.4.2075-2082.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oliver D. Protein secretion in Escherichia coli. Annu Rev Microbiol. 1985;39:615–648. doi: 10.1146/annurev.mi.39.100185.003151. [DOI] [PubMed] [Google Scholar]
- Pabo C. O., Sauer R. T. Protein-DNA recognition. Annu Rev Biochem. 1984;53:293–321. doi: 10.1146/annurev.bi.53.070184.001453. [DOI] [PubMed] [Google Scholar]
- Ralling G., Linn T. Relative activities of the transcriptional regulatory sites in the rplKAJLrpoBC gene cluster of Escherichia coli. J Bacteriol. 1984 Apr;158(1):279–285. doi: 10.1128/jb.158.1.279-285.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmeissner U., McKenney K., Rosenberg M., Court D. Removal of a terminator structure by RNA processing regulates int gene expression. J Mol Biol. 1984 Jun 15;176(1):39–53. doi: 10.1016/0022-2836(84)90381-4. [DOI] [PubMed] [Google Scholar]
- Sharp P. M., Li W. H. The codon Adaptation Index--a measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res. 1987 Feb 11;15(3):1281–1295. doi: 10.1093/nar/15.3.1281. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stader J., Matsumura P., Vacante D., Dean G. E., Macnab R. M. Nucleotide sequence of the Escherichia coli motB gene and site-limited incorporation of its product into the cytoplasmic membrane. J Bacteriol. 1986 Apr;166(1):244–252. doi: 10.1128/jb.166.1.244-252.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stallmeyer M. J., Aizawa S., Macnab R. M., DeRosier D. J. Image reconstruction of the flagellar basal body of Salmonella typhimurium. J Mol Biol. 1989 Feb 5;205(3):519–528. doi: 10.1016/0022-2836(89)90223-4. [DOI] [PubMed] [Google Scholar]
- Stern M. J., Ames G. F., Smith N. H., Robinson E. C., Higgins C. F. Repetitive extragenic palindromic sequences: a major component of the bacterial genome. Cell. 1984 Jul;37(3):1015–1026. doi: 10.1016/0092-8674(84)90436-7. [DOI] [PubMed] [Google Scholar]
- Suzuki T., Iino T., Horiguchi T., Yamaguchi S. Incomplete flagellar structures in nonflagellate mutants of Salmonella typhimurium. J Bacteriol. 1978 Feb;133(2):904–915. doi: 10.1128/jb.133.2.904-915.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suzuki T., Komeda Y. Incomplete flagellar structures in Escherichia coli mutants. J Bacteriol. 1981 Feb;145(2):1036–1041. doi: 10.1128/jb.145.2.1036-1041.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tabor S., Richardson C. C. DNA sequence analysis with a modified bacteriophage T7 DNA polymerase. Proc Natl Acad Sci U S A. 1987 Jul;84(14):4767–4771. doi: 10.1073/pnas.84.14.4767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tinoco I., Jr, Borer P. N., Dengler B., Levin M. D., Uhlenbeck O. C., Crothers D. M., Bralla J. Improved estimation of secondary structure in ribonucleic acids. Nat New Biol. 1973 Nov 14;246(150):40–41. doi: 10.1038/newbio246040a0. [DOI] [PubMed] [Google Scholar]
- Vogel H., Jähnig F. Models for the structure of outer-membrane proteins of Escherichia coli derived from raman spectroscopy and prediction methods. J Mol Biol. 1986 Jul 20;190(2):191–199. doi: 10.1016/0022-2836(86)90292-5. [DOI] [PubMed] [Google Scholar]
- Yamaguchi S., Aizawa S., Kihara M., Isomura M., Jones C. J., Macnab R. M. Genetic evidence for a switching and energy-transducing complex in the flagellar motor of Salmonella typhimurium. J Bacteriol. 1986 Dec;168(3):1172–1179. doi: 10.1128/jb.168.3.1172-1179.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamamori T., Ito K., Yura T., Suzuki T., Iino T. Ribonucleic acid polymerase mutant of Escherichia coli defective in flagella formation. J Bacteriol. 1977 Oct;132(1):254–261. doi: 10.1128/jb.132.1.254-261.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]


