Abstract
We describe a method for generating gene replacements and deletions in Escherichia coli. The technique is simple and rapid and can be applied to most genes, even those that are essential. What makes this method unique and particularly effective is the use of a temperature-sensitive pSC101 replicon to facilitate the gene replacement. The method proceeds by homologous recombination between a gene on the chromosome and homologous sequences carried on a plasmid temperature sensitive for DNA replication. Thus, after transformation of the plasmid into an appropriate host, it is possible to select for integration of the plasmid into the chromosome at 44 degrees C. Subsequent growth of these cointegrates at 30 degrees C leads to a second recombination event, resulting in their resolution. Depending on where the second recombination event takes place, the chromosome will either have undergone a gene replacement or retain the original copy of the gene. The procedure can also be used to effect the transfer of an allele from a plasmid to the chromosome or to rescue a chromosomal allele onto a plasmid. Since the resolved plasmid can be maintained by selection, this technique can be used to generate deletions of essential genes.
Full text
PDF
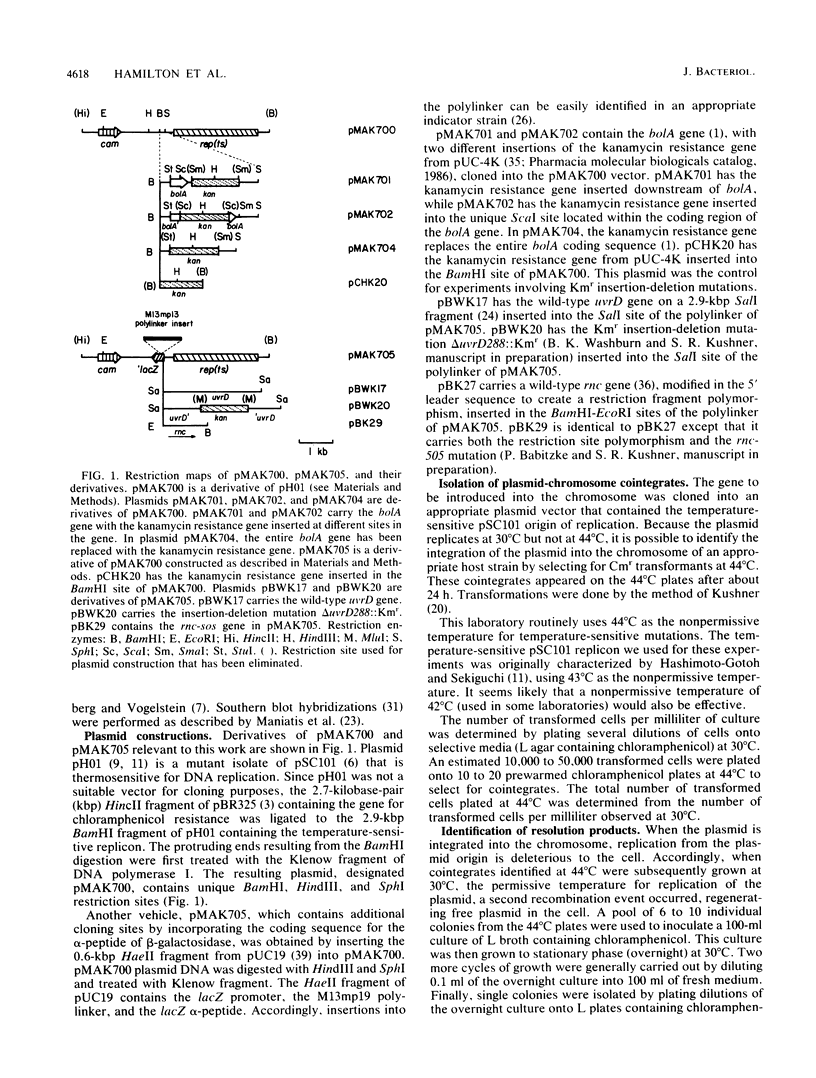
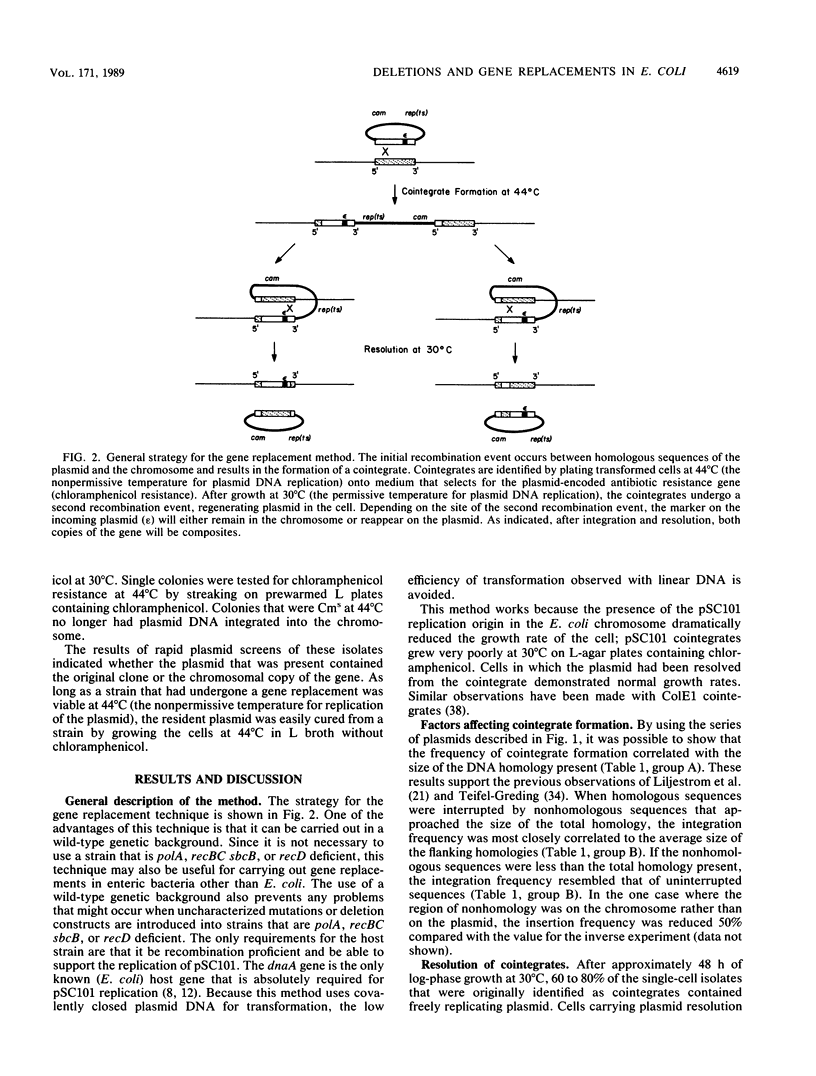





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Aldea M., Hernández-Chico C., de la Campa A. G., Kushner S. R., Vicente M. Identification, cloning, and expression of bolA, an ftsZ-dependent morphogene of Escherichia coli. J Bacteriol. 1988 Nov;170(11):5169–5176. doi: 10.1128/jb.170.11.5169-5176.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Balakrishnan R., Backman K. Controllable alteration of cell genotype in bacterial cultures using an excision vector. Gene. 1988 Jul 15;67(1):97–103. doi: 10.1016/0378-1119(88)90012-1. [DOI] [PubMed] [Google Scholar]
- Balbás P., Soberón X., Merino E., Zurita M., Lomeli H., Valle F., Flores N., Bolivar F. Plasmid vector pBR322 and its special-purpose derivatives--a review. Gene. 1986;50(1-3):3–40. doi: 10.1016/0378-1119(86)90307-0. [DOI] [PubMed] [Google Scholar]
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bochner B. R., Huang H. C., Schieven G. L., Ames B. N. Positive selection for loss of tetracycline resistance. J Bacteriol. 1980 Aug;143(2):926–933. doi: 10.1128/jb.143.2.926-933.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cohen S. N., Chang A. C. Revised interpretation of the origin of the pSC101 plasmid. J Bacteriol. 1977 Nov;132(2):734–737. doi: 10.1128/jb.132.2.734-737.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feinberg A. P., Vogelstein B. A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem. 1983 Jul 1;132(1):6–13. doi: 10.1016/0003-2697(83)90418-9. [DOI] [PubMed] [Google Scholar]
- Frey J., Chandler M., Caro L. The effects of an Escherichia coli dnaAts mutation on the replication of the plasmids colE1 pSC101, R100.1 and RTF-TC. Mol Gen Genet. 1979 Jul 13;174(2):117–126. doi: 10.1007/BF00268349. [DOI] [PubMed] [Google Scholar]
- Gamas P., Burger A. C., Churchward G., Caro L., Galas D., Chandler M. Replication of pSC101: effects of mutations in the E. coli DNA binding protein IHF. Mol Gen Genet. 1986 Jul;204(1):85–89. doi: 10.1007/BF00330192. [DOI] [PubMed] [Google Scholar]
- Gutterson N. I., Koshland D. E., Jr Replacement and amplification of bacterial genes with sequences altered in vitro. Proc Natl Acad Sci U S A. 1983 Aug;80(16):4894–4898. doi: 10.1073/pnas.80.16.4894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hashimoto-Gotoh T., Sekiguchi M. Mutations of temperature sensitivity in R plasmid pSC101. J Bacteriol. 1977 Aug;131(2):405–412. doi: 10.1128/jb.131.2.405-412.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hasunuma K., Sekiguchi M. Replication of plasmid pSC101 in Escherichia coli K12: requirement for dnaA function. Mol Gen Genet. 1977 Sep 9;154(3):225–230. doi: 10.1007/BF00571277. [DOI] [PubMed] [Google Scholar]
- Ish-Horowicz D., Burke J. F. Rapid and efficient cosmid cloning. Nucleic Acids Res. 1981 Jul 10;9(13):2989–2998. doi: 10.1093/nar/9.13.2989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jasin M., Schimmel P. Deletion of an essential gene in Escherichia coli by site-specific recombination with linear DNA fragments. J Bacteriol. 1984 Aug;159(2):783–786. doi: 10.1128/jb.159.2.783-786.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones I. M., Primrose S. B., Robinson A., Ellwood D. C. Maintenance of some ColE1-type plasmids in chemostat culture. Mol Gen Genet. 1980;180(3):579–584. doi: 10.1007/BF00268063. [DOI] [PubMed] [Google Scholar]
- Joyce C. M., Grindley N. D. Method for determining whether a gene of Escherichia coli is essential: application to the polA gene. J Bacteriol. 1984 May;158(2):636–643. doi: 10.1128/jb.158.2.636-643.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kiel J. A., Vossen J. P., Venema G. A general method for the construction of Escherichia coli mutants by homologous recombination and plasmid segregation. Mol Gen Genet. 1987 May;207(2-3):294–301. doi: 10.1007/BF00331592. [DOI] [PubMed] [Google Scholar]
- Kleckner N., Roth J., Botstein D. Genetic engineering in vivo using translocatable drug-resistance elements. New methods in bacterial genetics. J Mol Biol. 1977 Oct 15;116(1):125–159. doi: 10.1016/0022-2836(77)90123-1. [DOI] [PubMed] [Google Scholar]
- Kushner S. R. In vivo studies of temperature-sensitive recB and recC mutants. J Bacteriol. 1974 Dec;120(3):1213–1218. doi: 10.1128/jb.120.3.1213-1218.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liljeström P., Pirhonen M., Palva E. T. In vivo transfer of chromosomal mutations onto multicopy plasmids by transduction with bacteriophage P1. Gene. 1985;40(2-3):241–246. doi: 10.1016/0378-1119(85)90046-0. [DOI] [PubMed] [Google Scholar]
- Maloy S. R., Nunn W. D. Selection for loss of tetracycline resistance by Escherichia coli. J Bacteriol. 1981 Feb;145(2):1110–1111. doi: 10.1128/jb.145.2.1110-1111.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maples V. F., Kushner S. R. DNA repair in Escherichia coli: identification of the uvrD gene product. Proc Natl Acad Sci U S A. 1982 Sep;79(18):5616–5620. doi: 10.1073/pnas.79.18.5616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsuyama S., Mizushima S. Construction and characterization of a deletion mutant lacking micF, a proposed regulatory gene for OmpF synthesis in Escherichia coli. J Bacteriol. 1985 Jun;162(3):1196–1202. doi: 10.1128/jb.162.3.1196-1202.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Parker B., Marinus M. G. A simple and rapid method to obtain substitution mutations in Escherichia coli: isolation of a dam deletion/insertion mutation. Gene. 1988 Dec 20;73(2):531–535. doi: 10.1016/0378-1119(88)90517-3. [DOI] [PubMed] [Google Scholar]
- Shevell D. E., Abou-Zamzam A. M., Demple B., Walker G. C. Construction of an Escherichia coli K-12 ada deletion by gene replacement in a recD strain reveals a second methyltransferase that repairs alkylated DNA. J Bacteriol. 1988 Jul;170(7):3294–3296. doi: 10.1128/jb.170.7.3294-3296.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Summers D. K., Sherratt D. J. Multimerization of high copy number plasmids causes instability: CoIE1 encodes a determinant essential for plasmid monomerization and stability. Cell. 1984 Apr;36(4):1097–1103. doi: 10.1016/0092-8674(84)90060-6. [DOI] [PubMed] [Google Scholar]
- Sunshine M. G., Kelly B. Extent of host deletions associated with bacteriophage P2-mediated eduction. J Bacteriol. 1971 Nov;108(2):695–704. doi: 10.1128/jb.108.2.695-704.1971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vieira J., Messing J. The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene. 1982 Oct;19(3):259–268. doi: 10.1016/0378-1119(82)90015-4. [DOI] [PubMed] [Google Scholar]
- Watson N., Apirion D. Molecular cloning of the gene for the RNA-processing enzyme RNase III of Escherichia coli. Proc Natl Acad Sci U S A. 1985 Feb;82(3):849–853. doi: 10.1073/pnas.82.3.849. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Winans S. C., Elledge S. J., Krueger J. H., Walker G. C. Site-directed insertion and deletion mutagenesis with cloned fragments in Escherichia coli. J Bacteriol. 1985 Mar;161(3):1219–1221. doi: 10.1128/jb.161.3.1219-1221.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamaguchi K., Tomizawa J. Establishment of Escherichia coli cells with an integrated high copy number plasmid. Mol Gen Genet. 1980;178(3):525–533. doi: 10.1007/BF00337857. [DOI] [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]
- Zieg J., Maples V. F., Kushner S. R. Recombinant levels of Escherichia coli K-12 mutants deficient in various replication, recombination, or repair genes. J Bacteriol. 1978 Jun;134(3):958–966. doi: 10.1128/jb.134.3.958-966.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]