Abstract
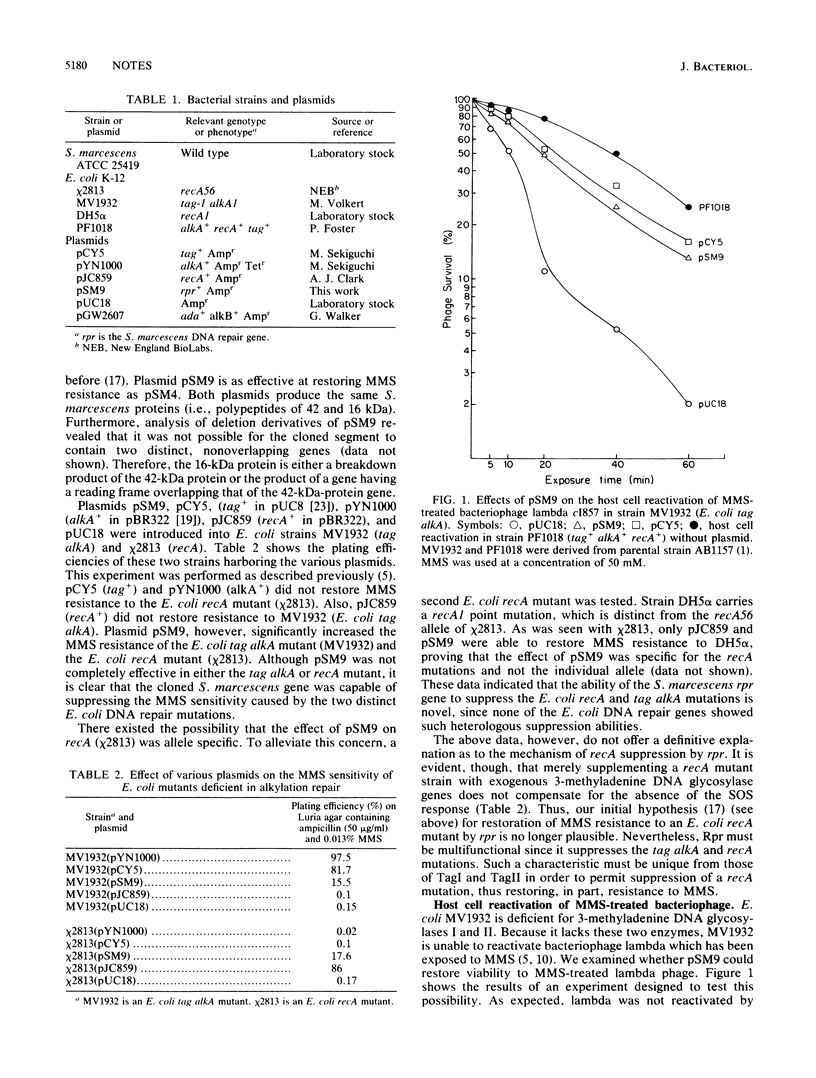
A recombinant plasmid containing a Serratia marcescens DNA repair gene has been analyzed biochemically and genetically in Escherichia coli mutants deficient for repair of alkylated DNA. The cloned gene suppressed sensitivity to methyl methanesulfonate of an E. coli strain deficient in 3-methyladenine DNA glycosylases I and II (i.e., E. coli tag alkA) and two different E. coli recA mutants. Attempts to suppress the methyl methanesulfonate sensitivity of the E. coli recA mutant by using the cloned E. coli tag and alkA genes were not successful. Southern blot analysis did not reveal any homology between the S. marcescens gene and various known E. coli DNA repair genes. Biochemical analysis with the S. marcescens gene showed that the encoded DNA repair protein liberated 3-methyladenine from alkylated DNA, indicating that the DNA repair molecular is an S. marcescens 3-methyladenine DNA glycosylase. The ability to suppress both types of E. coli DNA repair mutations, however, suggests that the S. marcescens gene is a unique bacterial DNA repair gene.
Full text
PDF

Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bachmann B. J. Pedigrees of some mutant strains of Escherichia coli K-12. Bacteriol Rev. 1972 Dec;36(4):525–557. doi: 10.1128/br.36.4.525-557.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beranek D. T., Weis C. C., Swenson D. H. A comprehensive quantitative analysis of methylated and ethylated DNA using high pressure liquid chromatography. Carcinogenesis. 1980 Jul;1(7):595–606. doi: 10.1093/carcin/1.7.595. [DOI] [PubMed] [Google Scholar]
- Clarke N. D., Kvaal M., Seeberg E. Cloning of Escherichia coli genes encoding 3-methyladenine DNA glycosylases I and II. Mol Gen Genet. 1984;197(3):368–372. doi: 10.1007/BF00329931. [DOI] [PubMed] [Google Scholar]
- Evensen G. Induction of 3-methyladenine DNA glycosylase II is recA+-independent. Mutat Res. 1985 Sep;146(2):143–147. doi: 10.1016/0167-8817(85)90004-5. [DOI] [PubMed] [Google Scholar]
- Evensen G., Seeberg E. Adaptation to alkylation resistance involves the induction of a DNA glycosylase. Nature. 1982 Apr 22;296(5859):773–775. doi: 10.1038/296773a0. [DOI] [PubMed] [Google Scholar]
- Hanawalt P. C., Cooper P. K., Ganesan A. K., Smith C. A. DNA repair in bacteria and mammalian cells. Annu Rev Biochem. 1979;48:783–836. doi: 10.1146/annurev.bi.48.070179.004031. [DOI] [PubMed] [Google Scholar]
- Haseltine W. A., Gordon L. K., Lindan C. P., Grafstrom R. H., Shaper N. L., Grossman L. Cleavage of pyrimidine dimers in specific DNA sequences by a pyrimidine dimer DNA-glycosylase of M. luteus. Nature. 1980 Jun 26;285(5767):634–641. doi: 10.1038/285634a0. [DOI] [PubMed] [Google Scholar]
- Kaasen I., Evensen G., Seeberg E. Amplified expression of the tag+ and alkA+ genes in Escherichia coli: identification of gene products and effects on alkylation resistance. J Bacteriol. 1986 Nov;168(2):642–647. doi: 10.1128/jb.168.2.642-647.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karran P., Hjelmgren T., Lindahl T. Induction of a DNA glycosylase for N-methylated purines is part of the adaptive response to alkylating agents. Nature. 1982 Apr 22;296(5859):770–773. doi: 10.1038/296770a0. [DOI] [PubMed] [Google Scholar]
- Karran P., Lindahl T., Ofsteng I., Evensen G. B., Seeberg E. Escherichia coli mutants deficient in 3-methyladenine-DNA glycosylase. J Mol Biol. 1980 Jun 15;140(1):101–127. doi: 10.1016/0022-2836(80)90358-7. [DOI] [PubMed] [Google Scholar]
- Laval J. Two enzymes are required from strand incision in repair of alkylated DNA. Nature. 1977 Oct 27;269(5631):829–832. doi: 10.1038/269829a0. [DOI] [PubMed] [Google Scholar]
- Lemotte P. K., Walker G. C. Induction and autoregulation of ada, a positively acting element regulating the response of Escherichia coli K-12 to methylating agents. J Bacteriol. 1985 Mar;161(3):888–895. doi: 10.1128/jb.161.3.888-895.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lindahl T. DNA glycosylases, endonucleases for apurinic/apyrimidinic sites, and base excision-repair. Prog Nucleic Acid Res Mol Biol. 1979;22:135–192. doi: 10.1016/s0079-6603(08)60800-4. [DOI] [PubMed] [Google Scholar]
- Lindahl T. New class of enzymes acting on damaged DNA. Nature. 1976 Jan 1;259(5538):64–66. doi: 10.1038/259064a0. [DOI] [PubMed] [Google Scholar]
- Male R., Helland D. E., Kleppe K. Purification and characterization of 3-methyladenine-DNA glycosylase from calf thymus. J Biol Chem. 1985 Feb 10;260(3):1623–1629. [PubMed] [Google Scholar]
- Murphy K. E., Braymer H. D. Molecular cloning and characterization of a genetic region from Serratia marcescens involved in DNA repair. Mol Microbiol. 1989 Feb;3(2):249–255. doi: 10.1111/j.1365-2958.1989.tb01814.x. [DOI] [PubMed] [Google Scholar]
- Nakabeppu Y., Kondo H., Sekiguchi M. Cloning and characterization of the alkA gene of Escherichia coli that encodes 3-methyladenine DNA glycosylase II. J Biol Chem. 1984 Nov 25;259(22):13723–13729. [PubMed] [Google Scholar]
- Nakabeppu Y., Miyata T., Kondo H., Iwanaga S., Sekiguchi M. Structure and expression of the alkA gene of Escherichia coli involved in adaptive response to alkylating agents. J Biol Chem. 1984 Nov 25;259(22):13730–13736. [PubMed] [Google Scholar]
- Radany E. H., Friedberg E. C. A pyrimidine dimer-DNA glycosylase activity associated with the v gene product of bacterophage T4. Nature. 1980 Jul 10;286(5769):182–185. doi: 10.1038/286182a0. [DOI] [PubMed] [Google Scholar]
- Riazuddin S., Lindahl T. Properties of 3-methyladenine-DNA glycosylase from Escherichia coli. Biochemistry. 1978 May 30;17(11):2110–2118. doi: 10.1021/bi00604a014. [DOI] [PubMed] [Google Scholar]
- Sakumi K., Nakabeppu Y., Yamamoto Y., Kawabata S., Iwanaga S., Sekiguchi M. Purification and structure of 3-methyladenine-DNA glycosylase I of Escherichia coli. J Biol Chem. 1986 Nov 25;261(33):15761–15766. [PubMed] [Google Scholar]
- Seeberg E., Nissen-Meyer J., Strike P. Incision of ultraviolet-irradiated DNA by extracts of E. coli requires three different gene products. Nature. 1976 Oct 7;263(5577):524–526. doi: 10.1038/263524a0. [DOI] [PubMed] [Google Scholar]
- Singer B., Brent T. P. Human lymphoblasts contain DNA glycosylase activity excising N-3 and N-7 methyl and ethyl purines but not O6-alkylguanines or 1-alkyladenines. Proc Natl Acad Sci U S A. 1981 Feb;78(2):856–860. doi: 10.1073/pnas.78.2.856. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Thomas L., Yang C. H., Goldthwait D. A. Two DNA glycosylases in Escherichia coli which release primarily 3-methyladenine. Biochemistry. 1982 Mar 16;21(6):1162–1169. doi: 10.1021/bi00535a009. [DOI] [PubMed] [Google Scholar]
- Walker G. C. Mutagenesis and inducible responses to deoxyribonucleic acid damage in Escherichia coli. Microbiol Rev. 1984 Mar;48(1):60–93. doi: 10.1128/mr.48.1.60-93.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]


