Abstract
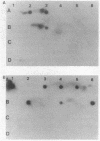
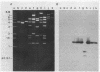
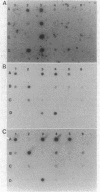
A series of intragenic DNA probes, encoding the major part of the transposase resolvase and inverted repeats of transposons Tn3, Tn21, and Tn2501, were used in hybridization assays for homologous DNA sequences in 18 transposons studied. The tnpA and tnpR probes detected extensive homology with Tn3-like and Tn21-like elements for 11 transposons. This high degree of homology was confirmed with the 38- and 48-base-pair inverted-repeat oligonucleotide probes of Tn3, Tn21, and Tn2501. The Southern-type gel hybridization experiments localized the tnpA-homologous sequences on the physical DNA maps constructed. The genetic and physical maps of the transposons were compared, as were their nucleic acid sequence homologies. These comparisons suggested a subfamily of mobile elements distinct from but related to the Tn21 group. Based on these results, an evolutionary model is proposed and a pedigree is presented for the genesis of multiresistance beta-lactamase transposons.
Full text
PDF






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Altenbuchner J., Schmid K., Schmitt R. Tn1721-encoded tetracycline resistance: mapping of structural and regulatory genes mediating resistance. J Bacteriol. 1983 Jan;153(1):116–123. doi: 10.1128/jb.153.1.116-123.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bennett P. M., Grinsted J., Choi C. L., Richmond M. H. Characterisation of Tn501, a transposon determining resistance to mercuric ions. Mol Gen Genet. 1978 Feb 7;159(1):101–106. doi: 10.1007/BF00401753. [DOI] [PubMed] [Google Scholar]
- Boissinot M., Mercier J., Levesque R. C. Development of natural and synthetic DNA probes for OXA-2 and TEM-1 beta-lactamases. Antimicrob Agents Chemother. 1987 May;31(5):728–734. doi: 10.1128/aac.31.5.728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boyer H. W., Roulland-Dussoix D. A complementation analysis of the restriction and modification of DNA in Escherichia coli. J Mol Biol. 1969 May 14;41(3):459–472. doi: 10.1016/0022-2836(69)90288-5. [DOI] [PubMed] [Google Scholar]
- Chang A. C., Cohen S. N. Construction and characterization of amplifiable multicopy DNA cloning vehicles derived from the P15A cryptic miniplasmid. J Bacteriol. 1978 Jun;134(3):1141–1156. doi: 10.1128/jb.134.3.1141-1156.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clerget M., Chandler M., Caro L. The structure of R1drd19: a revised physical map of the plasmid. Mol Gen Genet. 1981;181(2):183–191. doi: 10.1007/BF00268425. [DOI] [PubMed] [Google Scholar]
- Diver W. P., Grinsted J., Fritzinger D. C., Brown N. L., Altenbuchner J., Rogowsky P., Schmitt R. DNA sequences of and complementation by the tnpR genes of Tn21, Tn501 and Tn1721. Mol Gen Genet. 1983;191(2):189–193. doi: 10.1007/BF00334812. [DOI] [PubMed] [Google Scholar]
- Grinsted J., de la Cruz F., Altenbuchner J., Schmitt R. Complementation of transposition of tnpA mutants of Tn3, Tn21, Tn501, and Tn1721. Plasmid. 1982 Nov;8(3):276–286. doi: 10.1016/0147-619x(82)90065-8. [DOI] [PubMed] [Google Scholar]
- Guerry P., LeBlanc D. J., Falkow S. General method for the isolation of plasmid deoxyribonucleic acid. J Bacteriol. 1973 Nov;116(2):1064–1066. doi: 10.1128/jb.116.2.1064-1066.1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hedges R. W., Matthew M., Smith D. I., Cresswell J. M., Jacob A. E. Properties of a transposon conferring resistance to penicillins and streptomycin. Gene. 1977 May;1(3-4):241–253. doi: 10.1016/0378-1119(77)90048-8. [DOI] [PubMed] [Google Scholar]
- Heffron F., McCarthy B. J., Ohtsubo H., Ohtsubo E. DNA sequence analysis of the transposon Tn3: three genes and three sites involved in transposition of Tn3. Cell. 1979 Dec;18(4):1153–1163. doi: 10.1016/0092-8674(79)90228-9. [DOI] [PubMed] [Google Scholar]
- Humphreys G. O., Willshaw G. A., Anderson E. S. A simple method for the preparation of large quantities of pure plasmid DNA. Biochim Biophys Acta. 1975 Apr 2;383(4):457–463. doi: 10.1016/0005-2787(75)90318-4. [DOI] [PubMed] [Google Scholar]
- Hyde D. R., Tu C. P. tnpM: a novel regulatory gene that enhances Tn21 transposition and suppresses cointegrate resolution. Cell. 1985 Sep;42(2):629–638. doi: 10.1016/0092-8674(85)90120-5. [DOI] [PubMed] [Google Scholar]
- Kafatos F. C., Jones C. W., Efstratiadis A. Determination of nucleic acid sequence homologies and relative concentrations by a dot hybridization procedure. Nucleic Acids Res. 1979 Nov 24;7(6):1541–1552. doi: 10.1093/nar/7.6.1541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kahn M., Kolter R., Thomas C., Figurski D., Meyer R., Remaut E., Helinski D. R. Plasmid cloning vehicles derived from plasmids ColE1, F, R6K, and RK2. Methods Enzymol. 1979;68:268–280. doi: 10.1016/0076-6879(79)68019-9. [DOI] [PubMed] [Google Scholar]
- Kleckner N. Transposable elements in prokaryotes. Annu Rev Genet. 1981;15:341–404. doi: 10.1146/annurev.ge.15.120181.002013. [DOI] [PubMed] [Google Scholar]
- Levesque R. C., Jacoby G. A. Molecular structure and interrelationships of multiresistance beta-lactamase transposons. Plasmid. 1988 Jan;19(1):21–29. doi: 10.1016/0147-619x(88)90059-5. [DOI] [PubMed] [Google Scholar]
- Levesque R. C., Medeiros A. A., Jacoby G. A. Molecular cloning and DNA homology of plasmid-mediated beta-lactamase genes. Mol Gen Genet. 1987 Feb;206(2):252–258. doi: 10.1007/BF00333581. [DOI] [PubMed] [Google Scholar]
- Levesque R., Roy P. H., Letarte R., Pechère J. C. A plasmid-mediated cephalosporinase from Achromobacter species. J Infect Dis. 1982 May;145(5):753–761. doi: 10.1093/infdis/145.2.753. [DOI] [PubMed] [Google Scholar]
- Medeiros A. A. Beta-lactamases. Br Med Bull. 1984 Jan;40(1):18–27. doi: 10.1093/oxfordjournals.bmb.a071942. [DOI] [PubMed] [Google Scholar]
- Medeiros A. A., Cohenford M., Jacoby G. A. Five novel plasmid-determined beta-lactamases. Antimicrob Agents Chemother. 1985 May;27(5):715–719. doi: 10.1128/aac.27.5.715. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Michiels T., Cornelis G. Detection and characterization of Tn2501, a transposon included within the lactose transposon Tn951. J Bacteriol. 1984 Jun;158(3):866–871. doi: 10.1128/jb.158.3.866-871.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Michiels T., Cornelis G., Ellis K., Grinsted J. Tn2501, a component of the lactose transposon Tn951, is an example of a new category of class II transposable elements. J Bacteriol. 1987 Feb;169(2):624–631. doi: 10.1128/jb.169.2.624-631.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Philippon A. M., Paul G. C., Thabaut A. P., Jacoby G. A. Properties of a novel carbenicillin-hydrolyzing beta-lactamase (CARB-4) specified by an IncP-2 plasmid from Pseudomonas aeruginosa. Antimicrob Agents Chemother. 1986 Mar;29(3):519–520. doi: 10.1128/aac.29.3.519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Radloff R., Bauer W., Vinograd J. A dye-buoyant-density method for the detection and isolation of closed circular duplex DNA: the closed circular DNA in HeLa cells. Proc Natl Acad Sci U S A. 1967 May;57(5):1514–1521. doi: 10.1073/pnas.57.5.1514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rigby P. W., Dieckmann M., Rhodes C., Berg P. Labeling deoxyribonucleic acid to high specific activity in vitro by nick translation with DNA polymerase I. J Mol Biol. 1977 Jun 15;113(1):237–251. doi: 10.1016/0022-2836(77)90052-3. [DOI] [PubMed] [Google Scholar]
- Schmidt F., Klopfer-Kaul I. Evolutionary relationship between Tn21-like elements and pBP201, a plasmid from Klebsiella pneumoniae mediating resistance to gentamicin and eight other drugs. Mol Gen Genet. 1984;197(1):109–119. doi: 10.1007/BF00327930. [DOI] [PubMed] [Google Scholar]
- Sutcliffe J. G. Complete nucleotide sequence of the Escherichia coli plasmid pBR322. Cold Spring Harb Symp Quant Biol. 1979;43(Pt 1):77–90. doi: 10.1101/sqb.1979.043.01.013. [DOI] [PubMed] [Google Scholar]
- Tanaka M., Yamamoto T., Sawai T. Evolution of complex resistance transposons from an ancestral mercury transposon. J Bacteriol. 1983 Mar;153(3):1432–1438. doi: 10.1128/jb.153.3.1432-1438.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ward E., Grinsted J. The nucleotide sequence of the tnpA gene of Tn21. Nucleic Acids Res. 1987 Feb 25;15(4):1799–1806. doi: 10.1093/nar/15.4.1799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ward J. M., Grinsted J. Physical and genetic analysis of the Inc-W group plasmids R388, Sa, and R7K. Plasmid. 1982 May;7(3):239–250. doi: 10.1016/0147-619x(82)90005-1. [DOI] [PubMed] [Google Scholar]
- Yamamoto T., Tanaka M., Nohara C., Fukunaga Y., Yamagishi S. Transposition of the oxacillin-hydrolyzing penicillinase gene. J Bacteriol. 1981 Feb;145(2):808–813. doi: 10.1128/jb.145.2.808-813.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de la Cruz F., Grinsted J. Genetic and molecular characterization of Tn21, a multiple resistance transposon from R100.1. J Bacteriol. 1982 Jul;151(1):222–228. doi: 10.1128/jb.151.1.222-228.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]