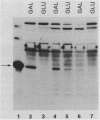
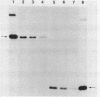
Abstract
We present the gene organization and DNA sequence of the Streptomyces lividans galactose utilization genes. Complementation of Escherichia coli galE, galT, or galK mutants and DNA sequence analysis were used to demonstrate that the galactose utilization genes are organized within an operon with the gene order galT, galE, and galK. Comparison of the inferred protein sequences for the S. lividans gal gene products to the corresponding E. coli and Saccharomyces carlbergensis sequences identified regions of structural homology within each of the galactose utilization enzymes. Finally, we discuss a potential relationship between the gene organization of the operon and the functional roles of the gal enzymes in cellular metabolism.
Full text
PDF
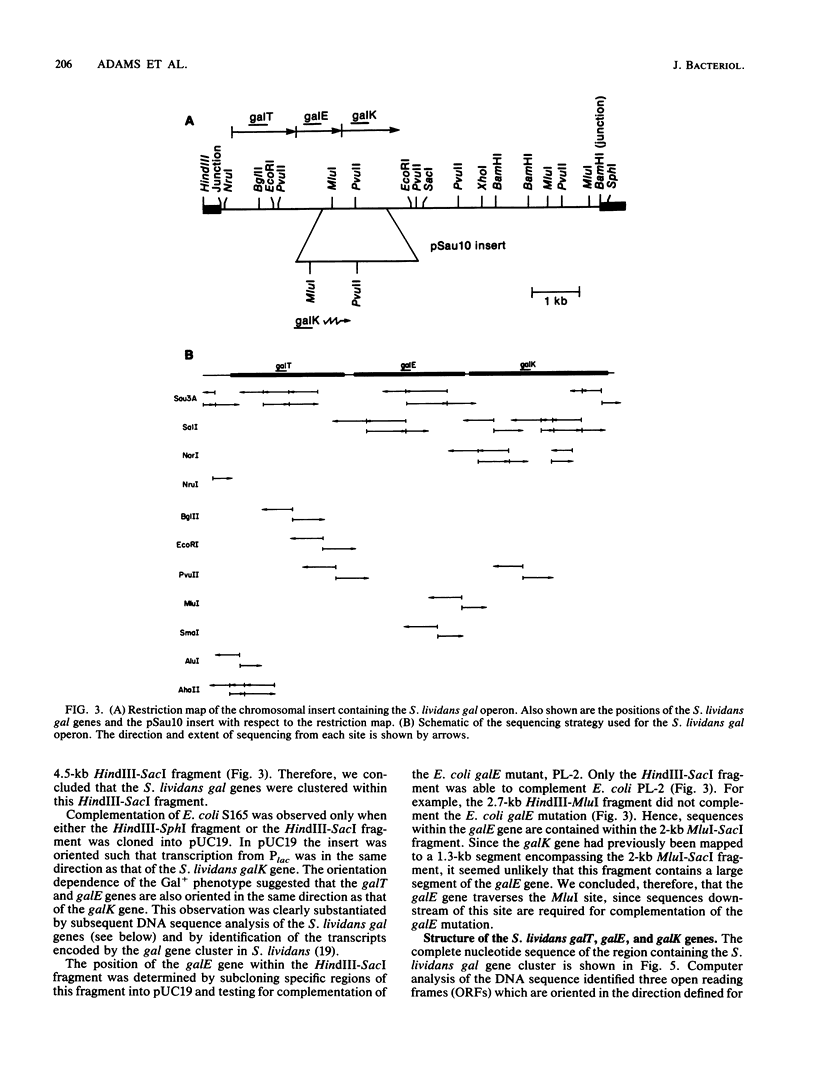
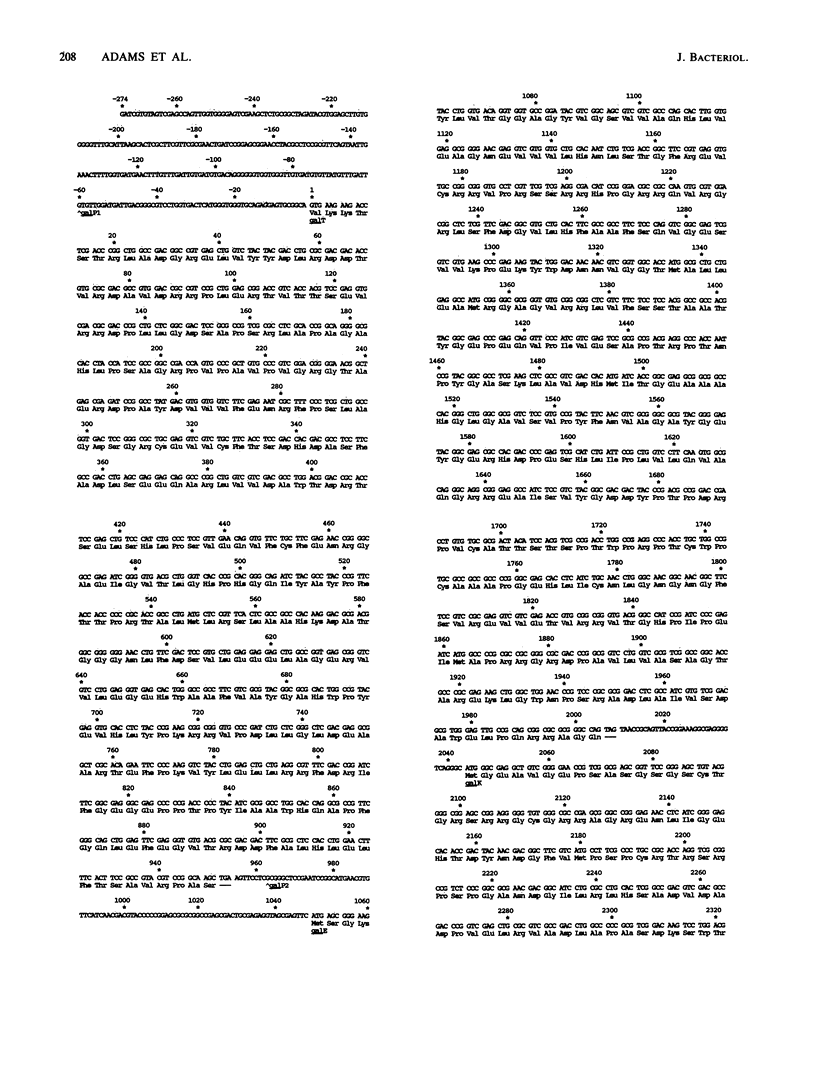
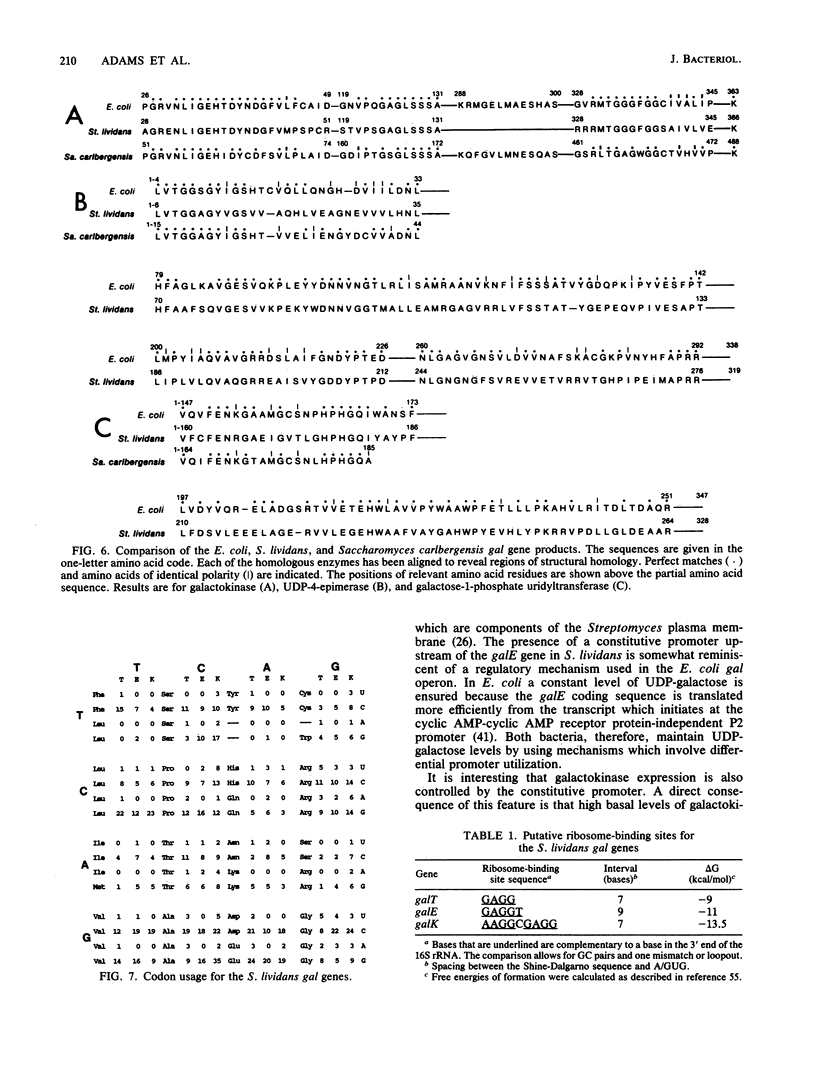








Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- ADLER J., TEMPLETON B. THE AMOUNT OF GALACTOSE GENETIC MATERIAL IN LAMBDA-DG BACTERIOPHAGE WITH DIFFERENT DENSITIES. J Mol Biol. 1963 Dec;7:710–720. doi: 10.1016/s0022-2836(63)80118-7. [DOI] [PubMed] [Google Scholar]
- Ambartsumyan N. S., Mazo A. M. Elimination of the secondary structure effect in gell sequencing of nucleic acids. FEBS Lett. 1980 Jun 2;114(2):265–268. doi: 10.1016/0014-5793(80)81130-6. [DOI] [PubMed] [Google Scholar]
- BUTTIN G. M'ECANISMES R'EGULATEURS DANS LA BIOSYNTH'ESE DES ENZYMES DU M'ETABOLISME DU GALACTOSE CHEZ ESCHERICHIA COLI K12. I. LA BIOSYNTH'ESE INDUITE DE LA GALACTOKINASE ET L'INDUCTION SIMULTAN'EE DE LA S'EQUENCE ENZYMATIQUE. J Mol Biol. 1963 Aug;7:164–182. doi: 10.1016/s0022-2836(63)80044-3. [DOI] [PubMed] [Google Scholar]
- Bernan V., Filpula D., Herber W., Bibb M., Katz E. The nucleotide sequence of the tyrosinase gene from Streptomyces antibioticus and characterization of the gene product. Gene. 1985;37(1-3):101–110. doi: 10.1016/0378-1119(85)90262-8. [DOI] [PubMed] [Google Scholar]
- Bibb M. J., Bibb M. J., Ward J. M., Cohen S. N. Nucleotide sequences encoding and promoting expression of three antibiotic resistance genes indigenous to Streptomyces. Mol Gen Genet. 1985;199(1):26–36. doi: 10.1007/BF00327505. [DOI] [PubMed] [Google Scholar]
- Bibb M. J., Cohen S. N. Gene expression in Streptomyces: construction and application of promoter-probe plasmid vectors in Streptomyces lividans. Mol Gen Genet. 1982;187(2):265–277. doi: 10.1007/BF00331128. [DOI] [PubMed] [Google Scholar]
- Bibb M. J., Janssen G. R., Ward J. M. Cloning and analysis of the promoter region of the erythromycin resistance gene (ermE) of Streptomyces erythraeus. Gene. 1985;38(1-3):215–226. doi: 10.1016/0378-1119(85)90220-3. [DOI] [PubMed] [Google Scholar]
- Brawner M. E., Auerbach J. I., Fornwald J. A., Rosenberg M., Taylor D. P. Characterization of Streptomyces promoter sequences using the Escherichia coli galactokinase gene. Gene. 1985;40(2-3):191–201. doi: 10.1016/0378-1119(85)90042-3. [DOI] [PubMed] [Google Scholar]
- Chater K. F., Bruton C. J. Resistance, regulatory and production genes for the antibiotic methylenomycin are clustered. EMBO J. 1985 Jul;4(7):1893–1897. doi: 10.1002/j.1460-2075.1985.tb03866.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Citron B. A., Donelson J. E. Sequence of the Saccharomyces GAL region and its transcription in vivo. J Bacteriol. 1984 Apr;158(1):269–278. doi: 10.1128/jb.158.1.269-278.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cohen S. N., Chang A. C., Hsu L. Nonchromosomal antibiotic resistance in bacteria: genetic transformation of Escherichia coli by R-factor DNA. Proc Natl Acad Sci U S A. 1972 Aug;69(8):2110–2114. doi: 10.1073/pnas.69.8.2110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DOUGLAS H. C., HAWTHORNE D. C. ENZYMATIC EXPRESSION AND GENETIC LINKAGE OF GENES CONTROLLING GALACTOSE UTILIZATION IN SACCHAROMYCES. Genetics. 1964 May;49:837–844. doi: 10.1093/genetics/49.5.837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- De Crombrugghe B., Perlman R. L., Varmus H. E., Pastan I. Regulation of inducible enzyme synthesis in Escherichia coli by cyclic adenosine 3', 5'-monophosphate. J Biol Chem. 1969 Nov 10;244(21):5828–5835. [PubMed] [Google Scholar]
- Debouck C., Riccio A., Schumperli D., McKenney K., Jeffers J., Hughes C., Rosenberg M., Heusterspreute M., Brunel F., Davison J. Structure of the galactokinase gene of Escherichia coli, the last (?) gene of the gal operon. Nucleic Acids Res. 1985 Mar 25;13(6):1841–1853. doi: 10.1093/nar/13.6.1841. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Denhardt D. T. A membrane-filter technique for the detection of complementary DNA. Biochem Biophys Res Commun. 1966 Jun 13;23(5):641–646. doi: 10.1016/0006-291x(66)90447-5. [DOI] [PubMed] [Google Scholar]
- Feitelson J. S., Hopwood D. A. Cloning of a Streptomyces gene for an O-methyltransferase involved in antibiotic biosynthesis. Mol Gen Genet. 1983;190(3):394–398. doi: 10.1007/BF00331065. [DOI] [PubMed] [Google Scholar]
- Fornwald J. A., Schmidt F. J., Adams C. W., Rosenberg M., Brawner M. E. Two promoters, one inducible and one constitutive, control transcription of the Streptomyces lividans galactose operon. Proc Natl Acad Sci U S A. 1987 Apr;84(8):2130–2134. doi: 10.1073/pnas.84.8.2130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gold L., Pribnow D., Schneider T., Shinedling S., Singer B. S., Stormo G. Translational initiation in prokaryotes. Annu Rev Microbiol. 1981;35:365–403. doi: 10.1146/annurev.mi.35.100181.002053. [DOI] [PubMed] [Google Scholar]
- Kieser T., Hopwood D. A., Wright H. M., Thompson C. J. pIJ101, a multi-copy broad host-range Streptomyces plasmid: functional analysis and development of DNA cloning vectors. Mol Gen Genet. 1982;185(2):223–228. doi: 10.1007/BF00330791. [DOI] [PubMed] [Google Scholar]
- Lemaire H. G., Müller-Hill B. Nucleotide sequences of the gal E gene and the gal T gene of E. coli. Nucleic Acids Res. 1986 Oct 10;14(19):7705–7711. doi: 10.1093/nar/14.19.7705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lomovskaya N. D., Mkrtumian N. M., Gostimskaya N. L., Danilenko V. N. Characterization of temperate actinophage phi C31 isolated from Streptomyces coelicolor A3(2). J Virol. 1972 Feb;9(2):258–262. doi: 10.1128/jvi.9.2.258-262.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Malpartida F., Hopwood D. A. Molecular cloning of the whole biosynthetic pathway of a Streptomyces antibiotic and its expression in a heterologous host. 1984 May 31-Jun 6Nature. 309(5967):462–464. doi: 10.1038/309462a0. [DOI] [PubMed] [Google Scholar]
- Malpartida F., Hopwood D. A. Physical and genetic characterisation of the gene cluster for the antibiotic actinorhodin in Streptomyces coelicolor A3(2). Mol Gen Genet. 1986 Oct;205(1):66–73. doi: 10.1007/BF02428033. [DOI] [PubMed] [Google Scholar]
- McLaughlin J. R., Murray C. L., Rabinowitz J. C. Unique features in the ribosome binding site sequence of the gram-positive Staphylococcus aureus beta-lactamase gene. J Biol Chem. 1981 Nov 10;256(21):11283–11291. [PubMed] [Google Scholar]
- Miller Z., Varmus H. E., Parks J. S., Perlman R. L., Pastan I. Regulation of gal messenger ribonucleic acid synthesis in Escherichia coli by 3',5'-cyclic adenosine monophosphate. J Biol Chem. 1971 May 10;246(9):2898–2903. [PubMed] [Google Scholar]
- Ohnuki T., Imanaka T., Aiba S. Self-cloning in Streptomyces griseus of an str gene cluster for streptomycin biosynthesis and streptomycin resistance. J Bacteriol. 1985 Oct;164(1):85–94. doi: 10.1128/jb.164.1.85-94.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piret J. M., Chater K. F. Phage-mediated cloning of bldA, a region involved in Streptomyces coelicolor morphological development, and its analysis by genetic complementation. J Bacteriol. 1985 Sep;163(3):965–972. doi: 10.1128/jb.163.3.965-972.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pustell J., Kafatos F. C. A convenient and adaptable package of computer programs for DNA and protein sequence management, analysis and homology determination. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 2):643–655. doi: 10.1093/nar/12.1part2.643. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pustell J., Kafatos F. C. A high speed, high capacity homology matrix: zooming through SV40 and polyoma. Nucleic Acids Res. 1982 Aug 11;10(15):4765–4782. doi: 10.1093/nar/10.15.4765. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Queen C., Rosenberg M. Differential translation efficiency explains discoordinate expression of the galactose operon. Cell. 1981 Jul;25(1):241–249. doi: 10.1016/0092-8674(81)90249-x. [DOI] [PubMed] [Google Scholar]
- Robbins P. W., Trimble R. B., Wirth D. F., Hering C., Maley F., Maley G. F., Das R., Gibson B. W., Royal N., Biemann K. Primary structure of the Streptomyces enzyme endo-beta-N-acetylglucosaminidase H. J Biol Chem. 1984 Jun 25;259(12):7577–7583. [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schümperli D., McKenney K., Sobieski D. A., Rosenberg M. Translational coupling at an intercistronic boundary of the Escherichia coli galactose operon. Cell. 1982 Oct;30(3):865–871. doi: 10.1016/0092-8674(82)90291-4. [DOI] [PubMed] [Google Scholar]
- Seno E. T., Chater K. F. Glycerol catabolic enzymes and their regulation in wild-type and mutant strains of Streptomyces coelicolor A3(2). J Gen Microbiol. 1983 May;129(5):1403–1413. doi: 10.1099/00221287-129-5-1403. [DOI] [PubMed] [Google Scholar]
- Shapiro J. A., Adhya S. L. The galactose operon of E. coli K-12. II. A deletion analysis of operon structure and polarity. Genetics. 1969 Jun;62(2):249–264. doi: 10.1093/genetics/62.2.249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Stormo G. D., Schneider T. D., Gold L. M. Characterization of translational initiation sites in E. coli. Nucleic Acids Res. 1982 May 11;10(9):2971–2996. doi: 10.1093/nar/10.9.2971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson C. J., Gray G. S. Nucleotide sequence of a streptomycete aminoglycoside phosphotransferase gene and its relationship to phosphotransferases encoded by resistance plasmids. Proc Natl Acad Sci U S A. 1983 Sep;80(17):5190–5194. doi: 10.1073/pnas.80.17.5190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson C. J., Ward J. M., Hopwood D. A. Cloning of antibiotic resistance and nutritional genes in streptomycetes. J Bacteriol. 1982 Aug;151(2):668–677. doi: 10.1128/jb.151.2.668-677.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson C. J., Ward J. M., Hopwood D. A. DNA cloning in Streptomyces: resistance genes from antibiotic-producing species. Nature. 1980 Jul 31;286(5772):525–527. doi: 10.1038/286525a0. [DOI] [PubMed] [Google Scholar]
- Timmis K. N., Cabello F., Cohen S. N. Cloning and characterization of EcoRI and HindIII restriction endonuclease-generated fragments of antibiotic resistance plasmids R6-5 and R6. Mol Gen Genet. 1978 Jun 14;162(2):121–137. doi: 10.1007/BF00267869. [DOI] [PubMed] [Google Scholar]
- Tinoco I., Jr, Borer P. N., Dengler B., Levin M. D., Uhlenbeck O. C., Crothers D. M., Bralla J. Improved estimation of secondary structure in ribonucleic acids. Nat New Biol. 1973 Nov 14;246(150):40–41. doi: 10.1038/newbio246040a0. [DOI] [PubMed] [Google Scholar]
- Vieira J., Messing J. The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene. 1982 Oct;19(3):259–268. doi: 10.1016/0378-1119(82)90015-4. [DOI] [PubMed] [Google Scholar]
- Yanisch-Perron C., Vieira J., Messing J. Improved M13 phage cloning vectors and host strains: nucleotide sequences of the M13mp18 and pUC19 vectors. Gene. 1985;33(1):103–119. doi: 10.1016/0378-1119(85)90120-9. [DOI] [PubMed] [Google Scholar]