Abstract
We have used Tn917lacZ to mutagenize the Bacillus subtilis chromosome and have isolated mutants that are defective in chemotaxis and motility. Mapping of the transposon inserts identified two new loci. Mutations in one of these loci generated mutants that had paralyzed flagella. Accordingly, we designate this a mot locus. The other locus is closely linked to the first and encodes proteins specifying chemotaxis functions. This locus is designated the cheX locus. Both the mot and cheX loci map close to ptsI. An additional transposon insert that maps in the hag locus was obtained. The pattern of beta-galactosidase expression from some of the transposons suggested that the mot locus is regulated by sigD, a minor sigma factor of B. subtilis. The cheX locus appeared to be under the control of vegetative sigA. Four transposon inserts were mapped to a previously characterized che locus near spcB. These mutants did not produce flagellin and were defective in the methylation of the methyl-accepting chemotaxis proteins. This locus probably encodes proteins required for flagellum biosynthesis and other proteins that are required for the methylation response.
Full text
PDF




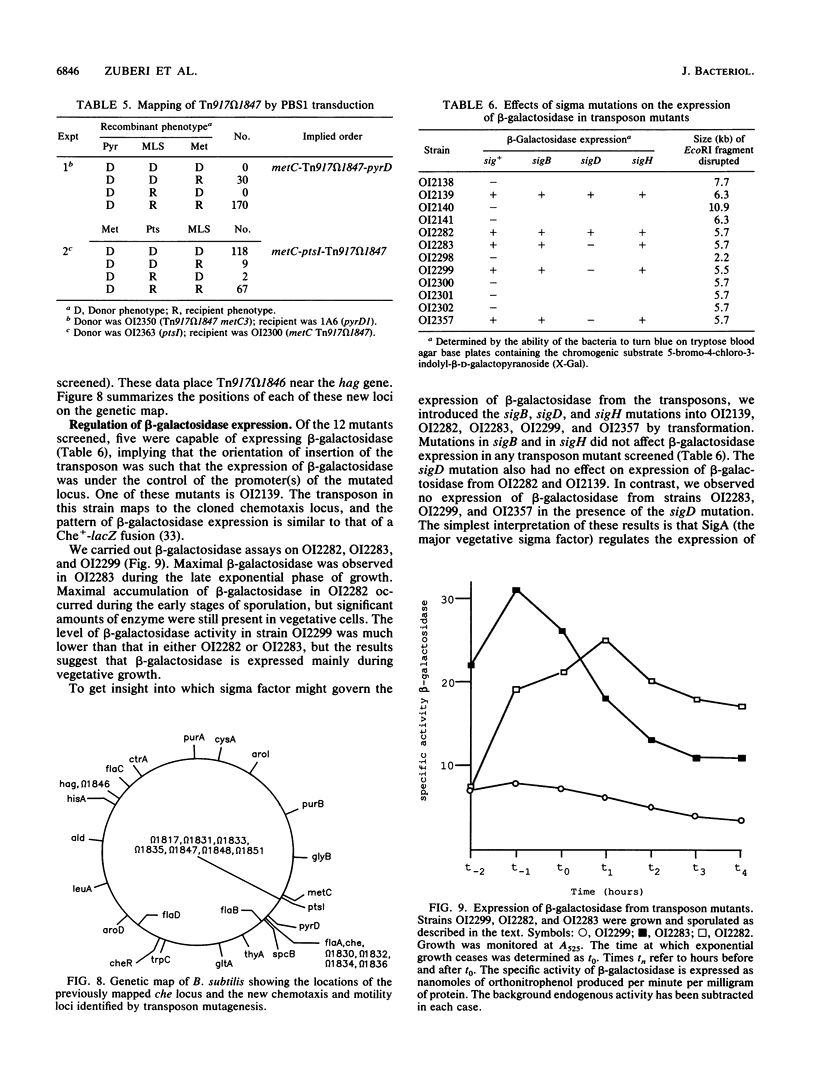


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alam M., Lebert M., Oesterhelt D., Hazelbauer G. L. Methyl-accepting taxis proteins in Halobacterium halobium. EMBO J. 1989 Feb;8(2):631–639. doi: 10.1002/j.1460-2075.1989.tb03418.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bourret R. B., Hess J. F., Simon M. I. Conserved aspartate residues and phosphorylation in signal transduction by the chemotaxis protein CheY. Proc Natl Acad Sci U S A. 1990 Jan;87(1):41–45. doi: 10.1073/pnas.87.1.41. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gaur N. K., Dubnau E., Smith I. Characterization of a cloned Bacillus subtilis gene that inhibits sporulation in multiple copies. J Bacteriol. 1986 Nov;168(2):860–869. doi: 10.1128/jb.168.2.860-869.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldman D. J., Ordal G. W. In vitro methylation and demethylation of methyl-accepting chemotaxis proteins in Bacillus subtilis. Biochemistry. 1984 Jun 5;23(12):2600–2606. doi: 10.1021/bi00307a010. [DOI] [PubMed] [Google Scholar]
- Kehry M. R., Doak T. G., Dahlquist F. W. Stimulus-induced changes in methylesterase activity during chemotaxis in Escherichia coli. J Biol Chem. 1984 Oct 10;259(19):11828–11835. [PubMed] [Google Scholar]
- Kutsukake K., Ohya Y., Iino T. Transcriptional analysis of the flagellar regulon of Salmonella typhimurium. J Bacteriol. 1990 Feb;172(2):741–747. doi: 10.1128/jb.172.2.741-747.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mesibov R., Adler J. Chemotaxis toward amino acids in Escherichia coli. J Bacteriol. 1972 Oct;112(1):315–326. doi: 10.1128/jb.112.1.315-326.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mirel D. B., Chamberlin M. J. The Bacillus subtilis flagellin gene (hag) is transcribed by the sigma 28 form of RNA polymerase. J Bacteriol. 1989 Jun;171(6):3095–3101. doi: 10.1128/jb.171.6.3095-3101.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Márquez L. M., Helmann J. D., Ferrari E., Parker H. M., Ordal G. W., Chamberlin M. J. Studies of sigma D-dependent functions in Bacillus subtilis. J Bacteriol. 1990 Jun;172(6):3435–3443. doi: 10.1128/jb.172.6.3435-3443.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohnishi K., Kutsukake K., Suzuki H., Iino T. Gene fliA encodes an alternative sigma factor specific for flagellar operons in Salmonella typhimurium. Mol Gen Genet. 1990 Apr;221(2):139–147. doi: 10.1007/BF00261713. [DOI] [PubMed] [Google Scholar]
- Ordal G. W., Goldman D. J. Chemotaxis away from uncouplers of oxidative phosphorylation in Bacillus subtilis. Science. 1975 Sep 5;189(4205):802–805. doi: 10.1126/science.808854. [DOI] [PubMed] [Google Scholar]
- Ordal G. W., Nettleton D. O., Hoch J. A. Genetics of Bacillus subtilis chemotaxis: isolation and mapping of mutations and cloning of chemotaxis genes. J Bacteriol. 1983 Jun;154(3):1088–1097. doi: 10.1128/jb.154.3.1088-1097.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ordal G. W., Parker H. M., Kirby J. R. Complementation and characterization of chemotaxis mutants of Bacillus subtilis. J Bacteriol. 1985 Nov;164(2):802–810. doi: 10.1128/jb.164.2.802-810.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ordal G. W., Villani D. P., Gibson K. J. Amino acid chemoreceptors of Bacillus subtilis. J Bacteriol. 1977 Jan;129(1):156–165. doi: 10.1128/jb.129.1.156-165.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sekiguchi J., Ezaki B., Kodama K., Akamatsu T. Molecular cloning of a gene affecting the autolysin level and flagellation in Bacillus subtilis. J Gen Microbiol. 1988 Jun;134(6):1611–1621. doi: 10.1099/00221287-134-6-1611. [DOI] [PubMed] [Google Scholar]
- Springer M. S., Goy M. F., Adler J. Sensory transduction in Escherichia coli: a requirement for methionine in sensory adaptation. Proc Natl Acad Sci U S A. 1977 Jan;74(1):183–187. doi: 10.1073/pnas.74.1.183. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thoelke M. S., Casper J. M., Ordal G. W. Methyl group turnover on methyl-accepting chemotaxis proteins during chemotaxis by Bacillus subtilis. J Biol Chem. 1990 Feb 5;265(4):1928–1932. [PubMed] [Google Scholar]
- Thoelke M. S., Kirby J. R., Ordal G. W. Novel methyl transfer during chemotaxis in Bacillus subtilis. Biochemistry. 1989 Jun 27;28(13):5585–5589. doi: 10.1021/bi00439a037. [DOI] [PubMed] [Google Scholar]
- Thoelke M. S., Parker H. M., Ordal E. A., Ordal G. W. Rapid attractant-induced changes in methylation of methyl-accepting chemotaxis proteins in Bacillus subtilis. Biochemistry. 1988 Nov 1;27(22):8453–8457. doi: 10.1021/bi00422a024. [DOI] [PubMed] [Google Scholar]
- Toews M. L., Goy M. F., Springer M. S., Adler J. Attractants and repellents control demethylation of methylated chemotaxis proteins in Escherichia coli. Proc Natl Acad Sci U S A. 1979 Nov;76(11):5544–5548. doi: 10.1073/pnas.76.11.5544. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tso W. W., Adler J. Negative chemotaxis in Escherichia coli. J Bacteriol. 1974 May;118(2):560–576. doi: 10.1128/jb.118.2.560-576.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ullah A. H., Ordal G. W. In vivo and in vitro chemotactic methylation in Bacillus subtilis. J Bacteriol. 1981 Feb;145(2):958–965. doi: 10.1128/jb.145.2.958-965.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang L. F., Doi R. H. Developmental expression of three proteins from the first gene of the RNA polymerase sigma 43 operon of Bacillus subtilis. J Bacteriol. 1987 Sep;169(9):4190–4195. doi: 10.1128/jb.169.9.4190-4195.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ward J. B., Jr, Zahler S. A. Genetic studies of leucine biosynthesis in Bacillus subtilis. J Bacteriol. 1973 Nov;116(2):719–726. doi: 10.1128/jb.116.2.719-726.1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zuberi A. R., Ying C. W., Weinreich M. R., Ordal G. W. Transcriptional organization of a cloned chemotaxis locus of Bacillus subtilis. J Bacteriol. 1990 Apr;172(4):1870–1876. doi: 10.1128/jb.172.4.1870-1876.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]