Abstract
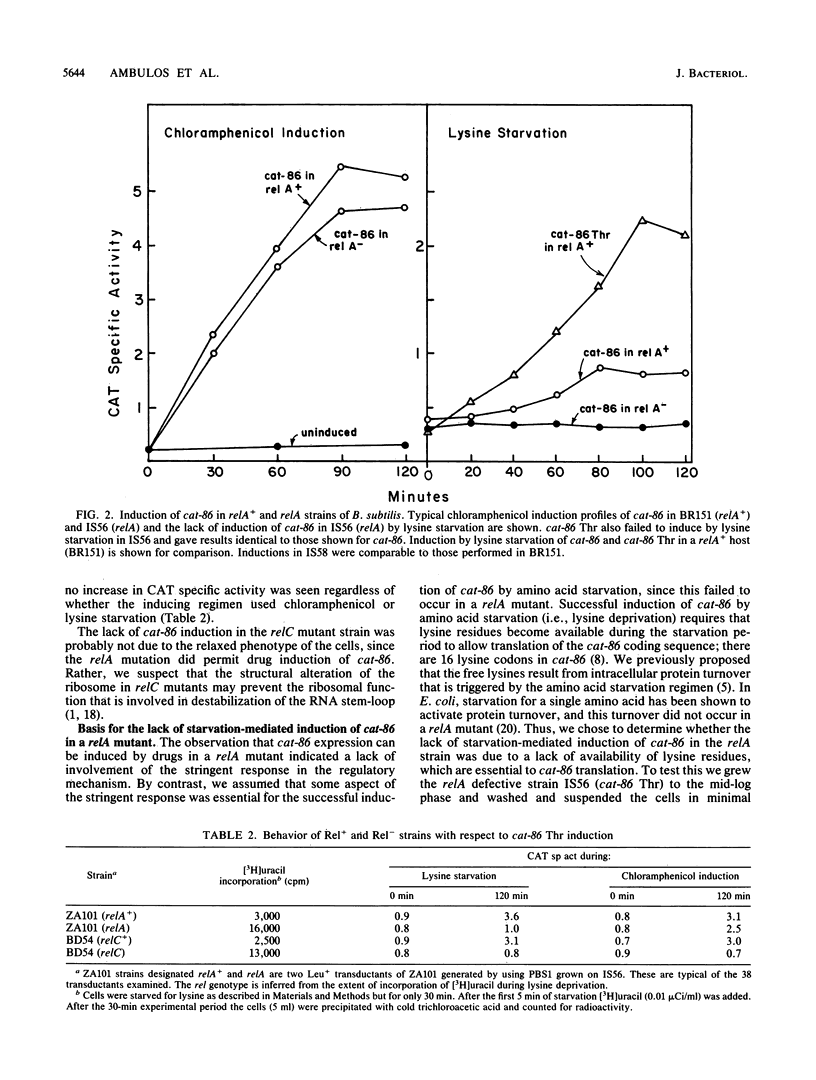
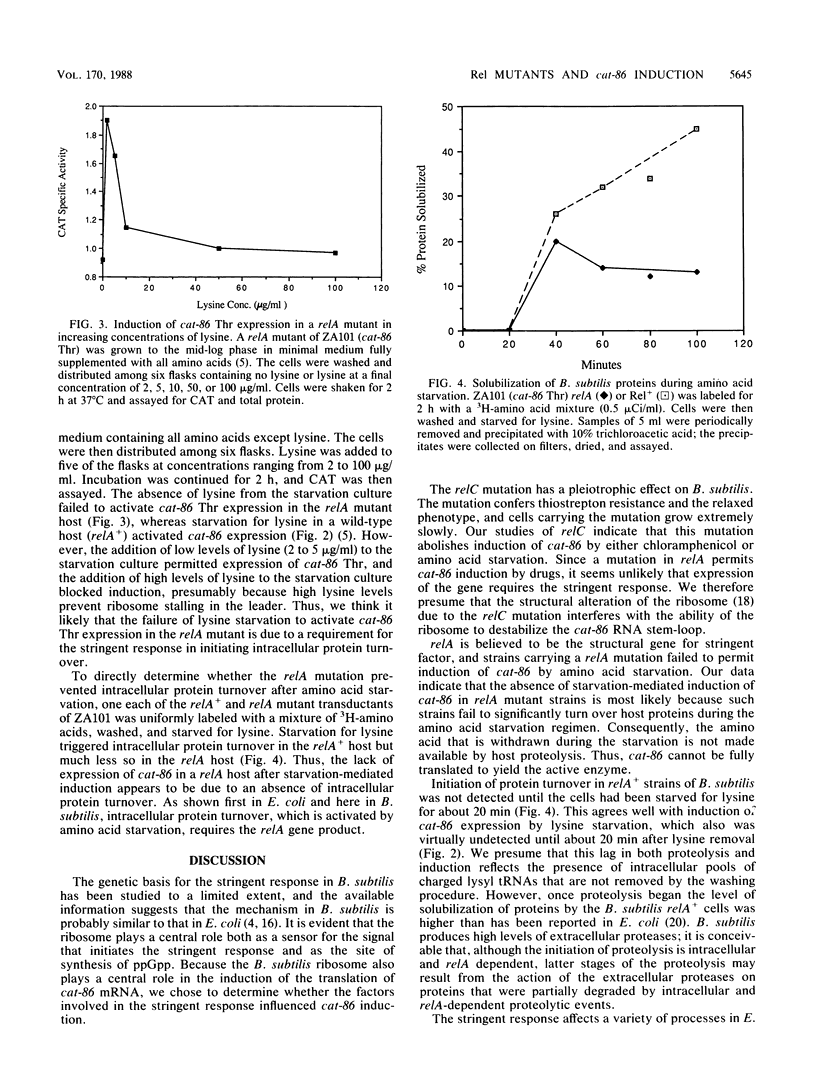
The chloramphenicol acetyltransferase gene cat-86 is induced through a mechanism that is a variation of classical attenuation. Induction results from the destabilization of an RNA stem-loop that normally sequesters the cat-86 ribosome-binding site. Destabilization of the stem-loop is due to the stalling of a ribosome in the leader region of cat-86 mRNA at a position that places the A site of the stalled ribosome at leader codon 6. Two events can stall ribosomes at the correct location to induce cat-86 translation: addition of chloramphenicol to cells and starvation of cells for the amino acid specified by leader codon 6. Induction by amino acid starvation is an anomaly because translation of the cat-86 coding sequence requires all 20 amino acids. To explain this apparent contradiction we postulated that amino acid starvation triggers intracellular proteolysis, thereby providing levels of the deprived amino acid sufficient for cat-86 translation. Here we show that a mutation in relA, the structural gene for stringent factor, blocks intracellular proteolysis that is normally triggered by amino acid starvation. The relA mutation also blocks induction of cat-86 by amino acid starvation, but the mutation does not interfere with chloramphenicol induction. Induction by amino acid starvation can be demonstrated in relA mutant cells if the depleted amino acid is restored at very low levels (e.g., 2 micrograms/ml). A mutation in relC, which may be the gene for ribosomal protein L11, blocks induction of cat-86 by either chloramphenicol or amino acid starvation. We believe this effect is due to a structural alteration of the ribosome resulting from the relC mutation and not to the relaxed phenotype of the cells.
Full text
PDF


Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alexieva Z., Duvall E. J., Ambulos N. P., Jr, Kim U. J., Lovett P. S. Chloramphenicol induction of cat-86 requires ribosome stalling at a specific site in the leader. Proc Natl Acad Sci U S A. 1988 May;85(9):3057–3061. doi: 10.1073/pnas.85.9.3057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bott K. F., Wilson G. A. Development of competence in the Bacillus subtilis transformation system. J Bacteriol. 1967 Sep;94(3):562–570. doi: 10.1128/jb.94.3.562-570.1967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bradford M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976 May 7;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Duvall E. J., Ambulos N. P., Jr, Lovett P. S. Drug-free induction of a chloramphenicol acetyltransferase gene in Bacillus subtilis by stalling ribosomes in a regulatory leader. J Bacteriol. 1987 Sep;169(9):4235–4241. doi: 10.1128/jb.169.9.4235-4241.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duvall E. J., Lovett P. S. Chloramphenicol induces translation of the mRNA for a chloramphenicol-resistance gene in Bacillus subtilis. Proc Natl Acad Sci U S A. 1986 Jun;83(11):3939–3943. doi: 10.1073/pnas.83.11.3939. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duvall E. J., Mongkolsuk S., Kim U. J., Lovett P. S., Henkin T. M., Chambliss G. H. Induction of the chloramphenicol acetyltransferase gene cat-86 through the action of the ribosomal antibiotic amicetin: involvement of a Bacillus subtilis ribosomal component in cat induction. J Bacteriol. 1985 Feb;161(2):665–672. doi: 10.1128/jb.161.2.665-672.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harwood C. R., Williams D. M., Lovett P. S. Nucleotide sequence of a Bacillus pumilus gene specifying chloramphenicol acetyltransferase. Gene. 1983 Oct;24(2-3):163–169. doi: 10.1016/0378-1119(83)90076-8. [DOI] [PubMed] [Google Scholar]
- Henner D. J., Hoch J. A. The Bacillus subtilis chromosome. Microbiol Rev. 1980 Mar;44(1):57–82. doi: 10.1128/mr.44.1.57-82.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lovett P. S., Keggins K. M. Bacillus subtilis as a host for molecular cloning. Methods Enzymol. 1979;68:342–357. doi: 10.1016/0076-6879(79)68025-4. [DOI] [PubMed] [Google Scholar]
- Lovett P. S., Young F. E. Genetic analysis in Bacillus pumilus by PBSI-mediated transduction. J Bacteriol. 1970 Feb;101(2):603–608. doi: 10.1128/jb.101.2.603-608.1970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Price V. L., Gallant J. A. Bacillus subtilis relG mutant: defect in glucose uptake. J Bacteriol. 1983 Jan;153(1):270–273. doi: 10.1128/jb.153.1.270-273.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shaw W. V. Chloramphenicol acetyltransferase from chloramphenicol-resistant bacteria. Methods Enzymol. 1975;43:737–755. doi: 10.1016/0076-6879(75)43141-x. [DOI] [PubMed] [Google Scholar]
- Smith I., Paress P., Cabane K., Dubnau E. Genetics and physiology of the rel system of Bacillus subtilis. Mol Gen Genet. 1980;178(2):271–279. doi: 10.1007/BF00270472. [DOI] [PubMed] [Google Scholar]
- Smith I., Paress P., Pestka S. Thiostrepton-resistant mutants exhibit relaxed synthesis of RNA. Proc Natl Acad Sci U S A. 1978 Dec;75(12):5993–5997. doi: 10.1073/pnas.75.12.5993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swanton M., Edlin G. Isolation and characterization of an RNA relaxed mutant of B. subtilis. Biochem Biophys Res Commun. 1972 Jan 31;46(2):583–588. doi: 10.1016/s0006-291x(72)80179-7. [DOI] [PubMed] [Google Scholar]
- Voellmy R., Goldberg A. L. Guanosine-5'-diphosphate-3'-diphosphate (ppGpp) and the regulation of protein breakdown in Escherichia coli. J Biol Chem. 1980 Feb 10;255(3):1008–1014. [PubMed] [Google Scholar]
- Yansura D. G., Henner D. J. Use of the Escherichia coli lac repressor and operator to control gene expression in Bacillus subtilis. Proc Natl Acad Sci U S A. 1984 Jan;81(2):439–443. doi: 10.1073/pnas.81.2.439. [DOI] [PMC free article] [PubMed] [Google Scholar]



