Abstract
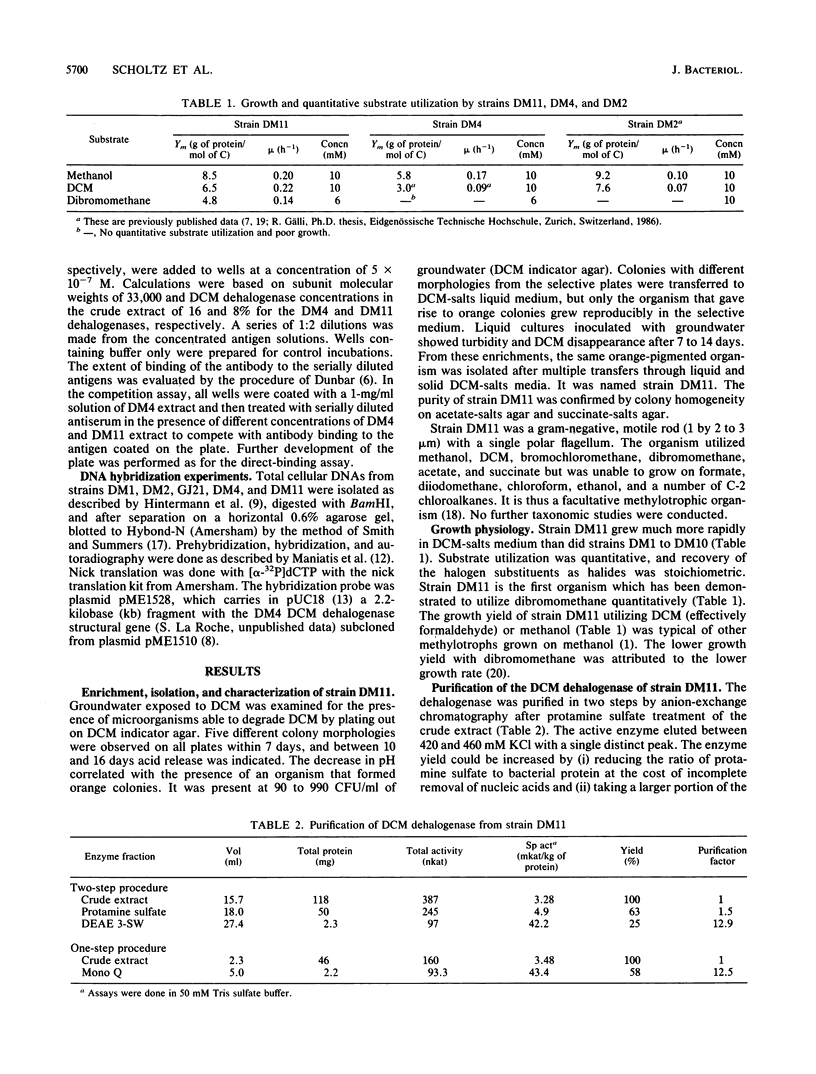
A methylotrophic bacterium, denoted strain DM11, was isolated from groundwater and shown to utilize dichloromethane or dibromomethane as the sole carbon and energy source. The new isolate grew at the high rate of 0.22 h-1 compared with 11 previously characterized dichloromethane-utilizing bacteria (micromax, 0.08 h-1). The dichloromethane dehalogenase from strain DM11 (group B enzyme) was purified by anion-exchange chromatography. It was shown to be substantially different from the set of dichloromethane dehalogenases from the 11 slow-growing strains (group A enzymes) that had previously been demonstrated to be identical. The Vmax for the group B enzyme was 97 mkat/kg of protein, some 5.6-fold higher than that of the group A enzymes. The group A dehalogenases showed hyperbolic saturation with the cosubstrate glutathione, whereas the group B enzyme showed positive cooperativity in glutathione binding. Only 1 of 15 amino acids occupied common positions at the N termini, and amino acid contents were substantially different in group A and group B dehalogenases. Immunological assays demonstrated weak cross-reactivity between the two enzymes. Despite the observed structural and kinetic differences, there is potentially evolutionary relatedness between group A and group B enzymes, as indicated by (i) hybridization of DM11 DNA with a gene probe of the group A enzyme, (ii) a common requirement for glutathione in catalysis, and (iii) similar subunit molecular weights of about 34,000.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bradford M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976 May 7;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Devereux J., Haeberli P., Smithies O. A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 1):387–395. doi: 10.1093/nar/12.1part1.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gälli R., Leisinger T. Plasmid analysis and cloning of the dichloromethane-utilization genes of Methylobacterium sp. DM4. J Gen Microbiol. 1988 Apr;134(4):943–952. doi: 10.1099/00221287-134-4-943. [DOI] [PubMed] [Google Scholar]
- Kohler-Staub D., Leisinger T. Dichloromethane dehalogenase of Hyphomicrobium sp. strain DM2. J Bacteriol. 1985 May;162(2):676–681. doi: 10.1128/jb.162.2.676-681.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Norrander J., Kempe T., Messing J. Construction of improved M13 vectors using oligodeoxynucleotide-directed mutagenesis. Gene. 1983 Dec;26(1):101–106. doi: 10.1016/0378-1119(83)90040-9. [DOI] [PubMed] [Google Scholar]
- Scholtz R., Leisinger T., Suter F., Cook A. M. Characterization of 1-chlorohexane halidohydrolase, a dehalogenase of wide substrate range from an Arthrobacter sp. J Bacteriol. 1987 Nov;169(11):5016–5021. doi: 10.1128/jb.169.11.5016-5021.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith G. E., Summers M. D. The bidirectional transfer of DNA and RNA to nitrocellulose or diazobenzyloxymethyl-paper. Anal Biochem. 1980 Nov 15;109(1):123–129. doi: 10.1016/0003-2697(80)90019-6. [DOI] [PubMed] [Google Scholar]
- Tempest D. W., Neijssel O. M. The status of YATP and maintenance energy as biologically interpretable phenomena. Annu Rev Microbiol. 1984;38:459–486. doi: 10.1146/annurev.mi.38.100184.002331. [DOI] [PubMed] [Google Scholar]




