Abstract
Heparan sulfate has been isolated for the first time from the mosquito Anopheles stephensi, a known vector for Plasmodium parasites, the causative agents of malaria. Chondroitin sulfate, but not dermatan sulfate or hyaluronan, was also present in the mosquito. The glycosaminoglycans were isolated, from salivary glands and midguts of the mosquito in quantities sufficient for disaccharide microanalysis. Both of these organs are invaded at different stages of the Plasmodium life cycle. Mosquito heparan sulfate was found to contain the critical trisulfated disaccharide sequence, →4)β-D-GlcNS6S(1 → 4)-α-L-IdoA2S(1→, that is commonly found in human liver heparan sulfate, which serves as the receptor for apolipoprotein E and is also believed to be responsible for binding to the circumsporozoite protein found on the surface of the Plasmodium sporozoite. The heparan sulfate isolated from the whole mosquito binds to circumsporozoite protein, suggesting a role within the mosquito for infection and transmission of the Plasmodium parasite.
Malaria infection is initiated with the bite of an infected anophelene mosquito, which injects Plasmodium sporozoites into the skin of the vertebrate host (Fig. 1A). Sporozoites migrate from the site of injection, enter the blood stream, and are carried to the liver, where they invade hepatocytes and develop into exoerythrocytic forms. Many lines of evidence suggest that clearance of sporozoites by the liver is mediated by the binding of the sporozoite major surface protein, the circumsporozoite protein (CSP),2 to heparan sulfate proteoglycans in the liver (1, 2). Liver heparan sulfate is more highly sulfated than heparan sulfate from other organs (3–6) and frequently contains the fully biosynthetically modified, highly sulfated sequence: →4)-GlcNS6S(1 →4)IdoA2S(1→(Fig. 2). Such fully modified sequences, while commonly found in the mast cell glycosaminoglycan, heparin, are infrequently found in heparan sulfate. Some heparan sulfates, such as those present in the liver, have a richer content of fully modified sequences in their high sulfate domains. These sequences have been shown to bind to apoE and facilitate the clearance of lipoprotein remnants from the blood by the liver (7, 8). ApoE containing remnant lipoproteins inhibits Plasmodium sporozoite infectivity and decreases clearance of CSP from the blood circulation, suggesting that sporozoites bind to the same hepatic heparan sulfate proteoglycans as remnant lipoproteins (9). This hypothesis is further supported by the finding that CSP binds to the same decasaccharide structure that binds to apoE (10, 11).
FIGURE 1. The life cycle of Plasmodium in the mammalian host (A) and the mosquito (B).

A, sporozoites are injected during blood feeding by infected anophelene mosquitoes, go to the liver, and invade hepatocytes, where they divide, rupture from the hepatocyte, and enter erythrocytes. The released merozoites invade erythrocytes, mature, divide, rupture from the cell, and enter new erythrocytes. The asexual erythrocytic cycle gives rise to the clinical symptoms associated with malaria. Some merozoites differentiate to gametocytes, which are infective for mosquitoes. This image is reprinted with permission from Ref. 51. B, mosquitoes become infected with Plasmodium when they ingest gametocytes (1) during blood feeding. In the midgut (MG) of the mosquito, these cells differentiate (2), fertilization occurs, and the resultant zygote differentiates into an ookinete (3), which traverses the midgut wall (4) and comes to rest extracellularly between the basal lamina and the midgut (5). The ookinete then differentiates into an oocyst, and sporozoites develop (6), and when mature, they emerge into the hemocoel (HC) of the mosquito and invade salivary glands (SG) (7). When the mosquito takes another blood meal, these salivary gland sporozoites are injected into the mammalian host (8), thus continuing the cycle. This image is reprinted with permission from Ref. 52.
FIGURE 2. Structure of heparan sulfate glycosaminoglycans.
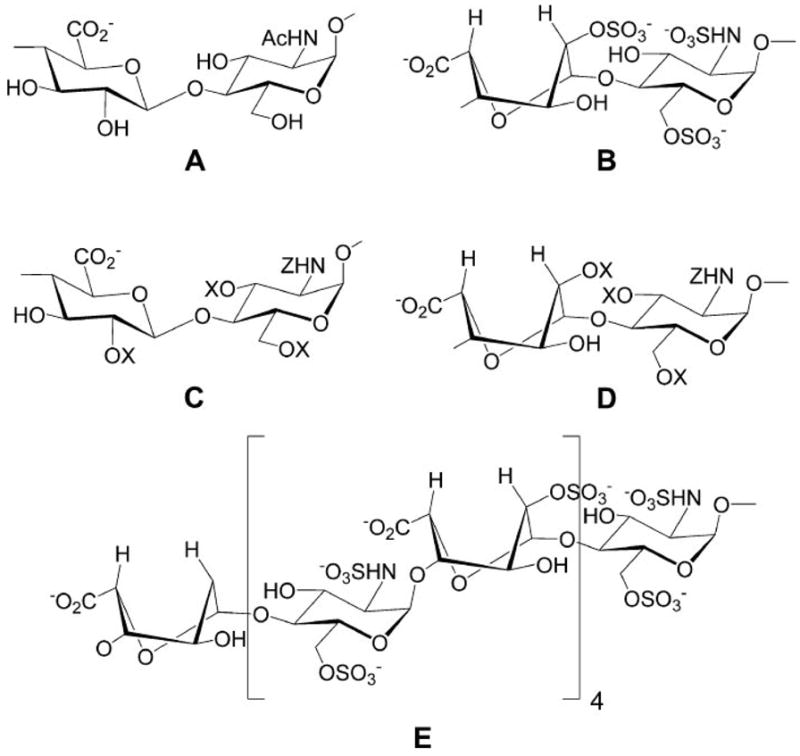
A, unsulfated disaccharide sequence prominent in heparan sulfate. B, fully sulfated disaccharide sequence prominent in heparin. C, structural variants of glucuronic acid containing disaccharide sequences found in heparan sulfate and heparin (X = SO3−, H). D, structural variants of iduronic acid containing disaccharide sequences found in heparan sulfate and heparin (X = SO3−, H; Z = SO3−, COCH3, H). E, proposed apoE and CSP binding domain.
Less is known about the role of proteoglycans in the mosquito host, where the parasite encounters a variety of mosquito matrices as it develops (Fig. 1B). When the mosquito takes a blood meal from an infected vertebrate host, gametocytes present in the blood meal differentiate, and fertilization occurs. The zygote develops into an ookinete, which crosses the midgut wall, stops at the basal lamina of the midgut, and develops into an oocyst. There is evidence that ookinetes recognize and adhere to specific sugars on the luminal side of the midgut (12, 13) and that recognition of the basal lamina by the migrating ookinete is a factor in its differentiation into an oocyst (14). Sporozoites develop in oocysts, emerge into the hemolymph, and invade salivary glands, where they wait for the mosquito to take another blood meal. Previous studies have shown that salivary gland recognition and invasion are probably receptor-mediated processes (15–17) and can be blocked by lectins (18), suggesting that carbohydrates are involved. We have previously shown that CSP binds specifically to salivary glands and not to other mosquito organs (19), raising the possibility that heparan sulfate proteoglycans are involved in target cell recognition in the mosquito just as they are in the vertebrate host.
Although the glycosaminoglycans in Drosophila melanogaster have been studied, there is currently no information on either the presence or the structure of glycosaminoglycans in the mosquito or any other insects of medical importance. The recent sequencing of the mosquito genome (20) as well as analysis of the salivary gland transcriptome and proteome (21) show the presence of both the core protein syndecan and the appropriate heparan sulfate biosynthetic enzymes (22), suggesting that heparan sulfate proteoglycans are present in mosquitoes. Anopheles stephensi was selected for the current study, because in addition to its being a vector of human malaria (23), it is widely used in the laboratory to study Plasmodium-vector interactions, since it can support the growth of both human and rodent malaria species. Here we determine the glycosaminoglycans present in this mosquito species using ultrasensitive disaccharide analysis (24) and examine their similarity to mammalian liver heparan sulfate and their binding to CSP.
EXPERIMENTAL PROCEDURES
Materials
Actinase E was obtained from Kaken Pharmaceutical Co. Ltd. (Tokyo, Japan). The following reagents were purchased from Sigma: urea, CHAPS, N-hydroxysulfosuccinimide sodium salt, N-(3-dimethylaminopropyl)-N′-ethylcarbodiimide, SDS, DNase I, endoglycosidase H (from Streptococcus plicatus, recombinant expressed in Escherichia coli), α-mannosidase (from Canavalia ensiformis), β-mannosidase (from Helix pomatia), β-N-acetylglucosaminidase (from C. ensiformis). Bovine serum albumin was from Amresco (Solon, OH). Chondroitinase ABC (EC 4.2.2.4), chondroitinase ACII (EC 4.2.2.5), chondro-4-sulfatase (EC 3.1.6.9), heparin lyase I (EC 4.2.2.7, heparinase I, heparinase), heparin lyase II (heparinase II, heparitinase II), and heparin lyase III (EC 4.2.2.8, heparinase III, heparitinase I) were obtained from Sigma and Seikagaku Kogyo. Heparin and heparan sulfate, both from porcine intestine, were from Celsus Laboratories. Heparan sulfate from porcine liver was prepared as previously described (4). Recombinant CSP was made in E. coli (DH5α) transfected with pCS27IVC. The plasmid contains the complete P. falciparum CSP sequence from the T4 isolate, except that the hydrophobic NH2- and COOH-terminal amino acids 1–26 and 412–424 have been deleted, and 5 histidine residues were added to the COOH terminus to facilitate purification. The protein used in this study was kindly provided by Dr. Bela Takacs (Hoffman-La Roche, Basel, Switzerland) (25).
Standard unsaturated heparan sulfate disaccharides were purchased from Sigma: 2-acetamido-2-deoxy-4-O-(4-deoxy-α-L-threo-hexenepyranosyluronic acid)-D-glucose (ΔUA-GlcNAc), 2-deoxy-2-sulfamido-4-O-(4-deoxy-α-L-threo-hexenepyranosyluronic acid)-D-glucose (ΔUA-GlcNS), 2-acetamido-2-deoxy-4-O-(4-deoxy-α-L-threo-hexenepyranosyluronic acid)-6-O-sulfo-D-glucose (ΔUA-GlcNAc6S), 2-acetamido-2-deoxy-4-O-(4-deoxy-2-O-sulfo-α-L-threo-hexenepyranosyluronic acid)-D-glucose (ΔUA2S-GlcNAc), 2-deoxy-2-sulfamido-4-O-(4-deoxy-2-O-sulfo-α-L-threo-hexenepyranosyluronic acid)-6-O-sulfo-D-glucose (ΔUA-GlcNS6S), 2-deoxy-2-sulfamido-4-O-(4-deoxy-2-O-sulfo-α-L-threo-hexenepyranosyluronic acid)-D-glucose (ΔUA2S-GlcNS), 2-acetamido-2-deoxy-4-O-(4-deoxy-2-O-sulfo-α-L-threo-hexenepyranosyluronic acid)-6-O-sulfo-D-glucose (ΔUA2S-GlcNAc6S), 2-deoxy-2-sulfamido-4-O-(4-deoxy-2-O-sulfo-α-L-threo-hexenepyranosyluronic acid)-6-O-sulfo-D-glucose (ΔUA2S-GlcNS6S), and heparan sulfate (from bovine kidney). Standard unsaturated disaccharides from chondroitin sulfate were obtained from Seikagaku Kogyo Co. (Tokyo): 2-acetamido-2-deoxy-3-O-(β-D-gluco-4-enepyranosyluronic acid)-D-glucose (ΔDi-HA), 2-acetamido-2-deoxy-3-O-(Δ-D-gluco-4-enepyranosyluronic acid)-D-galactose (ΔDi-0S), 2-acetamido-2-deoxy-3-O-(β-D-gluco-4-enepyranosyluronic acid)-4-O-sulfo-D-galactose (ΔDi-4S), 2-acetamido-2-deoxy-3-O-(β-D-gluco-4-enepyranosyluronic acid)-6-sulfo-D-galactose (ΔDi-6S), 2-acetamido-2-deoxy-3-O-(2-O-sulfo-β-D-gluco-4-enepyranosyluronic acid)-D-galactose (ΔDi-UA2S), 2-acetamido-2-deoxy-3-O-(2-O-sulfo-β-D-gluco-4-enepyranosyluronic acid)-4-O-sulfo-D-galactose (ΔDi-diSB), 2-acetamido-2-deoxy-3-O-(2-O-sulfo-β-D-gluco-4-enepyranosyluronic acid)-6-O-sulfo-D-galactose (ΔDi-diSD), 2-acetamido-2-deoxy-3-O-(β-D-gluco-4-enepyranosyluronic acid)-4,6-di-O-sulfo-D-ga-lactose (ΔDi-diSE), and 2-acetamido-2-deoxy-3-O-(2-O-sulfo-β-D-gluco-4-enepyranosyluronic acid)-4,6-di-O-sulfo-D-galactose (ΔDi-triS) were obtained from Sigma and Seikagaku Kogyo. Mr cut-off 3,000 dialysis tubing was purchased from Fisher. Bio-Gel P-10 polyacrylamide gels were purchased from Bio-Rad. The Vivapure Q column was purchased from Viva Science (Edgewood, NJ). CM5 sensor chip and Hepes-buffered saline (HBS-N; 0.01 M Hepes, pH 7.4, 0.15 M NaCl) were from Biacore. Senshu Pak Docosil (4.6 × 150 mm; particle size, 5 μm) was obtained from Senshu Scientific (Tokyo, Japan). Ultra-free-MC DEAE and Biomax-5 (5,000 normal Mr limit) were obtained from Millipore Corp. (Bedford, MA). All other chemicals used were of analytical reagent grade.
Apparatus
The chromatographic equipment included a gradient pump (L-7000), a chromato-integrator (D-7500) from Hitachi Instruments (Tokyo, Japan), a double plunger pump for the reagent solution and reaction heater (Shimamura Instruments, Tokyo, Japan), a sample injector with a 20-μl loop (model 7125; Reodyne, Rohnert Park, CA), and a fluorescence spectrophotometer (FP-1580; JASCO, Tokyo, Japan). SPR measurements were performed on a BIAcore 3000 operated using BIAcore 3000 control and BIAevaluation software (version 4.1). 1H NMR spectra were acquired on a Varian 500 MHz spectrometer at 25 °C in D2O.
Insects
Adult A. stephensi were raised as previously described (26). Our analysis was performed with 10–14-day-old adult female mosquitoes that had been allowed to feed for 10 min on an anesthetized rabbit 1 week prior to the analysis. Each mosquito imbibed, on average, between 1 and 2 μl of blood. As controls, we also used age-matched, naive, female A. stephensi mosquitoes that had never had a blood meal. For organ harvest, mosquitoes were placed on ice, rinsed briefly in 70% ethanol, and transferred to buffered saline (150 mM NaCl, 4 mM KCl, 3 mM CaCl2, 1.8 mM NaHCO3, 0.6 mM MgCl2, 25 mM Hepes, pH 7). Dissections were performed in buffered saline, and dissected organs were placed into glass tubes and kept on ice. After dissection, tubes were spun at 200 × g for 5 min to pellet the dissected organs, excess buffer was removed, and the organs were frozen at − 80 °C until use. Whole mosquitoes were counted, washed in ethanol as above, rinsed in buffered saline, and then frozen at − 80 °C. Weights of midguts and salivary glands were measured using a Cahn microbalance (model C-34; ATI). Organs were dissected dry and then transferred for weighing using siliconized capillary pipettes. Midguts were weighed in groups of two, and pairs of salivary glands were weighed in groups of five.
Preparation of Crude Glycosaminoglycans for Microanalysis—
Mosquito samples for disaccharide analysis were first lyophilized to dryness. Lyophilized sample (5 mg) was homogenized with acetone (1 ml), and the homogenate was washed with excess acetone and dried. The pellet was extracted with a solution containing SDS (1.0 ml, 0.5%), NaOH (0.1 M), NaBH4 (0.8%) with constant stirring for 16 h at room temperature. Sodium acetate (200 μl of 1 M solution) and HCl (300 μl of 1 M solution) were then added, the solution was filtered, and HCl (200 μl, 1 M) was added to the filtrate. Insoluble material was removed by centrifugation at 2,500 × g for 10 min at 4 °C. Ethanol (7 ml) was added to the supernatant and chilled for 2 h at 0 °C, and the crude glycosaminoglycan fraction was collected by centrifugation at 2,500 × g for 10 min at 4 °C. The resulting precipitate was dissolved in 250 μl of water, and the crude glycosaminoglycan solution was stored frozen (24).
Disaccharide Microanalysis—
Crude glycosaminoglycan solution (20 μl) was diluted to 100 μl with water and used as the sample solution for chondroitin disaccharide microanalysis. Chondroitinase digestion was performed using Tris-acetate buffer (5 μl, 0.2 M, pH 8.0), and 10 μl of an aqueous solution containing chondroitinase ABC (50 mIU) and chondroitinase ACII (50 mIU) were added to a 20-μl portion of the sample solution and incubated at 37 °C for 3 h. An 8-μl portion of this mixture was analyzed by high performance liquid chromatography. For heparan sulfate microanalysis, crude glycosaminoglycan sample (230 μl) and sodium phosphate buffer (50 μl, 0.3 M, pH 6.0) were added, and the solution was applied on an Ultrafree-MC DEAE membrane, which had been equilibrated with sodium phosphate buffer (pH 6.0) containing NaCl (0.15 M). The fractions eluted with NaCl (1 M) in the same buffer were collected, desalted with Biomax-5, evaporated, and resuspended in 12 μl of water in preparation of heparin lyase digestion. Next, sodium acetate buffer (5 μl, 0.1 M, pH 7.0) containing calcium acetate (10 mM) and 15 μl of an aqueous solution containing heparin lyase mixture (Seikagaku), heparin lyase I (1 mIU), heparin lyase II (1 mIU), and heparin lyase III (1 mIU) were added to a 5-μl portion of sample. The mixture was incubated at 37 °C for 16 h, and an 8-μl aliquot was loaded onto the high performance liquid chromatograph (24).
Determination of Unsaturated Disaccharides from Heparan Sulfate and Chondroitin Sulfate—
Unsaturated disaccharides produced enzymatically from heparan sulfate and chondroitin sulfate were analyzed by a reversed-phase ion pair chromatography with sensitive and specific postcolumn detection. A gradient was applied at a flow rate of 1.1 ml/min using a Senshu Pak Docosil column (4.6 × 150 mm) at 55 °C. The eluents used were as follows: A, H2O; B, 0.2 M sodium chloride; C, 10 mM tetra-n-butylammonium hydrogen sulfate; D, 50% acetonitrile. The gradient program was as follows: 0–10 min, 70–67% eluent A, 1–4% eluent B; 10–11 min, 67–56% eluent A, 4–15% eluent B; 11–20 min, 56–46% eluent A, 15–25% eluent B; 20–22 min, 46–18% eluent A, 25–53% eluent B; 22–29 min, 18% eluent A, 53% eluent B; equilibration with 70% A and 1% B for 20 min. The proportions of eluent C and D were constant at 12 and 17%, respectively. A solution of 2-cyanoacetamide (aqueous, 0.5% (w/v)) and sodium hydroxide (0.25 M) was added at the same flow rate (0.35 ml/min) to the effluent using a double plunger pump. The mixture passed through a reaction coil (internal diameter, 0.5 mm; length, 10 m) set in a dry reaction temperature-controlled bath at 125 °C and a following cooling coil (internal diameter, 0.25 mm; length, 3 m). The effluent was monitored fluorometrically (excitation, 346 nm; emission, 410 nm) (24).
Large Scale Preparation and Purification of Heparan Sulfate from Whole Mosquitoes for Surface Plasmon Resonance (SPR) Studies—
Approximately 8000 mosquitoes were ground with dry ice, washed with acetone three times to remove lipids, and then dried in vacuo. Mosquito acetone powder was dissolved in 100 ml of water, and 20 ml of 0.05 M Tris-acetate buffer (pH 8.0) containing 1% actinase E was added and incubated at 45 °C overnight. After actinase E digestion, 2 ml of 100 mM acetic acid was added and boiled for 5 min in water bath and centrifuged at 1,000 × g for 20 min. The supernatant was dialyzed against water in Mr cut-off 3,000 dialysis tubing overnight and freeze-dried. The resulting peptidoglycan (glycosaminoglycan linked to residual core peptide) from mosquitoes was dissolved in 8 M urea, 2% CHAPS buffer (pH 8.3) and loaded onto the Vivapure Q spin column and centrifuged at 500 × g for 15 min. The column was then washed with 8 M urea, 2% CHAPS buffer and then washed five times with 50 mM NaCl. The bound peptidoglycan was released by washing the spin column three times with 2.7 M (16%) NaCl at 500 × g. Methanol (final concentration 80%) was added to the eluent, and the peptidoglycan component was allowed to precipitate overnight at 4 °C. The peptidoglycan fraction was collected by centrifugation at 2,500 × g for 20 min. Core peptide was removed by β-elimination, by dissolving the isolated peptidoglycan in 2 ml of 0.5% SDS, 0.1 M NaOH, 0.8% NaBH4, and incubated at room temperature for 16 h with constant stirring. Sodium acetate (400 μl of 1 M) and HCl (600 μl of 1 M) were then added, the solution was filtered, and HCl (400 μl of 1 M) was added to the filtrate. Finally, the solution was dialyzed against Mr cut-off 3,000 dialysis tubing overnight and lyophilized. To remove chondroitin sulfate, the resulting glycosaminoglycan sample was redissolved in 0.2 M Tris-HCl buffer and incubated with a mixture of chondroitinase ABC and AC II overnight at 37 °C. After digestion, the sample was dialyzed against Mr cut-off 3,000 dialysis tubing overnight to remove the chondroitin disaccharide that had been formed and then freeze-dried. Residual DNA was removed by dissolving the sample in DNase I reaction buffer (10 mM Tris-HCl, 2.5 mM MgCl2, 0.5 mM CaCl2, pH 7.6), incubating with DNase I at 37 °C overnight. Chromatography on a P-10 column (1 × 50 cm) eluted with water removed DNase and nucleotides, and intact glycosaminoglycan was recovered and lyophilized. 1H NMR spectroscopy (not shown) suggested the presence of residual contaminating mannose-containing glycans. These contaminating glycans were removed by dissolving the sample in 250 mM sodium phosphate buffer (pH 5.5) and incubating with endoglycosidase H at 37 °C for 48 h followed by α-mannosidase, β-mannosidase, and β-N-acetylglucosaminidase for 24 h at 25 °C. The sample was dialyzed against water in Mr cut-off 3,000 dialysis tubing overnight and lyophilized. All of the enzymes (proteins) that had been added were then removed by dissolving the sample in 8 M urea, 2% CHAPS buffer (pH 8.3) and passing through a Vivapure Q spin column at 500 ×g for 15 min. The column was washed with 8 M urea, 2% CHAPS buffer, followed by washing five times with 50 mM NaCl. The heparan sulfate was released from the column with three washes of 2.7 M (16%) NaCl at 500 × g. The heparan sulfate was precipitated at 4 °C overnight by adding absolute methanol (final concentration 80%). The pure heparan sulfate was collected by centrifugation at 2,500 × g for 20 min. A carbazole assay (27) showed that 532 μg of heparan sulfate was obtained.
Surface Plasmon Resonance Studies—
HBS-N buffer was used for immobilization and interaction studies. CM5 chips were activated with 50 mM N-hydroxysulfosuccinimide sodium salt and 200 mM N-(3-dimethylaminopropyl)-N′-ethylcarbodiimide mixture for 7 min at a flow rate of 5 μl/min. CSP was immobilized at a concentration of ~700 nM. The surfaces were deactivated with 1 M ethanolamine hydrochloride, pH 8.5. The final response units after ethanolamine was 8,000–10,000. The control surface was prepared the same way, except ~6000 response units of bovine serum albumin was immobilized. At a flow rate of 5 μl/min and temperature of 25 °C, the interaction between CSP and various concentrations (1, 5, 10, 20, and 50 μM) of heparan sulfate was studied, selecting an interaction with an association time of 3 min and dissociation time of 4 min. Between each injection, the surface was regenerated with 2 M NaCl. The interaction was monitored by plotting the increase in response over time (sensorgram), and the control flow cell was subtracted from the experimental flow cell. The curves were evaluated with BIAevaluation version 4.1.
RESULTS
Preliminary studies on salivary glands, prepared from adult female A. stephensi mosquitoes, showed that they contained both chondroitin sulfate and heparan sulfate. The amount of chondroitin sulfate in the mosquito, however, appeared to be considerably lower than that reported in the whole body of D. melanogaster third stage larvae (24). Based on these preliminary findings, hundreds of adult female A. stephensi mosquitoes and selected tissues from these mosquitoes were collected. Salivary glands, midguts, and malpighian tubules with ovaries were collected in buffered saline, centrifuged to remove excess buffer, frozen, and then ground into a fine powder for analysis. The total dry mass, prior to removing salts and lipids, of each sample is presented in Table 1. The dry mass of individual mosquitoes or organs was obtained by either dividing the total mass by the number of individuals or by obtaining wet weights on a Cahn microbalance and correcting for water content. Salivary glands (in pairs) and midguts corresponded to 0.6 and 1.2% of the total insect body mass, respectively.
TABLE 1.
Samples prepared for analysis
| Sample | Total mass | Number | Mass/insect or organ | Percentage of insect body mass |
|---|---|---|---|---|
| mg | μg | % | ||
| Whole A. stephensi mosquito | 71 | 150 | 340a | 100 |
| Salivary glands | 4.3 | 486 | 2.0b | 0.6 |
| Midguts | 3.7 | 247 | 4.1b | 1.2 |
| Malpighian tubules and pairs of ovaries | 10 | 320 | NDc | ND |
Based on the dry weight of 1,000 insects multiplied by 0.001.
Calculated from six independent determinations of the wet weight of either two midguts or five pairs of salivary glands. The dry wet weight of each organ was calculated by multiplying the wet weight by 0.25 (corresponding to a water content of 75%).
ND, not determined.
The whole insect and these isolated organs were extracted by a method previously developed for use with D. melanogaster (24) to recover total glycosaminoglycans. A single mosquito was found to contain 6.0 ng of chondroitin sulfate and 11.3 ng of heparan sulfate (Tables 2 and 3). We used mosquitoes that had taken a blood meal 1 week prior to the analysis, because blood feeding initiates changes in the mosquito’s transcriptional repertoire (28), which could, in turn, alter the composition of the cell barriers encountered by malaria parasites (15). Analysis of whole rabbit blood, however, showed that it contained 3–4 ng/μl glycosaminoglycans, making it theoretically possible that the blood meal contributed to the chondroitin sulfate and heparan sulfate detected in the mosquitoes. To control for this possibility, we compared the chondroitin sulfate and heparan sulfate content of blood-fed mosquitoes (1 week after blood feeding) with age-matched naive mosquitoes and found no difference in disaccharide composition between the two groups (data not shown).
TABLE 2.
Unsaturated disaccharides from chondroitin sulfate (CS)
| Unsaturated disaccharide
|
Amount of chondroitin sulfate
|
||||
|---|---|---|---|---|---|
| ΔDi-0S | ΔDi-4S | Sulfo group/disaccharide | ng of CS/mg of tissue | ng of CS/insect or organ | |
| mol % | ng/mg | ||||
| Female A. stephensi mosquitoes | 73.4 | 26.6 | 0.3 | 17.6 | 6.0 |
| Mosquito salivary glands | 78.1 | 21.9 | 0.3 | 21.6 | 0.043 |
| Mosquito midgets | 69.2 | 30.8 | 0.3 | 6.4 | 0.026 |
| Mosquito malpighian tubules and ovaries | 54.1 | 45.9 | 0.5 | 25.2 | NDa |
| D. melanogaster (adult)b | 70.8 | 29.2 | 0.3 | 126 | 25 |
ND, not determined.
The mass of a dry fruit fly is 200 μg (see Ref. 24).
TABLE 3.
Unsaturated disaccharides from heparan sulfate
| Amount of heparan-sulfated unsaturated disaccharide
|
|||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Heparan sulfate source | ΔUA- GlcNAc |
ΔUA- GlcNS |
ΔUA- GlcNAc6S |
ΔUA2S- GlcNAc |
ΔUA- GlcNS6S |
ΔUA2S- GlcNS |
ΔUA2S- GlcNAc6S |
ΔUA2S- GlcNS6S |
Sulfo group/disaccharide | Dry tissue | HS/insect or organ |
| mol % | ng/mg | ng | |||||||||
| Female A. stephensi mosquitoes | 38.9 | 24.4 | 4.8 | NDa | 19.5 | 9.1 | ND | 3.3 | 1.0 | 33.1 | 11.3 |
| Mosquito salivary glands | 41.3 | 22.8 | 3.2 | ND | 19.4 | 10.3 | ND | 3.0 | 0.9 | 23.0 | 0.045 |
| Mosquito midguts | 35.3 | 37.8 | 1.1 | ND | 11.1 | 12.7 | ND | 2.0 | 0.9 | 50.4 | 0.2 |
| qMosquito malpighian tubules and ovaries | 40.0 | 29.6 | 3.0 | ND | 16.6 | 9.1 | ND | 1.7 | 0.9 | 42.3 | ND |
| D. melanogaster (adult)b | 42.6 | 18.8 | 4.4 | ND | 15.4 | 14.5 | ND | 4.3 | 1.0 | 19.7 | 3.9 |
| Human liverc | 37.3 | 15.5 | 9.8 | 1.2 | 7.8 | 5.9 | 0.7 | 21.8 | 1.2 | ~8 | |
Next we performed a disaccharide analysis of whole mosquitoes and mosquito organs. Treatment of total purified intact glycosaminoglycans with chondroitin ABC lyase afforded chondroitin disaccharides. The resulting disaccharide mixture was analyzed by reversed-phase ion pairing high performance liquid chromatography (HPLC) using ultrasensitive detection following postcolumn fluorescence derivatization (24, 29) (Fig. 3 and Table 2). The whole insects and insect organs contained 54–78% ΔUA (1 → 3) GalNAc (0S) and 22–46% ΔUA (1 → 3) GalNAc4S (4S) disaccharide. Six of the eight possible chondroitin disaccharides were not present, since no traces of 6-O-sulfo, 2-O-sulfo, disulfated, or trisulfated chondroitin disaccharides could be observed. The chondroitinase-sensitive glycosaminoglycans obtained from mosquitoes contained no hyaluronic acid or dermatan sulfate. The presence of hyaluronan was ruled out using a graphite column that can discriminate between glucosaminoglycans and galactosaminoglycans (29). The presence of dermatan sulfate was ruled out, since the disaccharide analysis with chondroitinase AC was identical to that with chondroitinase ABC (24). These data on mosquito and mosquito organs are consistent with the known absence of dermatan sulfate and hyaluronan in D. melanogaster (24). Interestingly, the whole mosquito contained only one-fourth as much chondroitin sulfate as the fly on a per insect basis.
FIGURE 3. Reversed-phase ion pairing-HPLC determination of chondroitin lyase-derived glycosaminoglycan disaccharides.

A, female mosquitoes; B, salivary glands; C, midguts; D, malpighian tubules and ovaries. Peak 1, ΔDi-0S; peak 2, ΔDi-4S. Other conditions were as described under “Experimental Procedures.”
In contrast, the whole mosquito contained ~3 times as much heparan sulfate as the fly on a per insect basis. The same glycosaminoglycan samples from whole mosquito and isolated organs were collected and treated with a mixture of heparin lyase I, II, and III, to convert the heparan sulfate into disaccharides. Disaccharide analysis (Fig. 4 and Table 3) revealed that all samples contained six of the eight common heparan sulfate disaccharides, with ΔUA2S (1 → 4) GlcNAc (2SNAc) and ΔUA2S (1 → 4) GlcNAc6S (2SNAc6S) conspicuously absent. The ΔUA2S (1 → 4) GlcNS6S trisulfated disaccharide, probably arising from heparin lyase cleavage of the →4)-GlcNS6S(1 → 4)IdoA2S(1→ sequence, was found in all of the mosquito samples. No clear patterns or differences in disaccharide composition between different insect organs could be discerned. Comparison of the composition of these samples with human liver heparan sulfate clearly showed a lower degree of sulfation, 0.9–1.0 sulfo groups/disaccharide for the insect samples compared with 1.2 and 1.0 sulfo groups/disaccharide for the human and porcine liver heparan sulfate (5).
FIGURE 4. RPIP-HPLC determination of heparin lyase-derived glycosaminoglycan disaccharides.

A, female mosquitoes; B, salivary glands; C, midguts; D, malpighian tubules and ovaries. Peak 1, ΔUA-GlcNAc; peak 2, ΔUA-GlcNS; peak 3, ΔUA-GlcNAc6S; peak 4, ΔUA-GlcNS6S; peak 5, ΔUA2S-GlcNS; peak 6, ΔUA2S-GlcNS6S. Other conditions were as described under “Experimental Procedures.”
Domain mapping of whole mosquito heparan sulfate was undertaken by treatment with heparin lyase III, followed by polyacrylamide gel electrophoresis (30) (not shown). The results of this analysis demonstrated the presence of heparin lyase III-resistant (high sulfate) domains ranging in size from hexasaccharide to hexadecasaccharide.
Next, ~8,000 mosquitoes were collected and used to prepare ~0.5 mg of heparan sulfate required for CSP binding studies. A number of additional steps were added to ensure the purity of the heparan sulfate and the absence of contaminating nucleic acids, proteins, and non-heparan sulfate glycans. The interaction of the purified mosquito heparan sulfate with immobilized CSP was studied using SPR and compared with previous results obtained on the interaction of CSP with porcine heparin and heparan sulfate (10, 11). The resulting sensorgrams all afforded excellent fitting to a 1:1 conformational change model with compensation for mass transfer limitations. The sensorgrams for mosquito heparan sulfate binding to immobilized CSP are shown in Fig. 5. A single concentration, corresponding to 10 μM, was selected to compare the binding to CSP of mosquito heparan sulfate with that of heparin and other heparan sulfates (Table 4). Heparin bound with a much higher affinity than all other ligands, with porcine liver heparan sulfate binding with the second highest affinity. Mosquito heparan sulfate showed a CSP binding affinity comparable with porcine intestinal heparan sulfate. It is noteworthy that the heparan sulfate from mosquitoes, while fitting the 1:1 conformational change model quite well, gave very different binding kinetics from the samples prepared from mammalian tissues. The kinetics of mosquito heparan sulfate was dominated by its slow on-rate and off-rate, followed by a very slow conformational change.
FIGURE 5. Surface plasmon resonance studies on whole mosquito heparan sulfate binding to CSP.

Shown are sensorgrams at heparan sulfate (solid lines) concentrations of (beginning from the bottom solid curve) 1, 5, 10, 20, and 50 μM. The start of injection begins at 0 s and ends at 180 s (the arrow indicates stop of injection). The dashed curve represents interaction between CSP and 10 μM porcine liver HS.
TABLE 4.
SPR data for a 1:1 conformational change model with mass transport limitation at 10 μM analyte glycosaminoglycan concentration binding to immobilized CSP
| Sample | ka1 | kd1 | k2 | k−2 | Rmax | χ2 | KD |
|---|---|---|---|---|---|---|---|
| M−1s−1 | s−1 | s−1 | s−1 | RU | μM | ||
| Heparin | 2.3 × 104 | 5.0 × 1032 | 1.5 × 1032 | 3.4 × 1034 | 216 | 0.356 | 0.05 |
| Porcine liver HS | 1.1 × 104 | 5.1 × 1032 | 1.2 × 1032 | 5.7 × 1034 | 149 | 0.876 | 0.2 |
| Porcine intestinal HS | 2.9 × 103 | 2.8 × 1032 | 1.1 × 1032 | 1.9 × 1033 | 102 | 0.211 | 2 |
| Mosquito HS | 5.0 × 102 | 5.8 × 1034 | 5.7 × 1036 | 1.0 × 1035 | 131 | 0.277 | 2 |
DISCUSSION
The proteoglycans chondroitin sulfate and heparan sulfate, as well as the structurally related heparin (31), have been observed in a wide variety of animals, ranging from mammals (3–6, 32, 33), to birds (34), clams (35), fruit flies, and worms (24). Nothing, however, is known about these molecules in insects of medical importance, where they may play a role in the transmission of pathogens. The recent genome sequencing of two medically important mosquito species, A. gambiae and Aedes aegypti demonstrates that the genes encoding the biosynthetic machinery necessary for synthesis of heparan and chondroitin sulfates are present in these species. Our current work in A. stephensi, a species closely related to A. gambiae, demonstrates that these molecules are indeed biosynthesized and elucidates their fine structure.
Heparan sulfate is found linked to a number of core proteins, including members of the syndecan family (integral membrane proteins), the glypican family (GPI-anchored proteins), and the perlecan proteoglycans (extracellular matrix proteins) (36). The heparan sulfate glycosaminoglycan side chains of these proteoglycans serve an essential role in cell-cell interactions and cellular communication (37, 38). Of relevance to our findings, heparan sulfate glycosaminoglycan chains are involved in cell-cell adhesion by binding to a variety of different proteins, such as selectins, fibronectin, laminin, and vitronectin (38, 39). The interaction of heparan sulfate with heparin-binding proteins is believed to be specific and related to the sequence within heparan sulfate (40). The sequences within a given heparan sulfate chain are the result of its biosynthesis in the Golgi through the sequential action of N-deacetylase, N-sulfotransferase, C5 epimerase, and O-sulfotransferases. Interestingly, some of these enzymes have multiple isoforms that are differentially expressed in a tissue-dependent and a time-dependent fashion, leading to a complex and little understood mixture of glycotypes.
A number of infectious agents are known to use the negative charge of cell surface glycosaminoglycans to localize, enter, and infect animal cells (41, 42). Viruses, including dengue virus (43), herpes simplex virus 1 (44), hepatitis C virus (45, 46), and human immunodeficiency virus-1 (47), are known to utilize heparan sulfate as a receptor. Herpes simplex virus 1 targets heparan sulfate having a unique sequence found in nervous tissue (44) and requires 3-O-sulfation of specific glucosamine residues of heparan sulfate to enter cells (48), whereas dengue virus and hepatitis C virus target the highly sulfated heparan sulfate in liver (43, 46).
Recent studies have demonstrated that in the mammalian host, Plasmodium sporozoites also target the highly sulfated heparan sulfate of the liver, and binding to these heparan sulfate chains is required for attachment to hepatocytes (2, 49). This binding is mediated by CSP, the major surface protein of the sporozoite, which is probably also involved in sporozoite attachment to mosquito salivary glands (16, 19). The role of CSP in targeting sporozoites to their target organs in both the mammalian and mosquito hosts prompted us to determine whether mosquito salivary glands have heparan sulfate and, if so, whether it is similar to that found in the vertebrate liver. This study clearly shows, for the first time, that mosquitoes contain both heparan sulfate and chondroitin sulfate proteoglycans. These findings raise the possibility that Plasmodium sporozoites use heparan sulfate proteoglycans in both the mosquito and mammalian hosts to localize to their target organs. In addition, they may also be of relevance to other vector-borne pathogens. For example, dengue virus, which enters mammalian cells using heparan sulfate proteoglycans (43), is transmitted to humans by mosquitoes and may utilize the heparan sulfate in the mosquito to localize to the salivary glands. Recent analysis of the salivary gland transcriptome (21) has shown that syndecan is one of the most abundant transcripts in mosquito salivary glands, highlighting the importance of this observation.
The disaccharides comprising the sequence of mosquito heparan sulfate are not as diverse as those found in higher animals, since the mosquito contains only six of the eight common disaccharides found in human heparan sulfate. The mosquito does, however, contain the critical trisulfated disaccharide sequence, which binds to apoE in the mammalian liver (7, 8) and which sporozoites use to specifically attach to hepatocytes (9). There are also extended high sulfate domains in the mosquito heparan sulfate that would correlate well with the suggested decasaccharide heparin-binding site for CSP (10). One caveat, however, is that we found no obvious differences in the amount or type of heparan sulfate in salivary glands and midguts. Since Plasmodium sporozoites develop in oocysts on the mosquito midgut wall, it is important that they do not bind to the midgut upon exit from the oocyst but instead float freely in the hemolymph so that they can contact salivary glands, to which they bind and enter. We cannot, at this point, explain this specificity, but our finding that the midgut contains a significantly lower density of chondroitin sulfate chains compared with the salivary gland may explain the differential binding of sporozoites to these organs. This hypothesis is supported by the previous demonstration that CSP also binds to chondroitin sulfate (50), and we are currently investigating whether distinct domains of CSP have different binding affinities for distinct proteoglycans.
Previous studies using SPR have shown that mammalian heparin and liver heparan sulfate bind CSP (10, 11). Here we used SPR to analyze, the interaction between CSP and heparan sulfate from whole mosquitoes. Compared with its mammalian counterpart, mosquito heparan sulfate binds to CSP but with reduced affinity and different kinetics. Since only ~1% of total mosquito heparan sulfate is derived from tissues infected by the malaria parasite (Table 3), it is difficult to draw firm conclusions regarding the role of the binding differences we observed between whole mosquito heparan sulfate and mammalian liver heparan sulfate in malaria infection and transmission. It will be important to examine the interaction between CSP and glycosaminoglycans isolated from individual mosquito organs; however, it has been difficult to isolate sufficient amounts of heparan sulfate required for SPR studies from individual mosquito organs. Further studies to isolate and characterize larger amounts of intact glycosaminoglycans and oligosaccharides from mosquito salivary glands and midgut are currently under way in our laboratories.
In conclusion, we have characterized both heparan sulfate and chondroitin sulfate from Anopheles mosquitoes. In addition, the major surface protein of the Plasmodium sporozoite, the infectious stage of the malaria parasite that is transmitted by anophelene mosquitoes, binds to mosquito heparan sulfate. These data suggest that the similarities we have observed between mosquito and mammalian glycosaminoglycans may facilitate transmission of the sporozoite from the mosquito to the mammalian host.
Footnotes
This work was supported in part by National Institutes of Health Grants HL62244, HL52622, and GM38060 (to R. J. L.) and AI056840 (to P. S.).
The abbreviations used are: CSP, circumsporozoite protein; apoE, apolipoprotein E; GlcN, 2-deoxy-2 aminoglucopyranose; IdoA, idopyranosyluronic acid; S, sulfate; CHAPS, 3-[(3-cholamidopropyl)dimethylammonio]-1-propanesulfonic acid; ΔUA, 4-deoxy-α-L-threo-hexenepyranosyluronic acid; SPR, surface plasmon resonance; HPLC, high performance liquid chromatography; ΔDi, 2-acetamido-2-deoxy-3-O-(β-D-gluco-4-enopyranosy-luronic acid)-D-galactose(glucose).
References
- 1.Frevert U, Sinnis P, Cerami C, Shreffler W, Takacs B, Nussenzweig V. J Exp Med. 1993;177:1287–1298. doi: 10.1084/jem.177.5.1287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Pinzon-Ortiz C, Friedman J, Esko J, Sinnis P. J Biol Chem. 2001;276:26784–26791. doi: 10.1074/jbc.M104038200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Lyon M, Dakin JA, Gallagher JT. J Biol Chem. 1994;269:11208–11215. [PubMed] [Google Scholar]
- 4.Hernáiz MJ, Yang HO, Gunay NS, Linhardt RJ. Carbohydr Polym. 2002;48:153–160. [Google Scholar]
- 5.Vongchan P, Warda M, Toyoda H, Toida T, Marks RM, Linhardt RJ. Biochim Biophys Acta. 2005;1721:1–8. doi: 10.1016/j.bbagen.2004.09.007. [DOI] [PubMed] [Google Scholar]
- 6.Warda M, Toida T, Zhang F, Sun P, Munoz E, Linhardt RJ. Glycoconj J. 2006;23:553–561. doi: 10.1007/s10719-006-7668-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Dong J, Peters-Libeu CA, Weisgraber KH, Segelke BW, Rupp B, Capila I, Hernáiz MJ, LeBrun LA, Linhardt RJ. Biochemistry. 2001;40:2826–2834. doi: 10.1021/bi002417n. [DOI] [PubMed] [Google Scholar]
- 8.Peters-Libeu C, Lund-Katz S, Phillips M, Wehrli S, Hernáiz MJ, Capila I, Linhardt RJ, Raffai R, Newhouse YM, Zhou MF, Weisgraber KH. J Biol Chem. 2001;276:39138–39142. doi: 10.1074/jbc.M104746200. [DOI] [PubMed] [Google Scholar]
- 9.Sinnis P, Willnow TE, Briones MRS, Herz J, Nussenzweig V. J Exp Med. 1996;184:945–954. doi: 10.1084/jem.184.3.945. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Rathore D, McCutchan TF, Hernáiz MJ, LeBrun LA, Lang SC, Linhardt RJ. Biochemistry. 2001;40:11518–11524. doi: 10.1021/bi0105476. [DOI] [PubMed] [Google Scholar]
- 11.Rathore D, Hrstka SCL, Sacci JB, de la Vega P, Linhardt RJ, Kumar S, McCutchan TF. J Biol Chem. 2003;278:40905–40910. doi: 10.1074/jbc.M306250200. [DOI] [PubMed] [Google Scholar]
- 12.Zieler H, Nawrocki JP, Shahabuddin M. J Exp Biol. 1999;202:485–495. doi: 10.1242/jeb.202.5.485. [DOI] [PubMed] [Google Scholar]
- 13.Siden-Kiamos I, Louis C. Insect Biochem Mol Biol. 2004;34:679–685. doi: 10.1016/j.ibmb.2004.03.026. [DOI] [PubMed] [Google Scholar]
- 14.Arrighi RBG, Hurd H. Int J Parasitol. 2002;32:91–98. doi: 10.1016/s0020-7519(01)00298-3. [DOI] [PubMed] [Google Scholar]
- 15.Brennan JDG, Kent M, Dhar R, Fujioka H, Kumar N. Proc Natl Acad Sci U S A. 2000;97:13859–13864. doi: 10.1073/pnas.250472597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Mo Myung J, Marshall P, Sinnis P. Mol Biochem Parasitol. 2004;133:53–591. doi: 10.1016/j.molbiopara.2003.09.002. [DOI] [PubMed] [Google Scholar]
- 17.Korochkina S, Barreau C, Pradel G, Jeffery E, Li J, Natarajan R, Shabanowitz J, Hunt D, Frevert U, Vernick KD. Cell Microbiol. 2006;8:163–175. doi: 10.1111/j.1462-5822.2005.00611.x. [DOI] [PubMed] [Google Scholar]
- 18.Barreau C, Touray M, Pimenta PF, Miller LH, Vernick KD. Exp Parasitol. 1995;81:332–343. doi: 10.1006/expr.1995.1124. [DOI] [PubMed] [Google Scholar]
- 19.Sidjanski SP, Vanderberg JP, Sinnis P. Mol Biochem Parasitol. 1997;90:33–41. doi: 10.1016/s0166-6851(97)00124-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Holt RA, Subramanian GM, Halpern A, Sutton GG, Charlab R, Nusskern DR, Wincker P, Clark AG, Ribeiro JM, Wides R, Salzberg SL, Loftus B, Yandell M, Majoros WH, Rusch DB, Lai Z, Kraft CL, Abril JF, Anthouard V, Arensburger P, Atkinson PW, Baden H, de Berardinis V, Baldwin D, Benes V, Biedler J, Blass C, Bolanos R, Boscus D, Barnstead M, Cai S, Center A, Chaturverdi K, Christophides GK, Chrystal MA, Clamp M, Cravchik A, Curwen V, Dana A, Delcher A, Dew I, Evans CA, Flanigan M, Grund-schober-Freimoser A, Friedli L, Gu Z, Guan P, Guigo R, Hillenmeyer ME, Hladun SL, Hogan JR, Hong YS, Hoover J, Jaillon O, Ke Z, Kodira C, Kokoza E, Koutsos A, Letunic I, Levitsky A, Liang Y, Lin JJ, Lobo NF, Lopez JR, Malek JA, McIntosh TC, Meister S, Miller J, Mobarry C, Mongin E, Murphy SD, O’Brochta DA, Pfann-koch C, Qi R, Regier MA, Remington K, Shao H, Sharakhova MV, Sitter CD, Shetty J, Smith TJ, Strong R, Sun J, Thomasova D, Ton LQ, Topalis P, Tu Z, Unger MF, Walenz B, Wang A, Wang J, Wang M, Wang X, Woodford KJ, Wortman JR, Wu M, Yao A, Zdobnov EM, Zhang H, Zhao Q, Zhao S, Zhu SC, Zhimulev I, Coluzzi M, della Torre A, Roth CW, Louis C, Kalush F, Mural RJ, Myers EW, Adams MD, Smith HO, Broder S, Gardner MJ, Fraser CM, Birney E, Bork P, Brey PT, Venter JC, Weissenbach J, Kafatos FC, Collins FH, Hoffman SL. Science. 2002;298:129–149. [Google Scholar]
- 21.Valenzuela JG, Francischetti IM, Pham VM, Garfield MK, Ribeiro JM. Insect Biochem Mol Biol. 2003;33:717–732. doi: 10.1016/s0965-1748(03)00067-5. [DOI] [PubMed] [Google Scholar]
- 22.Esko JD, Selleck SB. Annu Rev Biochem. 2002;71:435–471. doi: 10.1146/annurev.biochem.71.110601.135458. [DOI] [PubMed] [Google Scholar]
- 23.Reisen WK, Boreham PF. J Med Entomol. 1982;19:98–103. doi: 10.1093/jmedent/19.1.98. [DOI] [PubMed] [Google Scholar]
- 24.Toyoda H, Kinoshita-Toyoda A, Selleck SB. J Biol Chem. 2000;275:226–2775. doi: 10.1074/jbc.275.4.2269. [DOI] [PubMed] [Google Scholar]
- 25.Takacs BJ, Girard MF. J Immunol Methods. 1991;143:231–240. doi: 10.1016/0022-1759(91)90048-k. [DOI] [PubMed] [Google Scholar]
- 26.Foster WA. In: Malaria. Julius P, Kreier, editors. Vol. 2. Academic Press, Inc; New York: 1980. pp. 122–139. [Google Scholar]
- 27.Bitter T, Muir HM. Anal Biochem. 1962;4:330–334. doi: 10.1016/0003-2697(62)90095-7. [DOI] [PubMed] [Google Scholar]
- 28.Ribeiro JMC. Insect Biochem Mol Biol. 2003;33:865–882. doi: 10.1016/s0965-1748(03)00080-8. [DOI] [PubMed] [Google Scholar]
- 29.Mada A, Toyoda H, Imanari T. Anal Sci. 1992;8:793–797. [Google Scholar]
- 30.Edens RE, Al-Hakim A, Weiler JM, Rethwisch DG, Fareed J, Linhardt RJ. J Pharm Sci. 1992;81:823–827. doi: 10.1002/jps.2600810821. [DOI] [PubMed] [Google Scholar]
- 31.Linhardt RJ. J Med Chem. 2003;46:2551–2554. doi: 10.1021/jm030176m. [DOI] [PubMed] [Google Scholar]
- 32.Toida T, Yoshida H, Toyoda H, Koshiishi I, Imanari T, Hileman RE, Fromm JR, Linhardt RJ. Biochem J. 1997;322:499–506. doi: 10.1042/bj3220499. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Warda M, Gouda EM, Toida T, Chi L, Linhardt RJ. Comp Biochem Physiol C Toxicol Pharmacol. 2004;136:357–365. doi: 10.1016/j.cca.2003.10.009. [DOI] [PubMed] [Google Scholar]
- 34.Warda M, Mao W, Toida T, Linhardt RJ. Comp Biochem Physiol B. 2003;134:189–197. doi: 10.1016/s1096-4959(02)00250-6. [DOI] [PubMed] [Google Scholar]
- 35.Cesaretti M, Luppi E, Maccari F, Volpi N. Glycobiology. 2004;14:1275–1284. doi: 10.1093/glycob/cwh128. [DOI] [PubMed] [Google Scholar]
- 36.Park PW, Reizes O, Bernfield M. J Biol Chem. 2000;275:29923–29926. doi: 10.1074/jbc.R000008200. [DOI] [PubMed] [Google Scholar]
- 37.Linhardt RJ, Toida T. Acc Chem Res. 2004;37:431–438. doi: 10.1021/ar030138x. [DOI] [PubMed] [Google Scholar]
- 38.Ibrahimi OA, Yeh BK, Eliseenkova AV, Zhang F, Olsen SK, Igarashi M, Aaronson SA, Linhardt RJ, Mohammadi M. Mol Cell Biol. 2005;25:671–684. doi: 10.1128/MCB.25.2.671-684.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Edens RE, LeBrun LA, Weiler JM, Linhardt RJ, Kaul PR. Arch Biochem Biophys. 2001;391:278–285. doi: 10.1006/abbi.2001.2398. [DOI] [PubMed] [Google Scholar]
- 40.Capila I, Linhardt RJ. Angew Chem Int Ed Engl. 2002;41:390–412. doi: 10.1002/1521-3773(20020201)41:3<390::aid-anie390>3.0.co;2-b. [DOI] [PubMed] [Google Scholar]
- 41.Sawitzky D. Med Microbiol Immunol. 1996;184:155–161. doi: 10.1007/BF02456129. [DOI] [PubMed] [Google Scholar]
- 42.Rostand KS, Esko JD. Infect Immun. 1997;65:1–8. doi: 10.1128/iai.65.1.1-8.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Chen Y, Maguire T, Hileman RE, Fromm JR, Esko JD, Linhardt RJ, Marks RM. Nat Med. 1997;3:866–871. doi: 10.1038/nm0897-866. [DOI] [PubMed] [Google Scholar]
- 44.Liu J, Shriver Z, Pope RM, Thorp SC, Raska CS, Yoshida K, Eisenberg RJ, Cohen G, Linhardt RJ, Sasisekharan R. J Biol Chem. 2002;277:33456–33467. doi: 10.1074/jbc.M202034200. [DOI] [PubMed] [Google Scholar]
- 45.Barth H, Schäfer C, Adah MI, Zhang F, Linhardt RJ, Toyoda H, Kinoshita-Toyoda A, Toida T, van Kuppevelt TH, Depla E, von Weizsäcker F, Blum HE, Baumert TF. J Biol Chem. 2003;278:41003–41012. doi: 10.1074/jbc.M302267200. [DOI] [PubMed] [Google Scholar]
- 46.Barth H, Schnober EK, Zhang F, Linhardt RJ, Depla E, Boson B, Cosset FL, Patel A, Blum HE, Baumert TF. J Virol. 2006;80:10579–10590. doi: 10.1128/JVI.00941-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Rider CC. Glycoconj J. 1997;14:639–642. doi: 10.1023/a:1018596728605. [DOI] [PubMed] [Google Scholar]
- 48.Shukla D, Liu J, Blaiklock P, Shworak NW, Bai X, Esko JD, Cohen GH, Eisenberg RJ, Rosenberg RD, Spear PG. Cell. 1999;99:13–22. doi: 10.1016/s0092-8674(00)80058-6. [DOI] [PubMed] [Google Scholar]
- 49.Pradel G, Garapaty S, Frevert U. Mol Microbiol. 2002;45:637–651. doi: 10.1046/j.1365-2958.2002.03057.x. [DOI] [PubMed] [Google Scholar]
- 50.Gantt SM, Clavijo P, Bai X, Esko JD, Sinnis P. J Biol Chem. 1997;272:19205–19213. doi: 10.1074/jbc.272.31.19205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Miller L, Howard R, Carter R, Good M, Nussenzweig V, Nussenzweig R. Science. 1986;234:1349–1356. doi: 10.1126/science.2431481. [DOI] [PubMed] [Google Scholar]
- 52.Warburg A, Miller LH. Science. 1992;255:448–450. doi: 10.1126/science.1734521. [DOI] [PubMed] [Google Scholar]


