Abstract
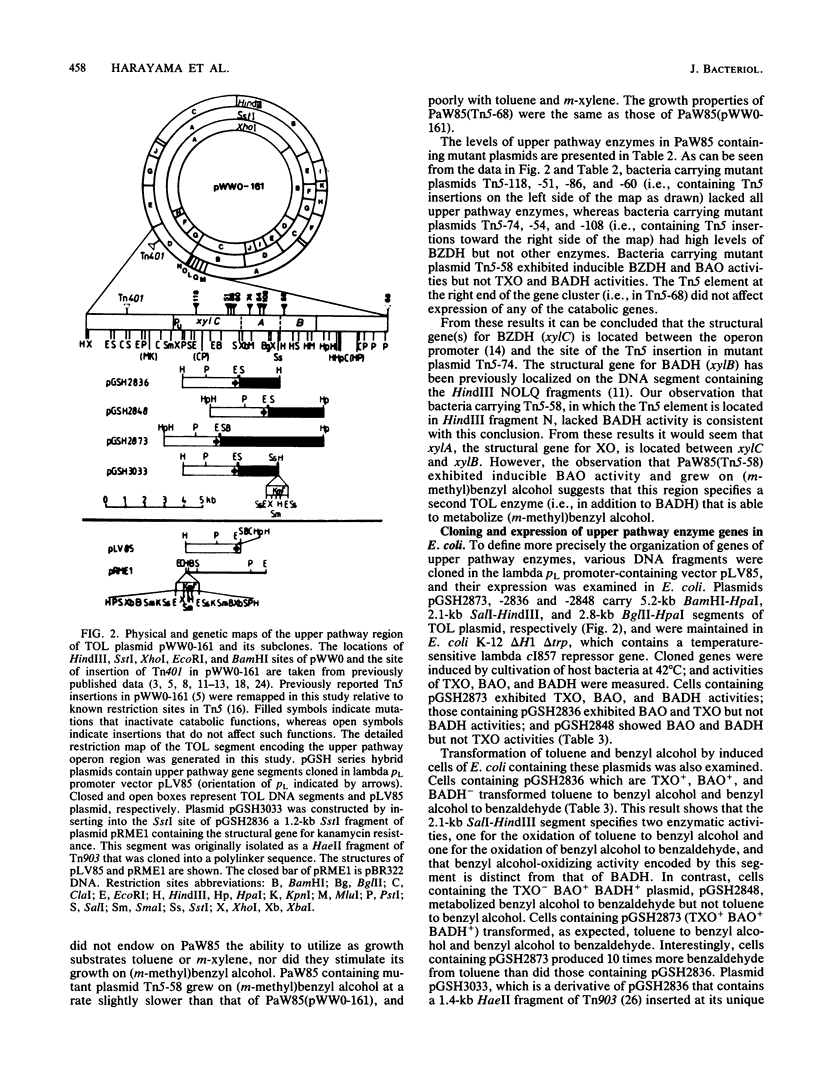
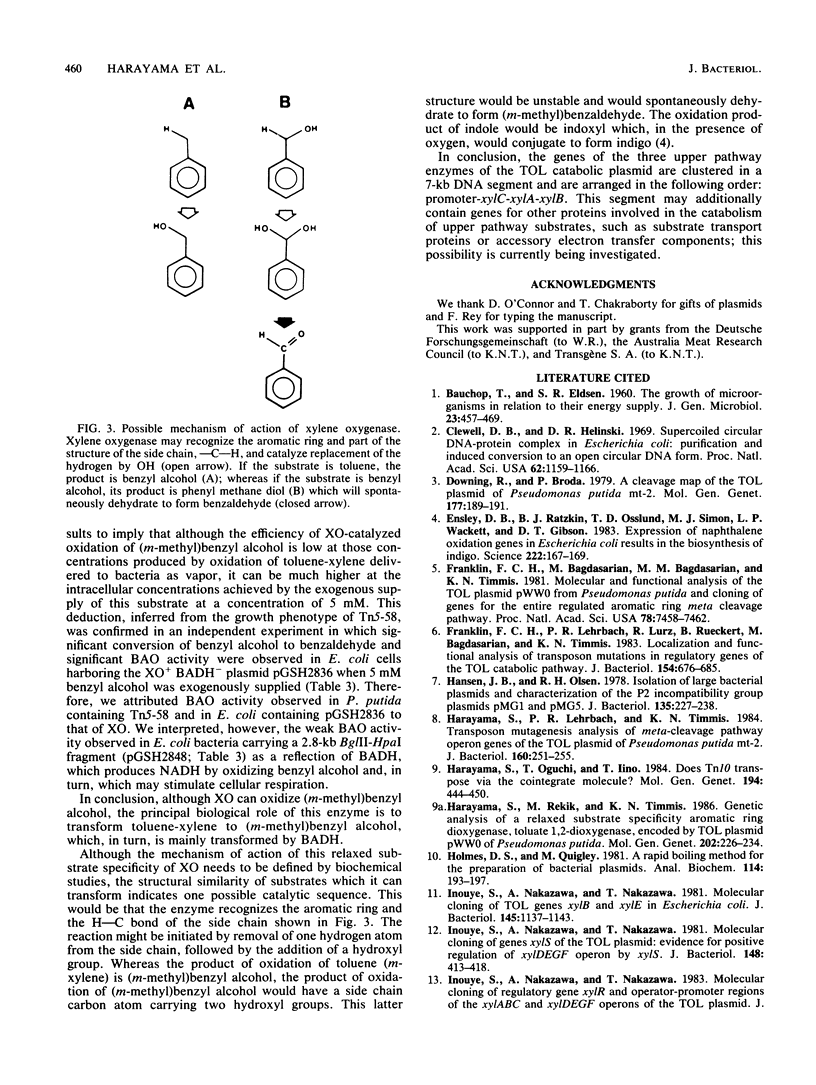
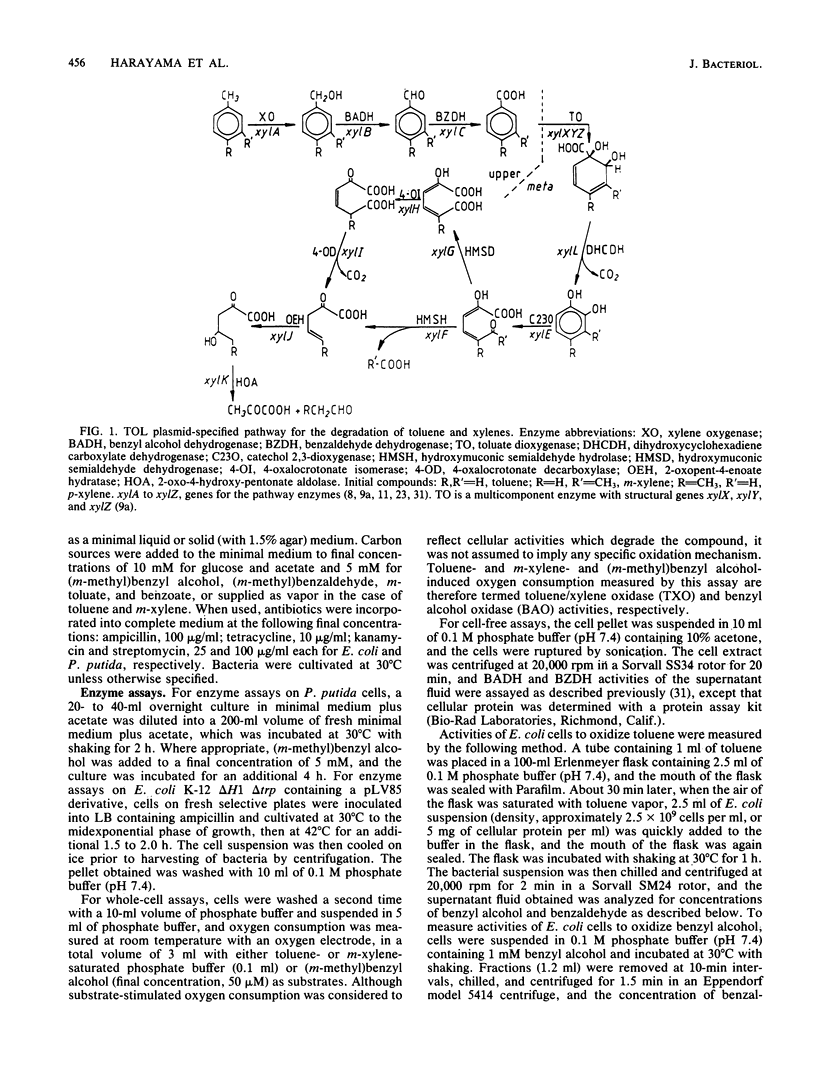
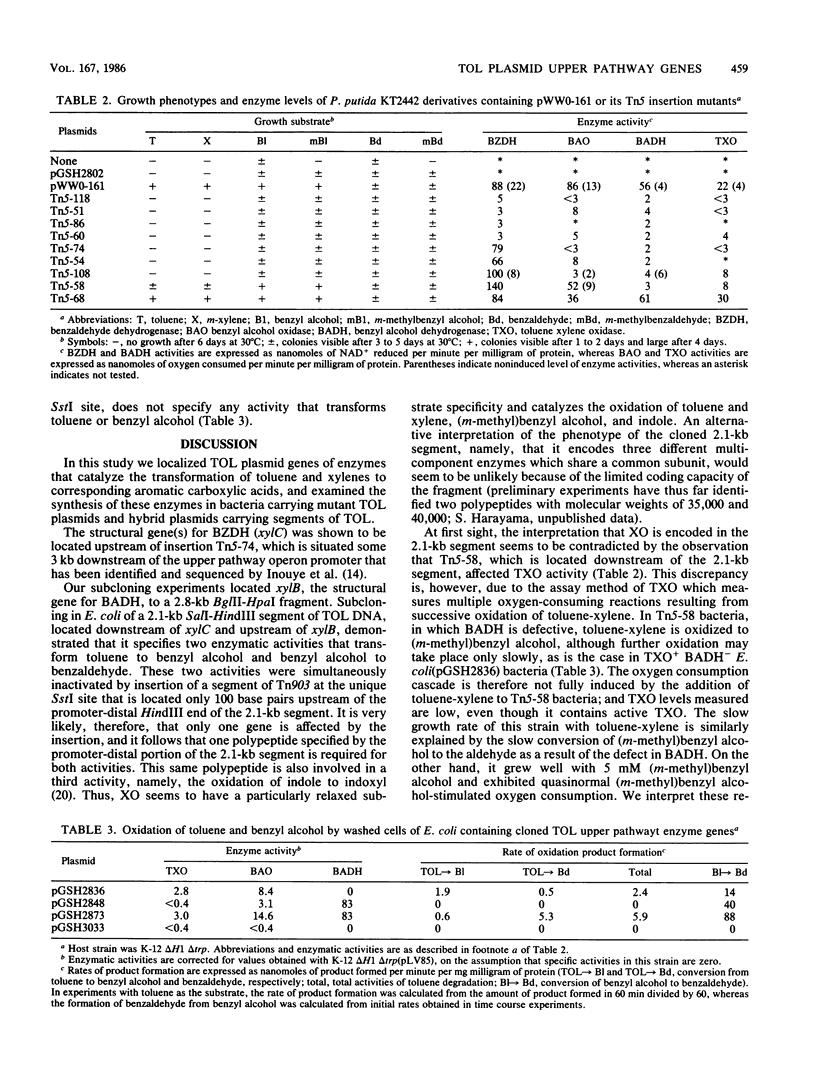
TOL plasmid pWW0 specifies enzymes for the oxidative catabolism of toluene and xylenes. The upper pathway converts the aromatic hydrocarbons to aromatic carboxylic acids via corresponding alcohols and aldehydes and involves three enzymes: xylene oxygenase, benzyl alcohol dehydrogenase, and benzaldehyde dehydrogenase. The synthesis of these enzymes is positively regulated by the product of xylR. Determination of upper pathway enzyme levels in bacteria carrying Tn5 insertion mutant derivatives of plasmid pWW0-161 has shown that the genes for upper pathway enzymes are organized in an operon with the following order: promoter-xylC (benzaldehyde dehydrogenase gene[s])-xylA (xylene oxygenase gene[s])-xylB (benzyl alcohol dehydrogenase gene). Subcloning of the upper pathway genes in a lambda pL promoter-containing vector and analysis of their expression in Escherichia coli K-12 confirmed this order. Two distinct enzymes were found to attack benzyl alcohol, namely, xylene oxygenase and benzyl alcohol dehydrogenase; and their catalytic activities were additive in the conversion of benzyl alcohol to benzaldehyde. The fact that benzyl alcohol is both a product and a substrate of xylene oxygenase indicates that this enzyme has a relaxed substrate specificity.
Full text
PDF




Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- BAUCHOP T., ELSDEN S. R. The growth of micro-organisms in relation to their energy supply. J Gen Microbiol. 1960 Dec;23:457–469. doi: 10.1099/00221287-23-3-457. [DOI] [PubMed] [Google Scholar]
- Clewell D. B., Helinski D. R. Supercoiled circular DNA-protein complex in Escherichia coli: purification and induced conversion to an opern circular DNA form. Proc Natl Acad Sci U S A. 1969 Apr;62(4):1159–1166. doi: 10.1073/pnas.62.4.1159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Downing R., Broda P. A cleavage map of the TOL plasmid of Pseudomonas putida mt-2. Mol Gen Genet. 1979;177(1):189–191. doi: 10.1007/BF00267270. [DOI] [PubMed] [Google Scholar]
- Ensley B. D., Ratzkin B. J., Osslund T. D., Simon M. J., Wackett L. P., Gibson D. T. Expression of naphthalene oxidation genes in Escherichia coli results in the biosynthesis of indigo. Science. 1983 Oct 14;222(4620):167–169. doi: 10.1126/science.6353574. [DOI] [PubMed] [Google Scholar]
- Franklin F. C., Bagdasarian M., Bagdasarian M. M., Timmis K. N. Molecular and functional analysis of the TOL plasmid pWWO from Pseudomonas putida and cloning of genes for the entire regulated aromatic ring meta cleavage pathway. Proc Natl Acad Sci U S A. 1981 Dec;78(12):7458–7462. doi: 10.1073/pnas.78.12.7458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Franklin F. C., Lehrbach P. R., Lurz R., Rueckert B., Bagdasarian M., Timmis K. N. Localization and functional analysis of transposon mutations in regulatory genes of the TOL catabolic pathway. J Bacteriol. 1983 May;154(2):676–685. doi: 10.1128/jb.154.2.676-685.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hansen J. B., Olsen R. H. Isolation of large bacterial plasmids and characterization of the P2 incompatibility group plasmids pMG1 and pMG5. J Bacteriol. 1978 Jul;135(1):227–238. doi: 10.1128/jb.135.1.227-238.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harayama S., Lehrbach P. R., Timmis K. N. Transposon mutagenesis analysis of meta-cleavage pathway operon genes of the TOL plasmid of Pseudomonas putida mt-2. J Bacteriol. 1984 Oct;160(1):251–255. doi: 10.1128/jb.160.1.251-255.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harayama S., Oguchi T., Iino T. Does Tn10 transpose via the cointegrate molecule? Mol Gen Genet. 1984;194(3):444–450. doi: 10.1007/BF00425556. [DOI] [PubMed] [Google Scholar]
- Harayama S., Rekik M., Timmis K. N. Genetic analysis of a relaxed substrate specificity aromatic ring dioxygenase, toluate 1,2-dioxygenase, encoded by TOL plasmid pWW0 of Pseudomonas putida. Mol Gen Genet. 1986 Feb;202(2):226–234. doi: 10.1007/BF00331641. [DOI] [PubMed] [Google Scholar]
- Holmes D. S., Quigley M. A rapid boiling method for the preparation of bacterial plasmids. Anal Biochem. 1981 Jun;114(1):193–197. doi: 10.1016/0003-2697(81)90473-5. [DOI] [PubMed] [Google Scholar]
- Inouye S., Ebina Y., Nakazawa A., Nakazawa T. Nucleotide sequence surrounding transcription initiation site of xylABC operon on TOL plasmid of Pseudomonas putida. Proc Natl Acad Sci U S A. 1984 Mar;81(6):1688–1691. doi: 10.1073/pnas.81.6.1688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inouye S., Nakazawa A., Nakazawa T. Molecular cloning of TOL genes xylB and xylE in Escherichia coli. J Bacteriol. 1981 Mar;145(3):1137–1143. doi: 10.1128/jb.145.3.1137-1143.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inouye S., Nakazawa A., Nakazawa T. Molecular cloning of gene xylS of the TOL plasmid: evidence for positive regulation of the xylDEGF operon by xylS. J Bacteriol. 1981 Nov;148(2):413–418. doi: 10.1128/jb.148.2.413-418.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inouye S., Nakazawa A., Nakazawa T. Nucleotide sequence of the promoter region of the xylDEGF operon on TOL plasmid of Pseudomonas putida. Gene. 1984 Sep;29(3):323–330. doi: 10.1016/0378-1119(84)90061-1. [DOI] [PubMed] [Google Scholar]
- Jorgensen R. A., Rothstein S. J., Reznikoff W. S. A restriction enzyme cleavage map of Tn5 and location of a region encoding neomycin resistance. Mol Gen Genet. 1979;177(1):65–72. doi: 10.1007/BF00267254. [DOI] [PubMed] [Google Scholar]
- Kunz D. A., Chapman P. J. Catabolism of pseudocumene and 3-ethyltoluene by Pseudomonas putida (arvilla) mt-2: evidence for new functions of the TOL (pWWO) plasmid. J Bacteriol. 1981 Apr;146(1):179–191. doi: 10.1128/jb.146.1.179-191.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lehrbach P. R., Ward J., Meulien P., Broda P. Physical mapping of TOL plasmids pWWO and pND2 and various R plasmid-TOL derivatives from Pseudomonas spp. J Bacteriol. 1982 Dec;152(3):1280–1283. doi: 10.1128/jb.152.3.1280-1283.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mermod N., Lehrbach P. R., Reineke W., Timmis K. N. Transcription of the TOL plasmid toluate catabolic pathway operon of Pseudomonas putida is determined by a pair of co-ordinately and positively regulated overlapping promoters. EMBO J. 1984 Nov;3(11):2461–2466. doi: 10.1002/j.1460-2075.1984.tb02156.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meulien P., Broda P. Identification of chromosomally integrated TOL DNA in cured derivatives of Pseudomonas putida PAW1. J Bacteriol. 1982 Nov;152(2):911–914. doi: 10.1128/jb.152.2.911-914.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murray K., Duggleby C. J., Sala-Trepat J. M., Williams P. A. The metabolism of benzoate and methylbenzoates via the meta-cleavage pathway by Pseudomonas arvilla mt-2. Eur J Biochem. 1972 Jul 24;28(3):301–310. doi: 10.1111/j.1432-1033.1972.tb01914.x. [DOI] [PubMed] [Google Scholar]
- Nakazawa T., Inouye S., Nakazawa A. Physical and functional mapping of RP4-TOL plasmid recombinants: analysis of insertion and deletion mutants. J Bacteriol. 1980 Oct;144(1):222–231. doi: 10.1128/jb.144.1.222-231.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakazawa T., Yokota T. Benzoate metabolism in Pseudomonas putida(arvilla) mt-2: demonstration of two benzoate pathways. J Bacteriol. 1973 Jul;115(1):262–267. doi: 10.1128/jb.115.1.262-267.1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oka A., Sugisaki H., Takanami M. Nucleotide sequence of the kanamycin resistance transposon Tn903. J Mol Biol. 1981 Apr 5;147(2):217–226. doi: 10.1016/0022-2836(81)90438-1. [DOI] [PubMed] [Google Scholar]
- Remaut E., Stanssens P., Fiers W. Inducible high level synthesis of mature human fibroblast interferon in Escherichia coli. Nucleic Acids Res. 1983 Jul 25;11(14):4677–4688. doi: 10.1093/nar/11.14.4677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Williams P. A., Murray K. Metabolism of benzoate and the methylbenzoates by Pseudomonas putida (arvilla) mt-2: evidence for the existence of a TOL plasmid. J Bacteriol. 1974 Oct;120(1):416–423. doi: 10.1128/jb.120.1.416-423.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wong C. L., Dunn N. W. Transmissible plasmid coding for the degradation of benzoate and m-toluate in Pseudomonas arvilla mt-2. Genet Res. 1974 Apr;23(2):227–232. doi: 10.1017/s0016672300014853. [DOI] [PubMed] [Google Scholar]
- Worsey M. J., Franklin F. C., Williams P. A. Regulation of the degradative pathway enzymes coded for by the TOL plasmid (pWWO) from Pseudomonas putida mt-2. J Bacteriol. 1978 Jun;134(3):757–764. doi: 10.1128/jb.134.3.757-764.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Worsey M. J., Williams P. A. Metabolism of toluene and xylenes by Pseudomonas (putida (arvilla) mt-2: evidence for a new function of the TOL plasmid. J Bacteriol. 1975 Oct;124(1):7–13. doi: 10.1128/jb.124.1.7-13.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]



